Abstract
Aims
Increasing nitrogen (N) deposition has considerable effects on soil organic matter (SOM) decomposition mediated by soil enzyme activities. Few studies, however, have explored how N addition shapes soil enzyme activity patterns by changing plants, soils and microbes.
Methods
We conducted a five-year field fertilization experiment (0, 5, 10, and 15 g N m−2 yr.−1) to study how N addition affected soil enzyme activity patterns in the topsoil (0–20 cm) and subsoil (20–40 cm) in a Tibetan alpine meadow. Enzyme activity patterns were calculated by the percentage of the sum of all measured enzyme activities. The composition of the plant and microbial communities were evaluated through measuring the abundance of plant functional groups and quantifying microbial phospholipid fatty acids (PLFAs), respectively. Soil pH and available N were also measured.
Results
We found that soil N availability primarily controlled plant community composition, but pH controlled the composition of the microbial community, irrespective of soil depth. Soil enzyme activity patterns differed between two soil depths and among N addition rates. Importantly, N addition shaped soil enzyme activity patterns through the changes in soil pH rather than via the composition of the plant and microbial communities.
Conclusions
Our findings indicate that N addition can affect components of plant-soil system and, in particular, weaken the linkages between plant and microbial communities and enzyme activity patterns. The work suggests that N enrichment-induced soil acidification plays a key role in SOM decomposition and nutrient cycling in the Tibetan meadow ecosystem.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nitrogen (N) addition impacts many ecosystem functions (Manning et al. 2006), for example, productivity and diversity of the plant and microbial communities (Bassin et al. 2007; LeBauer and Treseder 2008; Jing et al. 2016; Riggs and Hobbie 2016; Wardle et al. 2004), which have cascading effects on soil enzyme activities (Ge et al. 2017; Kjoller et al. 2012; Treseder 2008). Soil enzyme activity plays a vital role in understanding the biochemistry of decomposition and nutrient cycling, owing to decomposing soil organic matter (Treseder 2008). Although the influence of N addition on soil enzyme activities has been studied (Riggs and Hobbie 2016; Stone et al. 2014), the direct and indirect roles of N enrichment in modulating soil enzyme activities through the plant-soil system is poorly understood (Schnecker et al. 2015).
There is a complex relationship among N availability, acidification, the resulting composition of the plant and microbial communities and soil enzyme activity. Elevated N availability can stimulate hydrolytic enzyme activities, but depress oxidative enzyme activities directly by modifying stoichiometry of substrates for enzymes (Burns et al. 2013). N addition also indirectly affects soil enzyme activities through the changes in the composition of plant and microbial communities (Sinsabaugh et al. 2008; Kjoller et al. 2012). Furthermore, supplemental N can cause shifts in biomass of different plant functional groups (Bai et al. 2010), which may mediate soil enzyme activities differently, such as increased activity of cellulose enzymes in the presence of legumes (Sanaullah et al. 2011). A global scale meta-analysis study suggests that soil microbial communities are constrained by C and P limitation under N addition, which can change composition of microbial community and then produce more related soil hydrolytic enzyme activities to relax the limitation (Jian et al. 2016). Additionally, N enrichment generally reduces soil fungi biomass, which can suppress activity of soil oxidases (Stone et al. 2014; Jian et al. 2016). It is well-established that N enrichment leads to soil acidification due to increased nitrification (Wei et al. 2013; Riggs and Hobbie 2016), which can directly affect enzyme activities by altering the enzyme molecular conformation (Speir et al. 1999). As a consequence of increased toxic ions (e.g. NH4+, Al3+) and decreased base cations (e.g. Ca2+, Mg2+ and Na+), soil acidification also can affect the composition of plant and microbial communities through the effect on biomass and diversity (Chen et al. 2013; Liu and Greaver 2010). Therefore, the interrelated effects of N availability, acidification and plant and microbial community composition on soil enzyme activities following N addition are important but unresolved issues.
Many biotic and abiotic properties change with soil depth (Rumpel and Kögel-Knabner 2011; Stone et al. 2014), and the soil microenvironment and substrates for soil enzymes vary accordingly. For example, the biomass of plant roots and soil microbes decreases (Fierer et al. 2003; Lamb et al. 2010) and their composition changes with soil depth (Jackson et al. 2009; Schnecker et al. 2015). In addition, soil abiotic properties depend on the soil depth. Indeed, factors such as oxygen, temperature, and pH can decrease from the topsoil to the subsoil, whereas moisture and complex organic matter can increase with depth (Davidson et al. 2012; Wallander et al. 2003). Owing to the above factors, soil depth may affect distribution, production and activity of soil enzymes (Bending et al. 2002; Sinsabaugh et al. 2008). To date, the majority of studies on soil enzyme activities have focused exclusively on topsoil (Eskelinen et al. 2009; van der Putten et al. 2016).
The overarching goal of this study was to better understand how changes in N availability affected patterns in enzyme activity patterns in Tibetan meadows, we conducted a five-year N addition field experiment. Tibetan alpine meadows are characterized by cold temperatures, high SOM content and low soil N and P content (Cao and Zhang 2001). Large-scale N deposition (1.0–1.5 g N m−2 yr.−1) has occurred during recent decades in the Tibetan grasslands (Jia et al. 2014). We intended to explore the following hypotheses: (1) Supplemental N will result in a change in the composition of the plant and microbial communities. Specifically, there will be a shift in biomass of different plant functional groups, increased bacterial biomass, and decreased fungal biomass; (2) N addition will increase soil hydrolytic enzyme activities associated with C, N and P acquisitions while also depressing oxidative enzyme activities, and thus affect soil enzyme activity patterns; (3) All biological production (i.e. plant belowground biomass, soil microbial biomass and enzyme activities) will be greater in topsoil compared to subsoil; (4) Soil enzyme activity patterns in topsoil and subsoil are both related to a set of key parameters (e.g. N availability, pH and composition of plant and microbial communities), and will be mainly driven by microbial community composition, because enzyme activities are generally related to microbial community composition (Sinsabaugh et al. 2008; Nannipieri et al. 2012; Riggs and Hobbie 2016). Structural equation modeling (SEM) is used to reveal the pathway through which N addition influences soil N availability, pH, the composition of plant and microbial communities and enzyme activity patterns.
Materials and methods
Study site
The study was conducted at the Alpine Meadow and Wetland Ecosystems Research Station of Lanzhou University (Azi branch station), in Maqu, Gansu Province, China (101°51′E, 33°40′N, 3500 m a.s.l). The study site has a mean annual temperature of 2.75 °C and precipitation of 601 mm for the period of 2011–2015 (Fig. S1). The soil type is defined as typical alpine meadow soil. Species richness is relatively high, with 25–40 species per 0.25 m2. The plant community is dominated by Kobresia capillifolia, Carex kansuensis, Elymus nutans, Anemone rivularis and Saussurea nigrescens, and the mean aboveground biomass is 440 g m−2 (dry mass) (Zhang et al. 2015).
Experimental design
The experimental site (100 × 100 m) has been fenced since April 2011 with grazing allowed during the winter. We used a randomized block design with five replicates, and four 10 × 20 m plots composed of four fertilization treatments distributed in each block (60 × 80 m). Nitrogen fertilizer (NH4NO3) was applied annually in early May at the following rates: 0 g N m−2 yr.−1 (Control), 5 g N m−2 yr.−1 (N5), 10 g N m−2 yr.−1 (N10), and 15 g N m−2 yr.−1 (N15).
Plant sampling and measurements
In late July 2015, a 0.5 × 0.5 m subplot was harvested for each treatment within a block. We clipped all aboveground vegetation and separated it into four functional groups: grasses, legumes, sedges and forbs. After clipping, we collected topsoil (0–20 cm) and subsoil (20–40 cm) samples in each subplot with three soil cores (4 cm inner diameter). The roots in the two soil depths were separated from soil by running water. Aboveground plant vegetation and roots were both dried at 70 °C for 48 h and weighed as aboveground biomass (biomass of each functional group and total biomass of the four functional groups) and belowground biomass (root biomass in topsoil and subsoil), respectively (g m−2).
Soil sampling and measurements
For each subplot, five additional soil cores (4 cm inner diameter) were collected and pooled together for topsoil and subsoil, respectively. After removal of visible organic debris, each soil sample was sieved (< 2 mm) and divided into three subsamples: 1) the first was air-dried at room temperature for soil physicochemical analysis; 2) the second was stored at −20 °C for analysis of soil enzyme activities within one month; 3) the third was freeze-dried and stored at −80 °C for phospholipid fatty acid (PLFA) analysis.
Soil pH was measured with a soil:water (1:2.5) mixture using a pH meter. Soil NO3−-N and NH4+-N were extracted with 2 M KCl and then analyzed by a flow injection analyzer (San++, Skalar, Netherlands). The composition of the soil microbial community was evaluated by PLFA analysis according to Bossio and Scow (1998). Qualitative and quantitative PLFA analysis were performed by GC-MS (a gas chromatograph combined with a mass spectrometer, Thermo Scientific, Bremen, Germany). An internal standard of 19:0 methyl ester was used for assurance of quantification. The PLFAs i14:0, i15:0, a15:0, i16:0, i17:0 and a17:0 were used as indicators of gram-positive (G+) bacteria; 16:1ω7c, 17:1ω8c, 18:1ω7c, cy17:0 and cy19:0 were used as indicators of gram-negative (G−) bacteria; the PLFAs 10Me16:0, 10Me17:0 and 10Me18:0 were used as indicators of actinomycetes; the PLFAs16:1ω5c and 18:2ω6,9c were used as indicators of fungi (Frostegåd and Bååth 1996; Frostegård et al. 1993; Zelles 1999). Bacteria consisted of G+ and G− bacteria in this study. The PLFAs for soil microbes were expressed as nmol g−1 dry soil.
We measured the activities of β-1,4-glucosidase (EC 3.2.1.21), urease (EC 3.5.1.5), acid phosphomonoesterase (EC 3.1.3.1) and phenol oxidase (EC 1.10.3.2) in topsoil and subsoil. The β-1,4-glucosidase activity was measured using the substrate of ρ-nitrophenyl-β-D-glucopyranoside (Tabatabai 1994). The product ρ-nitrophenol (PNP) was measured colorimetrically at 410 nm with a UV-VIS spectrophotometer (Specord 200 Plus, Analytic Jena, Germany). The acid phosphomonoesterase assay was similar to the β-1,4-glucosidase assay, except that ρ-nitrophenyl phosphate was the substrate (Tabatabai and Bremner 1969). The activity of β-1,4-glucosidase and acid phosphatase was expressed as μmol PNP h−1 g−1 dry soil. For the urease assay, 4 mM urea was used as the substrate. The NH4+ produced was measured at 578 nm, and urease activity was expressed as μmol NH4+ h−1 g−1 dry soil (Kandeler and Gerber 1988). The assay for phenol oxidase activity used 50 mM pyrogallol as a substrate and was measured at 410 nm, and the phenol oxidase activity was expressed as μmol PYGL h −1 g−1 dry soil (Sinsabaugh 2010).
Data analysis
Prior to analysis, all variables were log-transformed to mitigate non-normality and linearity except for soil pH. One-way analysis of variance followed by Duncan’s multiple range test (P < 0.05) was used to evaluate the effects of N addition and soil depth on the measured variables. We also used mixed linear models with N addition and soil depths as fixed factors and block as a random factor to compare the responses of soil N availability, pH, the composition of microbial community and enzyme activities. Second, we calculated enzyme activity patterns (as described by Schnecker et al. 2015), and then obtained Euclidean distance matrixes. We used these matrixes to create Nonmetric Multidimensional Scaling (NMDS) plots to visualize the effects of N addition and soil depth on enzyme activity patterns. We used Permutational Multivariate Analysis of Variance (PERMANOVA) with Euclidean distance matrixes (Anderson 2001), to evaluate the differences in soil enzyme activity patterns between different depths and N addition rates. Third, we classified all variables into the following four groups (Table S1): 1) soil N availability, evaluated by PC1 scores based on NH4+-N and NO3−-N; 2) plant community composition, using PC1 scores based on the abundance of the four plant functional groups and root:shoot ratio; 3) microbial community composition, evaluated by PC1 scores using PLFAs of three microbial groups (bacteria, fungi and actinomycetes), G+:G− ratio and F:B ratio; 4) soil enzyme activity patterns, derived from PC1 scores using the community-weighted activities of β-1,4-D-glucosidase, urease, acid phosphatase and phenol oxidase. Principal component analysis (PCA) summarized each variable group for each soil depth (Table S1), and four PC1 scores explained 50.5–97.8% of the total variance for each group. SEM was performed to analyze hypothetical pathways that may explain how N availability affects soil enzyme activity patterns following N addition. The initial structural equation model was shown in Fig. S2. Most variables or categories examined in our N addition experiment were correlated with one another, making these data sets appropriate for SEM analysis (Table S2).
Statistical analysis was performed using R version 3.3.2 (R Core Team 2016). The mixed linear model was implemented using the “lme4” package (Bates et al. 2018). The PCA, NMDS and PERMANOVA were conducted using the “vegan” package (Oksanen et al. 2018). The SEM analysis was performed using AMOS 21.0 (Amos Development, Spring House, Pennsylvania, USA).
Results
Plant biomass, soil N availability, pH and PLFAs
Aboveground biomass responded to N addition but the pattern varied among functional groups. The total aboveground biomass increased in the N5 and N10, but there was no significant difference in N15 compared to control (Fig. S3a). N addition increased sedge biomass, but there was no significant difference between N addition rates (Fig. S3a). Legume biomass declined in both N10 and N15 (Fig. S3a). Forb biomass tended to decrease with N addition rates and there was a significant difference between N5 and N10. In contrast, grass biomass did not vary significantly with N supplementation (Fig. S3a).
The topsoil biomass followed the same pattern as the total aboveground biomass with increasing N addition rates. The subsoil biomass decreased significantly in all N addition treatments compared to control (Fig. S3b). Additionally, the biomass in topsoil was far higher than that in subsoil (Fig. S3c).
N addition and soil depth had significant effects on pH, NH4+-N and NO3−-N individually, but their interaction only affected NO3−-N (Table 1). N addition strongly increased soil NO3−-N and NH4+-N, and decreased pH (Table 1). Soil pH was lower in topsoil than in subsoil, while soil NH4+-N and NO3−-N were higher in topsoil than subsoil (Table 1).
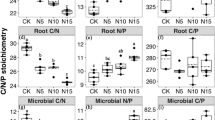
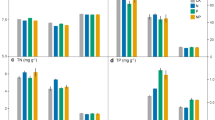
The mixed linear model showed that N addition and soil depth affected PLFAs of bacteria, G+, G−, actinomycetes, fungi and total PLFAs. The F:B ratio was affected by N addition but not by soil depth (Table 2). N15 significantly increased PLFAs of bacteria, G+, G−, actinomycetes and total PLFAs, but significantly decreased PLFAs of fungi and F:B ratio at both soil depths compared to control. For all measured microbial parameters, there was significant difference between N15 and N10 (Fig. 1a–c). All the soil PLFAs decreased with soil depth (Fig. 1d).
Responses of soil microbial community to N addition (a-c) and to soil depth (d), respectively. Values are means ± SE. Different letters above- and below- bars/dots denote significant differences along N addition rates and between soil depths. Actin, G+, G−, Total and F:B represent actinomycetes, gram-positive bacteria, gram-negative bacteria, total microbes and fungi:bacteria ratio, respectively. Control, 0 gN m−2yr.−1; N5, 5 g N m−2 yr.−1; N10, 10 g N m−2 yr.−1; N15, 15 g N m−2 yr.−1
Enzyme activity patterns
The activities of β-1,4-D-glucosidase, urease, phenol oxidase were affected by soil depth and N addition, whereas acid phosphomonoesterase activity was only affected by soil depth (Table 2). In the topsoil, β-1,4-glucosidase activity markedly increased compared to the control and there was no significant difference among N addition rates (Fig. 2a). Urease activity significantly increased with increasing N addition rates, and the phenol oxidase activity decreased in N15 compared to control (Fig. 2a). The effects of N addition on all of the enzyme activities in the subsoil were very similar with those in the topsoil (Fig. 2b). Generally, soil enzyme activity was lower in subsoil than in topsoil (Fig. 2c).
Soil enzyme activity patterns differed more among N addition rates than between soil depths (Fig. 3). Indeed, PERMANOVA pairwise comparisons of enzyme activity patterns between N addition rates in each depth showed significant differences, except that there was a similar pattern between N5 and N10 in the topsoil and between N10 and N15 in the subsoil (Table 3). In contrast, PERMANOVA pairwise comparisons between soil depths for each N addition rate showed significant differences (Table 3).
Non-metric Multidimensional Scaling (NMDS) ordination of soil enzyme activity patterns. NMDS plot of enzyme activity patterns calculated from distance matrices of standardized individual enzyme activities. Colors represent N addition rates (0 gN m−2yr.−1, pink; 5 gN m−2yr.−1, purple; 10 gN m−2yr.−1, green;15 gN m−2yr.−1, blue), and different symbols represent different soil depths (circles are topsoil; triangles are subsoil). The results of PERMANOVA show the effects of nitrogen addition (N) and soil depth (D), indicated by R2 and P value. Error bars represent standard error. N, N addition; D, soil depth; BG, β-1,4-D-glucosidase activity; UREA, urease activity; AP, acid phosphomoesterase activity; PHO, phenol oxidase activity
The SEMs implied by our data supported some of the hypothesized relationships among soil availability, pH, the composition of plant and microbial community and soil enzyme activity patterns (Fig. 4). N availability explained 84–86% of the variation in soil pH, 61–64% of the variation in plant community composition, and 61–74% of the variation in the composition of the soil microbial community through soil pH at both depths (Fig. 4). Surprisingly, the topsoil and subsoil model explained 82% and 87% of the variation in the enzyme activity patterns only by soil pH, respectively, but not by the other measured properties (Fig. 4). This suggests that soil pH played an important role in determining the soil enzyme activity patterns following N addition.
Structural equation model (SEM) of the relationships between soil N availability, pH, plant and microbial community and enzyme activity patterns following N addition in both soil depths. (a) represents topsoil, and model fit statistics for the model were χ2 = 3.799, P = 0.150, RMSEA = 0.021. (b) represents subsoil, and model fit statistics for the model: χ2 = 1.203, P = 0.273, RMSEA = 0.013. Gray dashed arrows show the paths removing from the conceptual model. Solid and dashed arrows indicate significant (P < 0.05) and non-significant relationships (P > 0.05) between the variable of onset arrow and the variable of terminated arrow, respectively. The thickness of the arrows reflects the degree of relationships, and red and green arrows indicate positive and negative relationships. Numbers at arrows are standardized path coefficients. r2 values indicate the variation of response variables explained by the model
Discussion
Our study represents a attempt to how N addition affects soil enzyme activity patterns through plant-soil system. We demonstrated soil pH rather than soil N availability and the composition of the plant and microbial communities was main predictor of changes in soil enzyme activity patterns in a Tibetan alpine meadow. Thus, our results provide empirical evidence of the importance of N enrichment-induced acidification for soil nutrient cycling.
Influence of N addition on the composition of plant and microbial community
In support of our first hypothesis, we found N addition altered the plant community composition with sedge biomass increasing and forbs biomass decreasing (Fig. S3a), probably because sedges are better adapted to high N availability and have higher N-use efficiency than forbs, which are more sensitive to acidification (Bai et al. 2010; van den Berg et al. 2005; Yang et al. 2011). For total aboveground biomass, our results showed that higher biomass occurred in N5 and N10 compared to control (Fig. S3a), which is consistent with previous reports (LeBauer and Treseder 2008; Xi et al. 2014; Xia and Wan 2008).
In our third hypothesis, we speculated that belowground biomass would be higher in topsoil than in subsoil. Consistent with our hypothesis, belowground biomass declined with soil depth (Fig. S3b, c). Indeed, most roots are present in the top 20 cm of alpine meadow soils (Tao et al. 2006), probably because the soil particle size increases and soil nutrient decreases with soil depth (Li et al. 2011). It is interesting to note that the topsoil biomass showed the same trend as the total aboveground biomass with increasing N addition rates (Fig. S3a, b), confirming what has already been reported in alpine grasslands (Yang et al. 2009). In contrast, the subsoil biomass declined with increasing N addition rates (Fig. S3c), probably because sedges, whose biomass increased significantly with N addition, had fibrous root systems, which cannot reach deeper soil layers. In contrast, the tap roots of forbs, such as Anemone rivularis and Saussurea nigrescens, can reach deeper soil layers than fibrous root systems. Moreover, the fibrous root system of sedges experiences rapid cycles of renewal and decay within the growing season, which also limits the access to deep soil layers (Vamerali et al. 2003).
In agreement with our first hypothesis, N addition increased soil PLFAs of gram-positive and gram-negative bacteria, actinomycetes and total PLFAs, but decreased fungal PLFAs and the F:B ratio, regardless of soil depth (Fig. 1a–c; Table 2), changing the composition of soil microbial community. Our findings confirmed the observations of Zhou et al. (2017), but contradicted the report of Canarini et al. (2016), who observed that fertilization did not significantly affect total soil PLFAs and the PLFAs of the microbial groups. Increased N availability can stimulate microbial growth (Zhou et al. 2017), but it can also inhibit microbial growth due to the decreased pH (Chen et al. 2015). Therefore, divergent responses of soil PLFAs to N addition can be the result of the relative role of soil N availability and soil pH. The decline in the F:B ratio coincided with the decline in soil pH (Fig. 4), confirming previous reports (Wei et al. 2013; Xiong et al. 2016; Zhou et al. 2017). This result arises likely because fungi are more sensitive to N addition and have lower nutrient demands than bacteria (Kjoller et al. 2012). Thus, N addition could cause asymmetric changes in the composition of soil microbial community (Zhao et al. 2014).
Consistent with our third hypothesis, we found all of the soil PLFAs declined with depth (Fig. 1d; Table 2), confirming previous reports (Heitkötter et al. 2017; Stone et al. 2014), probably due to the decline in energy sources from root exudation and to the decrease in nutrient availability with soil depth (Fierer et al. 2003; Lamb et al. 2010). In contrast to observations of the F:B ratio in forest soil (Jackson et al. 2009; Stone et al. 2014), the F:B ratio did not change with soil depth in our alpine meadow soil (Fig. 1d; Table 2). Changes in the F:B ratio are due to changes in soil micro-environmental conditions (Hu et al. 2014; Schnecker et al. 2015), and these changes probably did not occur in the alpine meadow because the biomass of plant litter is lower than in a forest (van der Heijden et al. 2008). It is possible that changes in the F:B ratio may occur in deeper soil layers than those we analyzed.
Response of soil enzyme activity patterns to N addition
In our second hypothesis, we speculated that N addition would increase soil hydrolytic enzyme activities associated with C, N and P acquisitions but depress oxidative enzyme activities, and thus influence soil enzyme patterns. Our results partially supported this expectation. Both β-1,4-glucosidase and urease activities increased, whereas acid phosphomonoesterase activity did not change in both soil depths, and phenol oxidase activity decreased at higher N addition rates (Fig. 2a, b). According to the resource allocation theory, sufficient N supply can stimulate C-acquisition enzyme activities, leading to faster decomposition of cellulose-rich litter (Allison et al. 2011; Carreiro et al. 2000). The increase in urease activity with increasing N addition rates confirmed the findings of Jian et al. (2016) and Shi et al. (2018), because soil microorganisms prefer low molecular weight organic N compounds, such as urea, to inorganic N (Dunn et al. 2006; Schimel and Weintraub 2003). The lack of a significant effect of N addition on acid phosphomonoesterase activity is in agreement with previous reports (Geisseler and Scow 2014; Jing et al. 2016). This result depends on the fact that this activity is not related to soil N reactions but to P reactions, given that enzyme synthesis is repressed by inorganic P (Nannipieri et al. 2011; Nannipieri et al. 2012). The phenol oxidase activity decreased at higher N addition rates in our study, possibly because the growth of some fungal species producing ligninolytic enzymes is depressed (Fog 1988). It is worth noting that most soil enzyme activity patterns were significantly different among N addition rates at both soil depths (Fig. 3; Table 3), possibly resulting from the obvious responses of phenol oxidase and urease to N addition rates (Fig. 2).
In support of our third hypothesis, our study revealed all enzyme activities decreased with soil depth (Fig. 2d; Table 2), which is consistent with numerous reports (Heitkötter et al. 2017; Šnajdr et al. 2008; Stone et al. 2014; Venkatesan and Senthurpandian 2006). Overall, the contribution of plant roots and litter to subsoil C pools is small, causing the reduction in soil enzyme activities with soil depth (Hernández and Hobbie 2010). We also found that topsoil enzyme activity patterns differed from subsoil enzyme activity patterns (Fig. 3; Table 3) due to the decrease in the activity of each enzyme with soil depth, as discussed above. Future studies that focus on detecting the expression of the relative enzyme-encoding genes are needed to better understand the underlying mechanisms of soil location-specific patterns of enzyme activity.
The role of abiotic and biotic factors in determining soil enzyme activity patterns following N addition
In contrast to our fourth hypothesis, we found that only soil pH but not N availability and the composition of plant and microbial communities affected enzyme activity patterns in both soil depths (Fig. 4). Manning et al. (2006) suggested that abiotic effects of N deposition on ecosystem functions could be more important than the changes in the composition of the plant community. Indeed, our results showed the variation of plant community following N addition did not affect enzyme activity patterns, which may be partially explained by removing substantial litter owing to severe winter grazing in our study. Another explanation might be that impact of N addition on the overall composition of the plant community was relatively small in our study. Although soil pH affected the composition of microbial community (Fig. 4), as indicated by other studies (Delgado-Baquerizo et al. 2016; Sun et al. 2018), Surprisingly, we found no effect of microbial community composition on soil enzyme activity patterns, which was conflicted with many previous reports (Nannipieri et al. 2012; Riggs and Hobbie 2016). In fact, the relative importance of microbial community composition vs environmental factors in regulating enzyme expression remains unclear (Allison et al. 2007; Frossard et al. 2012). Notably, some recent reports showed variation in soil enzyme activities was better accounted for with soil physicochemical characteristics than microbial community composition (Bowles et al. 2014; Schnecker et al. 2014). In this study, N availability did not directly affect soil enzyme activity patterns, but indirectly though soil pH. The probable explanation is that the primary controlling factor for soil microbes shifts from resource availability (e.g. soil N availability) to soil acidification following N addition (Wei et al. 2013). Among our measured variables, soil pH significantly negatively affected enzyme activity patterns, indicating soil pH was a dominant controller of soil microbial enzyme activities following N addition in alpine meadow soil.
Conclusion
In conclusion, our results showed that N addition shaped soil enzyme activity patterns mediated by increasing soil acidification but not the changes in the composition of the plant and microbial communities. Our study provides a new mechanistic understanding of how N addition affects soil enzyme activities through the plant-soil system in the Tibetan meadow. More research involving the detection and the expression of the enzyme-encoding genes and combined with the determination of the relative enzyme activities is needed to help expand our understanding of nutrient cycling in this high altitude ecosystem.
Abbreviations
- C:
-
Carbon
- N:
-
Nitrogen
- P:
-
Phosphorus
- SOM:
-
Soil organic matter
- SEM:
-
Structural equation model
- PLFAs:
-
Phospholipid fatty acids
- Control:
-
0 g N m−2 yr.−1
- N5:
-
5g N m−2 yr.−1
- N10:
-
10 g N m−2 yr.−1
- N15:
-
15 g N m−2 yr.−1
- G+:G− ratio:
-
Gram-positive bacteria:gram-negative bacteria
- F:B:
-
Fungi:bacteria
- NMDS:
-
Nonmetric multidimensional scaling
- PERMANOVA:
-
Permutational multivariate analysis of variance
- PCA:
-
Principal component analysis
- BG:
-
β-1,4-glucosidase
- UREA:
-
Urease
- AP:
-
Acid phosphomonoesterase
- PHO:
-
Phenol oxidase
References
Allison VJ, Condron LM, Peltzer DA, Richardson SJ, Turner BL (2007) Changes in enzyme activities and soil microbial community composition along carbon and nutrient gradients at the Franz Josef chronosequence, New Zealand. Soil Biol Biochem 39:1770–1781
Allison SD, Weintraub MN, Gartner TB, Waldrop MP (2011) Evolutionary-economic principles as regulators of soil enzyme production and ecosystem function. In: Shukla G, Varma A (eds) Soil enzymology, 1st edn. Springer, Berlin, pp 229–243
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46
Bai Y, Wu J, Clark CM, Naeem S, Pan Q, Huang J, Zhang L, Han X (2010) Tradeoffs and thresholds in the effects of nitrogen addition on biodiversity and ecosystem functioning: evidence from inner Mongolia grasslands. Global Chang Biol 16:358–372
Bassin S, Volk M, Suter M, Buchmann N, Fuhrer J (2007) Nitrogen deposition but not ozone affects productivity and community composition of subalpine grassland after 3 yr of treatment. New Phytol 175:523–534
Bates D, Maechler M, Bolker B, Walker S (2018) lme4: linear mixed-effects models using ‘Eigen’ and S4. R package version 1:1–17 http://cran.r-project.org/package=lme4. Accessed 3 Apr 2018
Bending GD, Turner MK, Jones JE (2002) Interactions between crop residue and soil organic matter quality and the functional diversity of soil microbial communities. Soil Biol Biochem 34:1073–1082
Bossio DA, Scow KM (1998) Impacts of carbon and flooding on soil microbial communities: phospholipid fatty acid profiles and substrate utilization patterns. Microb Ecol 35:265–278
Bowles TM, Acosta-Martínez V, Calderón F, Jackson LE (2014) Soil enzyme activities, microbial communities, and carbon and nitrogen availability in organic agroecosystems across an intensively-managed agricultural landscape. Soil Biol Biochem 68:252–262
Burns RG, DeForest JL, Marxsen J, Sinsabaugh RL, Stromberger ME, Wallenstein MD, Weintraub MN, Zoppini A (2013) Soil enzymes in a changing environment: current knowledge and future directions. Soil Biol Biochem 58:216–234
Canarini A, Carrillo Y, Mariotte P, Ingram L, Dijkstra FA (2016) Soil microbial community resistance to drought and links to C stabilization in an Australian grassland. Soil Biol Biochem 103:171–180
Cao GM, Zhang JX (2001) Soil nutrition and substance cycle of Kobresia meadow. In: Zhou XM (ed) Alpine Kobresia meadows in China. Science, Beijing, pp 58–147
Carreiro MM, Sinsabaugh RL, Repert DA, Parkhurst DF (2000) Microbial enzyme shifts explain litter decay responses to simulated nitrogen deposition. Ecology 81:2359–2365
Chen DM, Lan ZC, Bai X, Grace JB, Bai YF (2013) Evidence that acidification-induced declines in plant diversity and productivity are mediated by changes in below-ground communities and soil properties in a semi-arid steppe. J Ecol 101:1322–1334
Chen DM, Lan ZC, Hu SJH, Bai YF (2015) Effects of nitrogen enrichment on belowground communities in grassland: relative role of soil nitrogen availability vs. soil acidification. Soil Biol Biochem 89:99–108
Davidson EA, Samanta S, Caramori SS, Savage K (2012) The dual Arrhenius and Michaelis-Menten kinetics model for decomposition of soil organic matter at hourly to seasonal time scales. Global Chang Biol 18:371–384
Delgado-Baquerizo M, Grinyer J, Reich PB, Singh BK (2016) Relative importance of soil properties and microbial community for soil functionality: insights from a microbial swap experiment. Funct Ecol 30:1862–1873
Dunn RM, Mikola J, Bol R, Bardgett RD (2006) Influence of microbial activity on plant-microbial competition for organic and inorganic nitrogen. Plant Soil 289:321–334
Eskelinen A, Stark S, Mannisto M (2009) Links between plant community composition, soil organic matter quality and microbial communities in contrasting tundra habitats. Oecologia 161:113–123
Fierer N, Schimel JP, Holden PA (2003) Variations in microbial community composition through two soil depth profiles. Soil Biol Biochem 35:167–176
Fog K (1988) The effect of added nitrogen on the rate of decomposition of organic matter. Biol Rev 63:433–462
Frossard A, Gerull L, Mutz M, Gessner MO (2012) Disconnect of microbial structure and function: enzyme activities and bacterial communities in nascent stream corridors. ISME J 6:680–691
Frostegåd A, Bååth E (1996) The use of phospholipid fatty acid analysis to estimate bacterial and fungal biomass in soil. Biol Fertil Soils 22:59–65
Frostegård Å, Bååth E, Tunlio A (1993) Shifts in the structure of soil microbial communities in limed forests as revealed by phospholipid fatty acid analysis. Soil Biol Biochem 25:723–730
Ge T, Wei X, Razavi BS, Zhu Z, Hu Y, Kuzyakov Y, Jones DL, Wu J (2017) Stability and dynamics of enzyme activity patterns in the rice rhizosphere: effects of plant growth and temperature. Soil Biol Biochem 113:108–115
Geisseler D, Scow KM (2014) Long-term effects of mineral fertilizers on soil microorganisms – a review. Soil Biol Biochem 75:54–63
Heitkötter J, Niebuhr J, Heinze S, Marschner B (2017) Patterns of nitrogen and citric acid induced changes in C-turnover and enzyme activities are different in topsoil and subsoils of a sandy Cambisol. Geoderma 292:111–117
Hernández DL, Hobbie SE (2010) The effects of substrate composition, quantity, and diversity on microbial activity. Plant Soil 335:397–411
Hu YJ, Xiang D, Veresoglou SD, Chen FL, Chen YL, Hao ZP, Zhang X, Chen BD (2014) Soil organic carbon and soil structure are driving microbial abundance and community composition across the arid and semi-arid grasslands in northern China. Soil Biol Biochem 77:51–57
Jackson CR, Liew KC, Yule CM (2009) Structural and functional changes with depth in microbial communities in a tropical Malaysian peat swamp forest. Microb Ecol 57:402–412
Jia Y, Yu G, He N, Zhan X, Fang H, Sheng W, Zuo Y, Zhang D, Wang Q (2014) Spatial and decadal variations in inorganic nitrogen wet deposition in China induced by human activity. Sci Rep 4:3763
Jian S, Li J, Chen J, Wang G, Mayes MA, Dzantor KE, Hui D, Luo Y (2016) Soil extracellular enzyme activities, soil carbon and nitrogen storage under nitrogen fertilization: a meta-analysis. Soil Biol Biochem 101:32–43
Jing X, Yang X, Ren F, Zhou H, Zhu B, He JS (2016) Neutral effect of nitrogen addition and negative effect of phosphorus addition on topsoil extracellular enzymatic activities in an alpine grassland ecosystem. Appl Soil Ecol 107:205–213
Kandeler E, Gerber H (1988) Short-term assay of soil urease activity using colorimetric determination of dmmonium. Biol Fertil Soils 6:68–72
Kjoller R, Nilsson LO, Hansen K, Schmidt IK, Vesterdal L, Gundersen P (2012) Dramatic changes in ectomycorrhizal community composition, root tip abundance and mycelial production along a stand-scale nitrogen deposition gradient. New Phytol 194:278–286
Lamb EG, Kennedy N, Siciliano SD (2010) Effects of plant species richness and evenness on soil microbial community diversity and function. Plant Soil 338:483–495
LeBauer DS, Treseder KK (2008) Nitrogen limitation of net primary productivity in terrestrial ecosystems is globally distributed. Ecology 89:371–379
Li X, Zhang X, Wu J, Shen Z, Zhang Y, Xu X, Fan Y, Zhao Y, Yan W (2011) Root biomass distribution in alpine ecosystems of the northern Tibetan plateau. Environ Earth Sci 64:1911–1919
Liu L, Greaver TL (2010) A global perspective on belowground carbon dynamics under nitrogen enrichment. Ecol Lett 13:819–828
Manning P, Newington JE, Robson HR et al (2006) Decoupling the direct and indirect effects of nitrogen deposition on ecosystem function. Ecol Lett 9:1015–1024
Nannipieri P, Giagnoni L, Landi L, Renella G (2011) Role of phosphatase enzymes in soil. In: Bunemann EK et al (eds) Phosphorus in action. Springer Verlag, Berlin Heidelberg, pp 215–243
Nannipieri P, Giagnoni L, Renella G, Puglisi E, Ceccanti B, Masciandaro G, Fornasier F, Moscatelli MC, Marinari S (2012) Soil enzymology: classical and molecular approaches. Biol Fertil Soils 48:743–762
Oksanen J, Blanchet FG, Friendly M et al (2018) Vegan:community ecology package. R package version 2:5–2. https://CRAN.R-project.org/package=vegan. Accessed 17 May 2018
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/. Accessed 10 March 2016
Riggs CE, Hobbie SE (2016) Mechanisms driving the soil organic matter decomposition response to nitrogen enrichment in grassland soils. Soil Biol Biochem 99:54–65
Rumpel C, Kögel-Knabner I (2011) Deep soil organic matter—a key but poorly understood component of terrestrial C cycle. Plant Soil 338:143–158
Sanaullah M, Blagodatskaya E, Chabbi A, Rumpel C, Kuzyakov Y (2011) Drought effects on microbial biomass and enzyme activities in the rhizosphere of grasses depend on plant community composition. Appl Soil Ecol 48:38–44
Schimel JP, Weintraub MN (2003) The implications of exoenzyme activity on microbial carbon and nitrogen limitation in soil: a theoretical model. Soil Biol Biochem 35:549–563
Schnecker J, Wild B, Hofhansl F et al (2014) Effects of soil organic matter properties and microbial community composition on enzyme activities in cryoturbated arctic soils. PLoS One 9:e94076
Schnecker J, Wild B, Takriti M et al (2015) Microbial community composition shapes enzyme patterns in topsoil and subsoil horizons along a latitudinal transect in Western Siberia. Soil Biol Biochem 83:106–115
Shi B, Zhang J, Wang C, Ma J, Sun W (2018) Responses of hydrolytic enzyme activities in saline-alkaline soil to mixed inorganic and organic nitrogen addition. Sci Rep 8:4543
Sinsabaugh RL (2010) Phenol oxidase, peroxidase and organic matter dynamics of soil. Soil Biol Biochem 42:391–404
Sinsabaugh RL, Lauber CL, Weintraub MN et al (2008) Stoichiometry of soil enzyme activity at global scale. Ecol Lett 11:1252–1264
Šnajdr J, Valášková V, Merhautová V, Herinková J, Cajthaml T, Baldrian P (2008) Spatial variability of enzyme activities and microbial biomass in the upper layers of Quercus petraea forest soil. Soil Biol Biochem 40:2068–2075
Speir TW, Kettles HA, Percival HJ, Parshotam A (1999) Is soil acidifcation the cause of biochemical responses when soils are amended with heavy metal salts? Soil Biol Biochem 31:1953–1961
Stone MM, DeForest JL, Plante AF (2014) Changes in extracellular enzyme activity and microbial community structure with soil depth at the Luquillo critical zone observatory. Soil Biol Biochem 75:237–247
Sun YF, Shen JP, Zhang CJ, Zhang LM, Bai WM, Fang Y, He JZ (2018) Responses of soil microbial community to nitrogen fertilizer and precipitation regimes in a semi-arid steppe. J Soils Sediment 18:762–774
Tabatabai MA (1994) Soil enzymes. In: Mickelson SH, Bifham JM (eds) Methods of soil analysis, part 2: microbiological and biochemical properties. Soil Science Society of America, Madison, pp 775–833
Tabatabai MA, Bremner JM (1969) Use of p-nitrophenyl phosphate for assay of soil phosphatase activity. Soil Biol Biochem 1:301–307
Tao Z, Shen C, Gao Q, Sun Y, Yi W, Li Y (2006) Soil organic carbon storage and vertical distribution of alpine meadow on the Tibetan plateau. Acta Geograph Sin 61:720–728
Treseder KK (2008) Nitrogen additions and microbial biomass: a meta-analysis of ecosystem studies. Ecol Lett 11:1111–1120
Vamerali T, Ganis A, Bona S, Mosca G (2003) Fibrous root turnover and growth in sugar beet (Beta vulgaris var. saccharifera) as affected by nitrogen shortage. Plant Soil 255:169–177
van den Berg LJ, Dorland E, Vergeer P, Hart MA, Bobbink RJG (2005) Decline of acid-sensitive plant species in heathland can be attributed to ammonium toxicity in combination with low pH. New Phytol 166:551–564
van der Heijden MG, Bardgett RD, van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310
van der Putten WH, Bradford MA, Pernilla Brinkman E, van de Voorde TFJ, Veen GF, Bailey JK (2016) Where, when and how plant-soil feedback matters in a changing world. Funct Ecol 30:1109–1121
Venkatesan S, Senthurpandian VK (2006) Comparison of enzyme activity with depth under tea plantations and forested sites in South India. Geoderma 137:212–216
Wallander H, Nilsson LO, Hagerberg D, Rosengren U (2003) Direct estimates of C : N ratios of ectomycorrhizal mycelia collected from Norway spruce forest soils. Soil Biol Biochem 35:997–999
Wardle DA, Bardgett RD, Klironomos JN, Setala H, van der Putten WH, Wall DH (2004) Ecological linkages between aboveground and belowground biota. Science 304:1629–1633
Wei C, Yu Q, Bai E, Lu X, Li Q, Xia J, Kardol P, Liang W, Wang Z, Han X (2013) Nitrogen deposition weakens plant-microbe interactions in grassland ecosystems. Global Chang Biol 19:3688–3697
Xi N, Carrère P, Bloor JMG (2014) Nitrogen form and spatial pattern promote asynchrony in plant Soilresponses to nitrogen inputs in a temperate grassland. Soil Biol Biochem 71:40–47
Xia J, Wan S (2008) Global response patterns of terrestrial plant species to nitrogen addition. New Phytol 179:428–439
Xiong QL, Pan KW, Zhang L, Wang YJ, Li W, He XJ, Luo HY (2016) Warming and nitrogen deposition are interactive in shaping surface soil microbial communities near the alpine timberline zone on the eastern Qinghai-Tibet plateau, southwestern China. Appl Soil Ecol 101:72–83
Yang YH, Fang JY, Ji CJ, Han WX (2009) Above- and belowground biomass allocation in Tibetan grasslands. J Veg Sci 20:177–184
Yang ZL, van Ruijven J, Du GZ (2011) The effects of long-term fertilization on the temporal stability of alpine meadow communities. Plant Soil 345:315–324
Zelles L (1999) Fatty acid patterns of phospholipids and lipopolysaccharides in the characterisation of microbial communities in soil: a review. Biol Fertil Soils 29:111–129
Zhang P, Zhou X, Li J, Guo Z, Du G (2015) Space resource utilisation: a novel indicator to quantify species competitive ability for light. Sci Rep 5:16832
Zhao CZ, Zhu LY, Liang J, Yin HJ, Yin CY, Li DD, Zhang NN, Liu Q (2014) Effects of experimental warming and nitrogen fertilization on soil microbial communities and processes of two subalpine coniferous species in eastern Tibetan plateau, China. Plant Soil 382:189–201
Zhou Z, Wang C, Zheng M, Jiang L, Luo Y (2017) Patterns and mechanisms of responses by soil microbial communities to nitrogen addition. Soil Biol Biochem 115:433–441
Acknowledgements
We thank Paolo Nannipieri for valuable comments and assistance in improving this manuscript. This work was supported by National Natural Science Foundation of China (31870435 to S.T.Z) and the National Key Research and Development Program of China (2018YFD0502401 to S.T.Z).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Elizabeth M Baggs.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 550 kb)
Rights and permissions
About this article
Cite this article
Liu, X., Zhang, S. Nitrogen addition shapes soil enzyme activity patterns by changing pH rather than the composition of the plant and microbial communities in an alpine meadow soil. Plant Soil 440, 11–24 (2019). https://doi.org/10.1007/s11104-019-04054-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-019-04054-5