Abstract
Alpine meadows on the Tibetan Plateau have experienced severe degradation in recent decades. Although the effects of alpine meadow degradation on soil properties have been well documented, there is still a paucity of knowledge regarding the responses of nitrogen-cycling microbes (NCMs) to degradation and their links to the changes in soil properties. Here, we systematically determined the effects of degraded patch formation on soil properties (i.e., total carbon, total nitrogen, ammonium nitrogen, nitrate nitrogen, available phosphorus, dissolved organic carbon, moisture, δ15N, δ13C, and pH) and NCMs (based on nifH, amoA, narG, nirK, and nirS genes and their transcripts) across three Tibetan alpine meadows at different degradation stages. Results showed that compared to the original grassed patches, the contents of most soil nutrients (e.g., carbon, nitrogen, and phosphorus) were significantly decreased in the degraded patches across the study sites. Degraded patches also tended to have higher soil δ15N values and nitrate contents. Among the aforementioned NCMs, soil diazotrophs and denitrifiers only showed weak responses to the patch formation, while ammonia-oxidizing microbes showed the highest consistency and sensitivity in response to the patch formation across the study sites. The abundance of amoA gene and archaeal amoA mRNA significantly increased in the degraded patches, and they were positively correlated with soil δ15N values and nitrate nitrogen contents, but negatively correlated with soil total nitrogen and inorganic nitrogen contents. These results suggest that the increased ammonia-oxidizing microbial abundance may be an important driver of soil nitrogen loss during degraded patch formation in alpine meadows.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Approximately 23.5% of global land is experiencing ongoing degradation, causing 9.56 × 1011 kg carbon loss and severely threatening the lives of more than 1.5 billion people (Bai et al. 2008). This situation is even worse in the Tibetan alpine meadows which account for approximately half of the usable Tibetan grasslands, and are extremely sensitive to climate changes and human activities (Cui and Graf 2009; Wang et al. 2016; Zhao et al. 2015). Soil nitrogen contents have been shown to be susceptible to loss under grassland degradation (Dong et al. 2012; Li et al. 2014; Wen et al. 2013a; Zhang et al. 2017), which in turn may affect the resistance and resilience of grassland ecosystems, and determine the extent of further degradation (Dong et al. 2016). In natural ecosystems, soil nitrogen contents are essentially determined by the balance of the different nitrogen cycling processes that are mediated by a series of microbes (i.e., nitrogen cycling microbes, NCMs; Ollivier et al. 2011). Therefore, exploring the responses of NCMs to grassland degradation not only can contribute to predicting grassland development, but also can provide a theoretical basis for the restoration of degraded grasslands.
In recent decades, the soil NCMs, especially diazotrophs, nitrifiers, and denitrifiers, have been extensively investigated using the corresponding molecular markers (Levy-Booth et al. 2014; Thamdrup 2012). Diazotrophs are a group of prokaryotes that can reduce N2 to NH3. The nifH gene, encoding dinitrogenase reductase, is widely employed to identify diazotrophs by virtue of its ubiquity and phylogeny consistency in diazotrophs (Gaby and Buckley 2011; Raymond et al. 2004). Similarly, the amoA gene, encoding subunit A of ammonia monooxygenase, is commonly used as a molecular marker to investigate the ammonia-oxidizing archaea (AOA) and bacteria (AOB) (Stahl and de la Torre 2012a). As ammonia oxidization is often the rate determining step of nitrification, AOA and AOB are the most well studied nitrifiers over the past decades. Denitrification is a sequence of processes in which nitrate is reduced to N2 through several intermediate products including nitrite, nitric oxide (NO), and nitrous oxide (N2O). These processes are performed by a series of heterotrophic bacteria (i.e., denitrifiers), among which narG (encoding the catalytic subunit of nitrate reductase), nirS, and nirK (encoding nitrite reductase) can be used to investigate the microbes mediating the reduction process from nitrate to NO (the first gas product of denitrification) via nitrite (Levy-Booth et al. 2014). Nitrogen fixation can increase soil nitrogen contents, while nitrification may increase the risk of soil nitrogen loss, and denitrification can compromise soil nitrogen contents through gas emission (Galloway et al. 2004; Ollivier et al. 2011). Therefore, investigating soil NCMs, based on the aforementioned molecular markers (i.e., nifH, amoA, narG, nirS and nirK genes) and their transcripts can provide crucial information for understanding the responses of soil nitrogen contents to grassland degradation.
Soil NCMs are sensitive to grassland degradation and the reasons can be multifaceted. First, degradation usually decreases plant biomass and soil organic carbon (Lal 2003; Wen et al. 2013b). Typically, diazotrophs can be either heterotrophic or autotrophic, whereas most nitrifiers and denitrifiers are autotrophic and heterotrophic, respectively (Levy-Booth et al. 2014; Raymond et al. 2004; Stahl and de la Torre 2012a). Thus, the lower input of organic carbon under degradation should be less critical to soil nitrifiers and autotrophic diazotrophs, and more detrimental to denitrifiers and heterotrophic diazotrophs. Second, soil nitrogen has been shown to decrease under degradation (Dong et al. 2012; Li et al. 2014), which should promote the activities of diazotrophs but not nitrifiers and denitrifiers. Third, degradation also can decrease other soil nutrients (Ren et al. 2013; Wu et al. 2014; Wu and Tiessen 2002), altering plant community composition and soil physical properties (Tang et al. 2015; Wu and Tiessen 2002), which may exert dramatic effects on the soil NCMs.
However, there is still a paucity of data to elucidate the responses of soil NCMs to grassland degradation, especially in the Tibetan alpine meadows. To date, no publications have documented the responses of NCMs to degradation in this region. Moreover, alpine meadow degradation can be divided into six stages, among which stages three to five are critical and characterized by the formation and expansion of degraded patches (Fig. 1; Lin et al. 2015). At these three degradation stages, the characteristic mattic epipedons of alpine meadow are first eroded to form bare soil patches (Fig. 1, Site 1 and Site 2), which then expand and interconnect to dominate over the isolated meadow patches (Fig. 1, Site 3). Most existing studies focused only on the vegetation and soil property variations along the degradation gradients (e.g. Li et al. 2015, 2016), whereas very few studies have examined the effects of degraded patch formation on soil nitrogen contents and NCMs (e.g., Zhang et al. 2017).
To address the aforementioned knowledge gaps, we initiated a study across three degraded Tibetan alpine meadows corresponding to the three aforementioned critical degradation stages to investigate: (i) whether and how soil physicochemical properties and NCMs (based on the nifH, amoA, narG, nirS, and nirK genes and their transcripts) were affected by degraded patch formation of alpine meadows; (ii) whether the responses of soil physicochemical properties and NCMs to the patch formation were consistent across different degradation stages; and (iii) whether the changes in NCMs were correlated with the variations in soil physicochemical properties.
Materials and methods
Study sites and soil sampling
We collected soil samples from three degraded alpine meadows (Fig. 1), Site 1 (32°52.39′N, 96°54.78′E; asl 3 982 m), Site 2 (33°14.96′N, 97°27.62′E; asl 4332 m), and Site 3 (34°34.06′N, 97°5.88′E; asl 4314 m), in August 2014. The degradation extents of the alpine meadows increased from Site 1 to Site 3, corresponding to the third to fifth degradation stages of alpine meadows (Lin et al. 2015), respectively. At Site 1 and Site 2, around 20% of the grassland was degraded to bare patches, but the original mattic epipedons at Site 1 remained intact. At Site 2, the mattic epipedons were partly destroyed and eroded. At Site 3, about 60% of the mattic epipedons were eroded, and the bared patches were re-covered by secondary vegetation. Herein, the original meadow patches and bare soil patches are depicted as original patches and degraded patches, respectively (Fig. 1). The effect of degraded patch formation refers to the difference in soil physicochemical properties and NCM indices between the original and degraded patches.
At each site, we collected soil samples (0–10 cm) from four paired original-degraded patches (Fig. 1). After being sieved to 2 mm, we treated the soils (2.0 g) used for RNA extraction with 4 mL LifeGuard™ Soil Preservation Solution (MO BIO Laboratories, Carlsbad, CA, USA) following the instruction manual. The treated and 100 g nontreated fresh soil (used for DNA extraction and soil property analysis) was immediately placed into the − 20 °C freezer, transported to the laboratory within 2 weeks, and then preserved at − 80 °C. We also air dried some soil subsamples (100 g) for the analysis of soil pH values and available phosphorus contents.
Analysis of soil physicochemical properties
Soil gravimetric moisture content was determined by drying at 105 °C for 24 h. Soil pH was determined with the soil mass to water ratio of 1:5. Soil dissolved organic carbon (DOC) content was analyzed using the cold water method (soil mass to water ratio of 1:3), and measured using a TOC Analyzer (Liqui TOC II; Elementar Analysensysteme GmbH, Hanau, Germany). Soil ammonium and nitrate nitrogen (AN and NN) contents were determined using 2 M KCl extraction (soil mass to extractant ratio of 1:5) and an auto flow analyzer (Auto Analyzer 3 System; SEAL Analytical GmbH, Norderstedt, Germany). Soil available phosphorus (AP) content was analyzed using the method depicted by Olsen et al. (1954). Soil total carbon (TC) contents, total nitrogen (TN) contents, and values of δ13C and δ15N were determined using an isotope ratio mass spectrometer coupled with an auto elemental analyzer (NA1500, Fisons Instruments, Milano, Italy). The soil gravimetric moisture, DOC, AN, and NN contents were determined using the fresh soils, while the values of soil pH, δ13C and δ15N values, and the contents of AP, TC, and TN were analyzed using the air-dried soils.
Nucleic acid extraction and cDNA synthesis
We used 0.30 and 2.0 g soil to extract DNA and RNA, respectively. The extractions were separately conducted using a PowerSoil™ DNA Isolation Kit and PowerSoil® Total RNA Isolation Kit (MO BIO Laboratories, Carlsbad, CA, USA), as described by suppliers. RNA extracts were treated with DNase I (MO BIO Laboratories, Carlsbad, CA, USA) to remove DNA residuals. The RNA was then reverse-transcribed into cDNA using a PrimeScript™ II 1st Strand cDNA Synthesis Kit with random hexamers (Takara Bio Inc., Shiga, Japan). Next, the DNA solutions were diluted 1:3 with water for the subsequent polymerase chain reaction (PCR) and real-time PCR amplifications, to attenuate the potential effects of PCR inhibitors.
Real-time PCR
We quantified the copy numbers of nifH, archaeal amoA, bacterial amoA, narG, nirK, and nirS genes and mRNAs using the 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA), with the degenerated primer sets as described in Table S1. The 20 μL reaction mixtures contained the following: 1 μL template DNA (DNA, cDNA, or serially diluted standards), 10 μL Maxima™ SYBR Green or ROX (2×, Thermo Fisher Scientific, Waltham, MA, USA), 0.5 μL forward primer (20 μmol L−1), 0.5 μL reverse primer (20 μmol L−1), and 8.0 μL nuclease-free water. Standard curves were constructed using plasmids harboring the corresponding DNA fragments. We analyzed all DNAs and cDNAs in triplicate and utilized three no template controls to check for reagent contamination. PCR runs started with an initial denaturation and enzyme activation step for 10 min at 95 °C, which was followed by 40 cycles of 15 s at 95 °C, 30 s at the annealing temperatures depicted in Table S1, 40 s at 72 °C, and 30 s at 80 °C. We recorded the fluorescence signals at 80 °C to attenuate influences of primer dimers. We tested the specificities of the PCR products using melting curve analysis. The amplification efficiencies and R2 values of the real-time PCRs are detailed in Table S2.
Terminal restriction fragment length polymorphism (T-RFLP)
The six nitrogen-cycling genes were separately amplified using PCR with labeled degenerate primer sets (Table S1). The forward primers were labeled with 6-carboxyfluorescein (FAM) at the 5′ ends. The 50 μL reaction mixture contained: 25 μL Premix Taq™ Hot Start Version (Takara Bio Inc., Shiga, Japan), 1 μL template DNA, 1 μL forward primer (20 μmol L−1), 1 μL reverse primer (20 μmol L−1), and 22 μL nuclease-free water. The PCR programs consisted of the initial denaturation and enzyme activation at 95 °C for 10 min, followed by 38 cycles of 45 s at 95 °C, 30 s at the annealing temperatures depicted in Table S1, and 45 s at 72 °C, and then with a final extension at 72 °C for 10 min. We conducted three technical replicates for each sample and then separately pooled the technical replicates into one tube. Subsequently, the PCR products were purified using a GeneJET Gel Extraction Kit (Thermo Scientific, Lithuania), after which the purified PCR products were completely digested with the restriction endonucleases listed in Table S1 (New England Biolabs Inc. Beverly, MA, USA). Terminal restriction fragments (T-RFs) were size-separated by Beijing Ruibo BioTech Co. Ltd (Beijing, China) using an ABI 3730 system (Applied Biosystems, Foster City, CA, USA). Electropherograms of T-RFLPs were retrieved using PeakScanner 1.0 (Applied Biosystems, Foster City, CA, USA). The threshold for peak assignment was ± 2 bp. After normalization, we removed unqualified peaks with a size less than the length of forward primers, more than the theoretical length, or a peak height lower than 50. Proportions of the qualified peaks were calculated based on their peak areas.
Statistical analysis
The effects of degraded patch formation (differences between the original and degraded patches) and study site on the soil physicochemical properties and copies of nitrogen-cycling functional genes and transcripts were determined using two-way nested ANOVA and Duncan’s post hoc tests. The effects of degraded patch formation and study site on the NCM community structures were revealed using non-metric multidimensional scaling (NMDS) and permutational multivariate analysis of variance (PERMANOVA). We also conducted Pearson’s correlation, redundancy analysis (RDA), and the envfit based on NMDS to analyze the relationships between NCM indices and soil physicochemical properties. We transformed some data to meet the requirements of analysis. All statistical analysis was conducted using R (R Development Core Team 2017) with the vegan package.
Results
Soil physicochemical properties
All the measured soil properties (i.e., TN, TC, C:N, inorganic nitrogen (IN), AN, NN, DOC, δ15N, δ13C, moisture, AP, and pH) differed significantly among the three study sites (P < 0.006, Fig. 2; Table S3). The highest soil TN, TC, IN, AN, DOC, AP and moisture contents all were recorded in the original patches of Site 1 and showed a decreasing trend from Sites 1 to 3 (Fig. 2). However, the soil NN contents, δ15N and δ13C values tended to increase with the degradation extent (Fig. 2). The soil C:N ratios were similar in the original patches across the three study sites, but they were significantly lower in the degraded patches at Sites 2 and 3 (Fig. 2h). Soil pH values were highest at Site 3 and lowest at Site 2 (Fig. 2l).
Effects of the study site and degraded patch formation on soil properties at 0–10 cm depth. IN inorganic nitrogen, DOC dissolved organic carbon, S1 Site 1, S2 Site 2, S3 Site 3, S effect of study site, D effect of degraded patch formation; *P < 0.05, **P < 0.01, ***P < 0.001. All data were presented as the mean ± SE (n = 4); bars with different letters indicate significant differences across the study sites
Although the soil properties were significantly different among the study sites, the effects of degraded patch formation on most soil properties were consistent. When compared with the original patches, soil TN, TC, IN, AN, DOC, AP, and moisture contents were significantly lower in the degraded patches (P < 0.001, Fig. 2; Table S3). However, there were also some inconsistent responses of soil properties to the patch formation across the study sites. The patch formation significantly increased the δ15N values only at Sites 1 and 2, whereas it significantly decreased soil C:N ratios only at Sites 2 and 3. Additionally, soil NN contents in the degraded patches were significantly lower than those in the original patches at Site 1, but this trend was reversed at Sites 2 and 3 (Fig. 2d).
Copies of nitrogen-cycling genes and transcripts
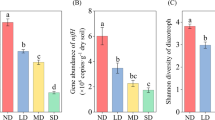
The copies of most nitrogen-cycling genes and transcripts under examination also significantly differed among the study sites (Table S3; Figs. 3, 4). The most obvious variation was that copies of archaeal amoA DNA at Site 1 were significantly lower than at Sites 2 and 3 (Fig. 3). Interestingly, the amoA gene also showed the highest sensitivity in response to the degraded patch formation. The copies of archaeal and bacterial amoA genes increased significantly in the degraded patches (Table S3; Fig. 3b). At Site 3, the amoA gene copies in the degraded patches were more than five times higher than in the original patches (Fig. 3b). The transcription of archaeal amoA genes was also greatly enhanced in the degraded patches (P = 0.032, Fig. 4b), especially at Site 3. However, the bacterial amoA mRNA copies remained unaffected by the patch formation, and showed a decreasing trend from Sites 1 to 3.
Effects of the study site and degraded patch formation on soil nitrogen-cycling gene copies. S1 Site 1, S2 Site 2, S3 Site 3, S effect of study site, D effect of degraded patch formation, *P < 0.05, **P < 0.01, ***P < 0.001. All data were presented as the mean ± SE (n = 4); bars with different letters indicate significant differences across the study sites
Effects of the study site and degraded patch formation on soil nitrogen-cycling gene mRNA copies. S1 Site 1, S2 Site 2, S3 Site 3, S effect of study site, D effect of degraded patch formation, *P < 0.05, **P < 0.01, ***P < 0.001. All data were presented as the mean ± SE (n = 4); bars with different letters indicate significant differences across the study sites
The degraded patch formation also tended to decrease nifH DNA copies, but this effect was not statistically significant (P = 0.094, Fig. 3a). In addition, transcriptions of the nifH and narG genes were significantly suppressed in the degraded patches at Site 2 (Fig. 4a, d), and the nirS transcription was significantly enhanced at Site 3 (Fig. 4f).
As revealed by the RDA ordinations, soil properties could be classified into three categories, with the first category including the contents of most soil nutrient indices (i.e., TN, TC, C:N, IN, AN, DOC, and AP) and soil moisture (Fig. 5). The second category included the soil NN contents and the values of δ13C and δ15N values, whereas the third category only included the soil pH values (Fig. 5). As determined by the ordinations, the first and second categories showed obviously negative correlations (Fig. 5).
Relationships between soil properties and DNA (a) or mRNA (b) copies of the nitrogen cycling genes. TC total carbon contents, TN total nitrogen contents, C:N ratio of total carbon to total nitrogen contents, M soil moisture contents, AP available phosphorus contents, IN inorganic nitrogen contents, DOC dissolved organic carbon contents, AN ammonium nitrogen contents, NN nitrate nitrogen contents, A-amoA archaeal amoA, B-amoA bacterial amoA. The ordinations were based on redundancy analysis
Similarly, nitrogen cycling DNA copies also could be clustered into three groups corresponding to diazotrophs (nifH), nitrifiers (amoA), and denitrifiers (narG, nirK, and nirS). Diazotrophs were positively correlated with the first category of the soil properties, but they were negatively correlated with the other soil properties (Fig. 4a). Nitrifiers showed almost totally inverse correlations, and were negatively and positively correlated with the first and second categories of the soil properties, respectively (Fig. 5a). The denitrifiers were only negatively correlated with soil pH values and several soil properties in the first category (Fig. 5a).
Regarding the nitrogen cycling mRNAs, there were no obvious clusters in the RDA ordination plot (Fig. 5b). However, when compared with their DNA counterparts, the nifH and archaeal amoA mRNA copies still showed similar correlations with the soil properties (Fig. 5b). Conversely, the other nitrogen cycling mRNA copies (e.g., bacterial amoA mRNA copies) showed different or even completely inverse correlations with the soil properties, when compared with their DNA counterparts (Fig. 5b). The results of RDA also were confirmed by the Pearson correlation between soil NCMs and properties (Table S4).
The community structures of the nitrogen-cycling groups
As suggested by the NMDS and PERMANOVA, the community structures of all the nitrogen-cycling groups differed significantly among the three study sites (Table S5; Fig. 6). However, only the community structures based on the nifH, narG, and archaeal amoA genes were significantly altered by the degraded patch formation (Table S5; Fig. 6). As revealed by the envfit analysis, the variations in the community structures were significantly correlated with a series of environmental factors (Table S6; Fig. 6). In particular, the community structures based on the nifH, narG, and nirS genes showed significant correlations with almost all soil properties included in this study. In addition, the nirK community structures were significantly correlated with the soil TN, TC, DOC, AN, IN, and moisture contents. However, the nitrifier community structures showed much weaker correlations with the soil properties. The AOA community structure was only significantly correlated with the soil C:N ratios and pH values, whereas the AOB community structure only showed significant correlations with the soil C:N ratios, pH values, and AN contents.
NMDS ordinations of the DNA-based community structures of nitrogen cycling microbes. S1O original patches at Site 1, S1D degraded patches at Site 1, S2O original patches at Site 2, S2D degraded patches at Site 2, S3O original patches at Site 3, S3D degraded patches at Site 3, TC total carbon contents, TN total nitrogen contents, C:N the ratios of total carbon to total nitrogen, M soil moisture, AP available phosphorus contents, IN inorganic nitrogen contents, DOC dissolved organic carbon contents, AN ammonium nitrogen contents, NN nitrate nitrogen contents, S effect of study site, D effect of degraded patch formation, *P < 0.05, **P < 0.01, ***P < 0.001. Vectors represent the soil properties that were significantly correlated with the corresponding NCG community structures (P < 0.05); the directions of the vectors represent the increase in gradient of each soil property. All vectors were drawn using the envfit function in the vegan package
Discussion
Decreased soil nitrogen contents under grassland degradation have been observed in a number of studies (Dong et al. 2012; Li et al. 2014; Wang et al. 2005, 2015a; Wen et al. 2013a). Our study found that, when compared with the adjacent original patches, the nitrogen contents in the degraded patches declined dramatically, and the nitrogen contents in the original patches also decreased from Sites 1 to 3 (Fig. 2). Thus, the formation of degraded patches might decrease not only the nitrogen contents within the bared patches but also the contents of their adjacent original patches through the “barren patch effect” (Han et al. 2011). The 15N natural abundance is widely used to explore soil nitrogen dynamics (Almaraz and Porder 2016; Chalk et al. 2017; Hall et al. 2016; Wang et al. 2015b). As summarized by Wang et al. (2014), plant absorption and gas emissions from ammonia volatilization, nitrification, and denitrification tend to increase the δ15N values of the remaining N, whereas nitrogen fixation and deposition usually decrease soil δ15N values. However, leaching exerts only a slightly negative fractionation effect on soil 15N values (Mnich and Houlton 2016). Thus, at Sites 1 and 2, the relatively high δ15N values in the degraded patches may indicate an increased gas loss of nitrogen and (or) lowered biological nitrogen fixation in the degraded patches, whereas at Site 3, the almost identical δ15N values in the degraded and original patches suggest that the lower nitrogen contents in the degraded patches may be mainly attributed to nitrogen leaching.
Another intriguing finding of this study is that although the degraded patch formation significantly decreased the soil nitrogen contents, it tended to increase the soil NN contents (Fig. 2). Both nitrate and soil particles carry negative charges. Because of the “mutual repulsion effect”, nitrate is more susceptible to leaching from soil than ammonium (Di and Cameron 2002a; Kowalchuk and Stephen 2001). Furthermore, nitrate is the substrate of denitrification and therefore is prone to loss through gas emissions during denitrification (Levy-Booth et al. 2014). In this study, the soil nitrate contents were shown to be negatively correlated with soil TN, IN, and AN contents, whereas it was positively correlated with soil δ15N values (Fig. 4). Therefore, the increased soil nitrate contents might also contribute to the decreased nitrogen contents under the degraded patch formation.
In the degraded patches, the accumulation of NN could be attributed to the potentially enhanced production and depressed consumption processes of nitrate. First, as soil nitrate is mainly derived from nitrification (Ollivier et al. 2011), the dramatically increased soil nitrifier abundance (Fig. 3b, c) and the upregulated transcription of the archaeal amoA gene (Fig. 4b) should play a vital role in the accumulation of NN in the degraded patches. Indeed, this is also supported by the significant correlations between the abundance and transcription of the amoA gene and NN contents (Fig. 5). Moreover, the lack of (or reduced number of) plants in the degraded patches could lower the plant assimilation of ammonium, thereby allowing more ammonium to be nitrified to nitrate. Similarly, the decrease in plant nitrate uptake also could contribute to the higher soil NN contents in the degraded patches. Among these potential causes, decreased consumption of nitrate should benefit the retention of soil nitrogen fertility, while the potential enhancement of nitrification could result in increased nitrogen loss. Moreover, because of the low availability of nitrogen in the degraded patches, AOA should be more responsible for the nitrification than AOB (Prosser and Nicol 2012; Stahl and de la Torre 2012a).
According to the physiological properties of nitrifiers and nitrification, the increased nitrifier abundance and archaeal amoA mRNA copies in the degraded patches could be elicited in a number of ways. First, degraded patch formation significantly decreased the soil moisture and led to erosion and coarsening of surface soils (Figs. 1, 2i; He and Richards 2015; Lin et al. 2015), which might indirectly increase the soil aeration. In most cases, ammoxidation is an aerobic process (Zhalnina et al. 2012), and thus the increased soil aeration should promote soil nitrification. Second, as most AOA and AOB are chemoautotrophic (Kowalchuk and Stephen 2001; Zhalnina et al. 2012), the decreased soil DOC under degradation should increase their competitiveness and abundance. This is supported by the significantly negative correlations between soil DOC and the abundance of ammonia-oxidizing microbes in this study (Fig. 2a) and a number of other investigations (Bates et al. 2011; Wessen et al. 2010). Consistently, our unpublished data also showed that plant litter deprivation could significantly increase the abundance of ammonia oxidizing microbes. Third, as reviewed by Subbarao et al. (2015), roots of some plants can produce and release nitrification inhibitors; therefore, the elimination of vegetation in the degraded patches also should promote nitrification. Furthermore, the lack of (or reduced number of) plants in the degraded patches could result in lower plant–microbe competition for ammonium (Kuzyakov and Xu 2013), which supports the higher nitrifier abundance and the potentially enhanced nitrification. Although several other soil properties (e.g., soil available phosphorus contents) also showed significant correlations with AOA and AOB abundance, because of the paucity of theoretical supports, their contributions to the increases in nitrifier abundance are still difficult to justify.
On the basis of these findings, there are several potentially promising management practices to alleviate the accelerated nitrogen loss that occurs under degraded patch formation. First, grazing exclusion, pika control, and the establishment of artificial grassland can increase the carbon input from plants to soil (Chen et al. 2017; Dong et al. 2015; Li et al. 2013, 2014; Lu et al. 2015; Xiong et al. 2014), which can attenuate the competitive ability of nitrifiers, and thus lower nitrification and nitrogen loss risk. Second, application of nitrification inhibitors (e.g., 3, 4-dimethylpyrazole phosphate, dicyandiamide, and nitrapyrin) should be another potential measure, as they can directly inhibit soil nitrification (Di and Cameron 2002b; Zaman et al. 2008). However, the high cost of nitrification inhibitors may impede their current use in the restoration of degraded grassland. Accordingly, increased effort should be made to reduce the cost of nitrification inhibitors or incorporate plants capable of producing biogenic nitrification inhibitors into grassland restoration in the future.
In this study, we also examined the responses of diazotrophs and denitrifiers to degraded patch formation (Figs. 3, 4, and 6). However, the responses of the nitrogen-fixing and denitrifying gene copies and their transcriptions were generally weak and showed high inconsistency across the study sites (Figs. 3, 4; Table S3). This response could be elicited by the contrasting effects of the patch formation-induced variations. On one hand, the decreased soil IN and increased soil nitrate contents should benefit diazotrophs and denitrifiers, respectively (Che et al. 2017; Levy-Booth et al. 2014; Reed et al. 2011). On the other hand, the decrease in soil DOC, phosphorous, and other nutrient contents should negatively affect diazotrophs and denitrifiers (Levy-Booth et al. 2014; Reed et al. 2011). Nevertheless, the community structures based on nifH and narG genes were much more sensitive to the patch formation, and were significantly correlated with several different soil properties (Fig. 6; Table S5, S6), which deserves further exploration with more precise taxonomy techniques (e.g., MiSeq sequencing).
In conclusion, this study showed that degraded patch formation of the Tibetan alpine meadows significantly decreased most soil nutrient properties, while it tended to increase soil nitrate contents in heavily degraded alpine meadows. Among the NCMs included in this study, nitrifiers showed the highest sensitivity to the degraded patch formation. AOA and AOB abundance and archaeal amoA mRNA copies all significantly increased in the degraded patches. Moreover, amoA gene and archaeal amoA mRNA copies were positively correlated with soil nitrate nitrogen contents and δ15N values, but negatively correlated with soil total and inorganic nitrogen contents. These results suggest that the dramatically increased ammonia-oxidizing microbial abundance in the degraded patches may be a potentially important driver for the higher nitrogen loss in these patches. Therefore, grazing reduction, pika control, reseeding, and nitrification inhibitor applications could be potentially promising ways to alleviate nitrogen loss during degraded patch formation.
References
Almaraz M, Porder S (2016) Reviews and syntheses: measuring ecosystem nitrogen status—a comparison of proxies. Biogeosciences 13(18):5395–5403
Bai ZG, Dent DL, Olsson L, Schaepman ME (2008) Proxy global assessment of land degradation. Soil Use Manag 24(3):223–234
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N (2011) Examining the global distribution of dominant archaeal populations in soil. ISME J 5(5):908–917
Chalk PM, He JZ, Peoples MB, Chen D (2017) 15N2 as a tracer of biological N2 fixation: a 75-year retrospective. Soil Biol Biochem 106:36–50
Che RX, Deng YC, Wu YB, Zhang J, Wang F, Tang L, Li LF, Ma S, Liu HK, Zhao X, Wang YF, Hao YB, Cui XY (2017) Relationships between biological nitrogen fixation and available nitrogen at scales from molecular to community level. Chin J Ecol 36(1):224–232
Chen JJ, Yi SH, Qin Y (2017) The contribution of plateau pika disturbance and erosion on patchy alpine grassland soil on the Qinghai-Tibetan Plateau: implications for grassland restoration. Geoderma 297:1–9
Cui XF, Graf HF (2009) Recent land cover changes on the Tibetan Plateau: a review. Clim Change 94(1):47–61
Di HJ, Cameron KC (2002a) Nitrate leaching in temperate agroecosystems: sources, factors and mitigating strategies. Nutr Cycl Agroecosyst 64(3):237–256
Di HJ, Cameron KC (2002b) The use of a nitrification inhibitor, dicyandiamide (DCD), to decrease nitrate leaching and nitrous oxide emissions in a simulated grazed and irrigated grassland. Soil Use Manag 18(4):395–403
Dong SK, Wen L, Li YY, Wang XX, Zhu L, Li XY (2012) Soil-quality effects of grassland degradation and restoration on the Qinghai-Tibetan Plateau. Soil Sci Soc Am J 76(6):2256–2264
Dong SK, Wang XX, Liu SL, Li YY, Su XK, Wen L, Zhu L (2015) Reproductive responses of alpine plants to grassland degradation and artificial restoration in the Qinghai-Tibetan Plateau. Grass Forage Sci 70(2):229–238
Dong JF, Cui XY, Wang SP, Wang F, Pang Z, Xu N, Zhao GQ, Wang SP (2016) Changes in biomass and quality of alpine steppe in response to N & P fertilization in the Tibetan Plateau. PLoS ONE 11(5):e0156146
Gaby JC, Buckley DH (2011) A global census of nitrogenase diversity. Environ Microbiol 13(7):1790–1799
Galloway JN, Dentener FJ, Capone DG, Boyer EW, Howarth RW, Seitzinger SP, Asner GP, Cleveland CC, Green PA, Holland EA, Karl DM, Michaels AF, Porter JH, Townsend AR, Vorosmarty CJ (2004) Nitrogen cycles: past, present, and future. Biogeochemistry 70(2):153–226
Hall SJ, Weintraub SR, Bowling DR (2016) Scale-dependent linkages between nitrate isotopes and denitrification in surface soils: implications for isotope measurements and models. Oecologia 181(4):1221–1231
Han L, Shang Z, Ren G, Wang Y, Ma Y, Li X, Long R (2011) The responses of plant and soil in the “black soil beach” on the Tibetan Plateau to the changes of degradation patch areas. Acta Prataculturae Sin 20(1):1–6
He SY, Richards K (2015) Impact of meadow degradation on soil water status and pasture management—a case study in Tibet. Land Degrad Dev 26(5):468–479
Kowalchuk GA, Stephen JR (2001) Ammonia-oxidizing bacteria: a model for molecular microbial ecology. Annu Rev Microbiol 55(1):485–529
Kuzyakov Y, Xu XL (2013) Competition between roots and microorganisms for nitrogen: mechanisms and ecological relevance. New Phytol 198(3):656–669
Lal R (2003) Soil erosion and the global carbon budget. Environ Int 29(4):437–450
Levy-Booth DJ, Prescott CE, Grayston SJ (2014) Microbial functional genes involved in nitrogen fixation, nitrification and denitrification in forest ecosystems. Soil Biol Biochem 75:11–25
Li YY, Dong SK, Wen L, Wang XX, Wu Y (2013) The effects of fencing on carbon stocks in the degraded alpine grasslands of the Qinghai-Tibetan Plateau. J Environ Manag 128:393–399
Li YY, Dong SK, Wen L, Wang XX, Wu Y (2014) Soil carbon and nitrogen pools and their relationship to plant and soil dynamics of degraded and artificially restored grasslands of the Qinghai-Tibetan Plateau. Geoderma 213:178–184
Li YY, Dong SK, Liu SL, Zhou HK, Gao QZ, Cao GM, Wang XX, Su XK, Zhang Y, Tang L, Zhao HD, Wu XY (2015) Seasonal changes of CO2, CH4 and N2O fluxes in different types of alpine grassland in the Qinghai-Tibetan Plateau of China. Soil Biol Biochem 80:306–314
Li Y, Wang S, Jiang L, Zhang L, Cui S, Meng F, Wang Q, Li X, Zhou Y (2016) Changes of soil microbial community under different degraded gradients of alpine meadow. Agric Ecosyst Environ 222:213–222
Lin L, Li YK, Xu XL, Zhang FW, Du YG, Liu SL, Guo XW, Cao GM (2015) Predicting parameters of degradation succession processes of Tibetan Kobresia grasslands. Solid Earth 6(4):1237–1246
Lu XY, Yan Y, Sun J, Zhang XK, Chen YC, Wang XD, Cheng GW (2015) Carbon, nitrogen, and phosphorus storage in alpine grassland ecosystems of Tibet: effects of grazing exclusion. Ecol Evol 5(19):4492–4504
Mnich ME, Houlton BZ (2016) Evidence for a uniformly small isotope effect of nitrogen leaching loss: results from disturbed ecosystems in seasonally dry climates. Oecologia 181(2):323–333
Ollivier J, Towe S, Bannert A, Hai B, Kastl EM, Meyer A, Su MX, Kleineidam K, Schloter M (2011) Nitrogen turnover in soil and global change. FEMS Microbiol Ecol 78(1):3–16
Olsen SR, Cole CV, Watanabe FS, Dean LA (1954) Estimation of available phosphorus in soils by extraction with sodium bicarbonate. United States Department Of Agriculture Cirular 939, Washington
Prosser JI, Nicol GW (2012) Archaeal and bacterial ammonia-oxidisers in soil: the quest for niche specialisation and differentiation. Trends Microbiol 20(11):523–531
Raymond J, Siefert JL, Staples CR, Blankenship RE (2004) The natural history of nitrogen fixation. Mol Bio Evol 21(3):541–554
R Development Core Team (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, http://www.R-project.org
Reed SC, Cleveland CC, Townsend AR (2011) Functional ecology of free-living nitrogen fixation: a contemporary perspective. Annu Rev Ecol Evol Syst 42:489–512
Ren GH, Shang ZH, Long RJ, Hou Y, Deng B (2013) The relationship of vegetation and soil differentiation during the formation of black-soil-type degraded meadows in the headwater of the Qinghai-Tibetan Plateau, China. Environ Earth Sci 69(1):235–245
Stahl DA, de la Torre JR (2012) Physiology and diversity of ammonia-oxidizing archaea. Annu Rev Microbiol 66:83–101
Subbarao GV, Yoshihashi T, Worthington M, Nakahara K, Ando Y, Sahrawat KL, Rao IM, Lata JC, Kishii M, Braun HJ (2015) Suppression of soil nitrification by plants. Plant Sci 233:155–164
Tang L, Dong SK, Sherman R, Liu SL, Liu QR, Wang XX, Su XK, Zhang Y, Li YY, Wu Y, Zhao HD, Zhao C, Wu XY (2015) Changes in vegetation composition and plant diversity with rangeland degradation in the alpine region of Qinghai-Tibet Plateau. Rangel J 37(1):107–115
Thamdrup B (2012) New pathways and processes in the global nitrogen cycle. Annu Rev Ecol Evol Syst 43:407–428
Wang WY, Wang QJ, Wang CY, Shi HL, Li Y, Wang G (2005) The effect of land management on carbon and nitrogen status in plants and soils of alpine meadows on the Tibetan plateau. Land Degrad Dev 16(5):405–415
Wang C, Wang X, Liu D, Wu H, Lu X, Fang Y, Cheng W, Luo W, Jiang P, Shi J, Yin H, Zhou J, Han X, Bai E (2014) Aridity threshold in controlling ecosystem nitrogen cycling in arid and semi-arid grasslands. Nat Commun 5:4799
Wang XX, Dong SK, Sherman R, Liu QR, Liu SL, Li YY, Wu Y (2015a) A comparison of biodiversity-ecosystem function relationships in alpine grasslands across a degradation gradient on the Qinghai-Tibetan Plateau. Rangel J 37(1):45–55
Wang Y, Xu Z, Zheng J, Abdullah KM, Zhou Q (2015b) Delta 15N of soil nitrogen pools and their dynamics under decomposing leaf litters in a suburban native forest subject to repeated prescribed burning in southeast Queensland, Australia. J Soils Sediment 15(5):1063–1074
Wang ZQ, Zhang YZ, Yang Y, Zhou W, Gang CC, Zhang Y, Li JL, An R, Wang K, Odeh I, Qi JG (2016) Quantitative assess the driving forces on the grassland degradation in the Qinghai-Tibet Plateau, in China. Ecol Inform 33:32–44
Wen L, Dong SK, Li YY, Li XY, Shi JJ, Wang YL, Liu DM, Ma YS (2013a) Effect of degradation intensity on grassland ecosystem services in the alpine region of Qinghai-Tibetan Plateau, China. PLoS ONE 8(3):e58432
Wen L, Dong SK, Li YY, Wang XX, Li XY, Shi JJ, Dong QM (2013b) The impact of land degradation on the C pools in alpine grasslands of the Qinghai-Tibet Plateau. Plant Soil 368(1–2):329–340
Wessen E, Nyberg K, Jansson JK, Hallin S (2010) Responses of bacterial and archaeal ammonia oxidizers to soil organic and fertilizer amendments under long-term management. Appl Soil Ecol 45(3):193–200
Wu RG, Tiessen H (2002) Effect of land use on soil degradation in alpine grassland soil, China. Soil Sci Soc Am J 66(5):1648–1655
Wu GL, Ren GH, Dong QM, Shi JJ, Wang YL (2014) Above- and belowground response along degradation gradient in an alpine grassland of the Qinghai-Tibetan Plateau. Clean Soil Air Water 42(3):319–323
Xiong DP, Shi PL, Sun YL, Wu JS, Zhang XZ (2014) Effects of grazing exclusion on plant productivity and soil carbon, nitrogen storage in alpine meadows in northern Tibet, China. Chin Geogr Sci 24(4):488–498
Zaman M, Nguyen ML, Blennerhassett JD, Quin BF (2008) Reducing NH3, N2O and NO3–N losses from a pasture soil with urease or nitrification inhibitors and elemental S-amended nitrogenous fertilizers. Biol Fertil Soils 44(5):693–705
Zhalnina K, de Quadros PD, Camargo FAO, Triplett EW (2012) Drivers of archaeal ammonia-oxidizing communities in soil. Front Microbiol 3:210
Zhang L, Unteregelsbacher S, Hafner S, Xu X, Schleuss PM, Miehe G, Kuzyakov Y (2017) Fate of organic and inorganic nitrogen in crusted and non-crusted Kobresia grasslands. Land Degrad Dev 28(1):166–174
Zhao HD, Liu SL, Dong SK, Su XK, Wang XX, Wu XY, Wu L, Zhang X (2015) Analysis of vegetation change associated with human disturbance using MODIS data on the rangelands of the Qinghai-Tibet Plateau. Rangel J 37(1):77–87
Acknowledgements
This work was supported by the Strategic Priority Research Program (B) of the Chinese Academy of Sciences (XDB15010201), National Natural Science Foundation of China (41230750), National Basic Research Program (2013CB956000), and National Key Research and Development Program of China (2016YFC0501800). We sincerely appreciate the constructive suggestions from Prof. Zhongjun Jia and Dr. Yang Li regarding preparation of the manuscript. Rongxiao Che was also supported by a Griffith University Postgraduate Research Scholarship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Christine Hawkes.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Che, R., Wang, F., Wang, W. et al. Increase in ammonia-oxidizing microbe abundance during degradation of alpine meadows may lead to greater soil nitrogen loss. Biogeochemistry 136, 341–352 (2017). https://doi.org/10.1007/s10533-017-0399-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10533-017-0399-5