Abstract
This review examines the symbiotic, evolutionary, proteomic and genetic basis for a group of fungi that occupy a specialized niche as insect pathogens as well as endophytes. We focus primarily on species in the genera Metarhizium and Beauveria, traditionally recognized as insect pathogenic fungi but are also found as plant symbionts. Phylogenetic evidence suggests that these fungi are more closely related to grass endophytes and diverged from that lineage ca. 100 MYA. We explore how the dual life cycles of these fungi as insect pathogens and endophytes are coupled. We discuss the evolution of insect pathogenesis while maintaining an endophytic lifestyle and provide examples of genes that may be involved in the transition toward insect pathogenicity. That is, some genes for insect pathogenesis may have been co-opted from genes involved in endophytic colonization. Other genes may be multifunctional and serve in both lifestyle capacities. We suggest that their evolution as insect pathogens allowed them to effectively barter a specialized nitrogen source (i.e. insects) with host plants for photosynthate. These ubiquitous fungi may play an important role as plant growth promoters and have a potential reservoir of secondary metabolites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In terms of ecological interactions and evolutionary history, fungi that infect and kill insects are plainly fascinating. Most infect the host insect by transgressing the cuticle, that is, they do not have to be ingested to cause infection in the insect. This allows infection of insects with sucking mouthparts such as aphids (Chandler 1997) and adult mosquitoes (Blanford et al. 2005). The breadth of variety of insect pathogenic fungi is tremendous, a group that comprises over 100 fungal species. There are examples of insect pathogenic fungi found in most major fungal taxonomic groups from chytridiomycetes to basidiomycetes. However, there is no underlying phylogenetic relationship that suggests a basal group from which all insect pathogenic fungi arose. Even within the Hypocreales, the taxonomic group with the largest number of insect pathogenic fungi, there is little evidence to suggest that there is a singular common origin (Humber 2008). Some of these insect pathogenic fungi are obligate pathogens while many are facultative. Adding further intrigue into their ecology is a subset of insect pathogenic fungi that additionally function as endophytic symbionts of plants.
Two genera of insect pathogenic fungi that fall within the category of endophytes are Metarhizium and Beauveria. There is considerable divergence within the genus Metarhizium and some species (e.g. M. acridum) have restricted insect host ranges while other species (e.g. M. robertsii) have broad host ranges, and not all species show equivalent endophytic capabilities. The potential of Metarhizium and Beauveria to control insect pests in agroecosystems has been known since the early 20th century (Madelin 1963), and numerous formulations of Metarhizium and Beauveria have been approved for use in crop protection (Faria and Wraight 2007; Castrillo et al. 2011).
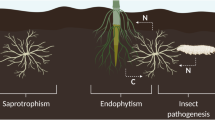
Phylogenetic analysis has shown that Metarhizium and Beauveria are related to the fungal grass endosymbionts Claviceps and Epichloë (Spatafora et al. 2007). Furthermore, comparative genomic analyses have shown that Metarhizium spp. are more closely related to endophytes and plant pathogens than to animal pathogens (Gao et al. 2011), and that Metarhizium lineages diverged from the lineage of the mutualistic plant endophyte Epichloë festucae approximately 88–114 MYA (Gao et al. 2011). This strongly suggests that Metarhizium evolved from fungi that were plant associates and that insect pathogenicity is a more recently acquired adaptation (Fig. 1). Metarhizium may have evolved from a plant symbiont lineage and subsequently acquired the ability to infect and kill insects. This is supported by genomic analysis that shows a large number of genes for plant degrading enzymes within Metarhizium genomes (Gao et al. 2011).
Genes involved in insect pathogenicity may have been co-opted from genes involved in plant colonization or from horizontal gene transfer (Screen and St. Leger 2000). Figure 2 shows hypothetical mechanisms by which genes involved in insect pathogenesis may have been co-opted, evolved, or acquired by horizontal gene transfer from a plant-associated fungus. The evolution of insect pathogenic Metarhizium spp. must have involved adaptations that enabled degradation of insect cuticle and host body components, as there exists numerous proteases, lipases, and chitinases within the Metarhizium and Beauveria genome. Furthermore, comparative analysis of plant and insect adhesin genes in Metarhizium spp. suggest that, while other abiotic and biotic factors cannot be excluded in contributing to divergence within the genus, plant relationships, rather than insect host, have been a driving factor in the divergence of the genus Metarhizium (Wyrebek and Bidochka 2013).
Potential model for the evolution (from top to bottom) of genetic events that lead to multifunctional lifestyles of EIPF. The accumulation of plant pathogenic genes potentially allowed for the root colonizing ability of saprotrophs; selection pressure could have then pushed plant pathogens toward symbiosis with the plant host, ultimately resulting in a mutualistic relationship; symbiotic pressure from the plant to receive nutrients (e.g. nitrogen) from the fungus could have lead to the evolution of insect pathogenesis—an extension of its ability to breakdown organically bound nutrients or the co-option of genes involved in plant pathogenesis/symbiosis. Proteases are illustrated as an example of one of the gene families that likely allowed for the evolution of EIPF. In Metarhizium spp. they are one of the most numerous and diverse enzymes that allow this fungus to adapt to various habitats and enables its multifunctional lifestyle. The occurrence of plant pathogenic homologous proteases in Metarhizium may be utilized in initial endophyte colonization and/or immune evasion but does not lead to pathogenesis. Coloured bars indicate genes involved in saprotrophic (orange) lifestyle, plant pathogenesis (green), plant symbiosis/endophytism (blue), and insect pathogenesis (red). The asterisk indicates a lifestyle not observed with the organisms discussed as the focus of this review (i.e. Metarhizium, Beauveria), but is a theoretical evolutionary transitional lifestyle
Why evolution toward insect pathogenicity? There is some speculation of interkingdom jumping by these fungi from plants back to arthropods and then back to plants (Humber 2008). We suggest that many of these fungi have never left the role as plant symbionts and subsequently gained the ability to infect insects. We hypothesize that insect pathogenicity is an adaptation that allowed certain species of endophytic fungi to access a specialized source of nitrogen (i.e. insects), or other insect derived nutrients, and effectively barter these insect-derived nutrients for access to plant carbohydrates.
Insect pathogenesis
Metarhizium, Beauveria, and related insect pathogenic fungi, transgress the insect cuticle and infect insect hosts through a combination of mechanical penetration and degradation of cuticular components by employing enzymes such as proteases, esterases, N-acetylglucosaminidases, chitinases and lipases (St. Leger et al. 1996; Schrank and Vainstein 2010; Pedrini et al. 2013). The infection strategies of insect pathogens, such as Metarhizium and Beauveria, are well documented and understood (Small and Bidochka 2005; Ortiz-Urquiza and Keyhani 2013). During the initial stages of infection, hydrophobic conidia adhere to the insect cuticle and germinate to form a germ tube and holdfast infection structures termed appressoria (Small and Bidochka 2005; Holder and Keyhani 2005; Holder et al. 2007), similar to those found in plant pathogenic fungi (Deising et al. 2000). Two of the critical genes involved in facilitating conidial adherence to the insect cuticle are identified as Metarhizium adhesion protein 1 (Mad1) and ssgA, a hydrophobin (Wang and St. Leger 2007a; St. Leger et al. 1992a), and the hydrophobin genes, hyd1 and hyd2, in Beauveria (Zhang et al. 2011). Fungal penetration of insect cuticle is primarily facilitated through enzymatic degradation by proteases, and by the expression of different isoforms of endoproteinases. During growth on insect cuticle, the major proteases produced by Metarhizium includes cuticle degrading subtilisin-like protease (Pr1A), a thermolysin-like metalloproteinase, a trypsin like serine protease (Pr2) and other exo-acting peptidases (St. Leger et al. 1998). Expressed sequence tag (EST) analysis showed that M. anisopliae expressed 11 subtilisin-like proteases during growth on insect cuticle (Bagga et al. 2004). Furthermore, proteases also play a significant role in nutrient acquisition, evasion of host defense by degrading antifungal proteins, and regulation of micro-environmental pH (St. Leger et al. 1999).
Beneath the appressoria, hyphae penetrate through the insect cuticle and once in the insect hemolymph, the hyphae differentiate into yeast-like bodies termed blastospores (Small and Bidochka 2005; Lewis et al. 2009; Wanchoo et al. 2009). Enzymatic degradation as well as mechanical pressure has been implicated in cuticular penetration. For example, the expression of Mpl1 (perilipin) is implicated in the transport of lipid bodies to the appressoria thereby increasing turgor pressure (Wang and St. Leger 2007b). Once in the hemocoel, Metarhizium evades the insect immune system by expression of a collagen-like protein (MCL1) (Wang and St. Leger 2006) and adapts to the osmotic pressure in the hemolymph through expression of Mos1, an osmosensor-like protein (Wang et al. 2008). The expression of genes involved in insect pathogenesis is coordinately expressed as microarray analysis on mutant strains lacking Metarhizium protein kinase A1 (MaPKA1) showed the down-regulation of 244 genes involved in cuticular infection processes (Fang et al. 2009). The complexities of signaling pathways and genes involved in stress and virulence responses in Beauveria, are examined by Ortiz-Urquiza and Keyhani (2015). Metarhizium kills insect hosts within 3–7 days by producing toxins and absorbing nutrients. Once hemocoelic nutrients are depleted, hyphae emerge from the insect cadaver and conidiate, resulting in the mummification of the insect host (Small and Bidochka 2005; Schrank and Vainstein 2010).
Metarhizium is an excellent example of a fungus with a multifunctional lifestyle. It is an insect pathogen, a saprobe, and an endophyte. Metarhizium displays genotypic plasticity when exposed to dissimilar environments, thereby enabling the fungus to effectively persist saprobically or as a colonizer of plant or insect hosts (Pava-Ripoll et al. 2011; Wang and St. Leger 2005; Wang et al. 2005). For instance, Metarhizium uses two different proteins, MAD1 and MAD2 to facilitate adherence on insect and plant surfaces, respectively (Wang and St. Leger 2007a). EST and cDNA microarray analysis revealed that Metarhizium expressed different, yet overlapping, subsets of genes when grown on different insect cuticles, insect hemolymph, or in root exudate media (Wang and St. Leger 2005; Wang et al. 2005; Freimoser et al. 2003).
Notable in Metarhizium is the large number of proteases that it produces, and of these, 18 out of 43 protease genes are differentially expressed on insect cuticle and in root exudate (Wang et al. 2005). One significant exception is the subtilisin-like protease, pr1A, which was highly expressed in insects and root exudate (Wang et al. 2005). This suggests that pr1A is an example of a gene for a multifunctional life style. Similar differential gene expression has been reported in B. bassiana (Luo et al. 2015). Ortiz-Urquiza et al. (2015) examine the multitude of genes implicated in insect pathogenesis of both Metarhizium and Beauveria from a mycoinsecticide perspective and highlight that the success and survivability of these biocontrol agents in the environment is tightly connected to their capacity as plant symbionts. The adaptation of these fungi to insect and/or plant hosts could be the result of gene duplication or horizontal gene transfer events (Fig. 2; Bagga et al. 2004; Screen and St. Leger 2000; Xiao et al. 2012). A detailed understanding of the molecular mechanisms involved in fungal colonization of plant roots could provide an overall description of genes required by a fungus as a plant symbiont as well as an insect pathogen. That is, it could provide evolutionary insight into genes used for plant colonization that have been co-opted for insect pathogenicity.
Plant root colonization by insect pathogenic fungi
A number of insect pathogenic fungi are also plant endophytes (Vega 2008; Sasan and Bidochka 2012; Ownley et al. 2010). We term these fungi endophytic, insect pathogenic fungi (EIPF). Colonization of plants roots by EIPF may have evolved as a way to survive in soils, in the absence of an insect host (Hu and St. Leger 2002). However, a more likely explanation is that the Metarhizium evolutionary lineage initially adapted as an endophyte (>100 MYA), and insect pathogenesis is more recently acquired trait (<100 MYA). While there is little information on the specific evolutionary history of EIPF, recent work has suggested evolutionary pressure to maintain broad-range insect pathogenesis, indicating strong selection by the plant host to acquire nutrients from the broadest range of soil insects (Hu et al. 2014). However, under certain environmental conditions several Metarhizium spp. have evolved as specialists to certain insect species (i.e. M. acridum).
The ability of EIPF to infect insects is predicated on adherence to the insect cuticle (Wang and St. Leger 2007a), a mechanism that holds true for plant colonization as well, as successful association is dependent on adherence to the plant surface (Nicholson and Epstein 1991). For example, in M. robertsii the gene Mad2 encodes a plant adhesin that is crucial for attachment to plant roots and is up regulated when Metarhizium is grown in root exudate media (Wang and St. Leger 2007a). An orthologue of Mad2 was found within the genome of B. bassiana, suggesting Mad2 plays a role in Beauveria plant adhesion as well (Xiao et al. 2012).
Metarhizium is capable of growing internally within plant tissue (Sasan and Bidochka 2012), and evidence has shown M. robertsii endophytically colonizes the roots of switchgrass (Sasan and Bidochka 2012) as well as wheat, haricot bean, and soybean (Behie et al. 2015). In field conditions in Ontario, three species of Metarhizium (M. robertsii, M. brunneum, and M. guizhouense) were found to associate with grasses, shrubs, and trees, respectively (Wyrebek et al. 2011). B. bassiana is capable of endophytic colonization of roots, stems, and leaf tissues of tomato, cotton, snap bean, and haricot bean (Ownley et al. 2008; Behie et al. 2015). Typically, it has been thought that B. bassiana gains entry through naturally occurring openings (e.g. stomata), however evidence of distortions in the cell wall of corn (Zea mays) around penetration sites suggests enzymatic activity, similar to that observed during cuticular invasion of corn earworm (Heliothis zea), may play a role in plant invasion (Wagner and Lewis 2000; Pekrul and Grula 1979).
Proteases are a key component of cuticular penetration during insect infection but may also play a role in plant colonization. The Pr1 subtilisin-like protease of Metarhizium and a protease produced by B. bassiana are homologous to the fungal protease, At1, from Acremonium typhinum, a grass endophyte (Reddy et al. 1996). At1 is believed to facilitate symbiotic development by aiding in the degradation of the plant cell wall and/or apoplastic proteins, to allow fungal colonization (Reddy et al. 1996). Homologs of At1 are also observed in the mycoparasite Trichoderma harzianum and the nematode-trapping fungus Arthrobotrys oligospora (Geremia et al. 1993; Tunlid et al. 1994), both of which are also endophytes (Bordallo et al. 2002). Sequence variations in regions coding for substrate specificity would yield proteases with differing substrate specificities and may reflect evolutionary changes allowing fungi to adapt to various lifestyles as pathogens or endophytes (St. Leger et al. 1992b).
Protease functionality is highly dependent on environmental pH (Mayerhofer et al. 2015) and may help partition fungi into different ecological niches based on utilizable proteins. Metarhizium is able to grow over a wide range of pH (2.5–10.5) (Hallsworth and Magan 1996), and can modulate the pH of its immediate environment through the production of ammonia (St. Leger et al. 1999). The alteration of environmental pH allows for optimum activity of the extracellular subtilisin-like proteases (St. Leger et al. 1999), and may be a factor in the success of this ubiquitous EIPF.
Once inside the plant, endosymbiotic EIPF must avoid plant host defense. Plants are able to detect the presence of a pathogen and increase defense pathways, resulting in the expulsion, suppression, or death of the invading fungus (Dangl and Jones 2001). Fungal endophytes and other fungal root colonizers however, are able to communicate with the plant, indicating they are not pathogens. The arbuscular mycorrhizal fungus, Glomus intraradices releases a diffusible factor that primes the plant for root colonization. This diffusible communication molecule, named myc (mycorrhizal) factor, was identified as a lipochitooligosaccharide (LCO), and resembles Nod factors found in rhizobia (Maillet et al. 2011). Myc factors have been shown to prepare the root for fungal colonization by inducing transcriptional changes, such as those that activate the SYM (symbiotic) signaling pathway, and by inducing morphological changes that increase contact between plant roots and hyphae, such as increased root hair growth (Oldroyd et al. 2009; Kosuta et al. 2003). M. robertsii root colonization causes extensive root hair development in switchgrass, a notable indication of root priming (Sasan and Bidochka 2012), suggesting Metarhizium releases a myc-like factor prior to root colonization.
Nutrient exchange between EIPF and their plant hosts
In most ecosystems, soil nutrients are limited, thus competition among plants for nutrients is high (Clark and Zeto 2000). The majority of plant species are able to overcome this insufficient supply of nutrients by forming symbiotic associations with soil bacteria, mycorrhizal fungi, and fungal endophytes (Clark and Zeto 2000). Typically, in plant-fungal symbioses, fungal partners transfer limiting soil nutrients, such as phosphorus and nitrogen (Behie and Bidochka 2014; Guether et al. 2009; Govindarajulu et al. 2005) to their host plant in exchange for plant-derived carbohydrates (Bonfante and Genre 2010).
The dual life cycle of Metarhizium and Beauveria, suggested that insect pathogenesis and plant root colonization are coupled to provide plant hosts with insect derived nitrogen. EIPF are able to infect insects and subsequently translocate insect derived nitrogen to a host plant (Behie et al. 2012; Behie and Bidochka 2014). In this manner, plants colonized by EIPF have access to a specialized nitrogen reservoir present in soil ecosystems, organically bound in insects, and are able to reacquire nitrogen lost through insect herbivory.
What is the fungus gaining in return for providing insect-derived nitrogen to the plant? Freely available carbon is very difficult to access in the soil and is generally bound into complex carbohydrates such as cellulose and lignin. We hypothesized that EIPF gain access to simple plant carbohydrates in return for nitrogen. To confirm this we used plants colonized with EIPF and tracked 13C, through the introduction of 13CO2 in plant growth chambers, into plant carbohydrates and ultimately into fungal carbohydrates in the root/endophyte complex (trehalose and chitin; unpublished data). Metarhizium mutants deficient in a raffinose transporter gene (mrt) showed reduced rhizosphere competency, suggesting that fungal carbon acquisition is critical to Metarhizium/plant symbioses, and that MRT is a potential a route for the uptake of plant derived carbohydrates (Fang and St. Leger 2010).
EIPF as plant growth promoters
Not only have EIPF been shown to be involved in plant acquisition of nitrogen, plants grown in the presence of these fungal partners have shown increased growth and productivity (Behie and Bidochka 2014). Plants colonized by Metarhizium showed significantly greater number of lateral roots and root hair formations when compared to untreated plants (Sasan and Bidochka 2012), and increased leaf collar formation and foliage biomass have been reported in corn seeds treated with different Metarhizium strains (Liao et al. 2014). The effect of Metarhizium on the growth of tomato plants has also been evaluated, and Metarhizium-colonized plants showed significantly greater plant height, root length, and shoot/root dry weights (Elena et al. 2011). Kabaluk and Ericsson (2007) reported highest corn yield when corn plants were treated with both Metarhizium and a conventional insecticide as compared to corn treated with just insecticide. These studies suggested that Metarhizium confers plant growth promotion properties beyond protection from insects, and help increase primary production in agricultural systems.
EIPF also confer protection to plants against microbial pathogens. Metarhizium is antagonistic towards the bean root rot fungus Fusarium solani (Sasan and Bidochka 2013) and Beauveria was found to stimulate plant defense responses (Ownley et al. 2008, 2010). Metarhizium also promoted soybean growth during salt stress, where Metarhizium treated plants showed significantly greater biomass and comparatively lower levels of the plant stress hormone abscisic acid (Khan et al. 2012).
Potential source of economically important secondary metabolites
Genomic data of Metarhizium and Beauveria showed that these EIPF are rich in secondary metabolite gene clusters when compared to fungi with other trophic associations (Gibson et al. 2014). Metarhizium spp. produce secondary metabolites that are toxic to insects as well as other microbes (Carollo et al. 2010). There are 85 and 52 core genes putatively involved in secondary metabolite biosynthesis in M. robertsii and M. acridum, respectively (Gibson et al. 2014). Both pharmaceutically active and insecticidal secondary metabolites are reported from Metarhizium and include compounds such as destruxins, fusarin-like compounds (NG39x), cytochalasin, and swainsonine (Gibson et al. 2014). In Beauveria, the best-studied secondary metabolite is beauvericin, which was shown to have antibacterial, anti-tumor, antifungal and insecticidal activities (Wang and Xu 2012). Furthermore, in vitro studies revealed beauvericin has cytotoxic activity against human cell lines, reverses multidrug resistance phenotype in yeast, and flucanazole resistance phenotype in Candida albicans (Gibson et al. 2014).
Several EIPF have been identified in the biotransformation of various chemical substrates and of these, Beauveria spp. have been frequently used for such purposes (Grogan and Holland 2000). Specific gene clusters, predicted in Metarhizium spp. and Beauveria spp., show potential to be exploited for biotransformation, or as biocatalysts, that may be utilized in bioremediation and drug discovery (Gibson et al. 2014). There are also examples of endophytes capable of synthesizing pharmaceutically active metabolites when associated with plants. For example, taxol, is an anticancer drug produced by endophytic fungi when associated with yew trees (Taxus family) (Garyali and Reddy 2013). Hence, more focus on metabolite production analysis during plant root colonization is needed, as the specific interaction between fungus and plant may yield commercially relevant bioactive metabolites.
Conclusions
Traditional research on insect pathogenic fungi has focused almost exclusively on insect virulence, and on increasing virulence, focusing on a faster acting biological control agent. Recent advances, however, have indicated that a number of insect biocontrol agents, such as Beauveria and Metarhizium, are also plant colonizers, and may play a larger role in the ecosystem than previously realized. These ubiquitous fungi may be critical to processes such as soil nutrient cycling. It has also been suggested that a number of other insect pathogenic fungi have a plant colonization life cycle, and therefore, the complete role of these fungi must be further elucidated in order to develop strong, industry applicable, biological control agents and pharmaceuticals. Future research into EIPF as biological control agents requires consideration of their role in the ecosystem, in order to exploit the genetic history of these fungi, leading to more powerful research and industrial applications that could potentially be exploited in a myriad of positive ways.
References
Bagga S, Hu G, Screen SE, St. Leger RJ (2004) Reconstructing the diversification of subtilisins in the pathogenic fungus Metarhizium anisopliae. Gene 324:159–169
Behie SW, Bidochka MJ (2014) Ubiquity of insect-derived nitrogen transfer to plants by endophytic insect-pathogenic fungi: an additional branch of the soil nitrogen cycle. Appl Environ Microbiol 80:1553–1560
Behie SW, Zelisko PM, Bidochka MJ (2012) Endophytic insect-parasitic fungi translocate nitrogen directly from insects to plants. Science 336:1576–1577
Behie SW, Jones SJ, Bidochka MJ (2015) Plant tissue localization of the endophytic insect pathogenic fungi Metarhizium and Beauveria. Fungal Ecol 13:112–119
Blanford S, Chan BHK, Jenkins N, Sim D, Turner RJ, Read AF, Thomas MB (2005) Fungal pathogen reduces potential for malaria transmission. Science 308:1638–1641
Bonfante P, Genre A (2010) Mechanisms underlying beneficial plant–fungus interactions in mycorrhizal symbiosis. Nat Commun 1:48
Bordallo JJ, Lopez-Llorca LV, Jansson HB, Salinas J, Persmark L, Asensio L (2002) Colonization of plant roots by egg-parasitic and nematode-trapping fungi. New Phytol 154:491–499
Carollo CA, Calil ALA, Schiave LA, Guaratini T, Roberts DW, Lopes NP, Braga GU (2010) Fungal tyrosine betaine, a novel secondary metabolite from conidia of entomopathogenic Metarhizium spp. fungi. Fungal Biol 114:473–480
Castrillo LA, Griggs MH, Ranger CM, Reding ME, Vandenberg JD (2011) Virulence of commercial strains of Beauveria bassiana and Metarhizium brunneum (Ascomycota: Hypocreales) against adult Xylosandrus germanus (Coleoptera: Curculionidae) and impact on brood. Biol Control 58:121–126
Chandler D (1997) Selection of an isolate of the insect pathogenic fungus Metarhizium anisopliae virulent to the lettuce root aphid, Pemphigus bursarius. Biocontrol Sci Technol 7:95–104
Clark RB, Zeto SK (2000) Mineral acquisition by arbuscular mycorrhizal plants. J Plant Nutr 23:867–902
Dangl JL, Jones JDG (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833
Deising HB, Werner S, Wernitz M (2000) The role of fungal appressoria in plant infection. Microbes Infect 13:1631–1641
Elena GJ, Beatriz PJ, Alejandro P, Lecuona RE (2011) Metarhizium anisopliae (Metschnikoff) Sorokin promotes growth and has endophytic activity in tomato plants. Adv Biol Res 5:22–27
Fang W, St. Leger RJ (2010) Mrt, a gene unique to fungi, encodes an oligosaccharide transporter and facilitates rhizosphere competency in Metarhizium robertsii. Plant Physiol 154:1549–1557
Fang W, Pava-Ripoll M, Wang S, Leger RS (2009) Protein kinase A regulates production of virulence determinants by the entomopathogenic fungus, Metarhizium anisopliae. Fungal Genet Biol 46:277–285
Faria MR, Wraight SP (2007) Mycoinsecticides and mycoacaricides: a comprehensive list with worldwide coverage and international classification of formulation types. Biol Control 43:237–256
Freimoser FM, Screen S, Bagga S, Hu G, St. Leger RJ (2003) Expressed sequence tag (EST) analysis of two subspecies of Metarhizium anisopliae reveals a plethora of secreted proteins with potential activity in insect hosts. Microbiology 149:239–247
Gao Q, Jin K, Ying SH et al (2011) Genome sequencing and comparative transcriptomics of the model entomopathogenic fungi Metarhizium anisopliae and M. acridum. PLoS Genet. doi:10.1371/journal.pgen.1001264
Garyali S, Reddy MS (2013) Taxol production by an endophytic fungus, Fusarium redolens, isolated from Himalayan yew. J Microbiol Biotechnol 23:1372–1380
Geremia RA, Goldman GH, Jacobs D, Ardiles W, Vila SB, Van Montagu M, Herrera-Estrella A (1993) Molecular characterization of the proteinase-encoding gene, prbl, related to mycoparasitism by Trichoderma harzianum. Mol Microbiol 8:603–613
Gibson DM, Donzelli BG, Krasnoff SB, Keyhani NO (2014) Discovering the secondary metabolite potential encoded within entomopathogenic fungi. Nat Prod Rep 31:1287–1305
Govindarajulu M, Pfeffer P, Philip E et al (2005) Nitrogen transfer in the arbuscular mycorrhizal symbiosis. Nature 435:819–823
Grogan GJ, Holland HL (2000) The biocatalytic reactions of Beauveria spp. J Mol Catal B Enzym 9:1–32
Guether M, Neuhäuser B, Balestrini R et al (2009) A mycorrhizal specific ammonium transporter from Lotus japonicus acquires nitrogen. Plant Physiol 150:73–83
Hallsworth JE, Magan N (1996) Culture age, temperature, and pH affect the polyol and trehalose contents of fungal propagules. Appl Environ Microbiol 62:2435–2442
Holder DJ, Keyhani NO (2005) Adhesion of the entomopathogenic fungus Beauveria (Cordyceps) bassiana to substrata. Appl Environ Microbiol 71:5260–5266
Holder DJ, Kirkland BH, Lewis MW, Keyhani NO (2007) Surface characteristics of the entomopathogenic fungus Beauveria (Cordyceps) bassiana. Microbiology 153:3448–3457
Hu G, St. Leger RJ (2002) Field studies using a recombinant mycoinsecticide (Metarhizium anisopliae) reveal that it is rhizosphere competent. Appl Environ Microbiol 68:6383–6387
Hu X, Xiao G, Zheng P et al (2014) Trajectory and genomic determinants of fungal-pathogen speciation and host adaptation. PNAS 111:16796–16801
Humber RA (2008) Evolution of entomopathogenicity in fungi. J Invertebr Pathol 98:262–266
Kabaluk JT, Ericsson JD (2007) Seed treatment increases yield of field corn when applied for wireworm control. Agron J 99:1377–1381
Khan AL, Hamayun M, Khan SA, Kang SM, Shinwari ZK, Kamran M, Rehman S, Kim JG, Lee IJ (2012) Pure culture of Metarhizium anisopliae LHL07 reprograms soybean to higher growth and mitigates salt stress. World J Microbiol Biotechnol 28:1483–1494
Kosuta S, Chabaud M, Lougnon G, Gough C, Dénarié J, Barker DG, Bécard G (2003) A diffusible factor from arbuscular mycorrhizal fungi induces symbiosis-specific MtENOD11 expression in roots of Medicago truncatula. Plant Physiol 131:952–962
Lewis MW, Robalino IV, Keyhani NO (2009) Uptake of the fluorescent probe FM4-64 by hyphae and haemolymph-derived in vivo hyphal bodies of the entomopathogenic fungus Beauveria bassiana. Microbiology 155:3110–3120
Liao X, O’Brien TR, Fang W, St. Leger RJ (2014) The plant beneficial effects of Metarhizium species correlate with their association with roots. Appl Microbiol Biotechnol 98:7089–7096
Luo F, Wang Q, Yin C et al (2015) Differential metabolic responses of Beauveria bassiana cultured in pupae extracts, root exudates and its interactions with insect and plant. J Invertebr Pathol. doi:10.1016/j.jip.2015.01.003
Madelin MF (1963) Diseases caused by hyphomycetous fungi. In: Steinhaus E (ed) Insect pathology: an advanced treatise, vol 2. Academic Press, New York, pp 233–272
Maillet F, Poinsot V, André O et al (2011) Fungal lipochitooligosaccharide symbiotic signals in arbuscular mycorrhiza. Nature 469:58–63
Mayerhofer MS, Fraser E, Kernaghan G (2015) Acid protease production in fungal root endophytes. Mycologia 107:1–11
Nicholson RL, Epstein L (1991) Adhesion of fungi to the plant surface: prerequisite for pathogenesis. In: Cole GT, Hoch HC (eds) The fungal spore and disease initiation in plants and animals. Plenum Press, New York, pp 3–23
Oldroyd GED, Harrison MJ, Paszkowski U (2009) Reprogramming plant cells for endosymbiosis. Science 324:753–754
Ortiz-Urquiza A, Keyhani NO (2013) Action on the surface: entomopathogenic fungi versus the insect cuticle. Insects 4:357–374
Ortiz-Urquiza A, Keyhani NO (2015) Stress response signaling and virulence: insights from entomopathogenic fungi. Curr Genet 61:239–249
Ortiz-Urquiza A, Luo Z, Keyhani NO (2015) Improving mycoinsecticides for insect biological control. Appl Microbiol Biotechnol 99:1057–1068
Ownley BH, Griffin MR, Klingeman WE, Gwinn KD, Moulton JK, Pereira RM (2008) Beauveria bassiana: endophytic colonization and plant disease control. J Invertebr Pathol 98:267–270
Ownley BH, Gwinn KD, Vega FE (2010) Endophytic fungal entomopathogens with activity against plant pathogens: ecology and evolution. Ecol Fungal Entomopathog 55:113–128
Pava-Ripoll M, Angelini C, Fang W, Wang S, Posada FJ, St. Leger RJ (2011) The rhizosphere-competent entomopathogen Metarhizium anisopliae expresses a specific subset of genes in plant root exudate. Microbiology 157:47–55
Pedrini N, Ortiz-Urquiza A, Huarte-Bonnet C et al (2013) Targeting of insect epicuticular lipids by the entomopathogenic fungus Beauveria bassiana: hydrocarbon oxidation within the context of a host-pathogen interaction. Front Microbiol 4:1–18
Pekrul S, Grula EA (1979) Mode of infection of the corn earworm (Heliothus zea) by Beauveria bassiana as revealed by scanning electron microscopy. J Invertebr Pathol 34:238–247
Reddy PV, Lam CK, Belanger FC (1996) Mutualistic fungal endophytes express a proteinase that is homologous to proteases suspected to be important in fungal pathogenicity. Plant Physiol 111:1209–1218
Sasan RK, Bidochka MJ (2012) The insect-pathogenic fungus Metarhizium robertsii (Clavicipitaceae) is also an endophyte that stimulates plant root development. Am J Bot 99:101–107
Sasan RK, Bidochka MJ (2013) Antagonism of the endophytic insect pathogenic fungus Metarhizium robertsii against the bean plant pathogen Fusarium solani f. sp. phaseoli. Can J Plant Pathol 35:288–293
Schrank A, Vainstein MH (2010) Metarhizium anisopliae enzymes and toxins. Toxicon 56:1267–1274
Screen SE, St. Leger RJ (2000) Cloning, expression, and substrate specificity of a fungal chymotrypsin—evidence for lateral gene transfer from an actinomycete bacterium. J Biol Chem 275:6689–6694
Small CLN, Bidochka MJ (2005) Up-regulation of Pr1, a subtilisin-like protease, during conidiation in the insect pathogen Metarhizium anisopliae. Mycol Res 109:307–313
Spatafora JW, Sung GH, Sung HM, Hywel-Jones L, White JF (2007) Phylogenetic evidence for an animal pathogen origin of ergot and the grass endophytes. Mol Ecol 16:1701–1711
St. Leger RJ, Staples RC, Roberts DW (1992a) Cloning and regulatory analysis of starvation-stress gene, ssgA, encoding a hydrophobin-like protein from the entomopathogenic fungus, Metarhizium anisopliae. Gene 120:119–124
St. Leger RJ, Frank DC, Roberts DW, Staples RC (1992b) Molecular cloning and regulatory analysis of the cuticle-degrading protease structural gene from the entomopathogenic fungus Metarhizium anisopliae. Eur J Biochem 204:991–1001
St. Leger RJ, Joshi L, Bidochka MJ, Rizzo NW, Roberts DW (1996) Characterization and ultrastructural localization of chitinases from Metarhizium anisopliae, M. flavoviride, and Beauveria bassiana during fungal invasion of host (Manduca sexta) cuticle. Appl Environ Microbiol 62:907–912
St. Leger RJ, Joshi L, Roberts D (1998) Ambient pH is a major determinant in the expression of cuticle-degrading enzymes and hydrophobin by Metarhizium anisopliae. Appl Environ Microbiol 64:709–713
St. Leger RJ, Nelson JO, Screen SE (1999) The entomopathogenic fungus Metarhizium anisopliae alters ambient pH, allowing extracellular protease production and activity. Microbiology 145:2691–2699
Tunlid A, Rosen SEB, Rask L (1994) Purification and characterization of an extracellular serine protease from the nematode trapping fungus Arthrobotrys oligospora. Microbiology 140:1687–1695
Vega FE (2008) Insect pathology and fungal endophytes. J Invertebr Pathol 98:277–279
Wagner BL, Lewis LC (2000) Colonization of corn, Zea mays, by the entomopathogenic fungus Beauveria bassiana. Appl Environ Microbiol 66:3468–3473
Wanchoo A, Lewis MW, Keyhani NO (2009) Lectin mapping reveals stage-specific display of surface carbohydrates in in vitro and haemolymph-derived cells of the entomopathogenic fungus Beauveria bassiana. Microbiology 155:3121–3133
Wang C, St. Leger RJ (2005) Developmental and transcriptional responses to host and nonhost cuticles by the specific locust pathogen Metarhizium anisopliae var. acridum. Eukaryot Cell 4:937–947
Wang C, St. Leger RJ (2006) A collagenous protective coat enables Metarhizium anisopliae to evade insect immune responses. Proc Natl Acad Sci USA 103:6647–6652
Wang C, St. Leger RJ (2007a) The MAD1 adhesin of Metarhizium anisopliae links adhesion with blastospore production and virulence to insects, and the MAD2 adhesin enables attachment to plants. Eukaryot Cell 6:808–816
Wang C, St. Leger RJ (2007b) The Metarhizium anisopliae perilipin homolog MPL1 regulates lipid metabolism, appressorial turgor pressure, and virulence. J Biol Chem 282:21110–21115
Wang Q, Xu L (2012) Beauvericin, a bioactive compound produced by fungi: a short review. Molecules 17:2367–2377
Wang C, Hu G, St. Leger RJ (2005) Differential gene expression by Metarhizium anisopliae growing in root exudate and host (Manduca sexta) cuticle or hemolymph reveals mechanisms of physiological adaptation. Fungal Genet Biol 42:704–718
Wang C, Duan Z, St. Leger RJ (2008) MOS1 osmosensor of Metarhizium anisopliae is required for adaptation to insect host hemolymph. Eukaryot Cell 7:302–309
Wyrebek M, Bidochka MJ (2013) Variability in the insect and plant adhesins, Mad1 and Mad2, within the fungal genus Metarhizium suggest plant adaptation as an evolutionary force. PLoS One. doi:10.1371/journal.pone.0059357
Wyrebek M, Huber C, Sasan RK, Bidochka MJ (2011) Three sympatrically occurring species of Metarhizium show plant rhizosphere specificity. Microbiology 157:2904–2911
Xiao G, Ying SH, Zheng P et al (2012) Genomic perspectives on the evolution of fungal entomopathogenicity in Beauveria bassiana. Sci Rep 2:1–10
Zhang S, Xia YX, Kim B, Keyhani NO (2011) Two hydrophobins are involved in fungal spore coat rodlet layer assembly and each play distinct roles in surface interactions, development and pathogenesis in the entomopathogenic fungus, Beauveria bassiana. Mol Microbiol 80:811–826
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Barelli, L., Moonjely, S., Behie, S.W. et al. Fungi with multifunctional lifestyles: endophytic insect pathogenic fungi. Plant Mol Biol 90, 657–664 (2016). https://doi.org/10.1007/s11103-015-0413-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0413-z