Abstract
Prolonged exposure to abiotic stresses causes oxidative stress, which affects plant development and survival. In this research, the overexpression of ZmARF1 improved tolerance to low Pi, drought and salinity stresses. The transgenic plants manifested tolerance to low Pi by their superior root phenotypic traits: root length, root tips, root surface area, and root volume, compared to wide-type (WT) plants. Moreover, the transgenic plants exhibited higher root and leaf Pi content and upregulated the high affinity Pi transporters PHT1;2 and phosphorus starvation inducing (PSI) genes PHO2 and PHR1 under low Pi conditions. Transgenic Arabidopsis displayed tolerance to drought and salt stress by maintaining higher chlorophyll content and chlorophyll fluorescence, lower water loss rates, and ion leakage, which contributed to the survival of overexpression lines compared to the WT. Transcriptome profiling identified a peroxidase gene, POX, whose transcript was upregulated by these abiotic stresses. Furthermore, we confirmed that ZmARF1 bound to the auxin response element (AuxRE) in the promoter of POX and enhanced its transcription to mediate tolerance to oxidative stress imposed by low Pi, drought and salt stress in the transgenic seedlings. These results demonstrate that ZmARF1 has significant potential for improving the tolerance of crops to multiple abiotic stresses.
Key message
ZmARF1 is a pleiotropic gene which confers tolerance to multiple abiotic stresses in ZmARF1-OE transgenic Arabidopsis as revealed by stress-response phenotypes of the transgenic lines relative to WT. ZmARF1 confers stress tolerance through a mechanism involving its association with an antioxidant enzyme, AtPOX, whose expression was simultaneously up regulated by low Pi, drought and salt stresses, whiles its promoter activity was significantly activated by ZmARF1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants, as sessile organisms, are predominantly exposed to numerous abiotic stresses during their growth cycles, such as extreme temperatures, nutrient deficiencies, drought, salinity, and metal toxicity, which pose a serious threat on global food security (Dixit et al. 2018). Abiotic stresses induce major constraints on crop productivity and have become one of the leading limitations to maximizing crop yields globally, causing up to an average crop yield loss of up to 50% (Pandey et al. 2017). The limitations imposed by environmental stresses on crop productivity are expected to worsen because of changes in global climate. The effect of global climate change are projected to decrease global yields of major staple crops such as maize, rice, wheat, and soybean by 3–7% per °C rise in temperature (Zhao et al. 2017). Moreover, with the global population projected to reach 10 billion by 2050 (Rockstrom et al. 2017), competition for arable land for non-agricultural activities, will push agricultural production into environments less suited for crop cultivation (Bajaj & Mohanty 2005; Miao et al. 2016). Consequently, the development of stress-tolerant crops has become imperative to sustaining yield stability under adverse environmental conditions to minimize the impact of environmental stress on crop production (Zhang et al. 2018).
Drought stress, or water deficit, is one of the major abiotic stresses that have significantly limited crop survival, development, and yields worldwide over the last few decades (Wang et al. 2019). An estimated 1,820 million Mg of cereal (maize, rice, and wheat) production has been lost globally due to drought stress alone over the past four decades (Lesk et al. 2016). Drought stress is projected to impose major limitations on the productivity of more than half of the world’s arable lands in the next 50 years (Dhankher & Foyer 2018). Both drought and salinity stresses decrease the availability of water to plant cells by inducing lower water potential, which inhibits overall crop yield and quality. Salt stress causes ion imbalance and disrupts nutrient acquisition due to osmotic stress and Na+ toxicity, consequently inhibiting the normal physiological functioning of plants (Nan et al. 2018). Salt stress also stimulates swift the production and accumulation of reactive oxygen species (ROS) such as superoxides (O2−) and hydrogen peroxide (H2O2), as secondary stress responses, which are extremely toxic to plants (Mittler 2017; Yang & Guo 2018). Approximately 20% of global arable lands are affected by salt stress, a figure projected to increase in the immediate future due to climate change and anthropogenic activities (Shrivastava & Kumar 2015). Additionally, phosphate (Pi) is a key constituent of important cellular molecules such as ATP, nucleic acids, and phospholipids, and participates in numerous biochemical pathways such as energy transmission, gene expression and signal transduction. Pi deficiency limits plant growth and development, and remains one of the main challenges affecting crop productivity (Chen et al. 2018). An estimated 70% of global arable lands are reported to contain suboptimal Pi concentration, which restricts optimal plant growth and development (Herrera-Estrella & Lopez-Arredondo 2016). The reduction in crop yield and productivity occasioned by these abiotic stresses could be mitigated by developing broad-spectrum abiotic stress-tolerant plants that can yield better on lands affected by these stress conditions. However, the mechanisms of gene–gene networks governing tolerance to multiple abiotic stresses are complex and not well understood.
The auxin response factor (ARF) family proteins are a group of highly conserved plant specific transcription factors that play key regulatory roles in the expression of auxin-responsive genes (Liscum & Reed 2002). To date, numerous ARFs have been described in various plant species, such as Arabidopsis thaliana (Guilfoyle & Hagen 2007), Oryza sativa (Wang et al. 2007), Zea mays (Wang et al. 2012) and Musa acuminata (Hu et al. 2015). Besides mediating several aspects of plant growth and development, ARF proteins have been reported to participate in plant response to multiple abiotic stresses (Bouzroud et al. 2020, 2018; Chen et al. 2019; Song et al. 2023). The molecular mechanism by which auxin controls gene expression occurs either through canonical or non-canonical signaling pathways. The canonical pathway, also called the nuclear auxin pathway (NAP), involves three families of proteins: the TRANSPORT INHIBITOR RESPONSE1/AUXIN SIGNALING F-BOX (TIR1/AFB) family, which acts as the auxin receptor, AUXIN/INDOLE 3-ACETIC ACID INDUCIBLE (Aux/IAA) repressor proteins, and ARF transcription factors (Kong et al. 2016; Wright & Nemhauser 2015). When auxin is low or absent, Aux/IAA recruits the transcription co-repressor TOPLESS/TOPLESS-RELATED (TPL/TPR) and interacts with ARF transcription factors through the shared PB1 domains, thus repressing gene transcription and auxin responses (Szemenyei et al. 2008). In the presence of auxin, auxin binds TIR1/AFBs and increases their affinity toward Aux/IAA transcriptional repressors to facilitate their proteasome-dependent degradation by targeting them for ubiquitination, thereby relieving the repression of ARF transcription factors to regulate the expression of auxin-responsive genes (Dharmasiri et al. 2005; Kepinski & Leyser 2005). The non-canonical auxin pathway can occur through two distinct routes. The first pathway involves the receptor-like kinase TRANSMEMBRANE KINASE 1 (TMK1), which interacts with AUXIN BINDING PROTEIN 1 (ABP1) and activates Rho-like guanosine triphosphatases (GTPase) from plants (ROP2/ROP6) in an auxin-dependent manner (Xu et al. 2014). Moreover, after auxin sensing, TMK1 phosphorylates and stabilizes IAA32 and IAA34, thus regulating ARF transcription factors (Cao et al. 2019). Alternatively, auxin binding to the atypical ARF3/ETTIN(ETT) modulates chromatin and interacts with other transcriptional regulators (Simonini et al. 2016). Direct auxin binding to ARF3 leads to its release from co-repressors of the TOPLESS/TOPLESS-RELATED family, resulting in histone acetylation and induction of gene expression, creating a reversible switch between repressive and de-repressive chromatin states (Kuhn et al. 2020; Liu et al., 2024). Constitutive expression of ARF genes in various plant species modulates tolerance to environmental stresses such as drought, cold, salinity and low Pi stress (Jain & Khurana 2009; Shen et al. 2013). For instance, overexpression of a sweet potato auxin response factor (IbARF5) modulated carotenoid biosynthesis and conferred salt and drought tolerance in transgenic Arabidopsis (Kang et al. 2018). Overexpression of ZmARF4 conferred low Pi stress tolerance in Arabidopsis by enhancing lateral root development, Pi mobilization and expression of Pi responsive gene AtANS1 (Li et al. 2022).
Under the natural growing environment of plants, they encounter two or more stresses that affect their survival and limit their productivity. Researchers have outlined different approaches to achieving tolerance to multiple stresses in plants. It has been outlined that tolerance to multiple stresses can be achieved in plants by the gene pyramid approach that involves concurrent expression of multiple genes (Hema et al. 2014; Hong et al. 2016). In this study, we demonstrated that overexpression of ZmARF1 conferred tolerance to low Pi, drought, and salt stresses in transgenic Arabidopsis compared to wild-type (WT). Low Pi tolerance in ZmARF1-transgenic Arabidopsis was manifested by better root phenotypic traits, higher Pi mobilization and upregulation of Pi stress inducible genes under low Pi conditions. Meanwhile, transgenic Arabidopsis manifested tolerance to drought and salt stress by maintaining lower ion leakage and water loss rates, higher chlorophyll parameters and survival rates compared to WT respectively. Transcriptome profiling identified POX, a member of the peroxidase superfamily, whose transcript level was upregulated under low Pi, drought, and salt stresses. Furthermore, ZmARF1 bound to the AuxRE in the promoter of POX and enhanced its transcription, and this interaction is believed to have contributed to conferring tolerance to oxidative stress imposed by low Pi, drought and salt stress in the transgenic seedlings.
Materials and methods
Plant material and stress treatment
The Arabidopsis thaliana WT plants were of the Columbia-0 (Col-0) ecotype. Seeds of the WT and ZmARF1 overexpressing lines were surface sterilized with 75% (v/v) ethanol and 5% (v/v) NaOCl followed by five times rinse with sterilized distilled water as outlined by Hu et al. (2023). Seeds were subsequently placed on P-sufficient (HP) 1/2 MS medium and cold treated at 4 °C for 2 days. The 1/2 MS medium plates were then transferred to a growth chamber and kept upright at 22 °C and a 16 h light/8 h dark photoperiod for 7 days. Uniform seedlings were then transferred to P-sufficient (HP) and P-deficient (LP) 1/2 MS medium and cultured for 7 days. HP condition was made up of 1/2 MS medium (Phytotech, Cat#M519) with 5% (w/v) sucrose and 0.6% (w/v) agar powder (Solarbio, Cat#A8190). LP condition was made up of 1/2 MS without phosphate (Caisson Lab. REF: MSP11-10LT) with 5% (w/v) sucrose and 0.6% (w/v) agar powder, supplemented with HP (100:1). Root phenotypic traits, Pi accumulation and expression of Pi stress inducible genes were determined after HP and LP treatments.
For salt stress treatment, WT and transgenic seeds germinated on 1/2 MS medium were transplanted to pots containing equal volumes of peat soil and grown in growth chamber for 5 weeks at 22 °C and a 16 h light/8 h dark condition. Salt stress was imposed on seedlings by irrigating with 200 mM NaCl solution at 3 days interval for two weeks. Control treatment was made up of irrigation with water. For natural drought treatment, WT and transgenic seeds germinated on 1/2 MS medium were transplanted to pots containing equal volumes of peat soil and grown in growth chamber for 3 weeks at 22 °C and a 16 h light/8 h dark condition. Pots were fully watered for 3 weeks and then water was withdrawn for 10 days for drought treatment, whiles control plants were continuously watered. After drought treatment, the seedlings were rehydrated for 3 days. Moreover, rosette leaves under drought and salt stress, alongside the respective control treatments, were sampled for physiological measurements. The phenotypic changes and survival rates among the WT and transgenic lines under drought stress and normal conditions were recorded.
Plasmid construction and plant transformation
The full coding sequence (CDS) of ZmARF1 (without the stop codon) was amplified by PCR from the maize P178 inbred line and cloned into the BamH I and Spe I sites of the pCAMBIA2300 expression vector, driven by the cauliflower mosaic virus (CaMV) 35S promoter, with a GFP tag at the C terminus. Gene-specific primers used are stated in Table S1. The pCAMBIA2300-ZmARF1-GFP construct was confirmed by sequencing and transformed into Arabidopsis (Col-0) ecotype by the Agrobacterium tumefaciens (strain GV3101) mediated transformation using the floral dip method (Clough & Bent 1998). T0 seeds were harvested and screened on 1/2 MS medium supplemented with 50 μg ml−1 of G418, resistant seedlings were transplanted into peat soil for selfing. T1 seeds were harvested and screened on 1/2 MS medium containing 50 μg ml−1 of G418 antibiotic for separation ratio identification. Approximately, 100 T3 seeds of independent transgenic lines and WT were screened on half MS strength medium supplemented with 50 μg ml−1 of G418, homozygosity was accessed based on 100% germination, root growth and survival. T3 homozygous positive transgenic lines overexpressing ZmARF1 were selected for subsequent experiments.
RNA extraction and RT-qPCR
Total RNA was extracted from seedlings using the Trizol kit (Invitrogen, CA, USA) according to the manufacturer’s protocol. According to the manufacturer’s instructions of reagent kit for operation of mRNA reverse transcription (Vazyme Biotech, Nanjing, China) and qRT-PCR (TransGen, Beijing, China), respectively. The expression of the housekeeping AtActin was used as an internal control. qRT-PCR was performed in three technical replicates. Specific primers used are listed in Table S1.
Expression pattern of ZmARF1 in response to Pi stress
Responsiveness of ZmARF1 to Pi stress was evaluated through RT-qPCR and western blotting assay. RT-qPCR was performed as described above. Total protein was extracted from the wild-type (Col-0) and ZmARF1 overexpression seedlings (OE8# and OE17#) after high and low Pi treatments using the HEPES cell lysis buffer (1 M HEPES, 2 M KCl, 500 mM EDTA, 20% Triton X-100,1 mM PMSF). Seedlings were ground into fine powder in liquid nitrogen and approximately 0.05 g was homogenized in 100 μl of HEPES cell lysis buffer. The protein extract (10 μl) was separated by gel electrophoresis on 10% SDS–polyacrylamide gel and transferred to a polyvinylidene difluoride membrane (Bio-rad). The membranes were subjected to immunoblotting with a primary anti-GFP antibody (Sangon) overnight and a secondary anti-rabbit antibody (Sangon) for 2 h. Proteins were visualized using an enhanced chemiluminescence kit (Thermo Fisher Scientific).
Measurement of root phenotypic traits
The root traits of WT and ZmARF1-overexpresing seedlings were captured by a scanner (Epson expression 10000XL, Epson Corporation, Japan) and analyzed with WinRHIZO software (WinRHIZO Pro 2008a) after HP and LP treatments. The root traits, including root length, root surface area, root volume and root tips were measured. Means were represented in bar graphs using the GraphPad Prism 8.0.1v software.
Measurement of Pi content
Phosphate content was determined according to previous study (Chiou et al. 2006) with slight modification. Fresh samples were ground into fine powder in liquid nitrogen. Approximately 0.1 g of powdered sample was dissolved in 400 μl of extraction buffer [10 mM Tris–HCl, 1 mM EDTA, 10 mM NaCl, 1 mM Beta-ME, and 1 mM PMSF, pH 8.0] and kept at 25 °C for 30 min. Approximately 100 μl from the previous step and homogenized in 900 μl of 1% acetic acid and centrifuged at 13,000 rpm for 5 min. Then 300 μl of the supernatant was separated into a new EP tube, and 700 μl of analysis buffer [0.35% (NH4)2MoO4, 2.34% H2SO4, and 1.4% VC] was added and incubated in a water bath at 42 °C for 30 min. Pi content was determined by measuring the absorbance of A820.
Calculation of water loss rate
WT and transgenic plants were grown under well-watered condition for 3 weeks. Healthy and uniform seedlings with intact rosette leaves were sampled from each genotype with sharp scissors. The rosette leaves were placed on dry filter papers in a light incubator with constant temperature and humidity and allowed to dehydrate naturally under room temperature. The leaves were weighed at 1, 2, 3, 4, 6, 8 and 10 h after separation, respectively, and the rate of water loss from the excised leaves at each sampling period was obtained by expressing the weights as a fraction of the fresh weight prior to dehydration and expressing it as a percentage.
Imaging and determination of chlorophyll fluorescence
Chlorophyll fluorescence was measured from rosette leaves of WT and ZmARF1 transgenic Arabidopsis under normal, drought or salt stress conditions. Rosette leaves were excised with sharp scissors and were immediately subjected to 20 min of dark acclimation. Initial fluorescence (F0), maximum fluorescence (Fm) and variable fluorescence (Fv) were determined with the FluorCam800MF system (Photon Systems Instruments, Ecotech). The experiment was conducted in triplicate, with three biological repeats each time. Chlorophyll fluorescence was calculated by the following formula.
Measurement of ion leakage
For membrane ion leakage measurement, five leaves disks (7 mm in diameter) of each treatment were picked from the fifth rosette leaf of the WT and transgenic seedlings under normal, drought and salt stress treatments. The leaf discs were collected in a 10 ml EP tube with deionized water for surface cleaning. Liquid was replaced with fresh 1 ml of deionized water and oscillated for 10 s, and the initial electrical conductivity (C1) was measured with an electro-conductivity meter. Samples were then placed in a vacuum pump for 30 min at a pressure of 0.8 to 1 MPa. After the vacuum treatment, the electrical conductivity was measured before (C2) and after (C3) boiling for 15 min with an electro-conductivity meter again. Total ion leakage (C) was determined by the following formula and expressed as a percentage.
RNA-seq analysis
Two-week old seedlings of WT and ZmARF1 transgenic Arabidopsis were transplanted in liquid medium with 1/2 MS (normal condition as control), low Pi, salinity (200 mM NaCl), and drought (200 mM d-mannitol) stress conditions, respectively. After 3 days treatment, seedlings were collected and prepared in three biological replicates for RNA-seq analysis. Total RNA extraction, library construction, transcriptome sequencing, transcriptome assembly, and functional annotation were performed by OE Biotech (Shanghai, China). High-quality reads were aligned against the Arabidopsis thaliana reference genome sequence (TAIR10 Genome Release) using HISAT2. Differentially expressed genes (DEGs) were identified using DEseq software with corrected p-value < 0.05. GO functional classification enrichment was performed using the online tools (http://geneontology.org/). GO terms with q < 0.05 were considered to be significantly enriched. The Integrative Genomics Viewer (IGV) (http://software.broadinstitute.org/software/igv) was used to visualize the reads of the selected genes.
Electrophoretic mobility shift assay (EMSA)
The fusion construct of full-length CDS of ZmARF1 fused with MBP and an empty MBP plasmid were used to transform Escherichia coli strain DE3 (TransGen, Beijing, China) and purified using amylose resin (NEB, E8021S#) as previously described (Wu et al. 2024). We synthesized 5ʹ biotin labeled probes of the auxin response element (AuxRE: TGTCTC) motif, together with unlabeled and mutated competitor (Sangon Biotech, Shanghai). EMSA was performed using the MBP-ZmARF1 recombinant protein, a biotin-labeled AuxRE probe, an unlabeled AuxRE, and its mutated competitor. EMSA was performed using a LightShift Chemiluminescent EMSA Kit (Thermo Fisher Scientific, Prod# 89880) following the manufacturer’s instructions, with slight modifications as previously described (Wu et al. 2024). Briefly, ZmARF1 protein and the biotin-labeled probe, with or without the unlabeled WT or mutant unlabeled probes, were incubated in 10 μl reaction volume at 25 °C for 30 min. The reaction mixtures were separated on 7% polyacrylamide gel and transferred to a nylon membrane. Signals were visualized using an imaging system (E-blot, Shanghai, China). All probes used are listed in Table S1.
Dual-luciferase assay
The fusion construct of full-length CDS of ZmARF1 fused with GFP in pGreenII62SK was used as effector. A ~ 2 kb promoter region of AtPOX was amplified and cloned into pGreenII0800-LUC to obtain ProAtPOX::LUC reporter construct. The reporter construct was co-expressed with effector either ZmARF1-GFP or pGreenII62SK-GFP as control in N. benthamiana leaves, as previously described (Hellens et al. 2005). The luciferase activity assay was performed using a dual luciferase reporter assay kit (Vazyme, Nanjing, China) according to the manufacturer’s protocol. Relative LUC activity was calculated as the ratio of LUC to Ren activity. Each sample was tested in triplicate. All primers used are listed in Table S1.
Results
Overexpression of ZmARF1 confers tolerance to low Pi stress via the regulation of root development
We had previously conducted a gene-based association analysis in 356 maize inbred lines and identified that ZmARF1 is associated with Pi stress tolerance in maize (Wu et al. 2024). To determine the potential role of ZmARF1 in low Pi stress response, we generated and selected three independent homozygous T3 ZmARF1-overexpressing lines with a high level of transgene expression (Fig. S1). For phenotypic detection, 7-day-old seedlings grown in 1/2 MS were transferred to either HP (high phosphate, 1/2 MS) or LP (low phosphate) conditions for 7 days (Fig. 1a, b). Ensuing results revealed that overexpression of ZmARF1 promotes root development (Fig. 1a, c–f), especially increasing root densities near the soil surface (Fig. 1a). Generally, low Pi stress reduced root development in all genotypes, but the ZmARF1 overexpressing transgenic plants recorded significantly (P < 0.05) increased root phenotypic traits, including root length, surface area, root volume and root tips compared to the WT (Fig. 1b–f). The results were in accordance with the theory that superior root morphology enhances phosphorus acquisition and promotes low Pi stress tolerance (Strock et al. 2018), suggesting that ZmARF1 confers low Pi stress tolerance by promoting root architecture reconstruction.
Overexpression of ZmARF1 promoted root development and conferred low Pi tolerance. Seven-day-old T3 transgenic and WT plants were transferred to a High Pi and b Low Pi 1/2 MS medium. Root phenotypes including c Root length, d Root surface area, e Root volume, and f Root tips, of WT and transgenic seedlings under high Pi and low Pi conditions were detected. Three biological repeats were performed and eight plants of each genotype were used for each biological repeat. Means are presented as bar graphs and error bars represent SEM. Two OE8# and OE17# represent the two transgenic lines overexpressing ZmARF1 and WT represents the wild-type
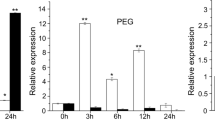
To further evaluate the responsiveness of ZmARF1 to low Pi stress, RT-qPCR and western blotting assays were performed to determine the transcript and protein abundance in ZmARF1-transgenic seedlings under HP and LP conditions, respectively. The results showed that transcript levels of ZmARF1 in the transgenic lines showed an undiscernible difference between the HP and LP conditions (Fig. 2a). On the contrary, protein abundance of ZmARF1 was significantly increased in the transgenic plants after LP treatment compared to HP (Fig. 2b). In addition to the initial comparison, a specific transgenic line (OE8#) with relatively lower ZmARF1 protein abundance was selected and used to carefully assess the accumulation dynamic of ZmARF1 protein following low Pi treatment. The results showed that ZmARF1 protein steadily accumulated after low Pi treatment and continued to increase throughout the entire day of treatment (Fig. 2c). This pattern of continuous increase in protein levels suggests that ZmARF1 protein abundance is directly induced by Pi starvation and that the response is time-dependent. The disparity between transcript levels and protein abundance of ZmARF1 due to low Pi treatment suggests that ZmARF1 might be exclusively regulated at the post-transcriptional level in response to low Pi stress. Moreover, protein abundance of ZmARF1 stabilized with MG132 treatment compared to the mock treatment (Fig. 2d), suggesting that ZmARF1 was regulated by the 26S proteasome mechanism and that addition of the proteasome inhibitor MG132 protected it from ubiquitination. This further indicates that ZmARF1 could potentially be exposed to some level of post-translational regulation.
ZmARF1 responded to low-Pi stress in overexpressed Arabidopsis. Seven-day-old T3 transgenic and WT seedlings were transferred to high Pi (HP) and low Pi medium (LP) at varying treatment times. a The transcript level of ZmARF1 in transgenic overexpression lines OE8# and OE17# and WT under HP and LP conditions. Data represent means ± SEM. b ZmARF1 protein levels increased under LP compared to HP conditions. c ZmARF1 protein abundance is directly induced by Pi starvation and that the response is time-dependent. d Addition of the proteasome inhibitor MG132 mitigated ubiquitination of ZmARF1 and stabilized its protein accumulation under high Pi condition. OE8# and OE17# represent the two transgenic lines overexpressing ZmARF1 and WT represents the wild-type
ZmARF1 positively modulates Pi homeostasis by regulating the expression of PSI genes
Plants require sufficient phosphorus for optimal growth and development. Phenotypes analysis revealed that the transgenic plants adapted better to low Pi stress compared to the WT (Fig. 1). We hypothesized that the transgenic plants might be better at mobilizing, translocating, and utilizing Pi. Therefore, we quantified the Pi concentration in roots and leaves of the transgenic plants and WT. Interestingly, there was no difference in the leaf Pi concentration among all the genotypes, and the root Pi concentration was significantly (P < 0.05) higher in WT than in transgenic lines under HP condition (Fig. 3a). However, under LP condition, the transgenic lines had significantly higher Pi concentration compared to WT in both leaf and root tissues (Fig. 3b). Low Pi stress resulted in a dramatic decrease in Pi concentration, especially in leaves. The Pi concentration ratio under LP to HP in the root was significantly higher than in leaves, with 2.74, 1.96 and 1.25 folds increases in OE17#, OE8# and WT, respectively (Fig. 3c). Since root system architecture (RSA) is critical for Pi uptake, the lower Pi concentration in transgenic roots and the indistinguishable measure in leaves could herald a more efficient phosphorus transport than the wild type under HP. Under LP conditions, the transgenic lines could maintain higher phosphorus uptake and recycling efficiency compared to WT to sustain growth. This may also be the reason why the transgenic lines manifested favorable RSA-related traits (Fig. 1b). These results suggest that ZmARF1 plays a regulatory role in Pi homeostasis under low Pi stress, probably by influencing the mobilization and translocation of phosphates from source (roots) to sink (shoot).
Overexpression of ZmARF1 enhances phosphorus starvation inducing gene expression in Arabidopsis. Quantification of inorganic phosphate content under High Pi (HP, a) and Low Pi condition (LP, b). c Fold change in Pi content due to Pi stress imposition. Inorganic phosphate quantification was determined after 7 days of low Pi treatment. d–f Differential expression of Pi stress inducible genes, d PHR1, e PHT1;2, f PHO2 under high and low Pi conditions. Three biological repeats were used; night plants of each genotype were used for each biological repeat. Significance difference was tested at P < 0.05. Means are presented as bar graphs and error bars represent SEM
To verify this hypothesis, we further investigated the expression of phosphorus starvation inducing (PSI) genes such as PHR1, PHT1;2, and PHO2 in different genotypes after high and low Pi treatments by RT-qPCR. Under HP conditions, the transcript abundance of the three target genes showed no noted differences between transgenic plants and WT (Fig. 3d–f). The expression levels of PHR1 and PHT1;2 even trended to decrease in two overexpressing lines (Fig. 3d, e). However, after LP stress treatment, all the detected PSI genes were dramatically upregulated, and the mRNA levels of PHR1, PHT1;2, and PHO2 in the transgenic plants were significantly higher than in WT (Fig. 3d–f). These results suggest that ZmARF1 modulates Pi homeostasis via the regulation of PSI genes expression, downregulating their transcription under sufficient phosphorus conditions, and upregulated when phosphorus is insufficient. The reduced expression of PSIs under HP conditions might be due to the superior RSA of transgenic plants, which increases the phosphorus absorption surface area. In contrast, the upregulated expression of PHR1, PHT1;2, and PHO2 under LP induction helps promote phosphorus absorption and transport from roots to shoots.
Overexpression of ZmARF1 confers tolerance to drought stress in transgenic Arabidopsis
Since ZmARF1 promotes root development under normal conditions, we hypothesized that ZmARF1 might be involved in other abiotic stress response processes, especially dehydration stress which is related to root function. Both drought and salinity initially result in osmotic stress, directly affecting the acquisition of water resources. To investigate the potential role of ZmARF1 in drought stress regulation, 3-week-old seedlings of the transgenic plants and WT were subjected to dehydration for 10 days. There was no discernible difference in the aboveground phenotype between the WT and transgenic plants prior to drought treatment (Fig. 4a). However, after drought treatment, the transgenic plants displayed strong tolerance to dehydration, while the WT showed susceptibility, as manifested by wilting of rosette leaves (Fig. 4a). Moreover, virtually all the WT plants could not recover after 3 days of rehydration, with survival rates below 15% (Fig. 4b). In contrast, the transgenic plants swiftly recovered after rehydration, with OE8# and OE17# recording more than 90% recovery rates, while OE22# recorded over 85% recovery rates (Fig. 4b). Analysis of the chlorophyll fluorescence, as indicated by Fv/Fm values showed no significant difference between the WT and transgenic plants under normal well-watered conditions (Fig. 4c). The Fv/Fm values of the WT plants decreased from ~ 0.8 under normal conditions to ~ 0.5 after drought treatment and rehydration. By contrast, Fv/Fm values of the transgenic lines remained greater than or equal to 0.79, which is indicative of healthy plants (Fig. 4c). The role of ZmARF1 in drought tolerance response was further examined by transferring the WT and transgenic plants to dry filter papers and allowing them to dehydrate at room temperature (Fig. 4d). After 10 h of dehydration, the rosette leaves of the WT plants were almost completely wilted, while the transgenic plants manifested a more stretched leaf morphology. After 12 h rehydration, the leaves of the transgenic lines showed be little or no wilting, while the damage to the WT plants appeared to be irreversible (Fig. 4d). Consistent with the drought stress response phenotype, the water loss and ion leakage rates of the WT plants were significantly higher than those of the transgenic plants (Fig. 4e, f). This suggests that overexpression of ZmARF1 confers tolerance to drought stress in transgenic Arabidopsis.
ZmARF1 enhances tolerance to drought stress. a Image of WT and transgenic plants under different water treatments. Normal cultivation of 3-week-old plants subjected to consecutive 10 days of drought treatment (DDT) and then rehydrated for 3 days. b Survival rate of WT and transgenic plants after 3 days of rehydration. c Chlorophyll fluorescence (Fv/Fm) of WT and transgenic plants measured under normal and dehydrated conditions. d The 3-week-old plants were placed on dry filter paper (upper panel), dehydrated for 10 h (middle panel), and then rehydrated for 12 h (lower panel). e Water loss rate among the WT and transgenic plants after natural dehydration. f Ion leakage measured from leaves of WT and transgenic plants under normal and dehydration conditions. These experiments were replicated three times with similar results. Student’s t test P value, *P < 0.05; **P < 0.01; ***P < 0.001; ns, not significantly different
ZmARF1 is a positive regulator of salt stress response in Arabidopsis
To elucidate the function of ZmARF1 in regulating salt stress tolerance, 5-week-old WT and ZmARF1 overexpressing plants were irrigated with either normal water (Fig. 5a) or 200 mM NaCl solution (Fig. 5b) for two weeks and evaluated for salt stress tolerance phenotypes. There were no discernible differences in the phenotypic and physiological indices between the ZmARF1 transgenic plants and WT under normal growth condition (Fig. 5). On the contrary, salt stress impaired multiple aspects of plant growth, resulting in significant reduction in plant height, leaf color, flowering time, and overall plant vitality (Fig. 5a–d). Despite this, the transgenic plants overexpressing ZmARF1 displayed stronger tolerance, with significantly increased plant height (Fig. 5c) and chlorophyll content (Fig. 5d) compared to WT under salt stress. Consistently, the chlorophyll fluorescence (Fv/Fm) of the WT plants was about 2.5 folds lower under salt stress treatment compared to the transgenic plants (Fig. 5f, g). Moreover, salt stress caused approximately a fold higher ion leakage in the WT plants compared to the transgenic lines (Fig. 5e). Taken together, overexpression of ZmARF1 contribute to salt stress tolerance by reducing ionic stress and preventing chlorophyll degradation.
ZmARF1 transgenic Arabidopsis displayed enhanced tolerance to salt stress compared to WT plants. WT and transgenic plants were grown in growth chamber for five weeks and irrigated with a Water or b 200 mM NaCl solution for two weeks. c Plant height, d chlorophyll content, e ion leakage, f and g chlorophyll fluorescence (Fv/Fm) of WT and transgenic plants measured under normal and salt stress conditions. These experiments were replicated three times with similar results. Student’s t test P values, ***P < 0.001; ns, not significantly different
Transcription profiling revealed that POX is a downstream target of ZmARF1
To investigate the mechanism governing the role of ZmARF1 in regulating various abiotic stresses, transgenic plants overexpressing ZmARF1 were subjected to multiple stresses: low Pi, salt (200 mM NaCl) and osmotic (200 mM d-Mannitol) stresses. RNA-seq analysis showed that different stress treatments induced specific transcriptomic response profiles. There were 674/210, 1540/513, and 4067/3364 genes identified as specifically upregulated/downregulated by osmotic stress, low Pi stress, and salt stress, respectively (Fig. 6a). Each treatment had significantly more upregulated genes than downregulated genes (Figs. 6a and S2a–f), suggesting a positive role of ZmARF1 in enhancing stress tolerance in Arabidopsis. Notably, the number of gene expression changes induced by salt stress was significantly higher compared to the changes caused by low Pi and osmotic stress (Fig. 6a, b). This suggests that ZmARF1 plays a more dominant role in the response to salt stress compared to its involvement in low Pi and osmotic stress responses. Gene ontology enrichment analysis revealed an array of signaling-mediated biological processes under the respective stress treatments: response to oxidative stress, response to salt stress, response to osmotic stress under salt stress, root morphogenesis, response to osmotic stress and root development under low Pi stress, and root morphogenesis, response to oxidative stress and response to water deprivation under osmotic stress (Fig. S3).
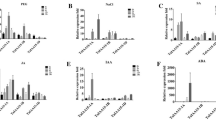
Transcriptome profiling revealed commonly induced DEGs under multiple abiotic stresses. a Differentially expressed gene analysis showing up- and down- regulated genes across the respective abiotic stress treatments. The transgenic and WT seedlings were treated to salt (200 mM NaCl), low Pi and osmotic stress (200 mM mannitol) and harvested for RNA-seq analysis. b Venn diagram analysis revealing upregulated DEGs expressed between the WT and transgenic lines under three different abiotic stresses. c,d Hierarchical clustering of DEGs from transgenic and WT seedlings under the three abiotic stresses and normal condition. Rows represent individual upregulated DEGs, and columns represent respective treatments. e Visualization of RNA-seq coverage profile for POX, as revealed by the integrated genome viewer browser, illustrated the transcript levels of POX in transgenic and WT seedlings under the three abiotic stress treatments. Scale represents normalized read counts. f RT-qPCR was used to confirm the transcript levels of POX (AT4G30170) between transgenic and WT seedlings under the three abiotic stress treatments. Actin was used as internal control
Moreover, we identified 18/3 commonly upregulated /downregulated genes among the three abiotic stresses (Figs. 6b, c and S2g–j). Among these commonly upregulated DEGs, a peroxidase gene POX (AT4G30170) showed significant induction under all three stress conditions, with the highest relative expression level observed under salt stress compared to low Pi and d-Mannitol (salt > low Pi > d-Mannitol > normal) (Fig. 6d). Additionally, the IGV visualizated the differential transcript levels using the RNA-seq data (Fig. 6e), and the upregulated expression of POX under salt, low Pi and osmotic stress was corroborated by an RT-qPCR analysis (Fig. 6f). The consistent trend of induced expression of POX under the tested abiotic stresses suggests that it may be a potential downstream target of ZmARF1 and function as an intersection in the ZmARF1s regulated pathway for multiple abiotic stress response.
ZmARF1 positively regulates POX by directly binding to AuxRE in its promoter
To further investigate how ZmARF1 regulates the expression of POX, we carried out a transient expression assay using Nicotiana benthamiana leaves. Co-expression of ZmARF1-GFP with Pro-POX-LUC recorded the highest luciferase activity compared to co-expression of GFP with Pro-POX-LUC or Pro-POX-LUC alone (Fig. 7a–c). Members of the ARF family protein have been reported to positively regulate abiotic stress responses in plants by binding to the auxin response element (AuxRE: TGTCTC) in the promoters of stress-responsive genes. Analysis of the 2 kb promoter region of POX identified a putative AuxRE motif, suggesting that ZmARF1 could directly bind to its downstream target POX (AT4G30170) to form a cascade in the stress response pathway. We investigated this possibility through an EMSA. The EMSA was conducted using an MBP-ZmARF1 recombinant protein and a biotin-labeled AuxRE: TGTCTC motif. The results showed that recombinant MBP-ZmARF1 could bind to the biotin-labeled AuxRE probe, which was revealed by the presence of a supershifted band when recombinant MBP-ZmARF1 was incubated with the labeled probe (Fig. 7d). Moreover, the unlabeled competitor probes effectively interfered and reduced the binding ability of ZmARF1 in a dose-dependent manner, while the mutant competitor did not significantly interfere with the binding ability of ZmARF1 (Fig. 7d). These results confirmed that ZmARF1 physically binds to AuxRE in the promoter of POX (AT4G30170) and activates its transcription. To detect whether the POX’s transcriptional activity affected by abiotic stresses is mediated by ZmARF1, we redetected the luciferase activity after co-injection of ZmARF1-GFP with Pro-POX-LUC, followed by the addition of 200 mM NaCl, 200 mM mannitol, and H2O. Fluorescence imaging and subsequent LUC/REN ratio estimation revealed that osmotic stress such as salt and mannitol enhanced the transcriptional activation ability of ZmARF1 on the target POX. Thus, we identified that ZmARF1 acts as a key positive regulatory factor in multiple abiotic stresses and clarified its potential molecular mechanisms in activating the stress response factors in the reactive oxygen species metabolism pathway.
ZmARF1 activates POX expression by binding to AuxRE in its promoter. a Schematic of the effector and reporter construct used for dual luciferase assay. Full length coding sequence of ZmARF1 with a C-terminus GFP protein or only a GFP protein, under the control of 35S promoter was used as the effector. The firefly luciferase gene (LUC) driven by POX promoter was used as the reporter construct. The Renilla luciferase gene (R-LUC) driven by the 35S promoter was used as the internal control. b The LUC/REN ratio representing the relative activity of POX promoter. Each bar represents the mean of three independent samples, and error bars represent the SEM. Significant differences between means were determined using students t-test. ***P < 0.001; ns, no significant difference. c Transient co-expression of ZmARF1-GFP or GFP effectors with the ProPOX::LUC reporter or expression of ProPOX::LUC alone in N. benthamiana leaves. d EMSA assay was performed to identify the interaction between ZmARF1-MBP recombinant protein and the POX promoter. e Osmotic stress for the transient co-expression of ZmARF1-GFP effectors with the ProPOX::LUC reporter in N. benthamiana leaves. f Upregulation of POX expression by ZmARF1 under stresses treatment. Relative LUC activity was determined by the LUC to REN ratio from (e)
Discussion
The inception of the green revolution birthed the development of many crop varieties with improved stress tolerance and enhanced yields (Bailey-Serres et al. 2019). Due to global warming and climate change, the limitations imposed by abiotic stresses on crop productivity have been more menacing (Mora et al. 2015; Sanchez-Bermudez et al. 2022). Plants display tailored physiological and molecular responses when exposed to multiple stresses (Bai et al. 2018; Mota et al. 2021). In this study, we characterized ZmARF1 and its biological function in response to multiple abiotic stresses. It has been reported that ARF transcription factors are involved in the abiotic stress response (Bouzroud et al. 2020; Du et al. 2013). In this study, athough the transcript level of ZmARF1 in the transgenic plants did not show any discernable difference between high Pi and low Pi treatment (Fig. 1a), low Pi treatment increased the protein abundance of ZmARF1 over time (Fig. 1b, c). This suggests that ZmARF1 is an early Pi stress responsive factor whose responsiveness to low Pi stress is regulated at the protein level, and that the stability of its protein is highly regulated by low Pi stress. A sharp disparity between mRNA and protein accumulation of WRKY46 in response to multiple abiotic stress has been reported (Chi et al. 2013). Protein blot assays showed that 26S proteasome inhibitor MG132 stabilized ZmARF1 and mitigated its ubiquitination under Pi-sufficient condition (Fig. 1d). This suggests that ZmARF1 protein accumulation is not exclusively regulated by Pi stress imposition but also by 26S proteasome -mediated post-translational regulation.
Furthermore, transgenic plants overexpressing ZmARF1 displayed significant increase in root phenotypic traits (root length, root surface area, root volume, and root tip number) compared to the WT under low Pi stress (Fig. 2). This is consistent with a previous study in which ZmARF1 was reported to confer low Pi stress tolerance in maize by promoting root development under low Pi stress (Wu et al. 2024). Overexpression of ZmARF1 increased Pi accumulation in transgenic plants compared to WT under low Pi stress (Fig. 3a–c), suggesting that ZmARF1 may help maintain Pi homeostasis under low Pi stress. We hypothesized that this homeostasis is governed by ZmARF1’s role in regulating the expression of some low Pi stress inducible genes. The results showed that transcriptional levels of PHR1, PHT1 and PHO2 increased significantly under low Pi stress in transgenic lines (Fig. 3d–f). The role of high affinity Pi transporters in maintaining Pi homeostasis under low Pi stress is well documented (Chen et al. 2009; Guo et al. 2015; Liu et al. 2018). Members of the ARF family have been reported to regulate the expression of some high-affinity Pi transporters under low Pi stress (Huang et al. 2018; Wang et al. 2014).
The ability of plants to maintain water balance is crucial for overcoming drought and salt stresses. In this study, overexpression of ZmARF1 resulted in better growth and higher survival rates of transgenic plants under drought and salt stress treatments compared to WT (Figs. 4 and 5), indicating a positive response of ZmARF1 to these stresses. In terms of physiological traits, the ZmARF1-transgenic seedlings had the least excised leaf water loss and ion leakage after drought stress treatment compared to the WT (Fig. 4e, f). Similarly, the transgenic plants recorded the least ion leakage after salt stress treatment compared to the WT (Fig. 5e). Water loss and ion leakage are important physiological indices related to osmotic stress imposed by drought and salinity and can be used as accurate metrics for assessing plant resistance to abiotic stress (Lu et al. 2019; Mehari et al. 2022). One of the most prominent phenotypic manifestations of salt stress is abnormal development and inhibition of vegetative growth (van Zelm et al. 2020). Although salt stress generally affected vegetative growth of all the genotypes, the transgenic seedlings displayed superior vegetative growth compared to the WT, which was manifested by relatively higher plant heights (Fig. 5c). Chlorophyll fluorescence, mostly quantified as Fv/Fm, is a true indicator of photosynthetic capability of plants under adverse environmental stresses (Baker 2008; Huang et al. 2021). The results of this study revealed that the ZmARF1-transgenic Arabidopsis had higher chlorophyll content and Fv/Fm values than the WT under drought (Fig. 4c) and salt stress (Fig. 5d, g) respectively. The decrease in chlorophyll content in the WT compared to the ZmARF1-transgenic lines under drought and salinity stress is likely due to the ability of ZmARF1 overexpression to mitigate pigment photo-oxidation and chlorophyll degradation caused by oxidative stress imposed by drought and salinity treatments (Ma et al. 2020). Drought and salinity stresses have been shown to trigger a reduction in leaf chlorophyll content (Taïbi et al. 2016).
The molecular mechanisms by which plants respond to abiotic stresses encompass the activation of signal pathways that mediate biochemical and physiological responses (Su et al. 2020). The class II peroxidases have been reported to respond to multiple abiotic stresses. To unravel the molecular mechanism underlying the role of ZmARF1 in multiple abiotic stress responses, ZmARF1 transgenic Arabidopsis and WT seedlings under low Pi, drought and salt stress treatment or normal conditions were subjected to RNA-seq analysis. Transcriptomic profiling identified a member of the peroxidase superfamily gene, POX, whose expression was upregulated by the three abiotic stresses (Fig. 6), suggesting its possible role in the response of transgenic Arabidopsis to low Pi, drought, and salt stresses. Members of the peroxidase family genes are upregulated under oxidative stress conditions (Jin et al. 2019; Kim et al. 2008; Su et al. 2023). When plants are exposed to prolonged oxidative stress, it triggers the induction of many ROS such as O2−, H2O2 and OH−, which can cause cellular injuries through protein oxidation, lipid peroxidation, and DNA damage that may eventually result in cellular death (Hasanuzzaman et al. 2019, 2020). Members of the class III peroxidases catalyze the reduction of H2O2 in the peroxidative cycle by transferring electrons from various donors (Dat et al. 2000; Gill & Tuteja 2010). Our results showed that POX was a downstream target of ZmARF1, and that ZmARF1 bound to the AuxRE in the promoter of POX and upregulated its transcription (Fig. 7). We belief that ARF-POX module contributes to the plants’ ability to manage and mitigate oxidative stress under these adverse conditions.
Conclusion
This study characterized ZmARF1, a member of the ARF protein family predominantly localized to the nucleus. ZmARF1 acted as an important regulator of phosphate starvation responses, such that overexpression of ZmARF1 resulted in enhanced tolerance to low Pi stress, with marked increased in root length, surface area, volume and root tip number, and increased expression of low Pi stress responsive genes in transgenic Arabidopsis compared to WT. Additionally, our results showed that ZmARF1 conferred tolerance to drought and salt stress, with transgenic plants exhibiting increased chlorophyll parameters, reduced water loss and ion leakage under drought and salt stress compared to WT. Transcriptome profiling suggested that AtPOX was multi-stress responsive gene whose expression was induced by low Pi, drought and salt stresses in the transgenic plants. EMSA and dual-luciferase assays suggested that AtPOX was a downstream gene of ZmARF1 and that its transcription was upregulated by ZmARF1, contributing to anti-oxidative stress tolerance in the transgenic plants and conferring tolerance to low Pi, drought and salt stress. The results of this study suggest that ZmARF1 is a pleiotropic gene conferring tolerance to multiple abiotic stresses in transgenic Arabidopsis.
References
Bai Y, Sunarti S, Kissoudis C, Visser RGF, van der Linden CG (2018) The role of tomato WRKY genes in plant responses to combined abiotic and biotic stresses. Front Plant Sci 9:801. https://doi.org/10.3389/fpls.2018.00801
Bailey-Serres J, Parker JE, Ainsworth EA, Oldroyd GED, Schroeder JI (2019) Genetic strategies for improving crop yields. Nature 575(7781):109–118. https://doi.org/10.1038/s41586-019-1679-0
Bajaj S, Mohanty A (2005) Recent advances in rice biotechnology—towards genetically superior transgenic rice. Plant Biotechnol J 3(3):275–307. https://doi.org/10.1111/j.1467-7652.2005.00130.x
Baker NR (2008) Chlorophyll fluorescence: a probe of photosynthesis in vivo. Annu Rev Plant Biol 59:89–113. https://doi.org/10.1146/annurev.arplant.59.032607.092759
Bouzroud S, Gouiaa S, Hu N, Bernadac A, Mila I, Bendaou N, Smouni A, Bouzayen M, Zouine M (2018) Auxin Response Factors (ARFs) are potential mediators of auxin action in tomato response to biotic and abiotic stress (Solanum lycopersicum). PLoS ONE 13(2):e0193517. https://doi.org/10.1371/journal.pone.0193517
Bouzroud S, Gasparini K, Hu G, Barbosa MAM, Rosa BL, Fahr M, Bendaou N, Bouzayen M, Zsogon A, Smouni A, Zouine M (2020) Down regulation and loss of auxin response factor 4 function using CRISPR/Cas9 alters plant growth stomatal function and improves tomato tolerance to salinity and osmotic stress. Genes (Basel). https://doi.org/10.3390/genes11030272
Cao M, Chen R, Li P, Yu Y, Zheng R, Ge D, Zheng W, Wang X, Gu Y, Gelová Z, Friml J (2019) TMK1-mediated auxin signalling regulates differential growth of the apical hook. Nature 568(7751):240–243. https://doi.org/10.1038/s41586-019-1069-7
Chen YF, Li LQ, Xu Q, Kong YH, Wang H, Wu WH (2009) The WRKY6 transcription factor modulates PHOSPHATE1 expression in response to low Pi stress in Arabidopsis. Plant Cell 21(11):3554–3566. https://doi.org/10.1105/tpc.108.064980
Chen Y, Wu P, Zhao Q, Tang Y, Chen Y, Li M, Jiang H, Wu G (2018) Overexpression of a phosphate starvation response AP2/ERF gene from physic nut in Arabidopsis alters root morphological traits and phosphate starvation-induced anthocyanin accumulation. Front Plant Sci 9:1186. https://doi.org/10.3389/fpls.2018.01186
Chen D, Wang W, Wu Y, Xie H, Zhao L, Zeng Q, Zhan Y (2019) Expression and distribution of the auxin response factors in Sorghum bicolor during development and temperature stress. Int J Mol Sci. https://doi.org/10.3390/ijms20194816
Chi Y, Yang Y, Zhou Y, Zhou J, Fan B, Yu JQ, Chen Z (2013) Protein-protein interactions in the regulation of WRKY transcription factors. Mol Plant 6(2):287–300. https://doi.org/10.1093/mp/sst026
Chiou T-J, Aung K, Lin S-I, Wu C-C, Chiang S-F, Su C-L (2006) Regulation of phosphate homeostasis by MicroRNA in Arabidopsis. Plant Cell 18(2):412–421. https://doi.org/10.1105/tpc.105.038943
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Dat J, Vandenabeele S, Vranova E, Van Montagu M, Inze D, Van Breusegem F (2000) Dual action of the active oxygen species during plant stress responses. Cell Mol Life Sci 57(5):779–795. https://doi.org/10.1007/s000180050041
Dhankher OP, Foyer CH (2018) Climate resilient crops for improving global food security and safety. Plant Cell Environ 41(5):877–884. https://doi.org/10.1111/pce.13207
Dharmasiri N, Dharmasiri S, Estelle M (2005) The F-box protein TIR1 is an auxin receptor. Nature 435(7041):441–445. https://doi.org/10.1038/nature03543
Dixit A, Tomar P, Vaine E, Abdullah H, Hazen S, Dhankher OP (2018) A stress-associated protein, AtSAP13, from Arabidopsis thaliana provides tolerance to multiple abiotic stresses. Plant Cell Environ 41(5):1171–1185. https://doi.org/10.1111/pce.13103
Du H, Liu H, Xiong L (2013) Endogenous auxin and jasmonic acid levels are differentially modulated by abiotic stresses in rice. Front Plant Sci 4:397. https://doi.org/10.3389/fpls.2013.00397
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48(12):909–930. https://doi.org/10.1016/j.plaphy.2010.08.016
Guilfoyle TJ, Hagen G (2007) Auxin response factors. Curr Opin Plant Biol 10(5):453–460. https://doi.org/10.1016/j.pbi.2007.08.014
Guo M, Ruan W, Li C, Huang F, Zeng M, Liu Y, Yu Y, Ding X, Wu Y, Wu Z, Mao C, Yi K, Wu P, Mo X (2015) Integrative comparison of the role of the PHOSPHATE RESPONSE1 subfamily in phosphate signaling and homeostasis in rice. Plant Physiol 168(4):1762–1776. https://doi.org/10.1104/pp.15.00736
Hasanuzzaman M, Bhuyan M, Anee TI, Parvin K, Nahar K, Mahmud JA, Fujita M (2019) Regulation of ascorbate-glutathione pathway in mitigating oxidative damage in plants under abiotic stress. Antioxidants (basel). https://doi.org/10.3390/antiox8090384
Hasanuzzaman M, Bhuyan M, Zulfiqar F, Raza A, Mohsin SM, Mahmud JA, Fujita M, Fotopoulos V (2020) Reactive oxygen species and antioxidant defense in plants under abiotic stress: revisiting the crucial role of a universal defense regulator. Antioxidants (basel). https://doi.org/10.3390/antiox9080681
Hellens RP, Allan AC, Friel EN, Bolitho K, Grafton K, Templeton MD, Karunairetnam S, Gleave AP, Laing WA (2005) Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1:13. https://doi.org/10.1186/1746-4811-1-13
Hema R, Vemanna RS, Sreeramulu S, Reddy CP, Senthil-Kumar M, Udayakumar M (2014) Stable expression of mtlD gene imparts multiple stress tolerance in finger millet. PLoS ONE 9(6):e99110. https://doi.org/10.1371/journal.pone.0099110
Herrera-Estrella L, Lopez-Arredondo D (2016) Phosphorus: the underrated element for feeding the world. Trends Plant Sci 21(6):461–463. https://doi.org/10.1016/j.tplants.2016.04.010
Hong Y, Zhang H, Huang L, Li D, Song F (2016) Overexpression of a stress-responsive NAC transcription factor gene ONAC022 improves drought and salt tolerance in rice. Front Plant Sci 7:4. https://doi.org/10.3389/fpls.2016.00004
Hu J, Chen H, Wang H, Zheng H, Cai W, Xu P (2023) A protocol for measuring the response of Arabidopsis roots to gravity and treatment for simulated microgravity. STAR Protoc 4(1):102099. https://doi.org/10.1016/j.xpro.2023.102099
Hu W, Zuo J, Hou X, Yan Y, Wei Y, Liu J, Li M, Xu B, Jin Z (2015) The auxin response factor gene family in banana: genome-wide identification and expression analyses during development, ripening, and abiotic stress. Front Plant Sci 6:742. https://doi.org/10.3389/fpls.2015.00742
Huang KL, Ma GJ, Zhang ML, Xiong H, Wu H, Zhao CZ, Liu CS, Jia HX, Chen L, Kjorven JO, Li XB, Ren F (2018) The ARF7 and ARF19 transcription factors positively regulate PHOSPHATE STARVATION RESPONSE1 in Arabidopsis roots. Plant Physiol 178(1):413–427. https://doi.org/10.1104/pp.17.01713
Huang D, Wang Q, Jing G, Ma M, Li C, Ma F (2021) Overexpression of MdIAA24 improves apple drought resistance by positively regulating strigolactone biosynthesis and mycorrhization. Tree Physiol 41(1):134–146. https://doi.org/10.1093/treephys/tpaa109
Jain M, Khurana JP (2009) Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J 276(11):3148–3162. https://doi.org/10.1111/j.1742-4658.2009.07033.x
Jin T, Sun Y, Zhao R, Shan Z, Gai J, Li Y (2019) Overexpression of peroxidase gene GsPRX9 confers salt tolerance in soybean. Int J Mol Sci. https://doi.org/10.3390/ijms20153745
Kang C, He S, Zhai H, Li R, Zhao N, Liu Q (2018) A sweetpotato auxin response factor gene (IbARF5) is involved in carotenoid biosynthesis and salt and drought tolerance in transgenic Arabidopsis. Front Plant Sci 9:1307. https://doi.org/10.3389/fpls.2018.01307
Kepinski S, Leyser O (2005) The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435(7041):446-451. https://doi.org/10.1038/nature03542
Kim YH, Kim CY, Song WK, Park DS, Kwon SY, Lee HS, Bang JW, Kwak SS (2008) Overexpression of sweetpotato swpa4 peroxidase results in increased hydrogen peroxide production and enhances stress tolerance in tobacco. Planta 227(4):867–881. https://doi.org/10.1007/s00425-007-0663-3
Kong X, Zhang L, Ding Z (2016) 26S Proteasome: Hunter and prey in auxin signaling. Trends Plant Sci 21(7):546–548. https://doi.org/10.1016/j.tplants.2016.05.007
Kuhn A, Ramans Harborough S, McLaughlin HM, Natarajan B, Verstraeten I, Friml J, Kepinski S, Ostergaard L (2020) Direct ETTIN-auxin interaction controls chromatin states in gynoecium development. Elife 9. https://doi.org/10.7554/eLife.51787
Lesk C, Rowhani P, Ramankutty N (2016) Influence of extreme weather disasters on global crop production. Nature 529(7584):84–87. https://doi.org/10.1038/nature16467
Li J, Wu F, He Y, He B, Gong Y, Yahaya BS, Xie Y, Xie W, Xu J, Wang Q, Feng X, Liu Y, Lu Y (2022) Maize transcription factor ZmARF4 confers phosphorus tolerance by promoting root morphological development. Int J Mol Sci. https://doi.org/10.3390/ijms23042361
Liscum E, Reed JW (2002) Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol Biol 49(3–4):387–400
Liu C, Su J, Stephen GK, Wang H, Song A, Chen F, Zhu Y, Chen S, Jiang J (2018) Overexpression of phosphate transporter gene CmPht1;2 facilitated Pi uptake and alternated the metabolic profiles of chrysanthemum under phosphate deficiency. Front Plant Sci 9:686. https://doi.org/10.3389/fpls.2018.00686
Liu L, Yahaya BS, Li J, Wu F (2024) Enigmatic role of auxin response factors in plant growth and stress tolerance. Front Plant Sci 15:1398818. https://doi.org/10.3389/fpls.2024.1398818
Lu P, Magwanga RO, Kirungu JN, Hu Y, Dong Q, Cai X, Zhou Z, Wang X, Zhang Z, Hou Y, Wang K, Liu F (2019) Overexpression of cotton a DTX/MATE gene enhances drought, salt, and cold stress tolerance in transgenic Arabidopsis. Front Plant Sci 10:299. https://doi.org/10.3389/fpls.2019.00299
Ma Y, Dias MC, Freitas H (2020) Drought and salinity stress responses and microbe-induced tolerance in plants. Front Plant Sci 11:591911. https://doi.org/10.3389/fpls.2020.591911
Mehari TG, Hou Y, Xu Y, Umer MJ, Shiraku ML, Wang Y, Wang H, Peng R, Wei Y, Cai X, Zhou Z, Liu F (2022) Overexpression of cotton GhNAC072 gene enhances drought and salt stress tolerance in transgenic Arabidopsis. BMC Genomics 23(1):648. https://doi.org/10.1186/s12864-022-08876-z
Miao L, Zhu F, Sun Z, Moore JC, Cui X (2016) China’s land-use changes during the past 300 years: a historical perspective. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph13090847
Mittler R (2017) ROS are good. Trends Plant Sci 22(1):11–19. https://doi.org/10.1016/j.tplants.2016.08.002
Mora C, Caldwell IR, Caldwell JM, Fisher MR, Genco BM, Running SW (2015) Suitable days for plant growth disappear under projected climate change: potential human and biotic vulnerability. PLoS Biol 13(6):e1002167. https://doi.org/10.1371/journal.pbio.1002167
Mota APZ, Brasileiro ACM, Vidigal B, Oliveira TN, da Cunha Quintana Martins A, Saraiva MAP, de Araujo ACG, Togawa RC, Grossi-de-Sa MF, Guimaraes PM (2021) Defining the combined stress response in wild Arachis. Sci Rep 11(1):11097. https://doi.org/10.1038/s41598-021-90607-7
Nan X, Huihui Z, Haixiu Z, Yining W, Jinbo L, Li X, Zepeng Y, Wenxu Z, Yi Q, Guangyu S (2018) The response of photosynthetic functions of F(1) cutting seedlings from Physocarpus amurensis Maxim (female symbol) x Physocarpus opulifolius “Diabolo” (male symbol) and the parental seedlings to salt stress. Front Plant Sci 9:714. https://doi.org/10.3389/fpls.2018.00714
Pandey P, Irulappan V, Bagavathiannan MV, Senthil-Kumar M (2017) Impact of combined abiotic and biotic stresses on plant growth and avenues for crop improvement by exploiting physio-morphological traits. Front Plant Sci 8:537. https://doi.org/10.3389/fpls.2017.00537
Rockstrom J, Williams J, Daily G, Noble A, Matthews N, Gordon L, Wetterstrand H, DeClerck F, Shah M, Steduto P, de Fraiture C, Hatibu N, Unver O, Bird J, Sibanda L, Smith J (2017) Sustainable intensification of agriculture for human prosperity and global sustainability. Ambio 46(1):4–17. https://doi.org/10.1007/s13280-016-0793-6
Sanchez-Bermudez M, Del Pozo JC, Pernas M (2022) Effects of combined abiotic stresses related to climate change on root growth in crops. Front Plant Sci 13:918537. https://doi.org/10.3389/fpls.2022.918537
Shen C, Wang S, Zhang S, Xu Y, Qian Q, Qi Y, de Jiang A (2013) OsARF16, a transcription factor, is required for auxin and phosphate starvation response in rice (Oryza sativa L.). Plant Cell Environ 36(3):607–620. https://doi.org/10.1111/pce.12001
Shrivastava P, Kumar R (2015) Soil salinity: a serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saudi J Biol Sci 22(2):123–131. https://doi.org/10.1016/j.sjbs.2014.12.001
Simonini S, Deb J, Moubayidin L, Stephenson P, Valluru M, Freire-Rios A, Sorefan K, Weijers D, Friml J, Ostergaard L (2016) A noncanonical auxin-sensing mechanism is required for organ morphogenesis in Arabidopsis. Genes Dev 30(20):2286–2296. https://doi.org/10.1101/gad.285361.116
Song X, Xiong Y, Kong X, Huang G (2023) Roles of auxin response factors in rice development and stress responses. Plant Cell Environ 46(4):1075–1086. https://doi.org/10.1111/pce.14494
Strock CF, Morrow de la Riva L, Lynch JP (2018) Reduction in root secondary growth as a strategy for phosphorus acquisition. Plant Physiol 176(1):691–703. https://doi.org/10.1104/pp.17.01583
Su P, Yan J, Li W, Wang L, Zhao J, Ma X, Li A, Wang H, Kong L (2020) A member of wheat class III peroxidase gene family, TaPRX-2A, enhanced the tolerance of salt stress. BMC Plant Biol 20(1):392. https://doi.org/10.1186/s12870-020-02602-1
Su P, Sui C, Niu Y, Li J, Wang S, Sun F, Yan J, Guo S (2023) Comparative transcriptomic analysis and functional characterization reveals that the class III peroxidase gene TaPRX-2A regulates drought stress tolerance in transgenic wheat. Front Plant Sci 14:1119162. https://doi.org/10.3389/fpls.2023.1119162
Szemenyei H, Hannon M, Long JA (2008) TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 319(5868):1384–1386. https://doi.org/10.1126/science.1151461
Taïbi K, Taïbi F, Abderrahim LA, Ennajah A, Belkhodja M, Mulet JM (2016) Effect of salt stress on growth chlorophyll content lipid peroxidation and antioxidant defence systems in Phaseolus vulgaris L. S Afr J Bot 105:306–312. https://doi.org/10.1016/j.sajb.2016.03.011
van Zelm E, Zhang Y, Testerink C (2020) Salt tolerance mechanisms of plants. Annu Rev Plant Biol 71:403–433. https://doi.org/10.1146/annurev-arplant-050718-100005
Wang D, Pei K, Fu Y, Sun Z, Li S, Liu H, Tang K, Han B, Tao Y (2007) Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 394(1–2):13–24. https://doi.org/10.1016/j.gene.2007.01.006
Wang Y, Deng D, Shi Y, Miao N, Bian Y, Yin Z (2012) Diversification, phylogeny and evolution of auxin response factor (ARF) family: insights gained from analyzing maize ARF genes. Mol Biol Rep 39(3):2401–2415. https://doi.org/10.1007/s11033-011-0991-z
Wang S, Zhang S, Sun C, Xu Y, Chen Y, Yu C, Qian Q, Jiang DA, Qi Y (2014) Auxin response factor (OsARF12), a novel regulator for phosphate homeostasis in rice (Oryza sativa). New Phytol 201(1):91–103. https://doi.org/10.1111/nph.12499
Wang NN, Xu SW, Sun YL, Liu D, Zhou L, Li Y, Li XB (2019) The cotton WRKY transcription factor (GhWRKY33) reduces transgenic Arabidopsis resistance to drought stress. Sci Rep 9(1):724. https://doi.org/10.1038/s41598-018-37035-2
Wright RC, Nemhauser JL (2015) New tangles in the auxin signaling web. F1000Prime Rep 7:19. https://doi.org/10.12703/P7-19
Wu F, Yahaya BS, Gong Y, He B, Gou J, He Y, Li J, Kang Y, Xu J, Wang Q, Feng X, Tang Q, Liu Y, Lu Y (2024) ZmARF1 positively regulates low phosphorus stress tolerance via modulating lateral root development in maize. PLoS Genet 20(2):e1011135. https://doi.org/10.1371/journal.pgen.1011135
Xu T, Dai N, Chen J, Nagawa S, Cao M, Li H, Zhou Z, Chen X, De Rycke R, Rakusova H, Wang W, Jones AM, Friml J, Patterson SE, Bleecker AB, Yang Z (2014) Cell surface ABP1-TMK auxin-sensing complex activates ROP GTPase signaling. Science, 343(6174), 1025-1028. https://doi.org/10.1126/science.1245125
Yang Y, Guo Y (2018) Unraveling salt stress signaling in plants. J Integr Plant Biol 60(9):796–804. https://doi.org/10.1111/jipb.12689
Zhang H, Li Y, Zhu JK (2018) Developing naturally stress-resistant crops for a sustainable agriculture. Nat Plants 4(12):989–996. https://doi.org/10.1038/s41477-018-0309-4
Zhao C, Liu B, Piao S, Wang X, Lobell DB, Huang Y, Huang M, Yao Y, Bassu S, Ciais P, Durand JL, Elliott J, Ewert F, Janssens IA, Li T, Lin E, Liu Q, Martre P, Muller C, Peng S, Penuelas J, Ruane AC, Wallach D, Wang T, Wu D, Liu Z, Zhu Y, Zhu Z, Asseng S (2017) Temperature increase reduces global yields of major crops in four independent estimates. Proc Natl Acad Sci U S A 114(35):9326–9331. https://doi.org/10.1073/pnas.1701762114
Funding
This study was supported by Innovative Research Group Project of the National Natural Science Foundation of China (grant numbers 32030078, 32101785 and U21A20209), General Project of Natural Science Foundation of Sichuan Province (2022NSFSC0147 and 2023NSFSC0221), the key Research Program of the Department of Science and Technology of Sichuan province, China (2021YFN0034), and The Foundation of Major Science and Technology Application Demonstration Project of Chengdu, China (2022-YF09-00054-SN).
Author information
Authors and Affiliations
Contributions
F.W. and Y.L. conceived the research; L.L. and Y.G. participated in its design and coordination; Y.G., B.S.Y. Y.C., and D.S. performed the experiments; L.L., F.L., J.G., and Z.Z. collected and analyzed the data; L.L. and B.S.Y. drafted the manuscript; L.L., F.W., and Y.L. revised the manuscript. L.L., Y.G. and B.S.Y. contributed equally to this study. F.W. advised on all the experiments and revised the manuscript. All the authors have read and approved the final version of this manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, L., Gong, Y., Yahaya, B.S. et al. Maize auxin response factor ZmARF1 confers multiple abiotic stresses resistances in transgenic Arabidopsis. Plant Mol Biol 114, 75 (2024). https://doi.org/10.1007/s11103-024-01470-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11103-024-01470-9