Abstract
Background and aims
Plant growth is frequently limited by the availability of inorganic phosphorus (P) in the soil. In most soils, a considerable amount of the soil P is bound to organic molecules. Of these, phytate is the most abundant identifiable organic P form, but is not readily available to plants. In contrast, microorganisms have been shown to degrade phytate with high efficiency. The current study aims to characterize the members of the phytate-hydrolysing bacterial community in rhizosphere, and the molecular and enzymatic ability of these bacteria to degrade phytate.
Methods and results
The phytate-hydrolysing bacterial community was characterized from the rhizosphere of plants cultivated in the presence or absence of phytate supplementation. Major changes in the bacterial community structure were observed with both culture-dependent and -independent methods, which highlighted the predominance of Proteobacteria and Actinobacteria. Phytase activity was detected for a range of rhizobacterial isolates as well as the presence of, β-propeller phytases (BPP) for both isolates and directly in a soil sample.
Conclusion
A wide taxonomic range of functional phytate utilizers have been discovered, in soil bacterial taxa that were previously not well known for their ability to utilise phytate as P or C sources. This study provides new insights into microbial carbon and phosphorus cycling in soil.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In the rhizosphere, the supply of inorganic phosphorus (P) constitutes one of the major limiting factors for plant growth (Hammond et al. 2004; Lambers et al. 2006). Although different types of phosphatases and RNases are produced by plants, inorganic phosphate remains the main source in plant P assimilation (Raghothama and Karthikeyan 2005). The phosphate concentration generally observed in the soil solution is below 10 μM (Hinsinger 2001). In addition, phosphate mobility is very poor and is highly influenced by soil composition and pH (Hinsinger 2001). A major proportion of the total soil P content is present in organic forms [29 to 90 %; (Richardson et al. 2001b; Turner et al. 2002)]. myo-Inositol hexakisphosphate (or myo-IP6) generally represents between 10 and 60 % of total organic P in soil, but may constitute almost all the organic P in some calcareous soils (Turner et al. 2002). Agriculture constitutes a major source of myo-IP6 to soils, because of seed sowing and the extensive use of monogastric manures (Turner et al. 2002; Turner and Leytem 2004). In addition, the formation of insoluble salts (named phytate) or clay- and organic matter-complexes leads to a high accumulation of phytate in soils (Hayes et al. 2000a; Turner et al. 2002). The solubility of these complexes may be increased in the rhizosphere by the release of organic acids in root exudates, enhancing the availability of phosphate and potentially phytate for mineralisation by rhizobacteria (Hayes et al. 2000b; Hinsinger 2001). Evidence of such processes has however mainly been provided by studies in vitro or in synthetic media, rather than directly in soils (Dao 2007; Giles et al. 2012). Importantly, it has also been suggested that phytase itself may be rapidly immobilised in soil environment, inhibiting its activity or limiting its mobility (George et al. 2005).
The hydrolysis of phytate is achieved by the phytase enzyme, named 3-phytase (EC 3.1.3.8) or 6-phytase (EC 3.1.3.26) depending on the position of the first phosphate group removed. The term 4-phytase (EC 3.1.3.26) found in the literature corresponds to 6-phytase when based on 1D-numbering system instead of 1 L-numbering. Successive dephosphorylation of myo-IP6 to yield myo-IP1 can be achieved by phytase, but the rate of dephosphorylation differs for the intermediate forms (Greiner et al. 2007). Phytase does not dephosphorylate myo-IP1, but this step can be catalysed by widely distributed phosphomonoesterase enzymes (acid and alkaline phosphatases) in the environment (Turner et al. 2002).
Phytases have been divided into four classes according to their structures and the mechanism implied in the cleavage of phosphate groups. These are (i) histidine acid phosphatase (HAP), (ii) β-propeller phytase (BPP), (iii) purple acid phosphatase (PAP) and (iv) cysteine phytase (CPhy; alternative name PTP-like, protein tyrosine phosphatase-like) (Lim et al. 2007). All four classes of phytase have been identified in terrestrial environments but only BPP has been retrieved in aquatic environment (Cheng and Lim 2006). In addition, BPP is the only class that exhibits phytase activity at neutral and alkaline pH (Greiner et al. 2007). The first phytases described were HAPs and were originally characterized from fungi (Mullaney et al. 2000). Expression of HAP in transgenic plants has been studied in some detail (Richardson et al. 2001a; Zimmermann et al. 2003). The bacterial diversity and the ecology of the other phytase classes remain poorly characterized (Mullaney and Ullah 2003; Hill and Richardson 2006; Lim et al. 2007; Huang et al. 2011). The BPP class of phytase enzymes were first isolated from Bacillus strains (Kerovuo et al. 1998; Shin et al. 2001), and have since been shown to occur in a wide range of environments (Cheng and Lim 2006; Greiner et al. 2007; Lim et al. 2007; Huang et al. 2009; Jorquera et al. 2012, 2014). Phylogenetic analysis of BPP genes in a range of bacteria, mostly from aquatic environments (Hill et al. 2007; Lim et al. 2007; Huang et al. 2009), revealed suitable DNA homology for the design of BPP-specific probes (Huang et al. 2009). BPP genes previously characterized from rhizosphere are mainly affiliated to Firmicutes (Jorquera et al. 2011, 2012, 2014).
Plant species lack extracellular phytase or display only low phytase activity (Li et al. 1997; Richardson et al. 2001b; Lambers et al. 2006) and therefore cannot readily use soil phytate as a P source. However, a variety of microorganisms have been shown to hydrolyse phytate in various ecosystems (Richardson and Hadobas 1997; Yanke et al. 1998; Idriss et al. 2002; Hill and Richardson 2006; Hill et al. 2007; Nakashima et al. 2007; Jorquera et al. 2008a; Huang et al. 2009). Many organisms harbour a phytase gene in their genomes (Lim et al. 2007; Jorquera et al. 2008b, 2012, 2014), though the catalytic efficiency of these phytase homologues has not yet been demonstrated biochemically. Pseudomonas and Bacillus strains have been isolated from soils and studied for their ability to hydrolyse phytate and to enhance the P nutrition of plants (Richardson and Hadobas 1997; Richardson et al. 2001b; Idriss et al. 2002). By contrast, the phytate-hydrolysing community in the rhizosphere remains poorly characterized and has primarily been assessed using culture-dependent methods, based on the use of phytate-containing media. Phytate-hydrolysing bacteria belonging to the Pseudomonas, Enterobacter and Pantoea genera have been identified in various plant rhizospheres (Jorquera et al. 2008a), while Burkholderia spp. have been shown to be the major component of the phytate-hydrolysing bacterial community in Lupinus albinus rhizosphere (Unno et al. 2005). Phytate-hydrolysing bacteria in Lolium perenne rhizosphere are thought to make up ~40 % of the total heterotrophic bacteria (Jorquera et al. 2008a). L. perenne is a key forage grass worldwide representing 70 % of agricultural land in United Kingdom (Sharma and Sahi 2005; King et al. 2008), but it shows limited growth with phytate as sole P source (Martin et al. 2004), and is therefore a useful model to assess the phytate-hydrolysing bacterial community in the rhizosphere.
In this study, we investigated the phytate-hydrolysing bacterial community in L. perenne rhizosphere, by combining 16S rRNA- and BPP-based molecular approaches and enzymatic activity.
Material and methods
Soils, media for plant and bacterial growth
Two sandy loam soils characterized by low-phosphorus content were selected for the experiment. Lindow soil was an unsterilized topsoil purchased from a commercial supplier (Lindow Turf company, Wilmslow, United Kingdom). The Warwick soil was obtained from a soil phosphate series located at Warwick HRI (Wellesbourne, United Kingdom: 52°12′46N, 1°36′21W), which had been previously cultivated with oilseed rape. This second soil has had no added phosphate for 40 years. Warwick soil was sampled from the top 15 cm in early December 2007. Soil characteristics for the two soils were determined by Eurofins laboratories (Wolverhampton, United Kingdom) and are reported in Table 1.
Nutrient media used for plant growth were modified versions of 0.25× Hoagland’s solution (Zysko et al. 2012). The solution was modified to contain 0.210 mM CaCl2 in order to avoid the formation of calcium-phytate precipitate at the pH required (pH 5.6). Two phosphorus levels were used, a ‘low-P’ treatment modified to contain 0.030 mM phosphate, and a ‘high-P’ treatment, supplied as phytate, containing 0.030 mM phosphate and 0.167 mM phytic acid sodium salt (Sigma-Aldrich, Gillingham, United Kingdom).
Two types of phytate-containing media were used for bacterial isolation, corresponding to a solid minimal medium (MM) (50 mM Tris base, 20 mM NH4Cl, 0.5 mM MgCl2, 20 mM NaCl, 20 mM KCl, 0.5 mM Na2SO4, 1.0 ml/L of a trace element solution (Kertesz et al. 1993)) supplemented either with 1 mM phytate (PMI; phytate as carbon (C) and (P) sources) or 1 mM phytate and D-glucose (0.72 g/L), Na succinate (1.52 g/L) and glycerol (0.734 ml/L) (PMII; phytate only as a P source). The pH of each medium was adjusted to 7.0 with HCl and 1.5 % agar (Bacto-Agar, Difco, <0.005 % P) was added to prepare solid media where required. R2A agar and liquid medium (Oxoid, Basingstoke, United Kingdom) (Reasoner and Geldreich 1985) were used for phytate-independent cultivation. All glassware was acid-washed (3 M HCl) and all medium components were orthophosphate-free.
Plant experimental set up and growth conditions
Lolium perenne L. ‘Kent’ (Emorsgate seeds, Norfolk, United Kingdom) seeds were washed in 70 % (v/v) ethanol, surface sterilised for 15 min in 1 % (v/v) peracetic acid (Sigma-Aldrich) and washed five times with sterile water (floating seeds were discarded) (Zysko et al. 2012). Surface-sterilised seeds were sown on 0.5× Murashige and Skoog basal medium (MS) (Murashige and Skoog 1962) (Duchefa Biochemie, Haarlem, the Netherlands) with 1.5 % (w/v) agar and germinated in the dark at 25 °C for 4 days.
Pots (10 × 10 cm) were filled with 200 g (dry weight equivalent) of each soil mixed with sand in a 9:1 sand:soil ratio. Ten 4-day old seedlings were transplanted into each pot, and the plants were incubated in a growth chamber (PlantMaster, CLF Plant Climatics, Emersacker, Germany) under controlled conditions (16 h light/8 h dark, 21 °C day/night; light intensity 1250 μmol.m−2.s−1). There were four experimental replicates for each treatment in each soil. Modified Hoagland’s medium was added to each pot (11 ml.day−1), using a multichannel, low-flow peristaltic pump (Watson-Marlow 250U, Falmouth, United Kingdom). All pots were initially treated for 5 days with a ‘low-P’ treatment, and replicate sets of pots were then subjected either to a continuation of the ‘low-P’ treatment or ‘high P’ for a further 23 days.
Isolation of phytate-hydrolysing bacteria from L. perenne rhizospheres
The 28-day old plants were carefully removed from the pots and the root system was shaken gently to separate loosely adhering sand and soil. For bacterial isolation, two grams of each rhizosphere sample was suspended in a sterile tube containing 20 ml of 10 mM MgCl2 and five glass beads (0.5 mm diameter). Tubes were vortexed at maximum speed for 30 s to remove bacteria, and the resultant suspension was diluted in 10 mM MgCl2. To isolate phytate-hydrolysing bacteria, serial dilutions were spread on PMI and PMII media, and then incubated at 20 °C for 14 days. A selection of all morphologically diverse colonies (3–5 colonies per morphotype) was re-purified by streaking on fresh PMI and PMII plates, and bacterial growth on the phytate-containing media (PMI and PMII) was confirmed. Purified cultures were re-grown on R2A medium (Oxoid, Basingstoke, United Kingdom) for storage at −80 °C.
Utilisation of phytate
The ability of selected isolates to utilize phytate as C source and/or P source was estimated in phytate-containing liquid media. Growth rates and phytate disappearance were estimated in liquid MM medium supplemented either with 1 mM phosphate (K2HPO4), or with 200 μM phytate (1.2 mM P) as P sources, and Na succinate (1.52 g/L) or glucose (0.72 g/L) as C sources. Phytate concentration in the growth supernatant was quantified by ion chromatography using a Dionex system controlled by Chromeleon software (Dionex), and equipped with Omnipac PAX-100 analytical (4 × 250 mm) and guard (4 × 50 mm) columns (Dionex). Phytate was eluted from the column using a multistep gradient of 0–120 mM sodium hydroxide in 6 % (v/v) aqueous isopropanol and using a flow rate of 1 ml min−1 at 25 °C. Compounds eluted from the column were detected by conductimetry, using an ED50 electrochemical detector coupled to an ASRS300 micromembrane suppressor (Dionex).
Isolates showing reproducible phytate disappearance were grown in liquid MM supplemented either with 10 mM phytate (as C and P sources), or with 10 mM phytate (as C and/or P sources) and 10 mM inositol (as C source). The liquid MM supplemented with inositol was used in order to specify the phytate catabolism of each isolate, since complete dephosphorylation of phytate leads to inositol. The isolates were pre-cultivated in Luria-Bertani (LB) medium overnight (20 °C, 200 r.p.m) and then washed twice with 10 mM MgCl2. The bacterial suspension was adjusted to an OD600 of 0.5 (~2 × 108 cells/mL; Pharmacia Ultrospec spectrophotometer), inoculated (1 % v/v) into 200 μl of the different liquid media in 96 microwell plates, and incubated at 25 °C for 48 h, with shaking. Bacterial growth was measured by periodic determinations (every 10 min) of OD600 during this period.
DNA extraction and 16S rRNA PCR conditions
Bacterial DNA extraction from rhizosphere samples (four replicates of 0.5 g per sample) was performed using soil FastDNA® SPIN Kit for Soil (QBiogene, Cambridge, United Kingdom) according to the manufacturer’s instructions. DNA was quantified with a Nanodrop ND100 spectrophotometer (Thermo-scientific, Waltham, MA, USA) and stored at −20 °C. Genomic DNA from individual isolates was obtained by suspending a colony in 100 μl sterile water and heating at 95 °C for 5 min.
The 16S rRNA genes from rhizobacterial isolates were amplified using the bacterial universal primers 27F and 1492R (Lane 1991). Polymerase chain reaction (PCR) amplifications were performed in a T1 cycler (Biometra, Goettingen, Germany), as follows (50 μl reaction): 1× reaction buffer, 1.5 mM MgCl2, 0.5 μM of each primer, 50 μM of each dNTP, 2.5 U of Taq DNA polymerase (BIOTAQ; Bioline, London, United Kingdom), 1 μl of genomic DNA. Thermal cycling was carried out with a denaturation step of 94 °C for 3 min, 30 cycles with 45 s denaturation at 94 °C, 45 s at annealing temperature (AT) 56 °C, 90 s elongation at 72 °C, and a final elongation step for 5 min at 72 °C. PCR products were purified with a QIAquick PCR purification column (QIAGEN) according to the manufacturer’s instructions.
PCR for 16S rRNA gene-based denaturing gradient gel electrophoresis (16S-DGGE) were carried out using universal and group-specific primers (Table 2). A 2- or 3-step nested PCR approach was used for 16S-DGGE as described in Muhling et al. (2008). A 16S rRNA gene PCR amplification using the bacterial universal primers 27F and 1492R was done for the first step, according to the PCR protocol described above. 16S-DGGE on the total bacterial community was done using PCR product (1 μl) from the first step as template and the bacterial universal primers 341F-GC and 518R (Muyzer et al. 1993). Group-specific PCR amplification was done using group-specific primers (Table 2) and PCR product (1 μl) from the first step as template. All group-specific PCRs used the following protocol: initial denaturation step at 94 °C for 3 min, 30 cycles with 1 min denaturation at 94 °C, 1 min at the respective annealing temperature (Table 2), 1 min elongation at 72 °C, and a final elongation step for 5 min at 72 °C. Finally, 16S group-specific DGGE were done using the second primer pair described in Table 2.
For all 16S PCR-DGGE, a touchdown PCR protocol using the DGGE primers (see Table 2) was done as follows: initial denaturation step at 95 °C for 5 min, 10 cycles with 1 min at 95 °C, 1 min at 65 °C (reducing by 1 °C/cycle to 55 °C), 1 min elongation at 72 °C and a further 20 cycles with a fixed annealing temperature at 55 °C [modified from Cunliffe and Kertesz (2006)]. PCR products for 16S-DGGE were purified through a QIAquick PCR purification column (QIAGEN) according to manufacturer and quantified with a Nano-drop ND100 spectrophotometer.
16S rRNA-based denaturing gradient gel electrophoresis (16S-DGGE)
Denaturing gradient gel electrophoresis was carried out on 20 × 16 cm gels in a Dcode electrophoresis chamber (Bio-Rad, Hercules, CA) with a denaturant gradient of 30–70 % (total bacterial community) or 40–60 % (group-specific community) and electrophoresis for 1000 Vh as previously described (Cunliffe and Kertesz 2006). Profiles from L. perenne rhizosphere samples were prepared with 300 ng of DNA, while samples with defined, mixed species contained 50 ng of DNA per species/band. Gels were stained for 30 min with SybrGold (Invitrogen, Carlsbad, CA), rinsed briefly with dH2O and scanned with an UVItec trans-illuminator (UVitec, Cambridge, United Kingdom).
DGGE profiles were analysed using Gelcompar II software (Applied Maths, Kortrijk, Belgium) and subjected to principal component analysis (PCA) using the R statistical computing environment (http://www.r-project.org). The significance of the differences between soils and treatments derived from PCA was evaluated using analysis of variance (ANOVA) tests, followed by Tukey’s honestly significant different (HSD) tests. Statistics were performed at P < 0.05 using R.
Restriction fragment length polymorphism analysis (16S-RFLP) and sequencing
The 16S rRNA gene of individual isolates was amplified as above, and aliquots of the purified PCR products (10 μl) were digested separately with MspI and HhaI (2.5 U; Fermentas, York, United Kingdom) at 37 °C for 2 h, following the manufacturer’s instructions. 16S-RFLP were resolved on 2 % (w/v) agarose gels and grouped into operational taxonomic units (OTUs), based on the restriction pattern obtained. Representative clones for OTUs of interest were sequenced with the 27F primer.
16S rRNA sequence affiliation was performed using the NAST alignment tool (DeSantis et al. 2006b) and the Classify tool (http://greengenes.lbl.gov/cgi-bin/nph-classify.cgi) from the online ribosomal RNA database Greengenes (DeSantis et al. 2006a). Phylogenetic analysis was performed in the MEGA 6.06 software package (Tamura et al. 2013), using the maximum likelihood method, and Bayesian estimation of the best-fitting model of molecular evolution. Nodal robustness of the tree was assessed by bootstrapping (1000 replicates). All of the sequences described here have been submitted to the EMBL database under accession numbers LN812266-LN812290.
β-propeller phytase (BPP) gene PCR amplification and sequencing
The BPP sequences reported in a previous analysis (Lim et al. 2007) were used for the design of phytase-specific PCR primers. The protein BPP sequences were aligned using the multiple alignment CLUSTALW algorithm implemented in the BioEdit Sequence Alignment Editor software (http://www.mbio.ncsu.edu/BioEdit/BioEdit.html). Alignments were performed with BPP sequences from groups I, II, IIIa, IIIb, and IIIc defined in Lim et al. (2007). Phytase-specific primers were designed using the Consensus Degenerate Hybrid Oligonucleotide Primers software [CODEHOP; (Rose et al. 2003)] as follows. Highly conserved regions (blocks) within the BPP alignments were identified using the BLOCKS multiple alignment processor and the generated blocks were applied to the CODEHOP algorithm to design several sets of degenerate primers (see Table 3 for details).
Three different protocols of PCR amplification were tested. In all protocols, the PCR mixture (50 μl) contained 1× reaction buffer, 1.5 mM MgCl2, 0.5 μM of each primer, 50 μM of each dNTP, 2.5 U of Taq DNA polymerase (BIOTAQ). Dimethyl sulfoxide (DMSO) was added at 10 % and 5 % in PCR mixture containing phytate-hydrolysing isolate DNA and rhizosphere DNA, respectively. For isolate DNA, two different one-step PCR protocols were tested. The first, using PhyblockR-f/PhyblockW-r primers, consisted in an initial denaturation step at 94 °C for 3 min, 30 cycles with 30 s at 94 °C, 30 s at 60 °C, 30 s elongation at 72 °C, and a final elongation step for 5 min at 72 °C. The second, using PhyblockC-f/PhyblockW-r primers, was based on a touchdown PCR protocol, as follows: initial denaturation step at 94 °C for 3 min, 10 cycles with 1 min at 94 °C, 1 min at 65 °C (reducing by 0.5 °C/cycle), 1 min elongation at 72 °C and a further 30 cycles with a fixed annealing temperature at 60 °C. For rhizosphere DNA, a 2-step nested PCR protocol was used, based on a touchdown PCR for each step, as follows: initial denaturation step at 94 °C for 3 min, 10 cycles with 1 min at 94 °C, 1 min at 65 °C (reducing by 0.5 °C/cycle), 1 min elongation at 72 °C and a further 30 cycles with a fixed annealing temperature at 60 °C. Primers PhyblockC-f/PhyblockW-r were used for the first step and primers PhyblockR-f/PhyblockW-r for the second.
PCR products were purified either with a QIAquick PCR purification column (QIAGEN) or with a QIAquick Gel Extraction Kit (QIAGEN), following the manufacturer’s instructions. PCR products were quantified with a Nanodrop ND100 spectrophotometer. Cloning of purified PCR products was done with the pGEM-T easy vector system (System I) (Promega Ltd UK, Southampton, United Kingdom) according to the manufacturer’s instructions. For rhizosphere, 96 clone inserts were subjected to RFLP with MspI and AluI (2.5 U; Fermentas, York, United Kingdom) at 37 °C for 2 h. BPP-RFLP were resolved on 2 % (w/v) agarose gels and grouped into operational taxonomic units (OTUs), based on the restriction pattern obtained. Representative clones for OTUs were sequenced with the T7 primer.
Phylogenetic analyses of BPP encoded peptide sequences were performed using the MEGA 6.06 software package (Tamura et al. 2013), as described above. All of the sequences described here have been submitted to the EMBL database under accession numbers (LN812291-LN812320).
Results
Effect of phytate input on rhizobacterial community structure
Because agricultural practices constitute an important source of phytate input in soil, which may affect soil functioning, the impact of such input on soil-inhabiting microorganisms was evaluated in rhizospheres of Lolium perenne growing in two low P soils (Table 1).
The bacterial community structure in the treated and untreated rhizospheres was examined by 16S-DGGE of total bacterial communities and of individual phyla and classes (α-, β-, γ-Proteobacteria, Actinobacteria and Firmicutes) that have been suggested to play a role in phytate cycling in soils (Idriss et al. 2002; Lim et al. 2007; Jorquera et al. 2008a). PCA of 16S-DGGE profiles showed a clear difference between the bacterial populations of rhizospheres derived from the two soils studied (along the first axis), both in the overall bacterial community and in the group-specific populations (Fig. 1). The effect of phytate treatment was less pronounced and mostly affected bacterial communities in ‘Warwick’ soil. Indeed, significant changes (P < 0.05) were only observed in ‘Warwick’ soil, primarily affecting the total bacterial community (Fig. 1a) and the γ-Proteobacteria community (Fig. 1d) structures, and to a lesser extent α-Proteobacteria and Actinobacteria (Fig. 1b and f). The γ-Proteobacteria in ‘Warwick’ soil were the bacterial population most affected by the phytate treatment. The β-Proteobacteria and Firmicutes (Fig. 1c and e) were also significantly affected by the addition of phytate in both soils, with greatest effects seen in ‘Warwick’ and ‘Lindow’ soils, respectively.
Principal component analysis of 16S-DGGE profiles for bacterial communities obtained from L. perenne rhizospheres. The profiles shown are total bacteria (a) and group-specific populations, α-Proteobacteria (b), β-Proteobacteria (c), γ-Proteobacteria (d), Firmicutes (e) and Actinobacteria (f), and were generated with group-specific primers, as detailed in Material and methods. For each condition, the mean and standard deviation of DGGE profiles (N = 4) are represented. Lindow and Warwick soils are indicated by red and green symbols, respectively. Untreated and treated samples are indicated by triangles and circles, respectively. The statistics (P < 0.05) are indicated along axes PC1 (by letters A and B) and PC2 (by letters a and b)
Effect of phytate input on cultivable phytate-hydrolysing rhizobacterial community
In order to isolate and characterize the bacteria involved in rhizosphere phytate cycling, rhizobacteria were extracted from L. perenne rhizosphere. Phytate treatment had a significant effect on the total phytate-hydrolysing bacterial count only for the ‘Warwick’ soil (Table 4). Although the differences were significant (P < 0.05), the fold-change on enrichment was small, and while phytate hydrolysers were indeed enriched, phytate treatment led to a decrease in the number of phytate P and C utilizers compared to control rhizospheres.
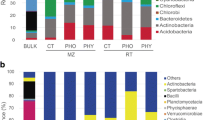
16S-RFLP analysis was performed for 217 morphologically diverse isolates, allowing their classification into 65 OTUs whose abundance and diversity in the two rhizosphere soils was clearly influenced by phytate treatment (Fig. S1). The dominant groups of PMI and PMII isolates obtained from ‘Lindow’ rhizospheres without phytate supplementation were OTU2 and OTU1, respectively (Fig. 1a and b). After phytate supplementation, the dominant groups were OTU5 (PMI) and OTU2 (PMII). In unsupplemented ‘Warwick’ rhizospheres (Fig. 1c and d), OTU2 was dominant on PMI and PMII media, whereas phytate treatment lead to an increase in the relative abundance of OTU30 in both cases, as well as of OTU5 on PMII. For the two types of rhizospheres, OTU abundance and the diversity of less common OTUs changed due to phytate treatment, and several OTUs represented by one or two isolates were obtained only in one of the two conditions. Among the phytate C and P utilizers (PMI isolates), 32 % were specific to phytate-supplemented condition in ‘Lindow’ rhizospheres and 18 % in ‘Warwick’ rhizospheres.
Assessment of phytate-hydrolysing ability of rhizobacteria and taxonomic identification
Thirty-five representatives of 30 OTUs were tested for their ability to hydrolyse phytate. These OTUs represented the most dominant OTUs or those most strongly affected by phytate supplementation (Fig. S1; solid and open triangles). Table 5 summarizes the results for the 24 isolates for which phytate disappearance was reliably determined by HPLC. All belonged to the Proteobacteria (α, β, γ) and the Actinobacteria (Fig. 2).
Phylogenetic relationship of phytate-hydrolysing bacteria isolated from L. perenne rhizosphere. Phytate-hydrolysing bacteria isolated in this study are indicated in bold. The soil from which each isolate was obtained is indicated as L (Lindow soil) or W (Warwick soil), respectively. Numbers I and II indicate on which phytate-containing medium the isolate was obtained, PMI and PMII, respectively, and the phytate treatment used for the plant growth experiments, is shown as (+) or (−). For each isolate, the corresponding OTU is indicated. The colour code represents the ability of phytate-hydrolysing isolates to use phytate as P and C sources (green) or only as P source (blue). The phylogenetic tree was constructed using the maximum likelihood method and the Kimura-two parameter with a discrete gamma distribution for distance correction. Levels of bootstrap value (1000 resamplings) are indicated by black circles (if >80 %) or open circles (if between 50 and 80 %). The scale bar shows the number of base changes per sequence position
In order to specify the metabolism of phytate by the rhizobacterial isolates, and to characterize their ability to degrade phytate, bacterial growth experiments were performed in liquid MM either supplemented with phytate as C and P sources or supplemented with phytate and inositol as P and C sources, respectively. Eighteen isolates were able to use phytate only as a P source, while six isolates could use phytate as C and P source (Table 5). Among the isolates using phytate only as P source, the use of inositol as C source by ten of them may indicate that their inability to assimilate phytate C for growth is linked to an inability to completely dephosphorylate phytate. For the others, it seems they did not have the enzymatic ability to degrade inositol.
BPP-specific rhizobacterial diversity
New primer sets were designed to explore BPP diversity of well known soil bacteria because of limitations observed in previous studies (Jorquera et al. 2011, 2012, 2014). The wide sequence diversity within the BPP class made primer design difficult, but three degenerate BPP-specific primers were designed by aligning the BPP sequences belonging to groups I, II, IIIb and IIIc genes, as defined in Lim et al. (2007), and using the CODEHOP software to predict suitable primers (Table 3). PhyblockC-f primer was obtained from alignment of BPP sequences belonging to groups I, II, IIIa, IIIb and IIIc, and PhyblockR-f/PhyblockW-r primers from IIIb and IIIc groups. Conserved regions targeted by the primers are shown in Fig. S2. Group III phytases (largely affiliated with γ-Proteobacteria) were more specifically targeted because of the known role of γ-Proteobacteria in L. perenne rhizosphere (Marilley and Aragno 1999).
Non-specific amplification was initially observed for isolates and rhizosphere DNA, and considerable optimization was required to remedy this. Two Pseudomonas strains were used as controls: Pseudomonas syringae DC3000, which harbours a BPP gene (Lim et al. 2007) and Pseudomonas protegens CHA0, a well-known soil-borne bacterium. The latter strain has been extensively studied for its importance in plant-microbial interactions (Haas and Defago 2005) and is closely related to Pseudomonas protegens Pf-5, which also harbours a BPP gene (Lim et al. 2007). Positive BPP amplifications were obtained for the two Pseudomonas strains using the two one-step PCR protocols (described in the methods) and the presence of BPP gene for strain CHA0 was confirmed by sequencing.
The rhizobacterial isolates displaying phytase activity were tested for the presence of BPP genes using the new BPP primers (Table 3). Positive BPP amplifications were obtained for isolates of Pseudomonas putida, Variovorax sp., Sphingomonas sp. and Acinetobacter sp. Interestingly, many of the isolates did not yield an amplification product, suggesting that they harbour non-BPP genes or BPP genes that are unrelated to those targeted by the primers. In addition, whereas PhyblockC-f targeted a conserved region in BPP groups I to IIIc, the two other regions targeted were less conserved (Fig. S2).
BPP sequences were retrieved for representatives of these OTUs and phylogenetic analysis of the encoded BPP protein sequences was performed (Fig. 3), revealing an affiliation to BPP groups II and IIIc [described in Lim et al. (2007)]. As expected, the BPP sequence from P. protegens CHA0 was closely related to that of Pf-5, and surprisingly also to the BPP sequence of a Variovorax isolate. BPP sequences from P. putida isolates formed a separate cluster. The BPP sequence from the Acinetobacter isolate, also belonged to a distinct cluster formed by BPP sequences from Acinetobacter johnsonii SH046, Pseudomonas mendocina YMP and Pseudomonas stutzeri A1501. BPP sequences from Sphingomonas isolate appeared more related to Caulobacter sequences than to those from other Sphingomonadaceae strains.
Phylogenetic analysis of partial BPP amino acid sequences from potential phytate-hydrolysing isolates and L. perenne rhizosphere (supplemented with phytate). The phylogenetic tree was calculated with the maximum-likelihood method using WAG models with discrete gamma distribution and invariable site parameter for distance correction. Levels of bootstrap value (500 resamplings) are indicated by black circles (if >80 %) or open circles (if between 50 and 80 %). The scale bar shows the number of amino acid changes per sequence position. BPP sequences obtained in this study from phytate-hydrolysing isolates, Pseudomonas protegens CHA0 and directly from the rhizosphere sample (WP1 to WP25) are in bold type. The soil from which each isolate was obtained is indicated as L (Lindow soil) or W (Warwick soil), respectively. Numbers I and II indicate on which phytate-containing medium the isolate was obtained, PMI and PMII, respectively, and the phytate treatment used for the plant growth experiments, is shown as (+) or (−). For each isolate, the corresponding OTU is indicated. For rhizosphere sample, BPP sequences sharing more than 90 % identity were clustered (comma separated list). The BPP sequences used as reference correspond to the ones detailed in Fig. S2. The numbers (I to IIIc) next to the strain names indicate BPP groups as defined in Lim et al. (2007)
The BPP-specific primers were also used to assess BPP diversity in rhizosphere samples. A BPP amplicon was obtained from Lolium rhizospheres grown in ‘Warwick soil’ supplemented with phytate. Twenty-five percent of the clones were excluded from further analysis because they contained a considerably larger insert than expected for the BPP fragment amplified. Twenty-three OTUs were determined and 1 to 4 inserts per OTU were sequenced. Overall, 25 sequences corresponding to BPP genes were obtained and phylogenetic analysis of the encoded BPP protein sequences revealed two main clusters affiliated to BPP group IIIc and, to a lesser extent, to group IIIb (Fig. 3).
Discussion
Phytate represents a major source of phosphate in soils but is poorly available for the plant. Due to the intensification of agricultural practices and the formation of insoluble phytate complexes, its accumulation in soils may lead to significant environmental problems (Lei and Stahl 2001; Vats et al. 2005). Soil microorganisms are known to hydrolyse phytate (Hill and Richardson 2006) but their functional and taxonomic characterization remains a challenge that must be addressed in order to understand their role in phytate cycling in soil environments. The increased number of sequenced bacterial genomes has provided new information concerning the potential ability of microorganisms to hydrolyse phytic acid (Lim et al. 2007) but there is still a need to develop molecular approaches to unravel the complexity of phytate-hydrolysing bacterial community in soils and particularly in the rhizosphere. Molecular approaches to BPP-diversity in the rhizosphere have revealed a relatively low functional diversity until now (Jorquera et al. 2011, 2012, 2014) and in addition, all studies were culture-dependent and were not directly targeted at soils.
The importance of rhizobacteria as plant growth-promoting organisms (PGPR), has been extensively documented (Lugtenberg and Kamilova 2009; Vacheron et al. 2013). Nevertheless, their potential role in plant nutrition through phytate degradation has not been studied in depth (Richardson et al. 2009). PGPR containing BPP genes have been characterized (Jorquera et al. 2012, 2014) but the link between plant growth-promoting properties and phytate degradation remains unclear (Jorquera et al. 2012). PGPR effect due to phytase activity has been clearly observed for few strains and only in defined media or in plants genetically modified with microbial phytase (Richardson and Simpson 2011). Phytate supplementation of the two low-P soils studied here revealed changes of rhizobacterial structure at community and population levels, with the strongest effect on bacterial community structure in ‘Warwick’ soil, mainly on γ-Proteobacteria. Only β-Proteobacteria and Firmicutes were significantly affected in both soils. Interestingly, the particular status of “Warwick soil” was also emphasized by significant changes in phytate-hydrolysing bacterial abundance. The effect on Proteobacteria was confirmed by identification of Pseudomonas sp., Variovorax sp. and Rhizobiaceae as the main cultivable phytate-hydrolysing taxa affected by phytate supplementation. Phytate input has previously been reported to cause major changes in total community structure of pasture soils and an increase in Bacillus-related BPP (Jorquera et al. 2013). In the current study, the differential response of rhizobacterial community between the two soils could not be explained by differences in soil phosphate content, but may be due to the chemical characteristics of the soils. Indeed, the difference measured in magnesium content between the two soils may affect phytate availability because of insoluble complex formation between phytate and Mg2+ (Martin and Evans 1987), since phytate availability is strongly affected by a wide range of cations, including Mg2+, Mn2+, Fe2+, Ca2+, Al3+ and Fe3+ (Martin and Evans 1987). The diversity of these insoluble complexes might explain the rhizobacterial OTU-dependent responses after phytate enrichment. Indeed, it has been shown that bacterial phytate degradation is dependent on both the type of phytate complex and on the specific bacterial isolate (Unno et al. 2005). In addition, the ‘Warwick’ soil is an agricultural soil with a long cultivation history, and consequently more exposed to large amounts of phytate in seeds than the commercial ‘Lindow’ topsoil, suggesting that its bacterial population may be more adapted to phytate utilisation. Differences in nutrient input but also in soil management are major factors affecting bacterial community and its biological activity (Bissett et al. 2013). The recent development of methods based on a chromophore tethered phytic acid probe is an important advance in the field, which will allow the comparison of rhizobacterial phytate degradation in undisturbed ecosystems and agro-ecosystems (Berry et al. 2009).
The ability to dephosphorylate phytate was confirmed for a subset of rhizobacterial isolates belonging to the Proteobacteria and Actinobacteria. Apart from isolates belonging to the Pseudomonas genus, most of these have not previously been described for their ability to hydrolyse phytate or for harbouring a phytase gene sequence. The ability to hydrolyse phytate therefore appears to be far more widely distributed than previously recognized. The importance of Pseudomonas species as phytate hydrolysers in the L. perenne rhizosphere has been reported previously (Marilley and Aragno 1999; Jorquera et al. 2008a) and phytate-hydrolysing Pseudomonas isolates have been reported in various environments (Richardson and Hadobas 1997; In et al. 2004; Cho et al. 2005; Hill et al. 2007; Jorquera et al. 2008a). A range of other phytate-hydrolysing rhizosphere bacterial isolates, including Acinetobacter, Agrobacterium and Arthrobacter, have also been reported, but taxonomic identification was based purely on morphological and phenotypic characteristics, and has not been confirmed at a genetic level (Ahmed 1976; Barea et al. 1976). Nevertheless, evidence for the potential role of Arthrobacter sp. and Agrobacterium sp. in phytate cycling has been found recently in soil and marine environments (Unno et al. 2005; Hill et al. 2007) as well as in the genome sequence of Arthrobacter chlorophenolicus A6. The predominance of Rhizobiaceae in the phytate-hydrolysing bacterial community is of particular interest because of their significance in plant–bacteria interactions. The potential role of rhizobia in phytate cycling was suggested by Lopez-Lopez et al. (2010) but the method used to demonstrate the rhizobial ability to hydrolyse phytate was not conclusive (Hill and Richardson 2006). Unknown and poorly-described phytate-hydrolysing bacteria (Variovorax sp. and Acinetobacter sp.) are well-known for their ability to degrade a broad range of organic and inorganic compounds (Abdel-El-Haleem 2003; Barbe et al. 2004; Sorensen et al. 2008; Uhlik et al. 2009), and more recently for their ecological role in plant nutrition and plant growth (Schmalenberger et al. 2008; Belimov et al. 2009; Peix et al. 2009). The lack of phytase activity detection for a range of strains may be due to a certain strain-dependent phytate specificity (type of phytate complex, stereoisomeric forms) (Adelt et al. 2003; Unno et al. 2005), but also to our experimental conditions (pH, P content) (Konietzny and Greiner 2004; Fu et al. 2008). Indeed, whereas a certain P inorganic content can inhibit phytase synthesis, a minimum level could be necessary to reach a bacterial growth stage suitable for the production of phytase.
The present investigation has revealed the ability of many phytate-hydrolysing isolates to use phytate not only as P source, but also as C source (Agrobacterium sp., Ensifer sp., Variovorax sp., and Acinetobacter sp.). In addition, the bacterial growth of other isolates only when inositol and phytate were provided as C and P sources, respectively, suggests the inability of these isolates to completely dephosphorylate phytate. Whereas phytate could constitute a major proportion of soil organic P (Lim et al. 2007), inositol is one of the most abundant carbohydrates in terrestrial ecosystems (Turner et al. 2002). Consequently, the determination of bacterial isolates able to metabolise phytate and/or inositol constitutes an important step in understanding the significance of phytate-hydrolysing bacteria in plant status, and more broadly in the carbon and phosphorus cycles. This ability to use inositol or derivatives as carbon sources is widely studied in rhizobia because of its importance for plant colonization and in nodulation processes (Jiang et al. 2001; Fry et al. 2001), but remains poorly studied in other plant-associated bacteria.
In the present study, the metabolic potential of these isolates to hydrolyse phytic acid was supported by the amplification of BPP gene in their genomes. BPP genes belonging to group II and IIIc (Lim et al. 2007) were amplified from Pseudomonas putida, Sphingomonas sp., Variovorax sp. and Acinetobacter sp. isolates. Interestingly, phylogenetic analysis of BPP genes suggested potential horizontal BPP transfer among rhizobacteria (Variovorax isolate containing Pseudomonas-related BPP). The rhizosphere has been suggested to be a privileged microbial compartment for horizontal gene transfer due to the exudates and high microbial density and competition (Berg et al. 2005; Molbak et al. 2007), and the hypothesis that BPP genes are subject to HGT is also strengthened by the isolation of a rhizosphere Pseudomonas isolate containing Bacillus-related BPP (Jorquera et al. 2012). Evidence for the BPP gene in the Acinetobacter genus was found recently through the genome sequencing project of A. johnsonii SH046 (RefSeq assembly accession, GCF_000162055.1) and A. lwoffii WJ10621 (GCF_000219275.1). The non-amplification of the BPP gene from a number of phytate-hydrolysing isolates described in this study may revealed an uncovered BPP diversity by the BPP-specific primers, or the presence of the other classes of phytase. Similar conclusions were put forward by Jorquera et al. (2014) to explain their difficulty in accessing phytase genes from a wide diversity of phytate-hydrolysing isolates, since even when BPP-specific primers were used (Huang et al. 2009), only a few sequences affiliated to Bacillus were retrieved. Non-BPP are known to be present in Acidovorax and Mycobacterium, for example, which harbour a cysteine phytase (CPhy) and a purple acid phosphatase (PAP), respectively (Lim et al. 2007; Jorquera et al. 2008b). Similarly, only BPP genes belonging to group IIIb and IIIc were retrieved from L. perenne rhizosphere. A part of this problem is probably also due to the fact that the BPP-specific PCR primers were originally designed to target BPP sequences belonging to groups I to III, with a special focus on group III, excluding the other groups because their low levels of sequence identity made primers design difficult. The limitations in the molecular tools available to assess environmental phytase diversity mean that extensive development is still needed, to allow us to target the whole diversity of the different classes of phytases, and better characterize the biological potential of soils in phytate cycling.
References
Abdel-El-Haleem D (2003) Acinetobacter: environmental and biotechnological applications. Afr J Biotechnol 2:71–74
Adelt S, Podeschwa M, Dallmann G, Altenbach H-J, Vogel G (2003) Stereo- and regiospecificity of yeast phytases–chemical synthesis and enzymatic conversion of the substrate analogues neo- and l-chiro-inositol hexakisphosphate. Bioorg Chem 31:44–67
Ahmed D (1976) Mineralization of inositol hexaphosphate in soil at varying static moisture levels. Plant Soil 44:253–256
Barbe V, Vallenet D, Fonknechten N et al (2004) Unique features revealed by the genome sequence of Acinetobacter sp. ADP1, a versatile and naturally transformation competent bacterium. Nucleic Acids Res 32:5766–5779
Barea JM, Navarro E, Montoya E (1976) Production of plant growth regulators by rhizosphere phosphate-solubilizing bacteria. J Appl Bacteriol 40:129–134
Belimov AA, Dodd IC, Hontzeas N, Theobald JC, Safronova VI, Davies WJ (2009) Rhizosphere bacteria containing 1-aminocyclopropane-1-carboxylate deaminase increase yield of plants grown in drying soil via both local and systemic hormone signalling. New Phytol 181:413–423
Berg G, Eberl L, Hartmann A (2005) The rhizosphere as a reservoir for opportunistic human pathogenic bacteria. Environ Microbiol 7:1673–1685
Berry DF, Shang C, Zelazny LW (2009) Measurment of phytase activity in soil using a chromophoric tethered phytic acid probe. Soil Biol Biochem 41:192–200
Bissett A, Richardson AE, Baker G, Kirkegaard J, Thrall PH (2013) Bacterial community response to tillage and nutrient additions in a long-term wheat cropping experiment. Soil Biol Biochem 58:281–292
Cheng C, Lim BL (2006) Beta-propeller phytases in the aquatic environment. Arch Microbiol 185:1–13
Cho J, Lee C, Kang S et al (2005) Molecular cloning of a phytase gene (phy M) from Pseudomonas syringae MOK1. Curr Microbiol 51:11–15
Cunliffe M, Kertesz MA (2006) Effect of Sphingobium yanoikuyae B1 inoculation on bacterial community dynamics and polycyclic aromatic hydrocarbon degradation in aged and freshly PAH-contaminated soils. Environ Pollut 144:228–237
Dao TH (2007) Ligand effects on inositol phosphate solubility and bioavailability in animal manures. In: Turner BL, Richardson AE, Mullaney EJ (eds) Inositol phosphates: linking agriculture and environment. CAB International, Oxford, pp 169–185
DeSantis TZ, Hugenholtz P, Larsen N et al (2006a) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
DeSantis TZ, Hugenholtz P, Keller K et al (2006b) NAST: a multiple sequence alignment server for comparative analysis of 16S rRNA genes. Nucleic Acids Res 34:W394–W399
Fry J, Wood M, Poole PS (2001) Investigation of myo-inositol catabolism in Rhizobium leguminosarum bv. viciae and its effect on nodulation competitiveness. Mol Plant Microbe Interact 14:1016–1025
Fu S, Sun J, Qian L, Li Z (2008) Bacillus phytases: present scenario and future perspectives. Appl Biochem Biotechnol 151:1–8
George TS, Richardson AE, Simpson RJ (2005) Behaviour of plant-derived extracellular phytase upon addition to soil. Soil Biol Biochem 37:977–988
Giles CD, Richardson AE, Druschel GK, Hill JE (2012) Organic anion-driven solubilization of precipitated and sorbed phytate improves hydrolysis by phytases and bioavailability to Nicotiana tabacum. Soil Sci 177:591–598
Greiner R, Lim BL, Cheng C, Carlsson NG (2007) Pathway of phytate dephosphorylation by beta-propeller phytases of different origins. Can J Microbiol 53:488–495
Haas D, Defago D (2005) Biological control of soil-borne pathogens by fluorescent pseudomonads. Nat Rev Microbiol 3:307–319
Hammond JP, Broadley MR, White PJ (2004) Genetic responses to phosphorus deficiency. Ann Bot (Lond) 94:323–332
Hayes JE, Simpson RJ, Richardson AE (2000a) The growth and phosphorus utilisation of plants in sterile media when supplied with inositol hexaphosphate, glucose 1-phosphate or inorganic phosphate. Plant Soil 220:165–174
Hayes JE, Richardson AE, Simpson RJ (2000b) Components of organic phosphorus in soil extracts that are hydrolysed by phytase and acid phosphatase. Biol Fertil Soils 32:279–286
Hill JE, Richardson AE (2006) Isolation and assessment of microorganisms that utilize phytate. In: Turner BL, Richardson AE, Mullaney EJ (eds) Inositol phosphates: linking agriculture and environment. CAB International, Oxford
Hill JE, Kysela D, Elimelech M (2007) Isolation and assessment of phytate-hydrolysing bacteria from the DelMarVa Peninsula. Environ Microbiol 9:3100–3107
Hinsinger P (2001) Bioavailability of soil inorganic P in the rhizosphere as affected by root-induced chemical changes: a review. Plant Soil 237:173–195
Huang H, Shi P, Wang Y et al (2009) Diversity of beta-propeller phytase genes in the intestinal contents of grass carp provides insight into the release of major phosphorus from phytate in nature. Appl Environ Microbiol 75:1508–1516
Huang H, Zhang R, Fu D et al (2011) Diversity, abundance and characterization of ruminal cysteine phytases suggest their important role in phytate degradation. Environ Microbiol 13:747–757
Idriss EE, Makarewicz O, Farouk A et al (2002) Extracellular phytase activity of Bacillus amyloliquefaciens FZB45 contributes to its plant-growth-promoting effect. Microbiology 148:2097–2109
In MJ, Jang ES, Kim YJ, Oh NS (2004) Purification and properties of an extracellular acid phytase from Pseudomonas fragi Y9451. J Microbiol Biotechnol 14:1004–1008
Jiang G, Krishnan AH, Kim YW, Wacek TJ, Krishnan HB (2001) A functional myo-inositol dehydrogenase gene is required for efficient nitrogen fixation and competitiveness of Sinorhizobium fredii USDA191 to nodulate soybean (Glycine max [L.] Merr.). J Bacteriol 183:2595–2604
Jorquera MA, Hernandez MT, Rengel Z, Marschner P, de la Luz Mora M (2008a) Isolation of culturable phosphobacteria with both phytate-mineralization and phosphate-solubilization activity from the rhizosphere of plants grown in a volcanic soil. Biol Fertil Soils 44:1025–1034
Jorquera M, Martinez O, Maruyama F, Marschner P, de la Luz Mora M (2008b) Current and future biotechnological applications of bacterial phytases and phytase-producing bacteria. Microbes Environ 23:182–191
Jorquera MA, Crowley DE, Marschner P, Greiner R, Fernandez MT, Romero D, Menezes-Blackburn D, de la Luz Mora M (2011) Identification of β-propeller phytase-encoding genes in culturable Paenibacillus and Bacillus spp. from the rhizosphere of pasture plants on volcanic soils. FEMS Microbiol Ecol 75:163–172
Jorquera MA, Shaharoona B, Nadeem SM, de la Luz Mora M, Crowley DE (2012) Plant growth-promoting rhizobacteria associated with ancient clones of creosote bush (Larrea tridentata). Microb Ecol 64:1008–1017
Jorquera MA, Saavedra N, Maruyama F, Richardson AE, Crowley DE, Catrilaf RC, Henriquez EJ, de la Luz Mora M (2013) Phytate addition to soil induces changes in the abundance and expression of Bacillus β-propeller phytase genes in the rhizosphere. FEMS Microbiol Ecol 83:352–360
Jorquera MA, Inostroza NG, Lorena ML, Barra PJ, Marileo LG, Rilling JI, Campos DC, Crowley DE, Richardson AE, Mora ML (2014) Bacterial community structure and detection of putative plant growth-promoting rhizobacteria associated with plants grown in Chilean agro-ecosystems and undisturbed ecosystems. Boil Fertil Soils 50:1141–1153
Kerovuo J, Lauraeus M, Nurminen P, Kalkkinen N, Apajalahti J (1998) Isolation, characterization, molecular gene cloning, and sequencing of a novel phytase from Bacillus subtilis. Appl Environ Microbiol 64:2079–2085
Kertesz MA, Leisinger T, Cook AM (1993) Proteins induced by sulfate limitation in Escherichia coli, Pseudomonas putida, or Staphylococcus aureus. J Bacteriol 175:1187–1190
King J, Thorogood D, Edwards KJ, Armstead IP et al (2008) Development of a genomic microsatellite library in perennial ryegrass (Lolium perenne) and its use in trait mapping. Ann Bot (Lond) 101:845–853
Konietzny U, Greiner R (2004) Bacterial phytase: potential application, in vivo function and regulation of its synthesis. Braz J Microbiol 35:11–18
Lambers H, Shane MW, Cramer MD, Pearse SJ, Veneklaas EJ (2006) Root structure and functioning for efficient acquisition of phosphorus: matching morphological and physiological traits. Ann Bot 98:693–713
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–147
Lee SH, Malone C, Kemp PF (1993) Use of 16S ribosomal-RNA-targeted fluorescent-probes to increase signal strength and measure cellular RNA from natural planktonic bacteria. Mar Ecol Prog Ser 101:193–201
Lei XG, Stahl CH (2001) Biotechnological development of effective phytases for mineral nutrition and environmental protection. Appl Microbiol Biotechnol 57:474–481
Li M, Osaki M, Rao IM, Tadano T (1997) Secretion of phytase from the roots of several plant species under phosphorus-deficient conditions. Plant Soil 195:161–169
Lim BL, Yeung P, Cheng C, Hill JE (2007) Distribution and diversity of phytate-mineralizing bacteria. ISME J 1:321–330
Lopez-Lopez A, Rogel MA, Ormeno-Orrillo E, Martinez-Romero J, Martinez-Romero E (2010) Phaseolus vulgaris seed-borne endophytic community with novel bacterial species such as Rhizobium endophyticum sp. nov. Syst Appl Microbiol 33:322–327
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizobacteria. Annu Rev Microbiol 63:541–556
Marilley L, Aragno M (1999) Phylogenetic diversity of bacterial communities differing in degree of proximity of Lolium perenne and Trifolium repens roots. Appl Soil Ecol 13:127–136
Martin CJ, Evans WJ (1987) Phytic acid: divalent cation interactions. V. titrimetric, calorimetric, and binding studies with cobalt(ii) and nickel(ii) and their comparison with ions. J Inorg Biochem 30:101–119
Martin M, Celi L, Barberis E (2004) Desorption and plant availability of myo-inositol hexaphosphate adsorbed on goethite. Soil Sci 169:115–124
Molbak L, Molin S, Kroer N (2007) Root growth and exudates production define the frequency of horizontal plasmid transfer in the rhizosphere: plasmid transfer in the rhizosphere. FEMS Microbiol Ecol 59:167–176
Muhling M, Woolven-Allen J, Murrell JC, Joint I (2008) Improved group-specific PCR primers for denaturing gradient gel electrophoresis analysis of the genetic diversity of complex microbial communities. ISME J 2:379–392
Mullaney EJ, Ullah AH (2003) The term phytase comprises several different classes of enzymes. Biochem Biophys Res Commun 312:179–184
Mullaney EJ, Daly CB, Ullah AH (2000) Advances in phytase research. Adv Appl Microbiol 47:157–199
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nakashima BA, McAllister TA, Sharma R, Selinger LB (2007) Diversity of phytases in the rumen. Microb Ecol 53:82–88
Peix A, Lang E, Verbarg S, Sproer C, Rivas R, Santa-Regina I, Mateos PF, Martinez-Molina E, Rodriguez-Barrueco C, Velazquez E (2009) Acinetobacter strains IH9 and OCI1, two rhizospheric phosphate solubilizing isolates able to promote plant growth, constitute a new genomovar of Acinetobacter calcoaceticus. Syst Appl Microbiol 32:334–341
Raghothama K, Karthikeyan A (2005) Phosphate acquisition. Plant Soil 274:37–49
Reasoner DJ, Geldreich EE (1985) A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microbiol 49:1–7
Richardson AE, Hadobas PA (1997) Soil isolates of Pseudomonas spp. that utilize inositol phosphates. Can J Microbiol 43:509–516
Richardson AE, Simpson RJ (2011) Soil microorganisms mediating phosphorus availability update on microbial phosphorus. Plant Physiol 156:989–996
Richardson AE, Hadobas PA, Hayes JE (2001a) Extracellular secretion of Aspergillus phytase from Arabidopsis roots enables plants to obtain phosphorus from phytate. Plant J 25:641–649
Richardson AE, Hadobas PA, Hayes JE, O’hara CP, Simpson RJ (2001b) Utilization of phosphorus by pasture plants supplied with myo-inositol hexaphosphate is enhanced by the presence of soil micro-organisms. Plant Soil 229:47–56
Richardson AE, Barea JM, McNeill AM, Prigent-Combaret C (2009) Acquisition of phosphorus and nitrogen in the rhizosphere and plant growth promotion by microorganisms. Plant Soil 321:305–339
Rose TM, Henikoff JG, Henikoff S (2003) CODEHOP (COnsensus-DEgenerate Hybrid Oligonucleotide Primer) PCR primer design. Nucleic Acids Res 31:3763–3766
Sanguin H, Sarniguet A, Gazengel K, Moënne-Loccoz Y, Grundmann GL (2009) Rhizosphere bacterial communities associated with disease suppressiveness stages of take-all decline in wheat monoculture. New Phytol 184:694–707
Schmalenberger A, Hodge S, Bryant A, Hawkesford MJ, Singh BK, Kertesz MA (2008) The role of Variovorax and other Comamonadaceae in sulfur transformations by microbial wheat rhizosphere communities exposed to different sulfur fertilization regimes. Environ Microbiol 10:1486–1500
Sharma NC, Sahi SV (2005) Characterization of phosphate accumulation in Lolium multiflorum for remediation of phosphorus-enriched soils. Environ Sci Technol 39:5475–5480
Shin S, Ha NC, Oh BC, Oh TK, Oh BH (2001) Enzyme mechanism and catalytic property of beta propeller phytase. Structure 9:851–858
Sorensen SR, Albers CN, Aamand J (2008) Rapid mineralization of the phenylurea herbicide diuron by Variovorax sp. strain SRS16 in pure culture and within a two-member consortium. Appl Environ Microbiol 74:2332–2340
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6. Mol Biol Evol 30:2725–2729
Turner BL, Leytem AB (2004) Phosphorus compounds in sequential extracts of animal manures: chemical speciation and a novel fractionation procedure. Environ Sci Technol 38:6101–6108
Turner BL, Paphazy MJ, Haygarth PM, McKelvie ID (2002) Inositol phosphates in the environment. Philos Trans R Soc Lond B Biol Sci 357:449–469
Uhlik O, Jecna K, Mackova M, Vlcek C, Hroudova M, Demnerova K, Paces V, Macek T (2009) Biphenyl-metabolizing bacteria in the rhizosphere of horseradish and bulk soil contaminated by polychlorinated biphenyls as revealed by stable isotope probing. Appl Environ Microbiol 75:6471–6477
Unno Y, Okubo K, Wasaki J, Shinano T, Osaki M (2005) Plant growth promotion abilities and microscale bacterial dynamics in the rhizosphere of Lupin analysed by phytate utilization ability. Environ Microbiol 7:396–404
Vacheron J, Desbrosses G, Bouffaud ML, Touraine B, Moënne-Loccoz Y, Muller D, Legendre L, Wisniewski-Dyé F, Prigent-Combaret C (2013) Plant growth-promoting rhizobacteria and root system functioning. Front Plant Sci 4:356
Vats P, Bhattacharyya MS, Banerjee UC (2005) Use of phytases (myo-inositolhexakisphosphate phosphohydrolases) for combatting environmental pollution: a biological approach. Crit Rev Environ Sci Technol 35:469–486
Yanke LJ, Bae HD, Selinger LB, Cheng KJ (1998) Phytase activity of anaerobic ruminal bacteria. Microbiology 144:1565–1573
Zimmermann P, Zardi G, Lehmann M, Zeder C, Amrhein N, Frossard E, Bucher M (2003) Engineering the root-soil interface via targeted expression of a synthetic phytase gene in trichoblasts. Plant Biotechnol J 1:353–360
Zysko A, Sanguin H, Hayes A, Wardleworth L, Zeef LAH, Sim A, Paterson E, Singh BK, Kertesz MA (2012) Transcriptional response of Pseudomonas aeruginosa to a phosphate-deficient Lolium perenne rhizosphere. Plant Soil 359:25–44
Acknowledgments
We thank John Hammond and Gary Bending for assistance with soil sampling at Warwick HRI. This project was supported by the Natural Environment Research Council (NERC) and by a research grant from the University of Sydney. We thank John Morton for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Benjamin L. Turner.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Distribution of phytate-hydrolysing bacteria (OTUs) from L. perenne rhizosphere. Number of bacterial isolates (for each OTU) obtained on PMI (A and C) and PMII (B and D) with (black) or without (white) phytate supplemented is represented for Lindow soil (A and B) and Warwick soil (C and D). The symbols (▲) and (△) indicate the OTUs for which one or two isolates were selected to test their ability to utilize phytate as C source and/or P source. Solid and open triangles indicate the soil treatment, with or without phytate supplemented, respectively. (PPT 225 kb)
Fig. S2
Multiple sequence alignment of amino acids BPP sequences used for the design of phytase-specific PCR primers. The conserved sequence motifs targeted are highlighted in colours and the primer position is indicated at the bottom. The sequences used for the design of each primer are in enclosed boxes. The numbers (I to IIIc) next to the strain names indicate BPP groups defined in Lim et al. (2007). BAP, Bacillus amyloliquefaciens BAP (AAW28542); ATCC14580, Bacillus licheniformis ATCC14580 (YP_090097) ; DS11, Bacillus sp. DS11 (AAC38573); 168, Bacillus subtilis subsp. subtilis 168 (NP_389861); ATCC23134, Microscilla marina ATCC23134 (EAY24393); CB15, Caulobacter crescentus CB15 (AAK23276); RB2256, Sphingomonas alaskensis RB2256 (ABF54827.1); SKA58, Sphingomonas sp. SKA58 (EAT09404); RW1, Sphingomonas wittichii RW1 (YP_001261037); HTCC2633, Oceanicaulis alexandrii HTCC2633 (ZP_00953252); MCS10, Maricaulis maris MCS10 (ABI66660); ATCC15444, Hyphomonas neptunium ATCC15444 (ABI78101); HTCC2503, Parvularcula bermudensis HTCC2503 (EAQ17779); Deepecotype, Alteromonas macleodii ‘Deepecotype’ (YP_002126691); MED297, Reinekea sp. MED297 (EAR10111); S14, Vibrio angustum S14 (EAS63574); HTCC2207, marine gamma proteobacterium HTCC2207 (EAS47070); 2–40, Saccharophagus degradans 2–40 (ABD83254); TAC125, Pseudoalteromonas haloplanktis TAC125 (CAI85536); ANA-3, Shewanella sp. ANA-3 (ABK48437); MR-4, Shewanella sp. MR-4 (ABI39156); MR-7, Shewanella sp. MR-7 (ABI42881); MR-1, Shewanella oneidensis MR-1 (AAN55555); RED65, Oceanobacter sp. RED65 (EAT12633); OS145, Idiomarina baltica OS145 (EAQ32794); L2TR, Idiomarina loihiensis L2TR (AAV80945); Pf-5, Pseudomonas fluorescens Pf-5 (YP_260816); 1448A , Pseudomonas syringae pv. Phaseolicola 1448A (AAZ36343); DC3000, Pseudomonas syringae pv. Tomato DC3000 (AAO56720); ymp, Pseudomonas mendocina ymp (YP_001189542); DJ, Azotobacter vinelandii DJ (YP_002797726). (PPTX 114 kb)
Rights and permissions
About this article
Cite this article
Sanguin, H., Wilson, N.L. & Kertesz, M.A. Assessment of functional diversity and structure of phytate-hydrolysing bacterial community in Lolium perenne rhizosphere. Plant Soil 401, 151–167 (2016). https://doi.org/10.1007/s11104-015-2512-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-015-2512-7