Abstract
Key message
Two homologs PsnSuSy1 and PsnSuSy2 from poplar played largely similar but little distinct roles in modulating sink strength, accelerating vegetative growth and modifying secondary growth of plant. Co-overexpression of them together resulted in small but perceptible additive effects.
Abstract
Sucrose synthase (SuSy) acts as a crucial determinant of sink strength by controlling the conversion of sucrose into UDP-glucose, which is not only the sole precursor for cellulose biosynthesis but also an extracellular signaling molecule for plants growth. Therefore, modification of SuSy activity in plants is of utmost importance. We have isolated two SuSy genes from poplar, PsnSuSy1 and PsnSuSy2, which were preferentially expressed in secondary xylem/phloem. To investigate their functions, T2 tobacco transgenic lines of PsnSuSy1 and PsnSuSy2 were generated and then crossed to generate PsnSuSy1/PsnSuSy2 dual overexpression transgenic lines. SuSy activities in all lines were significantly increased though PsnSuSy1/PsnSuSy2 lines only exhibited slightly higher SuSy activities than either PsnSuSy1 or PsnSuSy2 lines. The significantly increased fructose and glucose, engendered by augmented SuSy activities, caused the alternations of many physiological, biochemical measures and phenotypic traits that include accelerated vegetative growth, thickened secondary cell wall, and increased stem breaking force, accompanied with altered expression levels of related pathway genes. The correlation relationships between SuSy activities and many of these traits were statistically significant. However, differences of almost all traits among three types of transgenic lines were insignificant. These findings clearly demonstrated that PsnSuSy1 and PsnSuSy2 had similar but little distinct functions and insubstantial additive effects on modulating sink strength and affecting allocation of carbon elements among secondary cell wall components.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In higher plants, assimilated carbon is transported as sucrose whose partition and metabolism play important roles in the growth and development of non-photosynthetic tissues (Asano et al. 2002). Sucrose cleavage initiates its utilization and is mainly catalyzed by two enzyme families with entirely different properties, invertase and sucrose synthase (SuSy). Invertase is a hydrolase that cleaves sucrose into glucose and fructose while SuSy is a glycosyl transferase and preferentially catalyzes the formation of UDP-glucose (UDPG), which not only is the sole precursor for cellulose and callose biosynthesis (Geigenberger and Stitt 1993; Heim et al. 1993) but also can be converted to starch biosynthesis precursor of ADP-glucose (Li et al. 2013). Recently studies also suggest a potential role of UDPG for acting as an extracellular signaling molecule for plant growth and development (Janse van Rensburg and Van den Ende 2017; Wai et al. 2017). All these suggest that SuSy activity is important for maintaining structural and storage carbohydrates and growth of plant cells and also signifies essential role of SuSy in sink tissue metabolism (Chen and Chourey 1989; Coleman et al. 2006, 2009). SuSy can exist in soluble and particulate forms, depending on their phosphorylation status (Komina et al. 2002). For example, phosphorylation was found to change Zea mays SuSy from a membrane-bound form to a soluble form (Winter et al. 1997), and this process seems to be reversible (Winter and Huber 2000). Cytosolic SuSy (soluble SuSy) is mainly involved in respiration and synthesis of starch (Salnikov et al. 2001), whereas particulate SuSy is linked to the synthesis of cell wall polysaccharides callose and cellulose by directly supplying UDP-glucose as substrate (Amor et al. 1995) and can accumulate to high levels along plasma membrane and in microtubules during secondary cell wall formation (Salnikov et al. 2001). For this reason, phosphorylation status of SuSy determines its exact functions in plant growth and development.
Many previous studies showed that SuSy activity modulations affect structural and storage carbohydrates, growth processes, biomass accumulation, and cellulose biosynthesis in plants. For example, the antisense suppression of SuSys has demonstrated significant changes of soluble carbohydrates in sink tissue of potato (Zrenner et al. 1995), carrots (Tang and Sturm 1999), tomato (D’Aoust et al. 1999), and poplar (Gerber et al. 2014). Silencing of SuSy in cotton impairs development of seed fibers through curtailing cellulose synthesis (Ruan et al. 2003). Suppression of SuSy led to reduced growth in carrot (Tang and Sturm 1999), but increased height in poplar (Gerber et al. 2014). In contrast to SuSys suppression, SuSys overexpression increases plant growth and soluble sugar contents in tobacco and poplar (Konishi et al. 2004; Coleman et al. 2006), increases starch level in potato (Baroja-Fernandez et al. 2009), accelerates leaf expansion and enhances fiber length and yield in cotton (Xu et al. 2012), and enhances secondary cell wall formation, biomass production and mechanical strength while reducing cellulose crystallinity in rice (Fan et al. 2017). Moreover, SuSys in woody plants have also been found to play important roles in wood formation as their expression levels were correlated with secondary thickening of xylem, wood strength and density (Salnikov et al. 2001; Nilsson et al. 2008; Gerber et al. 2014). In brief, the previous studies have clearly showed that SuSys play vital roles in modulating sink strength, vegetative growth, and wood formation.
The vast majority of SuSys characterized to date in both monocot and dicot species belong to multigene families (Komatsu et al. 2002; Bieniawska et al. 2007; Hirose et al. 2008), which have different and distinct developmental and organ-specific expression patterns (Fu and Park 1995; Sturm et al. 1999), thus leading to specialization and/or redundancy functions in plant growth and development (Goren et al. 2017). For example, the SuSy family in Arabidopsis thaliana consists of six distinct members that can be grouped into three distinct AtSuSy pairs (Baud et al. 2004; Bieniawska et al. 2007). Rice has six SuSy genes and four of them are expressed in a tissue- and stage-specific manner (Hirose et al. 2008). In maize, though two isoforms of SuSy, Shrunken1 (Sh1) and SuSy1, are expressed in the developmental endosperm, Sh1 preferentially functions in cellulose biosynthesis, whereas SuSy1 is mainly involved in starch biosynthesis (Chourey 1981). These studies suggest that some SuSys from the same family have developed differential functions. Therefore, we cannot conjecture the functions of SuSys based on other SuSys with known functions without experimental test and validation. It has been reported that SuSy in poplar is a multigene family of 15 members, each exhibits distinct but partially overlapping expression pattern (An et al. 2014), which indicates their functions have already differentiated. To date, we know little about the exact biological functions of poplar SuSys in regard to plant growth and development.
Besides the fibre and vessel characteristics, the most striking modification of tension wood is a unique cell wall layer internal to secondary cell wall of fibre, named the ‘gelatinous layer’ (G-layer) (Jourez et al. 2001; Pilate et al. 2004). PttSuSy1 and PttSuSy2 were listed at the top of all significantly up-regulated genes in tension wood of Populus tremula (Andersson-Gunneras et al. 2006), and their orthologous genes in Populus trichocarpa, PtrSuSy1 and PtrSuSy1, are also most abundantly expressed in xylem tissue of stems (Zhang et al. 2011). We have isolated two genes, PsnSuSy1 and PsnSuSy2, from Populus simonii × Populus nigra, a fast-growing and widely distributed tree species in Northern China. PsnSuSy2 was characterized previously and its functions in increasing vegetative growth, secondary cell wall thickness, cellulose contents, hemicellulose contents, and fibre lengths while decreasing lignin contents were substantiated (Wei et al. 2015). Nevertheless, we were still fascinated by learning PsnSuSy1’s functions in relationship with PsnSuSy2. In this study, we investigated the functions of PsnSuSy1 and PsnSuSy2 through independent overexpression of each gene and co-overexpression of both genes in transgenic tobacco, followed by characterization of their transgenic plants. The results from independent overexpression of each gene clearly demonstrated that PsnSuSy1 and PsnSuSy2 played largely similar but little distinct roles in modulating sink strength, accelerating vegetative growth, and modifying secondary cell wall and fibre formation of plant. Co-overexpression of them together resulted in small but perceptible additive effects.
Materials and methods
Plant materials
One-year-old Populus simonii × Populus nigra trees were propagated and planted in a mixture of turfy peat and sand (2:1 v/v) in the greenhouse. The primary shoot leaves, transition leaves, secondary leaves, primary xylem, transition xylem, secondary xylem, primary phloem, transition phloem, secondary phloem, and roots were collected and immediately frozen in liquid Nitrogen and stored at − 80 °C. The RNA was isolated according to a previously published method (Liao et al. 2004)and later treated with DNase I (Qiagen) to remove genomic DNA (Kolosova et al. 2004).
Cloning PsnSuSy1 from P. simonii × P. nigra
5 µg total RNA were used for the synthesizing cDNAs using SuperScript II Reverse Transcriptase (Invitrogen). The full PsnSuSy1 cDNA was amplified from P. simonii × P. nigra with gene-specific primers (Supplemental Table 1). The PCR product was cloned into pMD18-T vector (TaKaRa), and then transformed into Escherichia coli cells (DH5α) for validation by Sanger sequencing.
Sequence comparisons
BLASTP (http://ncbi.nlm.nih.gov) were used to analyze the sequence similarity of deduced proteins of PsnSuSy1 and PsnSuSy2. The conserved domains of PsnSuSy1 and PsnSuSy2 were searched by CDD algorithms (http://ncbi.nlm.nih.gov). Multiple sequence alignment was carried on using ClustalW2 (available in http://www.ebi.ac.uk/Tools/msa/clustalw2/) with default setting.
Subcellular localization
The full-length coding region of PsnSuSy1 and PsnSuSy2 without termination codon was amplified using specific primers (Supplemental Table 2) and then fused to the N-terminal of GFP driven by CaMV 35S promoter in pGWB5 vector. The two fusion constructs were delivered into onion epidermal cells via particle bombardment (GJ-1000). The GFP fluorescent images were photographed with confocal microscopy (Leica TCS SP5) at 24 h after bombardment.
Transformation of Nicotiana tabacum and production of single-and dual-transgenic lines
The PsnSuSy1 was amplified with specific primers (Supplemental Table 3), and then inserted into the pROKII vector at the position immediately downstream of CaMV 35S promoter. The pROKII-PsnSuSy1 was first transferred into Agrobacterium tumefaciens EHA105 using the freeze–thaw method. Tobacco plants (Nicotiana tabacum) were then transformed as described previously (Schroeder et al. 1993). Transgenic lines were selected on MS medium containing 250 µg/mL kanamycin and 500 µg/mL carbenicillin. The T1 seeds from self-pollinated lines were germinated on MS medium with kanamycin (25 mg/L) to produce T1 generation transgenic lines. We repeated this process to obtain the T2 generation seeds. The genomic DNA of T2 seedlings was amplified by regular PCR using the PROKII sequencing primers listed in Supplemental Table 3 to verify whether PsnSuSy1 was integrated into tobacco genome.
PsnSuSy1 single-transgene lines and PsnSuSy2 single-transgene lines (Wei et al. 2015) were grown to maturity in the greenhouse. Of these single-transgenic lines, the lines, with highest expression of PsnSuSy1 and PsnSuSy2 mRNA levels compared with other single-transgene lines were selectively crossed to generate PsnSuSy1 /PsnSuSy2 dual overexpression lines. The pods from each controlled crossed were collected, and the seeds were sterilized as described previously. The seeds were germinated on half-strength MS medium with 2% sucrose and kanamycin (25 mg/L). Transformants were confirmed for the presence of both PsnSuSy1 and PsnSuSy2 genes using genomic PCR screening.
All tested PsnSuSy1, PsnSuSy2, and PsnSuSy1 /PsnSuSy2 dual transgene lines, and wild-type (WT) were grown in the greenhouse and subsequently used for characterization.
Growth parameters measurement
The developmental stages of tissues were standardized by employing a plastichron index (PI) (PI = 0 was defined as the first leaf greater than 5 cm in length; PI = 1 was the leaf immediately below PI = 0). Stems spanning PI = 3 to PI = 5 were cut and frozen in liquid nitrogen, and retained for PsnSuSy1 and PsnSuSy2 transcript abundance, enzyme activity, soluble carbohydrates concentration, and transcript abundance of genes related to fibre and secondary cell wall formation analysis. Stems spanning PI = 5 to PI = 8 were cut and used for breaking force, fibre length and width, secondary cell wall thickness, and cell wall chemical composition analyses. Leaves from PI = 3 to PI = 5 were measured for length and width, chlorophyll content, and photosynthesis rate.
The plant height from base to tip of the highest bud and the base diameter above ~ 1 cm of soil were measured prior to harvest to measure biomass. The fresh weight was determined immediately. Then, the material was put into an oven and heated for 10 min at 100 °C. After that, the material was heated at 75 °C until the weight did not change. This final weight was taken as the dry weight. All experiments were conducted with three biological replicates.
Gene expression analysis
5 µg total RNA from multiple tissues of P. simonii × P. nigra and tobacco plants were used for synthesing cDNA. Samples of cDNA were run in triplicate with the SYBR premix ExTaq kit (TaKaRa) and an Applied Biosystems 7500 Real-Time PCR System to determine the critical threshold (Ct). The PsnSuSy1 and PsnSuSy2 expression levels in poplar were detected by the real-time quantitative PCR (qRT-PCR), and the primers used for qRT-PCR of PsnSuSy1 and PsnSuSy2 and reference gene, PsnACTIN1, are listed in Supplemental Table 4.
The expression levels of PsnSuSy1 in PsnSuSy1 lines, PsnSuSy2 in PsnSuSy2 lines, and PsnSuSy1 and PsnSuSy2 in the PsnSuSy1/PsnSuSy2 dual transgenic lines were determined by reverse transcription (RT)-PCR using NtACTIN2 as an internal reference. All the primers used are shown in Supplemental Table 4.
Analysis of expression levels of genes involved in cell expansion and elongation (ExpansinA, ExpansinB, TIP1;3, TIP1;4, XTH5, and XTH8) (Aspeborg et al. 2005), programmed cell death (XSP1, XCP2, SCPL45, and SCPL49) (Plavcova et al. 2013), cellulose biosynthesis (CesA4, CesA7, and CesA8) (Appenzeller et al. 2004), hemicellulose biosynthesis (FRA8, IRX9, and IRX10)(Wu et al. 2010), lignin biosynthesis (4CL1, PAL1, C3H1, CCoAOMT1, CCR1, C4H, CAD4, and CAD6) (Raes et al. 2003), and cellulose crystallinity (GH9B1, GH9B3, and GH9B16) (Xie et al. 2013; Li et al. 2015) in tobacco were performed using gene-specific primers (Supplemental Table 5). The primers of NtACTIN2, used as an internal control, were listed in Supplemental Table 4. Quantification of gene expression relative to PsnACTIN1 or NtACTIN2 was calculated using the delta–delta CT method (Livak and Schmittgen 2001). All experiments were conducted with three biological replicates.
Enzymatic activity analysis
The fresh materials (approximately 0.2 g) were ground in liquid nitrogen and then added 1 ml of extraction buffer that contained the following chemical compounds: 50 mM N-2-hydroxyethylpiperazine-N′-2-ethanesulphonic acid-KOH, pH 7.5, 10 mM MgCl2, 1 mM ethylene diamine tetraacetic acid, 2 mM dithiothreitol, 1 mM phenylmethylsulphonyl fluoride, 5 mM ε-amino-n-caproic acid, 0.1% v/v Triton X-100, 10% v/v glycerol. The samples were centrifuged at 10,000 g for 10 min at 4 °C. The extract was passed through a DG 10 desalting column and pre-equilibrated with ice-cold extraction buffer but without Triton X-100. A 100 µL aliquot was used for total protein content assay using BSA as the standard. The activities of SuSy, acid invertase, and alkaline invertase were measured immediately in a 250 µL reaction mixture for 0 and 30 min at pH 5.0, 6.5 and 7.5, respectively (Kennedy et al. 1983).
SuSy activity was assayed in the direction of sucrose breakdown using 50 µL extract. Each reaction contained 20 mM Pipes-KOH, pH 6.5, l00 mM sucrose, 2 mM UDP, and 20 µL of extract in a total volume of 250 µL. Control reactions lacked UDP. Reactions were started by the addition of extract and incubated at 25 °C for 30 min. The reactions were stopped with 250 µL of 0.5 M Tricine-KOH, pH 8.3 by boiling for 10 min. Fructose was measured spectrophotometrically, as described (King et al. 1997).
Soluble acid and alkaline invertases were measured by incubation of 20 µL of extract with 100 mM sucrose in 100 mM acetic acid-NaOH, pH 5.0 (acid invertase), or 100 mM sodium acetate-acetic acid, pH 7.5 (alkaline invertase), in a total volume of 250 µL. Reactions were started by the addition of extract and incubated at 25 °C for 30 min. The reactions were stopped with 250 µL of 0.5 M Tricine-KOH, pH 8.3 by boiling for 10 min. Control reactions contained only the boiled extract. Glucose in three biological replicates was measured at 520 nm on spectrophotometer as previously described (King et al. 1997).
Soluble carbohydrate content measurement
Approximately 50 mg of fresh stem tissue was ground in liquid nitrogen and, then, extracted with 1 mL of preheated 80% (v/v) ethanol for 5 min at 80 °C. Upon cooling, they were centrifuged at 12,000 g for 10 min. The supernatants were collected, and the pellets were resuspended in 0.5 mL of 50% (v/v) ethanol and spun again as described above. The resulting pellet was extracted with 0.5 mL of water and re-centrifuged. The 2 ml total supernatant was mixed with an equal volume of chloroform and shaken vigorously. The aqueous phase was collected, dried in a vacuum, and redissolved in 0.5 mL water. The total soluble sugar, sucrose, glucose, and fructose contents were measured enzymatically on a spectrophotometer at 340 nm as previously described (Lunn and Hatch 1995).
Determination of break forces
The breaking force, which has been reported to be correlated with the cellulose content (Dhugga 2007), refers to the tensile or bending strength used to break the stems. The breaking forces of stem segments in three biological replicates were analyzed using YYD-1 plant stalk analyzer according to the manufacturer’s instructions (Zhejiang Top Instrument Co., Ltd.)
Determination of chlorophyll contents and photosynthetic rates
To determine chlorophyll contents, three biological replicates of leaf samples were randomly selected and measured for total chlorophyll content as described previously (Qiu et al. 2013). The photosynthetic rate was acquired using Li-6400XT portable photosynthesis system (Li-COR) according to the manufacturer’s instructions.
Histological analysis
Six-month-old poplar basal stems were cut into 0.5 cm segments and submerged in 4% paraformaldehyde at 4 °C for 3 days, washed twice in 1 × PBS for 15 min, dehydrated in a graded ethanol series (2 h each), incubated sequentially in dimethylbenzene: ethanol 25: 75, 50: 50, 75: 25 and then incubated in100% dimethylbenzene twice, for 2 h each time. Stem sections were incubated in dimethylbenzene: paraplast 75: 25 overnight at 63 °C and then in pure paraplast overnight at 63 °C. The paraplast-embedded stems were sectioned to different thickness for different staining using a Leica RM 2235 microtome (Leica) and adhered to Super-frost Plus microscope slides (Thermo Fisher) overnight at 37 °C. Two 20 min incubations in dimethylbenzene were used to remove the paraplast from sections adhered to slides, followed by rehydration in a graded ethanol series. Stem sections (1 µm thick) cut with Leica EM UC6 microtome were stained with 0.01% Calcofluor White, and the cellulose was observed with an inverted UV fluorescence microscope. Under this condition, only secondary walls exhibited brilliant fluorescence. At the same time, some stem sections (50 µm thick) were stained with phloroglucinol-HCl for observing lignin, which takes on bright red color under a light microscope. The fluorescence-labeled xylan signals were visualized and imaged with an Olympus DX51 light microscope.
Scanning electron microscopy
Stem segments were prepared by freeze-drying for scanning electron microscopy (SEM) (S-4800, HITACHI). Dry segments were mounted on aluminum stubs using carbon tape with conductive silver paint applied to the sides to reduce sample charging. The segments were then sputter-coated with gold in an E-100 ion sputter. Imaging was performed at beam accelerating voltages from 12.5 to 25 kV. The secondary wall thicknesses of fibers in the SEM micrographs were quantified in a randomly selected area of 45 cells using Image J software (http://rsbweb.nih.gov/ij/).
Determination of cellulose, hemicellulose, and lignin contents
The determination of the content of lignin, cellulose, and hemicellulose was conducted with the ANKOM 2000i Automatic fiber analyzer. Briefly, the content of NDF, including hemicellulose, cellulose and lignin, was first measured according to the neutral detergent fiber (NDF) procedure. Then, the content of ADF, including cellulose and lignin, was measured according to the acid detergent fiber (ADF) measuring procedure. The hemicellulose content was the NDF content minus ADF content. At last, the rest material after determination of ADF and NDF, acid detergent lignin (ADL), was subjected to gravimetric analysis. The content of cellulose was the ADF minus ADL content. All experiments were conducted with three biological replicates.
Cellulose crystallinity measurement
The crystallinity index (CrI) has been widely used to account for cellulose crystallinity and can be detected by X-ray diffraction (XRD) patterns. In this paper, we used Rigaku-D/MAX instrument (Ultima III; Japan) to measure CrI of the crude cell wall materials in the stem of transgenic lines according to the previous described method (Xie et al. 2013). The raw material power was laid on glass sample holder and detected under plateau conditions. Ni-filtered Cu-Kα radiation (λ = 0.1541 nm) was generated and scanned at speed of 0.0197°/s from 10 to 45°. The CrI was estimated using the intensity of the 200 peak (I200, θ = 22.5°) and the intensity at the minimum between the 200 and 110 peaks (Iam, θ = 18.5°) as follows: CrI = 100 × (I200 − Iam)/I200. I200 represents both crystalline and amorphous materials while Iam represents amorphous materials. Stand error of CrI method was detected at ± 0.05–0.15 using five representative samples in triplicate.
Fiber length and width analysis
Stem segments with approximate dimension of 2 × 2 × 30 mm3 were harvested and immersed into Franklin solution (1:1 peroxide and glacial acetic acid) with 3.6% (g/v) sodium hypochlorite for 20 h at 70 °C. Upon decanting the solution, the materials were immersed in pure Franklin solution for 4 days at 70 °C, washed in a vacuum with deionized water until the materials reached a neutral pH, dried for 24 h at 105 °C, and then re-suspended in 10 mL of deionized water. The fiber length and width were obtained by counting 25–40 fibers per second on Fiber Quality Analyzer (FQA).
Statistical analysis
Differences of all measured traits were analyzed with one-way analysis of variance (ANOVA) using SPSS 21, with Duncan’s multiple range test being used for multiple comparisons. In addition, Mean values and standard deviations (SDs) were calculated from three biological replicates. Correlation coefficients were calculated by performing Spearman’s rank correlation analysis for pairs of measured traits. The significant levels were set to P < 0.05 and P < 0.01.
Results
Tissue-specific expression and subcellular location of PsnSuSy1 and PsnSuSy2
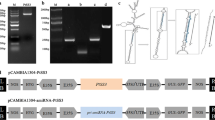
We obtained a full length PsnSuSy1 cDNA of 2405 bp from P. simonii × P. nigra. The deduced protein sequence of PsnSuSy1 is 92.7% identical to the protein encoded by PsnSuSy2 (Fig. 1a). Although PsnSuSy1 and PsnSuSy2 both possess the deduced phosphorylation regions of eight amino acids (Huber et al. 1996; Zhang et al. 1997), their phosphorylation sites are different in that the ser is located at position 11 in PsnSuSy1 in contrast to the position 9 in PsnSuSy2 (Fig. 1a). The domain analysis revealed that they both had two conserved domains responsible for sucrose breakdown (TIGR02470) and synthesis (pfam00862), respectively (Fig. 1a). These results indicated that their functions have some discrepancies on the premise that they are highly conserved.
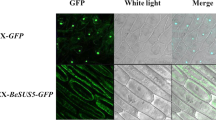
Sequence comparison and subcellular locations of PsnSuSy1 and PsnSuSy2 proteins, and tissue-specific expression patterns of PsnSuSy1 and PsnSuSy2. a Comparison of the amino acid sequences of PsnSuSy1 and PsnSuSy2 proteins. Arrows indicate conserved domains: (1) phosphorylation region (6–13 aa in PsnSuSy1 and 4–11 aa in PsnSuSy2); (2) Sucrose_synth domain(pfam00862, 16–553 aa in PsnSuSy1 and 14–551 aa in PsnSuSy2), which is responsible for catalyzing the synthesis of sucrose from UDP-glucose and fructose, and (3) Sucr_synth domain (TIGR02470, 22–800 aa in PsnSuSy1 and 24–802 aa in PsnSuSy2), which is responsible for catalyzing sucrose plus UDP (or ADP) to generate D-fructose and UDP-glucose (or ADP-glucose), which is the precursor for cell wall (or starch) biosynthesis. Residues are colored according to their polarity properties (neutral non-polar as black, neutral polar as green, acidic as red, and basic as blue). b Quantitative RT-PCR analysis of relative expression levels of PsnSuSy1 and PsnSuSy2 in ten tissues of 1-year-old P. simonii × P. nigra. The PsnACTIN2 was used as an internal control. Each error bar represents a standard deviation (SD) of three biological replicates. c Subcellular localization of PsnSuSy1 and PsnSuSy2 proteins in onion epidermal cells
Although PsnSuSy2 expression patterns in P. simonii × P. nigra have been analyzed in our previous study (Wei et al. 2015), we characterized PsnSuSy2 again for a more accurate comparison with PsnSuSy1. The quantitative real-time PCR (qRT-PCR) analysis revealed that PsnSuSy1 and PsnSuSy2 transcripts were detected in all examined tissues and predominantly expressed in transition and secondary xylem (Fig. 1b). It was noteworthy that the expression levels of PsnSuSy1 were consistently higher than that of PsnSuSy2 in all tissues examined, especially in secondary tissues.
In order to determinate the exact location of PsnSuSy1 and PsnSuSy2 at subcellular level, we performed in vivo localization experiments by transient overexpression PsnSuSy1 and PsnSuSy2 separately in onion epidermal cells through particle bombardment method. As shown in the Fig. 1c, the PsnSuSy1-GFP and PsnSuSy2-GFP fusion proteins were both detected in cytosol and plasma membrane, confirming that PsnSuSy1 and PsnSuSy2 co-existed in both soluble and particulate forms at same time. However, due to the limited resolution of this assay, we couldn’t further distinguish if they were significantly different at different subcellular locations.
Overexpression of PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 and SuSy enzyme activities in transgenic tobacco
To investigate the exact functions of PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 in plant growth and development, the three types of overexpression transgenic lines, PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2, were generated. The PsnSuSy2 transgenic lines generated in previous study (Wei et al. 2015) were characterized again along with PsnSuSy1 and PsnSuSy1/PsnSuSy2 transgenic lines for more accurate comparison.
The qRT-PCR analysis revealed that all verified transgenic lines of PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 showed obvious presence of the exogenous PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 simultaneously (Fig. 2a). We also found that there were conspicuous discrepancies of exogenous poplar SuSy expression levels in each type of transgenic lines. Based on exogenous PsnSuSy1, PsnSuSy2, and PsnSuSy1 / PsnSuSy2’s expression levels, we selected three transgenic lines per transgene or dual transgenes, and 15 plants per line for greenhouse experimental trials and further characterization. To ensure that there was no co-suppression of native SuSys in these selected transgenic tobacco lines, transcript levels of tobacco NtSuSy1 and NtSuSy2 were also determined by qRT-PCR. As shown in Fig. 2a, none of NtSuSy1 or NtSuSy2 exhibited an obvious alternation of transcript abundance in three types of transgenic lines as compared with WT. All selected transgenic plants, together with the WT, were grown in the greenhouse until the onset of floral buds, and then were subject to a destructive harvest for analysis and characterization.
Gene expression levels, enzymatic activities, and soluble carbohydrate contents of transgenic lines in comparison with WT. a Quantitative RT-PCR analysis of PsnSuSy1, PsnSuSy2, PsnSuSy1/PsnSuSy2, NtSuSy1, and NtSuSy1 expression levels. The PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic tobacco lines are denoted by PsnSuSy1, PsnSuSy2, and PsnSuSy × PsnSuSy2, respectively, WT refers to wild-type tobacco. NtSuSy1 and NtSuSy1 refer to endogenous SuSy1 and SuSy2 of Nicotiana tabacum. NtACTIN2 was used as an internal control. b SuSy and invertase activities. c Soluble carbohydrate contents included total soluble sugar, fructose, glucose, and sucrose contents. Each error bar represents SD of three biological replicates. Asterisks indicate levels of statistical significance (*P < 0.05, **P < 0.01)
Given the fact that enzyme activity assay is more sensitive for SuSy protein quantification than western blotting in screening SuSy transgenic lines (Xu et al. 2012), we acquired the SuSy activities of transgenic lines through a direct assay that quantified the production of fructose from the breakdown of sucrose. As shown in Fig. 2b, SuSy activities of PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic lines increased by 55.62%, 51.21%, and 60.47% as compared with WT, respectively. However, SuSy activities had no obvious differences among three types of transgenic lines (Supplemental Tables 6, 7). At same time, considering that invertase enzyme is one of two types of enzymes responsible for cleaving sucrose in plants, the activities of invertase, both soluble acid and alkaline invertases (Weber et al. 2005), were also examined. The results revealed that acid invertase activities in PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic lines increased though all changes were not beyond the statistically significant level as compared to WT (Fig. 2b). The alkaline invertase activities were much lower than acid invertase activities (Fig. 2b). Moreover, there were no significant differences in both acid and alkaline invertase activities among three types of transgenic lines (Supplemental Tables 6, 7).
Alternations of soluble carbohydrate contents in transgenic tobacco
To examine the effects on carbohydrate metabolism due to increased SuSy activities in three types of transgenic lines, the sucrose concentrations in PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic lines were measured and were found to decrease 6.56%, 9.83% and 6.34%, respectively (Fig. 2c). It was noteworthy that only the change of sucrose contents in PsnSuSy2’s lines exceeded significant level (Fig. 2c). Accordingly, the fructose contents in the three types of transgenic lines of above order increased significantly by 54.19%, 45.55%, and 55.5%, respectively, while glucose contents increased significantly by 16.9%, 18.87%, and 19.71%, respectively (Fig. 2c). As a result, the total soluble sugar contents significantly increased 42.8%, 35.51%, and 47.57% in the three types of transgenic lines of above order, respectively (Fig. 2c). There were also significantly positive correlations between SuSy activities and any one of total soluble carbohydrate contents, glucose contents, or fructose contents in three types of transgenic lines in above sequence at a significant level of P < 0.01, and R2 values of 0.91, 0.59, and 0.72, respectively (Supplemental Fig. 1A–C). However, there were no statistically significant differences in these above carbohydrate concentrations among three types of transgenic lines (Supplemental Tables 6, 7). These results indicated that the augmented SuSy activities resulting from PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 overexpression accelerated the breakdown metabolism of sucrose and then increased the carbohydrate contents in transgenic lines.
Increased growth and photosynthetic traits of transgenic tobacco
Since transgenic lines had exhibited vigorous growth since seed germination (Fig. 3a–d), we measured the root lengths of T2 seedlings on the tenth days after germination on selective MS medium. The results demonstrated that the root lengths of PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 lines were on overage 32.08%, 21.23%, and 24.15% longer than that of WT, respectively (Fig. 3e). Moreover, the average lengths of fully expanded leaves in the three transgenic lines of the above order increased 21.43%, 40.43% and 22.49%, while the average widths increased 39.45%, 22.26%, and 46.74%, respectively (Fig. 3f). At the onset time of floral buds, the heights of three types of transgenic lines in the above order increased 25.19%, 23.45%, and 25.99%, with no obvious changes in the diameters, respectively (Fig. 3g). In order to examine whether the photosynthesis was affected owning to leaf size enlargement, we measured the chlorophyll contents and photosynthetic rates. The results demonstrated that the chlorophyll contents increased 20.95%, 17.96%, and 25.15%, and photosynthetic rates increased 24.26%, 19.35%, and 28.31% in PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 lines, respectively (Fig. 3h). Furthermore, the three types of transgenic lines in above order increased 10.42%, 6.39%, and 11.7% in fresh weight and 11.34%, 6.88%, and 11.81% in dry weight, respectively (Fig. 3i). However, the differences of these above attributes among three types of transgenic lines were not significant in statistics (Supplemental Tables 6, 7).
Growth phenotypes of transgenic lines in comparison with WT. a–d Morphology of 10-day old roots (a), one-month old phenotype (b), three-month old leaves (c), and three-month old phenotype (d) in three types of transgenic lines and WT. e Root length. f. Leaf length and width. g Height and diameter. h Contents of chlorophyll and photosynthetic rate. i Fresh and dry weights. PsnSuSy1, PsnSuSy2, and PsnSuSy1 × PsnSuSy2 represent PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic tobacco, respectively. WT: wild-type tobacco. Bar in a = 1 cm. Each error bar represents standard deviation (SD) of three biological replicates. Asterisks indicate levels of statistical significance (*P < 0.05, **P < 0.01)
To further confirm that alternations of these traits in the transgenic lines were caused by the increased SuSy activities, we performed correlation analysis between SuSy activities and these traits above. We found that SuSy activities were positively correlated with these traits at the significant level of P < 0.01 levels, with R2 values varying from 0.5 to 0.87 (Supplemental Fig. 2A–H). These results demonstrated that PsnSuSy1 and PsnSuSy1/PsnSuSy2 overexpression could accelerate vegetative growth, enhance biomass accumulation, and improve photosynthesis through increasing SuSy activities in transgenic lines. And, PsnSuSy2 overexpression had similar effects on these traits except biomass accumulation,
Changes of secondary cell wall and fibre characteristics in transgenic tobacco
To examine the effects of PsnSuSy1 and PsnSuSy2 overexpression, and their co-overexpression on secondary cell wall formation, we further scrutinized the ultrastructure and composition of secondary cell wall in the stems of transgenic lines. Examination of scanning electron microscope (SEM) photographs revealed that PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic lines had significantly thicker secondary walls, which increased 17.58%, 15.68%, and 16.43% compared to WT, respectively (Figs. 4a–l, 5a). However, there was no evidence of the presence of cellulose-enriched G-layer in secondary cell wall, a typical characteristic of tension wood, in the three types of transgenic lines (Fig. 4j–l). To test whether the increased secondary cell wall thickness had impact on the stem strength of transgenic lines, we measured the stem breaking forces of all transgenic lines. As shown in Fig. 5b, the stem breaking forces increased significantly by 23.57%, 15.09%, and 20.78% in three types of transgenic lines of above order as compared with WT, respectively. However, the differences of these traits among three types of transgenic lines did not exceed the significant level in statistics (Supplemental Tables 6, 7).
Scanning electron microscope (SEM) images of transgenic lines in comparison with WT. a–d ×1000 magnification SEM. e–h ×3000 magnification SEM. i–l ×5000 magnification SEM. a, e, and i are WT. b, f, and j are PsnSuSy1 transgenic lines. c, g, and k are PsnSuSy2 transgenic lines. d, h, and l are PsnSuSy1 /PsnSuSy2 transgenic lines
Fibre secondary cell wall thickness and stem breaking forces of transgenic lines in comparison with WT. a Fibre secondary cell wall thickness. b Stem breaking force. PsnSuSy1, PsnSuSy2, and PsnSuSy1 × PsnSuSy2 represent PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 overexpression transgenic tobacco, respectively. WT is wild-type tobacco. Each error bar represents standard deviation (SD) of three biological replicates. Asterisks indicate levels of statistical significance (*P < 0.05, **P < 0.01)
Calcofluor and phloroglucinol-HCl were used to stain cellulose and lignin, respectively, to identify which component contributed to the secondary cell wall thickening. The results showed that the deposit of cellulose increased (Fig. 6a–d), whereas the lignin contents slightly decreased in three types of transgenic lines as compared to WT (Fig. 6e–h). Subsequent chemical analysis revealed that the contents of cellulose in PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic lines increased 11.82%, 9.36%, and 11.65%, hemicellulose increased 9.91%, 2.26%, and 8.25%, and lignin decreased 14.11%, 6.53%, and 15.58%, respectively (Fig. 6i). All these changes were beyond significant level except PsnSuSy2’s changes in hemicellulose and lignin contents. In order to know whether the cellulose crystallinity of transgenic lines was also impacted along with the increase of cellulose content, we further measured CrI that accounts for cellulose crystallinity using X-ray detection method (Tanaka et al. 2003). As shown in Fig. 6j, three types of transgenic lines had 10.7%, 9.63%, and 9.18% lower CrI values than WT, all beyond significant level. Moreover, we also found that the average lengths of fibres in three types of transgenic lines were about 12.39%, 11.16%, and 12.06% longer than that in WT (Fig. 6k), whereas the fibre width had no significant changes (Fig. 6k). Although most traits in above of three types of transgenic lines exhibited significant changes as compared with WT, the differences among themselves were not beyond the significant level (Supplemental Tables 6, 7). Furthermore, we examined the relationships between the SuSy activities and these traits with significant difference between the transgenic lines and WT through correlation analysis. The results demonstrated that SuSy activities were negatively or positively correlated with these traits at significant level P < 0.05 or 0.01, with the R2 values varying from 0.48 to 0.74, respectively (Supplemental Fig. 3A–G), which indicated that the alternations of above traits are corresponding to the higher SuSy activities engendered by PsnSuSy1, PsnSuSy2, or PsnSuSy1/PsnSuSy2 overexpression.
Cell wall compositions, secondary cell wall thickness and fibre characteristics of transgenic lines in comparison with WT. a–d Cellulose in stem sections stained with Calcofluor White (blue color). e–h Lignin (red color) in stem sections stained with Phloroglucinol-HCl. i Contents of cellulose, hemicellulose, and lignin. j Microcrystalline cellulose. k Fibre lengths and widths. PsnSuSy1, PsnSuSy2, and PsnSuSy1 × PsnSuSy2 represent to PsnSuSy1, PsnSuSy2, and dual PsnSuSy1/PsnSuSy2 transgenic tobacco, respectively. WT is wild-type tobacco. Each error bar represents standard deviation (SD) of three biological replicates. Asterisks indicate levels of statistical significance (*P < 0.05, **P < 0.01)
Alternations of gene expression in transgenic tobacco
To gain insight into the molecular events associated with the anatomical and compositional changes of secondary growth in transgenic lines, we measured the expression levels of genes related to these traits using qRT-PCR. The results demonstrated that the expression levels of genes involved in the biosynthesis of secondary wall components, including cellulose (CesA4, CesA7, and CesA8) and hemicellulose (FRA8, IRX9, and IRX10), were significantly up-regulated in PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic lines compared to those in WT (Fig. 7a). In contrast, the expression levels of lignin biosynthetic genes, such as 4CL1, PAL1, C3H1, CCoAOMT1, CCR1, C4H, CAD4, and CAD6, showed significant reductions (Fig. 7a). The genes involved in the fibre cell expansion and elongation, including ExpansinA, ExpansinB, TIP1;3, TIP1; 4, XTH5, and XTH8, were significantly up-regulated, whereas the genes including XSP1, XCP2, SCPL45, and SCPL49 that are involved in programmed cell death were notably down-regulated (Fig. 7b). In addition, GH9B1, GH9B3, and GH9B16, whose homologies in rice are involve in reducing cellulose crystallinity through increasing hemicellulosic arabinose (Xie et al. 2013; Li et al. 2015), were significantly up-regulated in three types of transgenic lines compared with WT (Fig. 7c). Above changes in pathway genes, regardless of up/down-regulation, are aligned well with the alternations of secondary cell wall and fibre traits. However, the comparisons of expression levels of these genes among three types of transgenic lines indicated that the differences did not exceed the significant level in statistics (Supplemental Tables 6, 7). These results suggested that the effects of PsnSuSy1, PsnSuSy2 overexpression, and their co-overexpression could affect expressions of related genes, leading to change secondary cell wall and fibre traits of transgenic lines.
Expression levels of genes involved in secondary cell wall and fibre formation of transgenic lines in comparison with WT. a Expression levels of genes that are involved in the biosynthesis of cellulose (CESA4, CESA7, and CESA8), hemicellulose (IRX8, IRX9, and IRX10), and lignin (4CL1, PAL1, C3H1, CCoAOMT1, CCR1, C4H, CAD4, and CAD6). b Expression levels of genes that are involved in cell expansion (ExpansinA, ExpansinB, TIP1;3, TIP1;4, XTH5, and XTH8) and programmed cell death (XSP1, XCP2, SCPL45, and SCPL49). c Expression levels of genes that are involved in cellulose crystallinity (GH9B1, GH9B3, and GH9B16). PsnSuSy1, PsnSuSy2, and PsnSuSy1 × PsnSuSy2 refer to PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 transgenic tobacco, respectively. WT refers to wild-type tobacco. NtACTIN2 was used as an internal control. The expression of level of each gene in the WT was set to 1. Each error bar represents standard deviation (SD) of three biological replicates. Asterisks indicate levels of statistical significance (*P < 0.05, **P < 0.01)
Discussion
In this study, we substantiated that overexpression of two poplar SuSy genes, PsnSuSy1 and PsnSuSy2, had significant impacts on fibre formation, secondary cell wall thickening, vegetative growth and development, and mechanical strength. We also for the first time generated dual transgenic tobacco lines of two SuSy genes, namely PsnSuSy1 and PsnSuSy2, and found their effects on above traits was perceptibly additive though not substantial. These effects can be directly attributed to the much higher SuSy activities engendered by PsnSuSy1, PsnSuSy2 overexpression, and their co-overexpression. The significance and implications of the findings are discussed and elaborated below.
The tissue-specific expression patterns of PsnSuSy1 and PsnSuSy2 were similar in all tissues we tested (Fig. 1b), supporting they might function similarly to enhance the plasticity of sucrolysis in poplar. However, the expression levels of PsnSuSy1 in secondary tissues were higher than those of PsnSuSy2 (Fig. 1b), indicating that PsnSuSy1 may play a greater role in sucrolysis than PsnSuSy2 in term of poplar wood formation. Both PsnSuSy1 and PsnSuSy2 proteins were found to exist simultaneously in cytosol and plasma membrane of onion epidermal cells (Fig. 1c), which is consistent with the previous conclusion that SuSy proteins exist in both soluble and particulate forms (Carlson and Chourey 1996; Komina et al. 2002). Moreover, the different phosphorylation sites between PsnSuSy1 and PsnSuSy2 proteins may lead to little differences in their subcellular compartment distribution, enzymatical activity, and involved biochemical processes, which may be worthy of further investigation in the future.
It was noteworthy that the PsnSuSy1/PsnSuSy2 dual transgenic lines did not exhibit significant increases in the transcript abundances and SuSy activities compared to either PsnSuSy1 or PsnSuSy2 transgenic lines (Fig. 2a, b), which may suggest there might exist a mechanism to exert some constraints on transcripts and enzyme activities of exogenous PsnSuSy1 and PsnSuSy2. Meanwhile, the transcript levels of native NtSuSy1 and NtSuSy2 in PsnSuSy1, PsnSuSy2, PsnSuSy1/PsnSuSy2 transgenic lines were not significantly affected by PsnSuSy1 and PsnSuSy2 overexpression, and their co-overexpression as well (Fig. 2a). Moreover, the alkaline and neutral invertase activities only exhibited a little increase compared with WT (Fig. 2b). Thus, we can preliminarily ascribe the alternations of above-mentioned traits in all three types of transgenic lines to exogenous PsnSuSy1’s and PsnSuSy2’s individual overexpression or their co-overexpression.
The significant increases of total soluble carbohydrate, fructose, and glucose arising from the overexpression of PsnSuSy1, PsnSuSy2 or both genes in tobacco transgenic lines were found to have significantly positive correlations with the increased SuSy activities, with the sucrose to be an exception (Supplemental Fig. 1A–C). Previous studies also reported that the elevated concentration of total soluble carbohydrate in SuSy overexpression plants is primarily caused by the increased fructose contents (Coleman et al. 2006, 2009). On the contrary, when SuSys are suppressed, there is a build-up of sucrose that is accompanied with a concurrent reduction in both glucose and fructose concentrations, for instance, in potatoes (Zrenner et al. 1995), carrots (Tang and Sturm 1999), and tomatoes (D’Aoust et al. 1999). It is noteworthy that the sucrose concentrations in three types of transgenic lines exhibited only a small decrease (Fig. 2c), which suggests that although an increase of SuSy activities in the sink tissues augmented sucrose degradation into fructose and glucose, sucrose was possibly transported quickly from source tissues and supplemented to sinks, making the reduction of sucrose reduction less perceivable (Fig. 3f, h). The slightly higher soluble carbohydrate contents in PsnSuSy1 and PsnSuSy2 dual transgenic lines support that PsnSuSy1 and PsnSuSy2 had similar functions and their effects was additive to a certain degree when they catalyzed sucrose degradation and determined the availability of carbohydrates in sink tissues. The small additive effect of two genes could also be caused by the constraint on the cellular levels of SuSy activities, and needless to mention that the reversible reactions of sucrose degradation and synthesis if the two genes performed the same function at the subcellular sites.
Conspicuous phenotypical and physiological changes, such as longer roots, increased heights, larger leaf sizes, increased chlorophyll contents, augmented photosynthesis, and improved biomass in three types of transgenic lines (Fig. 3a–i) and their correlations with SuSy activities (Supplemental Fig. 2A–G), are largely in agreement with the results obtained in other species, which include the increased heights, leaf areas, photosynthesis rate, and biomass (Hayashi et al. 2004; Coleman et al. 2006; Jiang et al. 2012; Xu et al. 2012; Poovaiah et al. 2015; Goren et al. 2017). In addition to the change of sink strength and the supply of photoassimilate (Tang and Sturm 1999), which can promote growth and biomass accumulation directly, recent studies also showed that UDPG can act as an extracellular signaling molecule to stimulate growth and biomass accumulation though the underlying mechanism is currently unclear (Janse van Rensburg and Van den Ende 2017). However, not all SuSys have the same functions as conflicting results on growth and biomass accumulation of some SuSys overexpression plants have been reported (Coleman et al. 2006, 2009; Bieniawska et al. 2007; Barratt et al. 2009). In current study, PsnSuSy2 transgenic lines exhibited no obvious increases of fresh and dry weights compared with WT though it had significant increased heights (Fig. 3i, g). These differences may be ascribed to the innate differences in mechanism of carbon translocation between herbaceous and woody plants (Burkle et al. 1998; Turgeon and Medville 1998) or functional differences among different SuSys. The novel finding of this study is the substantial effects on root length (Fig. 3e), which suggests that the increased SuSy activities and/or the subsequent formed UDPG might stimulate cell division and expansion, which in turn led to the lengthened roots similar to what was reported in GhVIN1 overexpression Arabidopsis (Wang et al. 2010).
The effects of PsnSuSy1, PsnSuSy2, and PsnSuSy1/PsnSuSy2 overexprssion on the cellulose deposition were reflected by a significant increase in the cellulose content and secondary cell wall thickenss (Figs. 5a, 6i), which is congruent to the results from previous studies of Gossypium hirsutum (Coleman et al. 2009) and rice (Fan et al. 2017). It is also noteworthy to mention that PsnSuSy1 and PsnSuSy1/PsnSuSy2 transgenic lines, but not PsnSuSy2 lines, showed a significant increase in hemicellulose contents and a significant decrease in the lignin contents when compared with WT (Fig. 6i). Given the lower expression levels but higher soluble carbohydrate contents and SuSy activities in PsnSuSy1’s transgenic lines than PsnSuSy2’s (Fig. 2a–c), we specutulate that the translation efficiencies of PsnSuSy1, PsnSuSy2 mRNAs were differentially different and/or their proteins had different SuSy activities. Correlation analysis further proved that SuSy activites were postively correlated with cellulose and hemicelluose contents but negatively correlated with lignin contents (Supplemental Fig. 3C–E). These results demonstrated that the increased SuSy activites in transgenic lines could allocate more carbon elements to both cellulose and hemicellulose synthesis, but much less for lignin synthesis. It is possible that the increased SuSy activities supplied more UDPG directly to cellulose synthesis, and more carbohydrate substrates to hemicellulose and many other non-cellulose cell wall component biosynthesis, as previously reported (Salnikov et al. 2001; Ruan et al. 2003; Coleman et al. 2009; Fujii et al. 2010).
Relationship of concodance between breakinig forces (Fig. 5b) and cellulose and hemicellulose contents as well as secondary cell wall thickness was corroborated by the correlation analysis (Supplemental Fig. 4A–C). This is consistent with previous conclusions that increased SuSy activities enhance cell wall thickness, cellulose content, and biomass yield, which in turn increase mechanical strength (Somerville 2006; Li et al. 2015; Fan et al. 2017, 2018). Additionally, we found that the fibre lengths were significantly elongated (Fig. 6k) in the way as reported in previous studies (Ruan and Chourey 1998; Ruan et al. 2005; Xu et al. 2012), and were also highly correlated with SuSy activies (Supplemental Fig. 3G). We spectulate that the increased soluble carbehydrate concentrations engendered by PsnSuSy1 and PsnSuSy2 overexpression decreased solute potentials, which then augmented water influx and turgor pressure, eventually giving rise to fibre elongation. However, this speculation fails to explain why fibre widths exhibated no signficant alternations in three types of transgenic lines as compared to WT. The strongly negative correlation between cellulose crystallinity and SuSy activities was observed (Supplemental Fig. 3F), which is in agreement with what was described for OsSUS3 overexpression transgenic rice (Fan et al. 2017), but was contrary to the change of cell wall crystallinity in GhSuSy poplar overexpression transgenic lines (Coleman et al. 2009). The different changing trends of cellulose crystallinity in various SuSs overexpression transgenic plants need to be further studied.
The expression levels of genes involved in cellulose, hemicellulose, and lignin biosynthesis, were significantly altered in transgenic lines (Fig. 7a), which were also concordant with the changes of secondary wall component contents (Fig. 6i). Needless to say, the expression levels of genes related to cell expansion and elongation of transgenic lines significantly increased, whereas the expression levels of genes about programmed cell death significantly decreased (Fig. 7b). It is need to be further studied that the alternations of these genes increased fibre lengths without influencing the fibre widths of transgenic lines (Fig. 6k). In addition, the increased expression levels of GH9B1, GH9B3, and GH9B, which is also observed in OsSUS3 rice (Fan et al. 2017), were a plausible explanation for cellulose crystallinity reductions in three types of transgenic lines. It has been revealed that the phosphoresced hexose originated from glucose can be readily phosphoresced by hexokinase, and then acts as important sugar signalling molecular to regulate genes expression (Gibson et al. 2006; Rolland et al. 2006). Recent study further have showed that UDPG can also acts as an extracellular signaling molecule to stimulate growth and biomass accumulation (Janse van Rensburg and Van den Ende 2017). These could be the molecular mechanisms underling the alternations of phenotypes of three types of transgenic lines.
Conclusions
The functional similarities and differences of PsnSuSy1 and PsnSuSy2 in plant growth and development were revealed and corroborated by using PsnSuSy1, PsnSuSy2, and PsnSuSy1 /PsnSuSy2 dual tobacco transgenic lines, which manifested significant and similar changes in both vegetative growth and secondary growth. These changes were caused directly by the increased SuSy activities, which modulated carbohydrate contents, carbon partition, and sink strength, and consequently, resulted in many salient changes such as the thickening cell wall and augmented mechanical strength. At the same time, the UDPG engendered by SuSy catalysis may act as a long-range extracellular signaling molecule to promote overall plants growth and development. Compared to either the PsnSuSy1 or PsnSuSy2 transgenic lines, PsnSuSy1/PsnSuSy2 dual transgenic lines displayed an insubstantial increase of SuSy activities and minor additive effects on some attributes and traits, which suggested that these two genes played largely the same but little distinct roles. Therefore, we concluded that the two genes function to augment the plasticity of secondary growth in P. simonii × P. nigra.
Change history
25 April 2023
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1007/s11103-023-01353-5
References
Amor Y, Haigler CH, Johnson S, Waincott M, Delmer PD (1995) A membrane-associated form of sucrose synthase and its potential role in synthesis of cellulose. Proc Natl Acad Sci USA 26(20):9353–9357
An X, Chen Z, Wang J, Ye M, Ji L, Wang J, Liao W, Ma H (2014) Identification and characterization of the Populus sucrose synthase gene family. Gene 539(1):58–67
Andersson-Gunneras S, Mellerowicz EJ, Love J, Segerman B, Ohmiya Y, Coutinho PM, Nilsson P, Henrissat B, Moritz T, Sundberg B (2006) Biosynthesis of cellulose-enriched tension wood in Populus: global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis. Plant J 45(2):144–165
Appenzeller L, Doblin M, Barreiro R, Wang HY, Niu XM, Kollipara K, Carrigan L, Tomes D, Chapman M, Dhugga KS (2004) Cellulose synthesis in maize: isolation and expression analysis of the cellulose synthase (CesA) gene family. Cellulose 11(3–4):287–299
Asano T, Kunieda N, Omura Y, Ibe H, Kawasaki T, Takano M, Sato M, Furuhashi H, Mujin T, Takaiwa F, Wu Cy CY, Tada Y, Satozawa T, Sakamoto M, Shimada H (2002) Rice SPK, a calmodulin-like domain protein kinase, is required for storage product accumulation during seed development: phosphorylation of sucrose synthase is a possible factor. Plant cell 14(3):619–628
Aspeborg H, Schrader J, Coutinho PM, Stam M, Kallas A, Djerbi S, Nilsson P, Denman S, Amini B, Sterky F, Master E, Sandberg G, Mellerowicz E, Sundberg B, Henrissat B, Teeri TT (2005) Carbohydrate-active enzymes involved in the secondary cell wall biogenesis in hybrid aspen. Plant Physiol 137(3):983–997
Baroja-Fernandez E, Munoz FJ, Montero M, Etxeberria E, Sesma MT, Ovecka M, Bahaji A, Ezquer I, Li J, Prat S, Pozueta-Romero J (2009) Enhancing sucrose synthase activity in transgenic potato (Solanum tuberosum L.) tubers results in increased levels of starch, ADPglucose and UDPglucose and total yield. Plant Cell Physiol 50(9):1651–1662
Barratt DH, Derbyshire P, Findlay K, Pike M, Wellner N, Lunn J, Feil R, Simpson C, Maule AJ, Smith AM (2009) Normal growth of Arabidopsis requires cytosolic invertase but not sucrose synthase. Proc Natl Acad Sci USA 106(31):13124–13129
Baud S, Vaultier MN, Rochat C (2004) Structure and expression profile of the sucrose synthase multigene family in Arabidopsis. J Exp Bot 55(396):397–409
Bieniawska Z, Paul Barratt D, Garlick AP, Thole V, Kruger NJ, Martin C, Zrenner R, Smith AM (2007) Analysis of the sucrose synthase gene family in Arabidopsis. Plant J 49(5):810–828
Burkle L, Hibberd JM, Quick WP, Kuhn C, Hirner B, Frommer WB (1998) The H+-sucrose cotransporter NtSUT1 is essential for sugar export from tobacco leaves. Plant Physiol 118(1):59–68
Carlson SJ, Chourey PS (1996) Evidence for plasma membrane-associated forms of sucrose synthase in maize. Mol Gen Genet 252(3):303–310
Chen YC, Chourey PS (1989) Spatial and temporal expression of the two sucrose synthase genes in maize: immunohistological evidence. Theor Appl Genet 78(4):553–559
Chourey PS (1981) Genetic control of sucrose synthetase in maize endosperm. Mol Gen Genet 184(3):372–376
Coleman HD, Ellis DD, Gilbert M, Mansfield SD (2006) Up-regulation of sucrose synthase and UDP-glucose pyrophosphorylase impacts plant growth and metabolism. Plant Biotechnol J 4(1):87–101
Coleman HD, Yan J, Mansfield SD (2009) Sucrose synthase affects carbon partitioning to increase cellulose production and altered cell wall ultrastructure. Proc Natl Acad Sci USA 106(31):13118–13123
D’Aoust M-A, Yelle S, Nguyen-Quoc B (1999) Antisense inhibition of tomato fruit sucrose synthase decreases fruit setting and the sucrose unloading capacity of young fruit. Plant cell 11(12):2407–2418
Dhugga KS (2007) Maize biomass yield and composition for biofuels. Crop Sci 47(6):2211–2227
Fan C, Feng S, Huang J, Wang Y, Wu L, Li X, Wang L, Tu Y, Xia T, Li J, Cai X, Peng L (2017) AtCesA8-driven OsSUS3 expression leads to largely enhanced biomass saccharification and lodging resistance by distinctively altering lignocellulose features in rice. Biotechnol Biofuels 10:221
Fan C, Li Y, Hu Z, Hu H, Wang G, Li A, Wang Y, Tu Y, Xia T, Peng L, Feng S (2018) Ectopic expression of a novel OsExtensin-like gene consistently enhances plant lodging resistance by regulating cell elongation and cell wall thickening in rice. Plant Biotechnol J 16(1):254–263
Fu H, Park WD (1995) Sink- and vascular-associated sucrose synthase functions are encoded by different gene classes in potato. Plant cell 7(9):1369–1385
Fujii S, Hayashi T, Mizuno K (2010) Sucrose synthase is an integral component of the cellulose synthesis machinery. Plant Cell Physiol 51(2):294–301
Geigenberger P, Stitt M (1993) Sucrose synthase catalyses a readily reversible reaction in vivo in developing potato tubers and other plant tissues. Planta 189(3):329–339
Gerber L, Zhang B, Roach M, Rende U, Gorzsas A, Kumar M, Burgert I, Niittyla T, Sundberg B (2014) Deficient sucrose synthase activity in developing wood does not specifically affect cellulose biosynthesis, but causes an overall decrease in cell wall polymers. New Phytol 203(4):1220–1230
Gibson TB, Lawrence MC, Gibson CJ, Vanderbilt CA, McGlynn K, Arnette D, Chen W, Collins J, Naziruddin B, Levy MF, Ehrlich BE, Cobb MH (2006) Inhibition of glucose-stimulated activation of extracellular signal-regulated protein kinases 1 and 2 by epinephrine in pancreatic beta-cells. Diabetes 55(4):1066–1073
Goren S, Lugassi N, Stein O, Yeselson Y, Schaffer AA, David-Schwartz R, Granot D (2017) Suppression of sucrose synthase affects auxin signaling and leaf morphology in tomato. PLoS ONE 12(8):e0182334
Hayashi T, Ohmiya Y, Konishi T (2004) Evidence that sucrose loaded into the phloem of a poplar leaf is used directly by sucrose synthase associated with xyloglucan synthase in the stem. Am Chem Soc 227:U292–U292
Heim U, Weber H, Baumlein H, Wobus U (1993) A sucrose-synthase gene of Vicia faba L.: expression pattern in developing seeds in relation to starch synthesis and metabolic regulation. Planta 191(3):394–401
Hirose T, Scofield GN, Terao T (2008) An expression analysis profile for the entire sucrose synthase gene family in rice. Plant Sci 174(5):534–543
Huber SC, Huber JL, Liao PC, Gage DA, McMichael RW Jr, Chourey PS, Hannah LC, Koch K (1996) Phosphorylation of serine-15 of maize leaf sucrose synthase. Occurrence in vivo and possible regulatory significance. Plant Physiol 112(2):793–802
Janse van Rensburg HC, Van den Ende W (2017) UDP-glucose: a potential signaling molecule in plants? Front Plant Sci 8:2230
Jiang Y, Guo W, Zhu H, Ruan YL, Zhang T (2012) Overexpression of GhSusA1 increases plant biomass and improves cotton fiber yield and quality. Plant Biotechnol J 10(3):301–312
Jourez B, Riboux A, Leclercq A (2001) Anatomical characteristics of tension wood and opposite wood in young inclined stems of poplar (Populus euramericana cvGhoy’). Iawa J 22(2):133–158
Kennedy A, Frank RN, Varma SD (1983) Aldose reductase activity in retinal and cerebral microvessels and cultured vascular cells. Invest Ophthalmol Vis Sci 24(9):1250–1258
King SP, Lunn JE, Furbank RT (1997) Carbohydrate content and enzyme metabolism in developing canola siliques. Plant Physiol 114(1):153–160
Kolosova N, Miller B, Ralph S, Ellis BE, Douglas C, Ritland K, Bohlmann J (2004) Isolation of high-quality RNA from gymnosperm and angiosperm trees. Biotechniques 2:353–359
Komatsu A, Moriguchi T, Koyama K, Omura M, Akihama T (2002) Analysis of sucrose synthase genes in citrus suggests different roles and phylogenetic relationships. J Exp Bot 53(366):61–71
Komina O, Zhou Y, Sarath G, Chollet R (2002) In vivo and in vitro phosphorylation of membrane and soluble forms of soybean nodule sucrose synthase. Plant Physiol 129(4):1664–1673
Konishi T, Ohmiya Y, Hayashi T (2004) Evidence that sucrose loaded into the phloem of a poplar leaf is used directly by sucrose synthase associated with various β-glucan synthases in the stem. Plant Physiol 134(3):1146–1152
Li J, Baroja-Fernandez E, Bahaji A, Munoz FJ, Ovecka M, Montero M, Sesma MT, Alonso-Casajus N, Almagro G, Sanchez-Lopez AM, Hidalgo M, Zamarbide M, Pozueta-Romero J (2013) Enhancing sucrose synthase activity results in increased levels of starch and ADP-glucose in maize (Zea mays L.) seed endosperms. Plant Cell Physiol 54(2):282–294
Li F, Zhang M, Guo K, Hu Z, Zhang R, Feng Y, Yi X, Zou W, Wang L, Wu C, Tian J, Lu T, Xie G, Peng L (2015) High-level hemicellulosic arabinose predominately affects lignocellulose crystallinity for genetically enhancing both plant lodging resistance and biomass enzymatic digestibility in rice mutants. Plant Biotechnol J 13(4):514–525
Liao Z, Chen M, Guo L, Gong Y, Tang F, Sun X, Tang K (2004) Rapid isolation of high-quality total RNA from taxus and ginkgo. Prep Biochem Biotechnol 34(3):209–214
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402–408
Lunn JE, Hatch MD (1995) Primary partitioning and storage of photosynthate in sucrose and starch in leaves of C4 plants. Planta 197:385–391
Nilsson J, Karlberg A, Antti H, Lopez-Vernaza M, Mellerowicz E, Perrot-Rechenmann C, Sandberg G, Bhalerao RP (2008) Dissecting the molecular basis of the regulation of wood formation by auxin in hybrid aspen. Plant cell 20(4):843–855
Pilate G, Dejardin A, Laurans F, Leple JC (2004) Tension wood as a model for functional genomics of wood formation. New Phytol 164(1):63–72
Plavcova L, Hacke UG, Almeida-Rodriguez AM, Li E, Douglas CJ (2013) Gene expression patterns underlying changes in xylem structure and function in response to increased nitrogen availability in hybrid poplar. Plant Cell Environ 36(1):186–199
Poovaiah CR, Mazarei M, Decker SR, Turner GB, Sykes RW, Davis MF, Stewart CN Jr (2015) Transgenic switchgrass (Panicum virgatum L.) biomass is increased by overexpression of switchgrass sucrose synthase (PvSUS1). Biotechnol J 10(4):552–563
Qiu Z, Wang L, Zhou Q (2013) Effects of bisphenol A on growth, photosynthesis and chlorophyll fluorescence in above-ground organs of soybean seedlings. Chemosphere 90(3):1274–1280
Raes J, Rohde A, Christensen JH, Van de Peer Y, Boerjan W (2003) Genome-wide characterization of the lignification toolbox in Arabidopsis. Plant Physiol 133(3):1051–1071
Rolland F, Baena-Gonzalez E, Sheen J (2006) Sugar sensing and signaling in plants: conserved and novel mechanisms. Annu Rev Plant Biol 57:675–709
Ruan YL, Chourey PS (1998) A fiberless seed mutation in cotton is associated with lack of fiber cell initiation in ovule epidermis and alterations in sucrose synthase expression and carbon partitioning in developing seeds. Plant Physiol 118(2):399–406
Ruan YL, Llewellyn DJ, Furbank RT (2003) Suppression of sucrose synthase gene expression represses cotton fiber cell initiation, elongation, and seed development. Plant cell 15(4):952–964
Ruan YL, Llewellyn DJ, Furbank RT, Chourey PS (2005) The delayed initiation and slow elongation of fuzz-like short fibre cells in relation to altered patterns of sucrose synthase expression and plasmodesmata gating in a lintless mutant of cotton. J Exp Bot 56(413):977–984
Salnikov VV, Grimson MJ, Delmer DP, Haigler CH (2001) Sucrose synthase localizes to cellulose synthesis sites in tracheary elements. Phytochemistry 57(6):823–833
Schroeder MR, Borkhsenious ON, Matsuoka K, Nakamura K, Raikhel NV (1993) Colocalization of barley lectin and sporamin in vacuoles of transgenic tobacco plants. Plant Physiol 101(2):451–458
Somerville C (2006) Cellulose synthesis in higher plants. Ann Rev Cell Dev Biol 22:53–78
Sturm A, Lienhard S, Schatt S, Hardegger M (1999) Tissue-specific expression of two genes for sucrose synthase in carrot (Daucus carota L.). Plant Mol Biol 39(2):349–360
Tanaka K, Murata K, Yamazaki M, Onosato K, Miyao A, Hirochika H (2003) Three distinct rice cellulose synthase catalytic subunit genes required for cellulose synthesis in the secondary wall. Plant Physiol 133(1):73–83
Tang G-Q, Sturm A (1999) Antisense repression of sucrose synthase in carrot (Daucus carota L.) affects growth rather than sucrose partitioning. Plant Mol Biol 41(4):465–479
Turgeon R, Medville R (1998) The absence of phloem loading in willow leaves. Proc Natl Acad Sci USA 95(20):12055–12060
Wai CM, Zhang J, Jones TC, Nagai C, Ming R (2017) Cell wall metabolism and hexose allocation contribute to biomass accumulation in high yielding extreme segregants of a Saccharum interspecific F2 population. BMC Genom 18(1):773
Wang L, Li XR, Lian H, Ni DA, He YK, Chen XY, Ruan YL (2010) Evidence that high activity of vacuolar invertase is required for cotton fiber and Arabidopsis root elongation through osmotic dependent and independent pathways, respectively. Plant Physiol 154(2):744–756
Weber H, Borisjuk L, Wobus U (2005) Molecular physiology of legume seed development. Annu Rev Plant Biol 56:253–279
Wei Z, Qu Z, Zhang L, Zhao S, Bi Z, Ji X, Wang X, Wei H (2015) Overexpression of poplar xylem sucrose synthase in tobacco leads to a thickened cell wall and increased height. PLoS ONE 10(3):e0120669
Winter H, Huber SC (2000) Regulation of sucrose metabolism in higher plants: localization and regulation of activity of key enzymes. Crit Rev Plant Sci 19(1):31–67
Winter H, Huber JL, Huber SC (1997) Membrane association of sucrose synthase: changes during the graviresponse and possible control by protein phosphorylation. FEBS Lett 420(2):151–155
Wu AM, Hornblad E, Voxeur A, Gerber L, Rihouey C, Lerouge P, Marchant A (2010) Analysis of the Arabidopsis IRX9/IRX9-L and IRX14/IRX14-L pairs of glycosyltransferase genes reveals critical contributions to biosynthesis of the hemicellulose glucuronoxylan. Plant Physiol 153(2):542–554
Xie G, Yang B, Xu Z, Li F, Guo K, Zhang M, Wang L, Zou W, Wang Y, Peng L (2013) Global identification of multiple OsGH9 family members and their involvement in cellulose crystallinity modification in rice. PLoS ONE 8(1):e50171
Xu SM, Brill E, Llewellyn DJ, Furbank RT, Ruan YL (2012) Overexpression of a potato sucrose synthase gene in cotton accelerates leaf expansion, reduces seed abortion, and enhances fiber production. Mol Plant 5(2):430–441
Zhang Q, Berggren PO, Tally M (1997) Glucose increases both the plasma membrane number and phosphorylation of insulin-like growth factor II/mannose 6-phosphate receptors. J Biol Chem 272(38):23703–23706
Zhang DQ, Xu BH, Yang XH, Zhang ZY, Li BL (2011) The sucrose synthase gene family in Populus: structure, expression, and evolution. Tree Genet Genomes 7(3):443–456
Zrenner R, Salanoubat M, Willmitzer L, Sonnewald U (1995) Evidence of the crucial role of sucrose synthase for sink strength using transgenic potato plants (Solanum tuberosum L.). Plant J 7(1):97–107
Funding
The work was funded by the National Nature Science Fund of China (31770640), the 111 Project (B16010), the Foundation Research Fund for the Central Universities of China (No.2572016EAJ2), and Returned Overseas Researcher Fund of Heilongjiang Province (LC2015012).
Author information
Authors and Affiliations
Contributions
ML and S Finished most of experiment and measurements; YY and MX participated in rising seeding of transgenic lines; Y participated in partly SEM analysis; LL participated in genes expression pattern of poplar; TT participated in Histochemical staining; HW: performed data analysis and wrote manuscript; ZG designed the experiments, performed data analysis, and wrote manuscript; All the authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been retracted. Please see the retraction notice for more detail: https://doi.org/10.1007/s11103-023-01353-5
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, M., Wang, S., Liu, Y. et al. RETRACTED ARTICLE: Overexpression of PsnSuSy1, 2 genes enhances secondary cell wall thickening, vegetative growth, and mechanical strength in transgenic tobacco. Plant Mol Biol 100, 215–230 (2019). https://doi.org/10.1007/s11103-019-00850-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-019-00850-w