Abstract
Aims
Ammonia oxidation and denitrification are crucial for nitrogen (N) availability and nitrous oxide production in N-limited permafrost soils. However, it remains unclear about the relative roles of abiotic and biotic properties in controlling the abundance and activity of ammonia-oxidizing and denitrifying microorganisms in permafrost-affected soils.
Methods
We analysed the potential ammonia oxidation and denitrification rates (PAO and PDR), the abundance of archaeal amoA, bacterial amoA, nirK, nirS and nosZ genes, soil characteristics, climatic and plant properties across two vegetation types in Tibetan permafrost-affected soils. The relative importance of abiotic and biotic properties in driving functional N gene abundance, PAO and PDR were assessed using variation partition analysis (VPA) and random forest (RF) model.
Results
The functional N gene abundance and PDR were lower in alpine steppe than in alpine meadow. Variations in the PAO and PDR and functional N gene abundance were mainly explained by abiotic variables such as organic carbon and total N, then by plant properties such as plant N concentration, plant species richness and productivity based on the VPA. The RF model showed that abiotic properties (e.g., precipitation) and plant properties (e.g., plant N concentration or plant productivity) predicted the PDR and the abundance of functional N genes. Both VPA and RF model showed that the PAO and PDR could be determined by the abundance of functional N genes such as archaeal amoA gene and nosZ gene, respectively.
Conclusions
Our study highlights that abiotic and plant properties are important predictors of the abundance and activity of ammonia-oxidizing and denitrifying communities in permafrost-affected regions, implying that plant properties, which were previously overlooked, should be incorporated into ecosystem models for improved prediction of belowground N process rates in a changing environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Permafrost-affected soils are large nitrogen (N) reservoirs, which stores more than 67 billion tons of total N within three metres from the surface (Harden et al. 2012). This substantial permafrost N storage may be altered by global warming, which is predicted to cause permafrost thawing. After the permafrost thaws, permafrost-affected soils may become important sources of nitrous oxide (N2O) (Elberling et al. 2010; Voigt et al. 2017a, b; Yang et al. 2018), leading to strong climate change feedbacks. This is because N2O has a global warming potential of approximately 300 times that of carbon dioxide (CO2) and is an important substance that damages the ozone layer (Ravishankara et al. 2009). Ammonia oxidation and denitrification are the two dominant biological processes that generate N2O from soils, and are mediated by the functional microorganisms involved in N cycling (Morales et al. 2010; Hu et al. 2015a). Therefore, it is of paramount importance to elucidate the factors that control the distribution patterns of these particular groups of microorganisms and their specialized functioning in permafrost-affected regions to predict the responses of permafrost N cycling to future global warming.
Over the past decades, numerous studies addressing functional N gene abundance and potential ammonia oxidation (PAO) and denitrification rate (PDR) have focused on various soils, including agricultural, grassland, forest and Arctic soils (Jia and Conrad 2009; Petersen et al. 2012; Shen et al. 2011; Pierre et al. 2017). Among such studies, the abundance of functional genes has been shown to be a good predictor of PAO and PDR (Petersen et al. 2012; Pierre et al. 2017). In general, ammonia-oxidizing archaea (AOA) are important drivers of autotrophic ammonia oxidation in acidic soils or those with a low ammonium concentration (Zhang et al. 2012; Alves et al. 2013, 2018), whereas significant activities of ammonia-oxidizing bacteria (AOB) have been found in soils with high ammonium concentrations and in response to N fertilization (Jia and Conrad 2009; Verhamme et al. 2011). Among denitrifiers, the nirS and nirK genes are linked to nitrite reduction, a key step in the denitrification pathway, and these genes encode functionally similar but structurally different nitrite reductases (Zumft 1997). The nosZ gene encodes the N2O reductase which catalyzes the reduction of N2O to N2, which is the final step of denitrification. In addition to functional N gene abundance, edaphic variables such as soil organic carbon (SOC), soil moisture (Attard et al. 2011; Banerjee and Siciliano 2012a, b), soil ammonium and nitrate availability (Enwall et al. 2010), have been described as being important regulators of potential process rates. These factors could indirectly affect N cycling processes through changes in functional N gene abundance. Although these studies have provided some evidence of the response patterns of N-cycling process rates to edaphic variables and the abundance of functional microorganisms, it remains unknown whether PAO and PDR are linked to the abundance of ammonia-oxidizing and denitrifying communities in permafrost-affected regions over large geographic scale.
Recently, a number of studies have focused on the influence of plant properties, such as plant diversity and productivity, on microbial communities (Prober et al. 2015; Chen et al. 2017a). In particular, multiple studies have examined the responses of N cycling to plant diversity and productivity in manipulative field experiments (Oelmann et al. 2011; Mueller et al. 2013). These studies have shown that higher plant diversity increases plant productivity and soil ammonium availability due to the increased mineralization of soil organic N but decreased soil nitrate content due to greater N uptake. Recent studies have also proposed that there is competition between plants and functional microorganisms for ammonia and nitrate and that the influence of plant species on microbial communities is dependent on soil N availability (Moreau et al. 2015). Therefore, plant properties may interact with available N to regulate the functional microorganisms involved in N cycling and further ammonia oxidation and denitrification. However, to date, it remains unclear the relative importance of plant properties and abiotic variables in controlling functional N gene abundance, PAO and PDR in permafrost-affected ecosystems at a broad scale.
In this study, we analysed the putative driving factors of the abundance and activity of N-cycling microbial communities across Tibetan permafrost-affected regions. We first hypothesized that the abundance of ammonia-oxidizing microorganisms and ammonia oxidation were greater in soils with higher plant diversity and productivity due to the greater soil ammonia availability, whereas the abundance of denitrifiers and denitrification were lower in soils with higher plant diversity and productivity due to more efficient N capture and lower nitrate availability (Mueller et al. 2013; Wei et al. 2017). We also hypothesized that the functional N gene abundance could be used to predict ammonia oxidation and denitrification over large scales (Petersen et al. 2012; Pierre et al. 2017).
Materials and methods
Study area and sampling
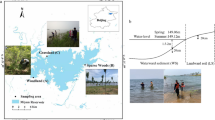
The Tibetan Plateau is the largest high-elevation permafrost region, with a total permafrost area of ~1.35 × 106 km2 (Ran et al. 2012). Of this permafrost, mountain permafrost and plateau permafrost are the two main permafrost types, with areas of 0.30 × 106 and 1.05 × 106 km2, respectively (Fig. 1) (Li and Cheng 1996; Ran et al. 2012). The mean annual temperature (MAT) and mean annual precipitation (MAP) vary from -4.9 to 6.1 °C and from 84.3 to 593.9 mm, respectively. In this region, there are two dominant vegetation types (alpine meadow and alpine steppe). The alpine meadow has higher plant diversity and productivity and is dominated by Kobresia pygmaea and K. humilis. The alpine steppe has lower plant diversity and productivity, with Stipa purpurea and Carex moorcroftii being the dominant species (Zhang et al. 1988).
Map of the sampling sites across the Tibetan permafrost-affected region. The background map shows the distributions of permafrost on the Tibetan Plateau, which was derived from Li and Cheng (1996)
Soil sampling was performed in July and August of 2013 and 2014 according to standardized sampling protocols (Ding et al. 2016). We collected soil samples from 149 sites along a transect covering strong climatic variability and two habitats across the central Tibetan Plateau (Ding et al. 2016). Among the sampling sites, 5 sites occurred in mountain permafrost, 56 sites occurred in plateau permafrost, and the remaining 88 sites occurred in seasonally frozen ground (Fig. 1). At each site, we established five independent 1 × 1 m2 squares (in the centre and four corners of the plot) in a randomly selected 10 × 10 m2 plot. In each square, we counted the plant species to calculate the plant species richness (SR) and harvested all grasses for the determination of aboveground biomass (AGB). We collected soil samples randomly from the surface layer (0-10 cm) within the three squares located along a diagonal line in the plot. Each soil sample was sieved and homogenized through a 2.0 mm mesh after removing roots and stones. The three replicates were mixed in equal proportions to form a composite sample per site, which was divided into two fractions. One was air dried for physiochemical analysis,and the other was frozen at -80 °C for DNA extraction.
Vegetation, soil and climatic variables
In the laboratory, the AGB was dried in an oven at 65 °C for 48 h and weighed. The plant material was ground and sieved through a 0.125 mm mesh to analyse the total N concentration through combustion in a C/N analyser (Vario EL Ш, Hanau, Germany). Based on the AGB, the belowground biomass (BGB) was calculated using an allometric function constructed with data from a field survey of AGB and BGB (Yang et al. 2009).
Soil moisture was calculated as the change in weight of 10 g fresh soil after drying at 105 °C for 48 h. Soil texture was determined using a Masterizer 2000 (Malvern Instruments, Malvern, UK) after discarding calcium carbonates and organic matter. Soil pH was measured using soil slurries with a soil-to-water ratio of 2.5 using a standard pH meter (PB-10, Sartorius, Germany). The total carbon (TC) and total nitrogen (TN) concentrations were measured using the same method as the plant N concentration. Soil inorganic carbon (SIC) was measured using a 08.53 calcimeter (Eijkelkamp, Giesbeek, Netherlands). The SOC concentration was calculated by subtracting the SIC from the TC. Total phosphorous (TP) was determined using acid-dissolved molybdenum, antimony and scandium colorimetry. Soil C:N ratio was calculated based on the ratio of SOC to TN, and C:P ratio was calculated as the quotient of SOC and TP. Total dissolved nitrogen (TDN) was extracted using a 0.5 M K2SO4 solution and measured with a dissolved organic carbon/total nitrogen (DOC/TN) analyser (Analytic Jena, Germany). Soil inorganic N (ammonium and nitrate) was measured in 2M KCl extracts using a flow injection analyser (Autoanalyser 3 SEAL, Bran and Luebbe, Norderstedt, Germany). Dissolved organic N (DON) was calculated by subtracting the ammonium and nitrate from the TDN.
To obtain climate data for each sampling site, we estimated the MAT and MAP across the study area using Kriging interpolation at a 10 km spatial resolution. Specifically, we first obtained the original meteorological data from 73 weather stations located on the Tibetan Plateau from the China Meteorological Data Sharing Service System (http://cdc.nmic.cn/home.do), and then performed the interpolation analyses with ARCMAP 10.0 (http://www.esri.com). We further exacted the MAT and MAP for each sampling site from the Kriging-based spatial distributions of climatic variables.
Potential ammonia oxidation rate (PAO) and potential denitrification rate (PDR)
The PAO was assessed using an incubation method (Kurola et al. 2005; Chen et al. 2013). For each sample, 5 g fresh soil was added to a 50 ml Falcon tube with 1 mM (NH4)2SO4 (20 ml). To inhibit nitrite oxidation, potassium chlorate at a final concentration of 10 mM was added to the tubes. The suspension was shaken at 120 r.p.m. at 25 °C for 24 hours in the dark. The nitrite was extracted with 2 M KCl (5 ml) and measured using a spectrophotometer. The PAO was calculated as the net increase in the nitrite concentration during the incubation.
The enhanced PDR was measured according to Patra et al. (2006). Briefly, 10 g of oven-dried equivalent soil was placed in a 135-ml serum bottle. Six milliliters of solution containing KNO3 (200 μg NO3-N g-1), glutamic acid (0.5 mg C g-1) and glucose (0.5 mg C g-1) were added to the bottles. Soil moisture was regulated to 100% of the water-holding capacity. We sealed the bottles with butyl stoppers, flushed the bottles with N2 for 30 min to remove oxygen and adjusted the atmospheric pressure using a glass syringe. We then removed fourteen millilitres of headspace and added 14 ml of acetylene, equalling 10% of the bottle, to inhibit N2O reduction. The soils were incubated at 20 °C for 6 hours and shaken on a rotary shaker. One millilitre of headspace was removed at the end of the incubation period using a pressure-lock syringe (VICI, Baton Rouge, LA, USA) and measured using a gas chromatograph (Agilent Technologies, Santa Clara, CA, USA). The PDR was calculated as the linear rate of N2O production during the incubation period. It should be noted that a preliminary assay was conducted to verify the incubation time for these soils. In particular, five time points was considered for each representative sample. The representative samples showed a linear increase in the N2O concentration during the incubation process, indicating that 6 hours was suitable for all soil samples.
Nucleic acid extraction and quantification of functional N gene abundance
For each sample, DNA was extracted from 500 mg of soil using the FastDNA® SPIN Kit for Soil (Q-Biogene, Carlsbad, CA, USA). The quantitative PCR (qPCR) was performed to evaluate the abundance of functional N genes including the bacterial and archaeal ammonia monoxygenase (amoA) genes involved in ammonia oxidation, the nitrite reductase genes (nirS, nirK) and the N2O reductase gene (nosZ) affiliated with denitrification. Each sample was analysed in triplicate using a StepOnePlusTM Real-time PCR system (Applied Biosystems, Foster City, CA, USA) using the primer sets and amplification conditions listed in Table S1. Each 20 μl reaction mixture included 10 μl of SYBR® Premix Ex TaqTM (TaKaRa Biotechnology, Dalian, China), 1 μl of each primer, 1 μl of bovine serum albumin (BSA), and 2 μl of diluted DNA template. Calibration curves were obtained using a 10-fold dilution series of plasmids containing the target genes. No-template control was also analysed, and this control had null or negligible values. The specificity of qPCR products was evaluated by melting curve analysis and agarose gel electrophoresis. For the different assays, the amplification efficiency ranged from 90% to 105%, and the r2 values were 0.97–0.99.
Statistical analyses
We tested the heterogeneity of the variances, and all data were log transformed when necessary to satisfy the normality assumption prior to statistical analysis. We used independent-samples t-tests to explore the differences in environmental properties (climatic, edaphic and plant properties), PAO, PDR, and functional N gene abundance between alpine steppe and alpine meadow. We performed Pearson’s correlation analysis to examine the relationships between the PAO, PDR, functional N gene abundance and the climatic, edaphic and plant properties. Based on this method, we also evaluated the associations among the PAO, PDR and the number of copies of functional genes. All statistical analyses were performed using SPSS 20.0 (IBM SPSS, Chicago, IL, USA), and the results were considered to be significant at P < 0.05. In addition, we estimated the spatial patterns of the functional N gene abundance, PAO and PDR across the Tibetan Plateau using Kriging interpolation.
We conducted variance partitioning analysis (VPA) to assess the relative contribution of climatic, edaphic and plant properties on the abundance of ammonia-oxidizing and denitrifying communities. We also used VPA to assess the relative contribution of abiotic and plant properties, functional N gene abundance on the PAO and PDR. The properties that significantly contributed to the abundance and activity of ammonia-oxidizing and denitrifying communities were included into VPA. This analysis was performed using the ‘vegan’ package (Oksanen et al. 2008) in the R software (http://cran.rproject.org/). To further evaluate the major predictors of soil functional N gene abundance, PAO and PDR, we performed a random forest analysis (Breiman 2001) using the importance function in the ‘randomForest’ package (Liaw and Wiener 2002) in the R software,version 3.3.3 (http://cran.rproject.org/). The importance of each predictor was assessed by evaluating the percentage of increase in the mean square when the data are randomly permuted (5000 permutations). In our RFs, the significant abiotic and plant properties that contributed to the copy numbers of functional N genes were included as predictors of the response variables. Similarly, the climatic, edaphic and plant properties, and functional N gene copy numbers that significantly contributed to the PAO and PDR were included as predictors of the response variables. We assessed the significance of the effect of each predictor on the response variable using the ‘rfPermute’ package (Archer 2016) in the R software (http://cran.rproject.org/).
Results
Abiotic and plant properties, PAO, PDR, and abundance of functional N genes
Both abiotic and plant properties exhibited significant differences between alpine steppe and alpine meadow (Table 1). Compared with alpine steppe, alpine meadow had higher MAP, soil moisture, silt content, SOC, TN, C:N ratio, C:P ratio, DON, soil ammonium and nitrate concentrations, SR, AGB, BGB and plant N concentration but had lower soil pH and sand content (Table 1). With exception, no significant difference was observed for MAT between these two vegetation types (Table 1).
The PAO varied from 0.15 to 10.35 μg N g-1 dry weight (DW) per day and showed no obvious difference between the two habitats (Fig. 2). The PDR varied from 0.10 to 38.8 ng N g-1 DW per hour, and was higher in alpine meadow than in alpine steppe. Based on the Kriging interpolation result, the PAO and PDR showed a decreasing trend from the east to the west across the Tibetan Plateau (Fig. 3).
Comparison of the potential ammonia oxidation (PAO) and potential denitrification rate (PDR) (a), and the number of copies of N-cycling genes (b) between alpine steppe and alpine meadow. Different lower-case letters above the boxes indicate significant differences between alpine steppe and alpine meadow (independent samples t test; P < 0.05).The gaps and squares in the boxes represent the median and mean values of the data, respectively, while the lower and upper edges of the boxes represent the 25th and 75th percentiles of the data, respectively. AOA, ammonia-oxidizing archaea; AOB, ammonia-oxidizing bacteria
Spatial distributions of potential process rates and functional N gene abundance in Tibetan permafrost-affected regions generated through Kriging interpolation. a potential ammonia oxidation (PAO, μg N g-1 DW day-1); b potential denitrification rate (PDR, ng N g-1 DW hour-1); c abundance of archaeal amoA gene (copy numbers g-1 dry soil); d abundance of bacterial amoA gene (copy numbers g-1 dry soil); enirK gene abundance (copy numbers g-1 dry soil); fnirS gene abundance (copy numbers g-1 dry soil); and (g) nosZ gene abundance (copy numbers g-1 dry soil)
The number of gene copies of amoA, nirS, nirK and nosZ was significantly higher in alpine meadow than in alpine steppe (all P < 0.05; Fig. 2). The ratio of the archaeal to bacterial amoA gene copies varied from 0.42 to 19,446, and showed no significant difference between the two vegetation types (P > 0.05; Fig. S1). The ratio of the number of (nirS + nirK) to nosZ gene copies ranged from 7.48 to 2,404 and showed no obvious distinction between alpine meadow and alpine steppe (P > 0.05; Fig. S1). The abundance of functional N genes showed diverse patterns across the study area based on the Kriging interpolation (Fig. 3).
Factors driving the activity and abundance of N-cycling microbial communities
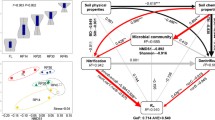
The PAO showed significant associations with most of the abiotic variables except MAP, soil pH, soil moisture, the ammonium concentration, the C:N ratio, and plant N concentration (Fig. 4). By contrast, the PDR was observed to be correlated with all the plant and abiotic properties except MAT (Fig. 4). Although there was not a significant linear correlation between the PAO and AOA abundance (Table S2), a significant nonlinear correlation was observed between them (Fig. 5). Significant correlations were also found between the PDR and the abundance of the nirS and nosZ genes (Fig. 5). VPA demonstrated that the PAO and PDR was mainly determined by abiotic properties, such as SOC and TN, then by plant properties, such as SR, AGB and BGB, and further by functional N gene abundance (Fig. 6). The amount of variation captured by explanatory variables was 35.8% and 65.1% for the PAO and PDR, respectively. Although the pure effects of plant properties were small, strong overlaps were observed between abiotic and plant properties for the PAO (12.1%) and PDR (47.5%), respectively. Our RF model identified abiotic factors (e.g., nitrate concentration, DON, SOC and TN) and the number of copies of N-cycling genes (e.g., archaral amoA gene and nosZ gene) as the main predictors of PAO and PDR (Fig. 7).
Correlogram representing the correlation coefficients between the potential ammonia oxidation (PAO), potential denitrification rate (PDR), functional N gene abundance and environmental variables. Significance is indicated as follows: *P < 0.05 and **P < 0.01. AOA, ammonia-oxidizing archaea; AOB, ammonia-oxidizing bacteria; MAT, mean annual temperature; MAP, mean annual precipitation; SOC, soil organic carbon; DON, dissolved organic nitrogen; TN, total nitrogen; SR, plant species richness; AGB, aboveground biomass; BGB, belowground biomass; Plant N, plant N concentration
Relationships between the potential ammonia oxidation (PAO), the potential denitrification rate (PDR) and the functional N gene abundance. The black and blue solid lines represent the fitted curves and their 95% confidence intervals, respectively. AOA, ammonia-oxidizing archaea; AOB, ammonia-oxidizing bacteria
The results of variance partitioning analysis for the potential ammonia oxidation (PAO) (a), potential denitrification rate (PDR) (b), archaeal amoA gene abundance (c), bacterial amoA gene abundance (d), nirK gene abundance (e), nirS gene abundance (f) and nosZ gene abundance (g). The variance partitioning analysis showed the percentages of variance in potential process rates explained by plant and abiotic properties, and functional N gene abundance (a, b), and illustrated the percentages of variance in functional N gene abundance explained by climatic, plant and edaphic properties (c-g)
The relative importance (percentage of increase in the mean square error) of the explanatory variables as drivers of the potential ammonia oxidation (PAO) (a), potential denitrification rate (PDR) (b), archaeal amoA gene abundance (c), bacterial amoA gene abundance (d), nirK gene abundance (e), nirS gene abundance (f) and nosZ gene abundance (g). The effects of all the shown predictors are significant at P < 0.05. The different colors were used to stand for various factors: yellow represents soil factors, red represent temperature, blue represents precipitation, green represents plant properties, and brown represents functional N gene abundance. AOA, ammonia-oxidizing archaea; MAT, mean annual temperature; MAP, mean annual precipitation; moisture, soil moisture; SOC, soil organic carbon; C:N, the ratio of SOC to TN; C:P, the ratio of SOC to total phosphorus; DON, dissolved organic nitrogen; TN, total nitrogen; SR, plant species richness; AGB, aboveground biomass; BGB, belowground biomass; Plant N, plant N concentration
The Pearson’s correlation analyses revealed significant correlations between the functional N gene abundance and climatic, edaphic and plant properties (Fig. 4). The variation captured by environmental variables was 37.1% for AOA abundance, 24.9% for AOB abundance, 21.5% for nirK gene abundance, 40.2% for nirS gene abundance and 52.1% for nosZ gene abundance (Fig. 6). The functional N gene abundance was mainly explained by abiotic factors, such as MAP, SOC and TN, followed by plant properties (Fig. 6). The RF models identified abiotic factors (e.g., MAP), and plant properties (e.g., plant productivity or plant N concentration) as the predictors of the abundance of functional N genes (Fig. 7).
Discussion
Roles of abiotic and plant properties in driving the abundance and activity of N-cycling microbial communities
In this study, the MAP and soil moisture significantly increased functional N gene abundance and PDR over a large scale. One possible explanation is that higher rainfall is linked to high SR and AGB, leading to increased SOC and N content, especially in the region with higher rainfall on the Tibetan Plateau (Chen et al. 2016a, b). In this region, the humid climatic conditions facilitate the build-up of organic matter (Yang et al. 2008). These changes in plant and edaphic properties along the precipitation gradient may lead to niche separation of N-cycling microorganisms and can have a profound effect on specialized functions such as PAO and PDR.
Our results showed that soil properties (e.g., SOC, nitrate and DON) played important roles in determining abundance and activity of ammonia-oxidizing microorganisms and denitrfiers across Tibetan permafrost regions. These results are supported by previous findings that soil properties mainly cause spatial variation in microbial community abundance (Bru et al. 2011) and process rates (Banerjee and Siciliano 2012a, b). These findings imply that alpine grassland ecosystems are faced with severe nutrient limitations, similar to what occurs in Arctic regions (Chapin III et al. 1993; Banerjee and Siciliano 2012a). This viewpoint is further supported by the data showing that the SOC and N concentrations on the Tibetan Plateau were lower than their counterparts in tundra or at the global scale (Chen et al. 2016a). Soil resources may exert their effects on PAO and PDR through ammonia-oxidizing and denitrifying microorganisms, and the influence of soil resources may mask that of the abundance of functional N gene in these nutrient-limited ecosystems.
We also observed important roles of plant properties (SR, AGB, BGB and plant N content) in determining functional N gene abundance and PDR. Our results contrasted with our first hypothesis that higher plant diversity and productivity could result in a lower abundance of denitrifiers and denitrification due to lower nitrate availability in grassland diversity experiments (Mueller et al. 2013; Wei et al. 2017). These comparisons indicate that the effects of plant properties on denitrification observed in manipulative field experiments do not hold true along natural environmental gradients. There are two possible explanations for the significant effects of plant properties on the abundance and activity of ammonia-oxidizing and denitrifying microorganisms over large scales. First, plant properties may indirectly affect the abundance and activity of N-cycling microbial communities through the availability of soil ammonium and nitrate, which are the substrates for the activity of ammonia-oxidizing microorganisms and denitrifiers, respectively (Tiedje 1988; Verhamme et al. 2011). Plant productivity has been reported to increase with SR in most diversity studies (Tilman et al. 2014). High plant diversity promotes niche differentiation or facilitation (Loreau and Hector 2001), and results in the increased exploitation of limiting soil resources and increased aboveground N uptake (Oelmann et al. 2011). Thus, more diverse plant communities can provide a more diverse and consistent long-term supply of organic N input in terms of the quantity, quality and timing of inputs into the soil (Knops et al. 2002), leading to higher soil inorganic N levels. Consistent with this assumption, we found positive correlations between plant properties (SR, AGB, BGB and plant N concentration) and DON, soil ammonium and nitrate concentrations in this study (Table S3). Therefore, higher levels of plant N concentration and plant productivity and more diverse plant communities may result in greater N inputs into the soil, leading to higher soil N availability, which stimulates the growth and activity of various functional microorganisms involved in N cycling.
Second, plant properties may affect functional N gene abundance and potential rates by altering the SOC availability. The positive effects of SOC on functional N gene abundance have been reported in agricultural ecosystems (Attard et al. 2011), Arctic ecosystems (Banerjee and Siciliano 2012a; Petersen et al. 2012), and tropical forests (Pajares et al. 2018). In general, the growth of soil microorganisms is believed to be limited by C, especially among microorganisms in alpine grasslands (Chen et al. 2016a, 2017b). The increased SOC availability under the higher plant diversity and productivity in this region (Chen et al. 2016b) can increase the supply of C for heterotrophic denitrifiers and may indirectly affect denitrification. Somehow, AOA can benefit from SOC (Tourna et al. 2011), indicating the potential influence of plant properties on the growth of ammonia-oxidizing microorganisms and PAO. Collectively, our results indicate that plant properties, which have been previously overlooked, are important predictors of the functional N gene abundance and N-cycling process rates over large scales.
Our results further indicated that plant properties strongly contributed to the PDR but weakly contributed to the PAO, which could be ascribed to the following two explanations. First, the ability of fungal denitrifiers to produce N2O may have contributed to the close correlation between plant properties and the PDR (Fig. 4). In contrast to bacteria, fungi have stronger biotrophic interactions with plants (Wardle 2006). Some fungal taxa were found to be the key sources of N2O production, which is a widespread trait in fungi (Maeda et al. 2015). For instance, Maeda et al. (2015) showed that 34% of 207 fungal isolates had the ability to produce N2O in pure culture, and the inoculation of these strains into pre-sterilized soils confirmed their N2O-producing ability. Recent studies reported that the fungal denitrification system was composed of nirK and P450nor (Shoun et al. 2012) and that fungal denitrifiers did not have the genetic capacity to transform N2O into N2 (Mothapo et al. 2015). By contrast, ammonia oxidation is mainly performed by autotrophic ammonia-oxidizing microorganisms rather than heterotrophic nitrifiers, such as fungi, in environments with limited C and water, as heterotrophic microbe activity and growth are less competitive than those of autotrophic ammonia-oxidizing microorganisms in such environments (Hart et al. 1994). Thus, fungal nitrifiers may play less important roles in ammonia oxidation, while fungal denitrifiers may contribute to the close correlations between plant properties and the PDR across Tibetan permafrost-affected regions. Second, the physiological differences between ammonia-oxidizing microorganisms and denitrifiers may also contribute to the different responses of the PAO and PDR to plant properties. The denitrifiers are heterotrophic microorganisms using C as their only energy source, whereas the nitrifiers are mainly autotrophic microorganisms that use ammonia as their source of energy. As stated above, high SR and AGB likely provide more C for denitrifiers, and thus the denitrifiers were more sensitive to plant properties than the ammonia-oxidizing microorganisms.
Influence of functional N gene abundances on PAO and PDR
Our results showed that AOA exhibited a weakly nonlinear association with the PAO, indicating that AOA are likely involved in ammonia oxidation in alpine grassland ecosystems. Our results agree with previous studies showing that AOA are involved in ammonia oxidation in semiarid and arid grasslands (Marusenko et al. 2013; Hu et al. 2015b; Kou et al. 2018) and Arctic ecosystems (Alves et al. 2013, 2018). In this study, we found that AOA outnumbered AOB by many orders of magnitude in most of the sites in alpine grasslands, similar to the findings of previous studies (Leininger et al. 2006; Shen et al. 2011; Pett-Ridge et al. 2013; Tian et al. 2014; Pajares et al. 2018). This is also consistent with the mounting evidence that AOA are predominant over AOB in many environments (Prosser and Nicol 2008), although AOB have been shown to drive PAO even when they are outnumbered by AOA (Di et al. 2009; Gleeson et al. 2010; Carey et al. 2016).
Our results also revealed significant correlations between nirS and nosZ gene abundance and the PDR, supporting our second hypothesis. These results are in line with previous results showing that denitrification is often correlated with the number of copies of the nirS gene (Čuhel et al. 2010) and nosZ gene (Hallin et al. 2009). These results collectively suggest that the abundance of the denitrifying community can predict the PDR, although this process is primarily driven by the C, nitrate, and oxygen availability (Del Grosso et al. 2000). In this study, the large differences in denitrification rates (Figs. 2 and 3) may contribute to the obvious relationship between denitrifier abundance and activity, as reported by Petersen et al. (2012). We also found that the nirK gene abundance was uncoupled from the PDR, which contrasted with the results of Attard et al. (2011), who reported an obvious correlation between the PDR and the abundance of the nirK gene. These discrepancies highlight the complexity of hierarchical and metabolic cellular regulation (Röling 2007). This discrepancy could also be due to choice of nirK primers in this study. We further found that the abundance of nirK gene was not correlated to abiotic factors such as temperature, soil texture and nitrate concentration in this study, suggesting that nirK denitrifiers are likely less sensitive to changes in the environment than nirS denitrifiers. Our results may support the viewpoint that niche differentiation between the two denitrifiers lead to two different types of nitrite reductases (Philippot et al. 2009).
Conclusions and implications
To the best of our knowledge, this study is the first to assess the large-scale patterns and driving factors of the abundance and activity of N-cycling microbial communities affiliated with nitrification and denitrification in permafrost-affected regions. We found that abiotic factors such as climate, soil moisture and soil resources appeared to have a more important role than biotic properties in determining the functional N gene abundance and the rate of potential N cycling process. We also found significant relationships between functional N gene abundance and potential process rates, implying that the functional microorganisms might regulate nitrification and denitrification in permafrost-affected regions. The large-scale patterns of functional N gene abundance and potential process rates could be predicted from plant properties such as SR, AGB and plant N concentration. Considering the role of plant attributes is often overlooked in current models, the role of plant properties can be incorporated into future ecosystem models to improve the prediction of ecosystem functioning responses to the changing environment.
References
Alves RJE, Wanek W, Zappe A, Richter A, Svenning MM, Schleper C, Urich T (2013) Nitrification rates in Arctic soils are associated with functionally distinct populations of ammonia-oxidizing archaea. ISME J 7:1620–1631
Alves RJE, Minh BQ, Urich T, von Haeseler A, Schleper C (2018) Unifying the global phylogeny and environmental distribution of ammonia-oxidising archaea based on amoA genes. Nat Commun 9:1517
Archer E (2016) Estimate permutation p-values for importance metrics. R package version 2(1):5 Retrieved from https://CRAN.R-project.org/package=rfpermute
Attard E, Recous S, Chabbi A, De Berranger C, Guillaumaud N, Labreuche J, Philippot L, Schmid B, Leroux X (2011) Soil environmental conditions rather than denitrifier abundance and diversity drive potential denitrification after changes in land uses. Glob Chang Biol 17:1975–1989
Banerjee S, Siciliano SD (2012a) Spatially tripartite interactions of denitrifiers in arctic ecosystems: activities, functional groups and soil resources. Environ Microbiol 14:2601–2613
Banerjee S, Siciliano SD (2012b) Factors driving potential ammonia oxidation in Canadian arctic ecosystems: does spatial scale matter? Appl Environ Microbiol 78:346–353
Breiman L (2001) Machine learning. Random Forests 45: 5
Bru D, Ramette A, Saby NP, Dequiedt S, Ranjard L, Jolivet C, Arrouays D, Philippot L (2011) Determinants of the distribution of nitrogen-cycling microbial communities at the landscape scale. ISME J 5: 532–542
Carey CJ, Dove NC, Michael Beman J, Hart SC, Aronson EL (2016) Meta-analysis reveals ammonia-oxidizing bacteria respond more strongly to nitrogen addition than ammonia-oxidizing archaea. Soil Biol Biochem 99:158–166
Chapin FS III, Moilanen L, Kielland K (1993) Preferential use of organic nitrogen for growth by a non-mycorrhizal arctic sedge. Nature 361:150–153
Chen YL, Xu ZW, Hu HW, Hu YJ, Hao ZP, Jiang Y, Chen BD (2013) Responses of ammonia-oxidizing bacteria and archaea to nitrogen fertilization and precipitation increment in a typical temperate steppe in Inner Mongolia. Appl Soil Ecol 68:36–45
Chen YL, Chen LY, Peng YF, Ding JZ, Li F, Yang GB, Kou D, Liu L, Fang K, Zhang BB, Wang J, Yang YH (2016a) Linking microbial C:N:P stoichiometry to microbial community and abiotic factors along a 3500-km grassland transect on the Tibetan Plateau. Global Ecol Biogeogr 25:1416–1427
Chen YL, Ding JZ, Peng YF, Li F, Yang GB, Liu L, Qin SQ, Fang K, Yang YH (2016b) Patterns and drivers of soil microbial communities in Tibetan alpine and global terrestrial ecosystems. J. Biogeogr 43:2027–2039
Chen YL, Xu TL, Veresoglou SD, Hu HW, Hao ZP, Hu YJ, Liu L, Hu YJ, Rillig MC, Chen BD (2017a) Plant diversity represents the prevalent determinant of soil fungal community structure across temperate grasslands in northern China. Soil Biol Biochem 110:12–21
Chen YL, Deng Y, Ding JZ, Hu HW, Xu TL, Li F, Yang GB, Yang YH (2017b) Distinct microbial communities in the active and permafrost layers on the Tibetan Plateau. Mol Ecol 26:6608–6620
Čuhel J, Šimek M, Laughlin RJ, Bru D, Chèneby D, Watson CJ, Philippot L (2010) Insights into the effect of soil pH on N2O and N2 emissions and denitrifier community size and activity. Appl Environ Microbiol 76:1870–1878
Del Grosso SJ, Parton WJ, Mosier AR, Ojima DS, Kulmala AE, Phongpan S (2000) General model for N2O and N2 gas emissions from soils due to denitrification. Global Biogeochem Cy 14:1045–1060
Di HJ, Cameron KC, Shen JP, Winefield CS, Callaghan MO, Bowatte S, He JZ (2009) Nitrification driven by bacteria and not archaea in nitrogen-rich grassland soils. Nat Geosci 2:621–624
Ding JZ, Li F, Yang GB, Chen LY, Zhang BB, Liu L, Fang K, Qin SQ, Chen YL, Peng YF, Ji CJ, He HL, Smith P, Yang YH (2016) The permafrost carbon inventory on the Tibetan Plateau: a new evaluation using deep sediment cores. Global Change Biol 22:2688–2701
Elberling B, Christiansen HH, Hansen BU (2010) High nitrous oxide production from thawing permafrost. Nat Geosci 3:332–335
Enwall K, Throback IN, Stenberg M, Soderstrom M, Hallin S (2010) Soil resources influence spatial patterns of denitrifying communities at scales compatible with land management. Appl Environ Microbiol 76:2243–2250
Gleeson DB, Müller C, Banerjee S, Ma W, Siciliano SD, Murphy SD (2010) Response of ammonia oxidizing archaea and bacteria to changing water filled pore space. Soil Biol Biochem 42:1888–1891
Hallin S, Jones CM, Schloter M, Philippot L (2009) Relationship between N-cycling communities and ecosystem functioning in a 50-year-old fertilization experiment. ISME J 3:597–605
Harden JW, Koven CD, Ping CL, Hugelius G, McGuire AD, Camill P, Jorgenson T, Kuhry P, Michaelson GJ, O’Donnell JA, Schuur EAG, Tarnocai C, Johnson K, Grosse G (2012) Field information links permafrost carbon to physical vulnerabilities of thawing. Geophys Res Lett 39:L15704
Hart SC, Stark JM, Davidson EA, Firestone MK (1994) Nitrogen mineralization, immobilization, and nitrification. Methods of soil analysis part 2: Microbiological and biochemical properties. Soil Sci Soc Am J:985–1018
Hu HW, Chen DL, He JZ (2015a) Microbial regulation of terrestrial nitrous oxide formation: understanding the biological pathways for prediction of emission rates. FEMS Microbiol Rev 39:729–749
Hu HW, Macdonald CA, Trivedi P, Holmes B, Bodrossy L, He JZ, Singh BK (2015b) Water addition regulates the metabolic activity of ammonia oxidizers responding to environmental perturbations in dry sub-humid ecosystems. Environ Microbiol 17:444–461
Jia ZJ, Conrad R (2009) Bacteria rather than Archaea dominate microbial ammonia oxidation in an agricultural soil. Environ Microbiol 11:1658–1671
Knops J, Bradley KL, Wedin DA (2002) Mechanisms of plant species impacts on ecosystem nitrogen cycling. Ecol Lett 5:454–466
Kou D, Peng YF, Wang GQ, Ding JZ, Chen YL, Yang GB, Fang K, Liu L, Zhang BB, Müller C, Zhang JB, Yang YH (2018) Diverse responses of belowground internal nitrogen cycling to increasing aridity. Soil Biol Biochem 116:189–192
Kurola J, Salkinoja-Salonen M, Aarnio T, Hultman J, Romantschuk M (2005) Activity, diversity and population size of ammonia-oxidising bacteria in oil contaminated land farming soil. FEMS Microbiol Lett 250:33–38
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW, Prosser JI, Schuster SC, Schleper C (2006) Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442:806–809
Li SD, Cheng GD (1996) Map of Frozen Ground on Qinghai-Xizang Plateau. Gansu Culture Press, Lanzhou
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2/3:18–22
Loreau M, Hector A (2001) Partitioning selection and complementarity in biodiversity experiments. Nature 412:72–76
Maeda K, Spor A, Edel-Hermann V, Heraud C, Breuil M, Bizouard F, Toyoda S, Yoshida N, Steinberg C, Philippot L (2015) N2O production, a widespread trait in fungi. Sci Rep 5:9697
Marusenko Y, Huber DP, Hall SJ (2013) Fungi mediate nitrous oxide production but not ammonia oxidation in aridland soils of the southwestern US. Soil Biol Biochem 63:24–36
Morales SE, Cosart T, Holben WE (2010) Bacterial gene abundances as indicators of greenhouse gas emission in soils. ISME J 4:799–808
Moreau D, Pivato B, Bru D, Busset H, Deau F, Faivre C, Matejicek A, Strbik F, Philippot L, Mougel C (2015) Plant traits related to nitrogen uptake influence plant-microbe competition. Ecology 96:2300–2310
Mothapo N, Chen HH, Cubeta MA, Grossman JM, Fuller F, Shi W (2015) Phylogenetic, taxonomic and functional diversity of fungal denitrifiers and associated N2O production efficacy. Soil Biol Biochem 83:160–175
Mueller KE, Hobbie SE, Tilman D, Reich PB (2013) Effects of plant diversity, N fertilization, and elevated carbon dioxide on grassland soil N cycling in a long-term experiment. Global Change Biol 19:1249–1261
Oelmann Y, Buchmann N, Gleixner G, Habekost M, Roscher C, Rosenkranz S, Schulze E, Steinbeiss S, Temperton VM, Weigelt A, Weisser WW, Wilcke W (2011) Plant diversity effects on aboveground and belowground N pools in temperate grassland ecosystems: Development in the first 5 years after establishment. Glob Biogeochem Cycles 25:415–421
Oksanen J, Kindt R, Legendre P, O’Hara B, Simpson GL, Solymos P, Steven MHH, Wagner H (2008) vegan: Community Ecology Package. R Package version 1.15–1
Pajares S, Campo J, Bohannan BJM, Etchevers JD (2018) Environmental controls on soil microbial communities in a seasonally dry tropical forest. Appl Environ Microbiol. https://doi.org/10.1128/AEM.00342-18
Patra AK, Abbadie L, Clays-Josserand A, Degrange V, Grayston SJ, Guillaumaud N, Loiseau P, Louault F, Mahmood S, Nazaret S, Philippot L, Poly F, Prosser JI, Le Roux X (2006) Effects of management regime and plant species on the enzyme activity and genetic structure of N-fixing, denitrifying and nitrifying bacterial communities in grassland soils. Environ Microbiol 8:1005–1016
Petersen DG, Blazewicz SJ, Firestone M, Herman DJ, Turetsky M, Waldrop M (2012) Abundance of microbial genes associated with nitrogen cycling as indices of biogeochemical process rates across a vegetation gradient in Alaska. Environ Microbiol 14:993–1008
Pett-Ridge J, Petersen DG, Nuccio E, Firestone MK (2013) Influence of oxic/anoxic fluenctuations on ammonia oxidizers and nitrification potential in a wet tropical soil. FEMS Microbiol Ecol 85:179–194
Philippot L, Čuhel J, Saby NPA, Chèneby D, Chroňáková A, Bru D, Arrouays D, Martin-Laurent F, Šimek M (2009) Mapping field-scale spatial patterns of size and activity of the denitrifier community. Environ Microbiol 11:1518–1526
Pierre S, Hewson I, Sparks JP, Litton CM, Giardina C, Groffman PM, Fahey TJ (2017) Ammonia oxidizer populations vary with nitrogen cycling across a tropical montane mean annual temperature gradient. Ecology 98:1896–1907
Prober SM, Leff JW, Bates ST, Borer ET, Firn J, Harpole WS, Lind EM, Seabloom EW, Adler PB, Bakker JD, Cleland EE, DeCrappeo NM, DeLorenze E, Hagenah N, Hautier Y, Hofmockel KS, Kirkman KP, Knops JMH, La Pierre KJ, MacDougall AS, McCulley RL, Mitchell CE, Risch AC, Schuetz M, Stevens CJ, Williams RJ, Fierer N (2015) Plant diversity predicts beta but not alpha diversity of soil microbes across grasslands worldwide. Ecol Lett 18: 85–95.
Prosser JI, Nicol GW (2008) Relative contributions of archaea and bacteria to aerobic ammonia oxidation in the environment. Environ Microbiol 10:2931–2941
Ran YH, Li X, Cheng GD, Zhang TJ, Wu QB, Jin HJ, Jin R (2012) Distribution of permafrost in China: An overview of existing permafrost maps. Permafrost Periglac 23:322–333
Ravishankara AR, Daniel JS, Portmann RW (2009) Nitrous oxide (N2O): the dominant ozone-depleting substance emitted in the 21st century. Science 326:123–125
Röling WFM (2007) Do microbial numbers count? Quantifying the regulation of biogeochemical fluxes by population size and cellular activity. FEMS Microbiol Ecol 62:202–210
Shen XY, Zhang LM, Shen JP, Li LH, Yuan CL, He JZ (2011) Nitrogen loading levels affect abundance and composition of soil ammonia oxidizing prokaryotes in semiarid temperate grassland. J Soils Sediments 11:1243–1252
Shoun H, Fushinobu S, Jiang L, Kim SW, Wakagi T (2012) Fungal denitrification and nitric oxide reductase cytochrome P450nor. Philos T R Soc B 367:1186–1194
Tian XF, Hu HW, Ding Q, Song MH, Xu XL, Zheng Y, Guo LD (2014) Influence of nitrogen fertilization on soil ammonia oxidizer and denitrifier abundance, microbial biomass, and enzyme activities in an alpine meadow. Biol Fertil Soils 50:703–713
Tiedje JM (1988) Ecology of denitrification and dissimilatory nitrate reduction to ammonium. In: Zaj B (ed) Biology of Anaerobic Microorganisms. John Wiley and Sons, New York, pp 179–244
Tilman D, Isbell F, Cowles JM (2014) Biodiversity and ecosystem functioning. Annu Rev Ecol Syst 45:471–493
Tourna M, Stieglmeier M, Spang A, Könneke M, Schintlmeister A, Urich T, Engel M, Schloter M, Wagner M, Richter A, Schleper C (2011) Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proc Natl Acad Sci USA 108:8420–8425
Verhamme DT, Prosser JI, Nicol GW (2011) Ammonia concentration determines differential growth of ammonia-oxidising archaea and bacteria in soil microcosms. ISME J 5: 1067–1071
Voigt C, Marushchak ME, Lamprecht RE, Jackowicz-Korczyński M, Lindgren A, Mastepanov M, Granlund L, Christensen TR, Tahvanainen T, Martikainen PJ, Biasi C (2017a) Increased nitrous oxide emissions from Arctic peatlands after permafrost thaw. Proc Natl Acad Sci USA 114:6238–6243
Voigt C, Lamprecht RE, Marushchak ME, Lind SE, Novakovskiy A, Aurela M, Martikainen PJ, Biasi C (2017b) Warming of subarctic tundra increases emissions of all three important greenhouse gases – carbon dioxide, methane, and nitrous oxide. Global Change Biol 23:3121–3138
Wardle DA (2006) The influence of biotic interactions on soil biodiversity. Ecol Lett 9:870–886
Wei XR, Reich PB, Hobbie SE, Kazanski CE (2017) Disentangling species and functional group richness effects on soil N cycling in a grassland ecosystem. Global Change Biol 23:4717–4727
Yang YH, Fang JY, Tang YH, He J-S, Ji CJ, Zheng CY, Zhu B (2008) Storage, patterns, and controls of soil organic carbon in the Tibetan grasslands. Global Change Biol 14:1592–1599
Yang YH, Fang JY, Ji CJ, Han WX (2009) Above- and belowground biomass allocation in Tibetan grasslands. J Veg Sci 20:177–184
Yang GB, Peng YF, Marushchak ME, Chen YL, Wang GQ, Li F, Zhang DY, Wang J, Yu JC, Liu L, Qin SQ, Kou D, Yang YH (2018) Magnitude and pathways of increased nitrous oxide emissions from uplands following permafrost thaw. Environ Sci Technol 52:9162–9169
Zhang JW, Wang JT, Chen W, Li B, Zhao K (1988) Vegetation of Xizang (Tibet). Science Press, Beijing
Zhang LM, Hu HW, Shen JP, He JZ (2012) Ammonia-oxidizing archaea have more important role than ammonia-oxidizing bacteria in ammonia oxidation of strongly acidic soils. ISME J 6:1032–1045
Zumft WG (1997) Cell biology and molecular basis of denitrification. Microbiol Mol Biol R 61:533–616
Acknowledgments
We appreciate the section editor (Dr. Feike A. Dijkstra) and two anonymous reviewers for their insightful comments on an early version of this manuscript. This work was supported by the National Key Research and Development Program of China (2017YFA0604803 and 2016YFC0500701), National Natural Science Foundation of China (41877046, 41501265 and 31670482), Key Research Program of Frontier Sciences, Chinese Academy of Sciences (QYZDB-SSW-SMC049), and Chinese Academy of Sciences-Peking University Pioneer Cooperation Team.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Feike A. Dijkstra.
Electronic Supplementary Material
ESM 1
(DOCX 75 KB)
Rights and permissions
About this article
Cite this article
Chen, Y., Kou, D., Li, F. et al. Linkage of plant and abiotic properties to the abundance and activity of N-cycling microbial communities in Tibetan permafrost-affected regions. Plant Soil 434, 453–466 (2019). https://doi.org/10.1007/s11104-018-3845-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-018-3845-9