Abstract
Non-target-site resistance (NTSR) to herbicides that disrupts agricultural weed control is a worldwide concern for food security. NTSR is considered a polygenic adaptive trait driven by differential gene regulation in resistant plants. Little is known about its genetic determinism, which precludes NTSR diagnosis and evolutionary studies. We used Illumina RNA-sequencing to investigate transcriptomic differences between plants from the global major weed rye-grass sensitive or resistant to the acetolactate-synthase (ALS) inhibiting herbicide pyroxsulam. Plants were collected before and along a time-course after herbicide application. De novo transcriptome assembly yielded a resource (LOLbase) including 92,381 contigs representing potentially active transcripts that were assigned putative annotations. Early effects of ALS inhibition consistent with the literature were observed in resistant and sensitive plants, proving LOLbase data were relevant to study herbicide response. Comparison of resistant and sensitive plants identified 30 candidate NTSR contigs. Further validation using 212 plants resistant or sensitive to pyroxsulam and/or to the ALS inhibitors iodosulfuron + mesosulfuron confirmed four contigs (two cytochromes P450, one glycosyl-transferase and one glutathione-S-transferase) were NTSR markers which combined expression levels could reliably identify resistant plants. This work confirmed that NTSR is driven by differential gene expression and involves different mechanisms. It provided tools and foundation for subsequent NTSR investigations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Agricultural weeds are currently a considerable threat to food security (Oerke 2006; Vigueira et al. 2013). Weeds are mostly short-lived plants that successfully adapted to agricultural ecosystems (Baker 1974; Vigueira et al. 2013). Understanding the mechanisms driving weed adaptation is crucial to devise strategies enabling more efficient weed control. One key step towards this goal is unravelling the genes driving traits crucial for weed success (Délye 2013; Vigueira et al. 2013). Prominent among these traits is resistance to herbicides that is weed adaptive response to herbicide applications (reviewed in Délye et al. 2013). Herbicide resistance is also an intriguing study system to decipher plant stress response: exposure to the stress (herbicide application) can be controlled and the adaptive phenotype is clear-cut: only adapted (resistant) plants survive herbicide application (Délye 2013).
Mechanisms of resistance to herbicides can be categorised into two classes. Target-site-based resistance is conferred by alleles of the herbicide target protein gene, with “allele” meaning a variant of a wild-type gene displaying differences in its nucleotide sequence and/or expression level. Target-site-based resistance is now well elucidated (Beckie and Tardif 2012; Délye et al. 2013). Non-target-site based resistance (NTSR) is endowed by alleles causing herbicide neutralisation or compensation for herbicide action, and is the most agronomically noxious type of resistance (Délye et al. 2013). NTSR is considered to be caused by constitutive or herbicide-induced mechanisms also involved in plant response to the herbicide stress, and thus would mostly be driven by inheritable differences in the expression of a set of genes between resistant and sensitive plants (Yuan et al. 2007; Délye 2013). Cytochromes P450, glycosyltransferases, glutathione-S-transferases and transporters are involved in NTSR, but very few genes have been identified to date (Yuan et al. 2007; Délye 2013; Gaines et al. 2014; Iwakami et al. 2014; Saika et al. 2014).
Unravelling the complex genetic control of herbicide response and NTSR can be achieved using whole-transcriptome sequencing (RNA-Seq) that is currently the most powerful tool to identify genes differentially regulated among experimental modalities (Martin and Wang 2011; Ozsolak and Milos 2011). RNA-Seq allows comprehensive transcriptome sequencing and yields qualitative data (transcript sequences) and quantitative data (transcript expression level) without the need for pre-existing genomic or transcriptomic resources. Accordingly, RNA-Seq is considered a highly promising way of unravelling the genetic control of complex traits in plants, but this approach is still fledging in weeds (Ward et al. 2012; Délye 2013; Vigueira et al. 2013).
Rye-grasses (Lolium spp.) are a group of outcrossing interfertile species. The three most common species in Europe are the diploid (2n = 14) perennial Lolium perenne L. and the annual Lolium multiflorum Lam. and Lolium rigidum Gaud. (Charmet et al. 1996). Because they can adapt to a broad range of ecological conditions, rye-grasses are major forage and turf grasses (Humphreys and O’Kiely 2006). Annual rye-grass species are also some of the world’s most troublesome and widespread agricultural weeds (Heap 2014). As weeds, rye-grasses are most efficiently controlled by herbicide applications. However, recurrent herbicide use has selected for resistance in numerous rye-grass populations worldwide (Beckie and Tardif 2012). Polygenic NTSR that confers an unpredictable resistance to a range of herbicides (Busi et al. 2011; 2013) is largely the most important type of resistance in rye-grass (Beckie and Tardif 2012) and can cause severe control failures. Several contigs potentially involved in rye-grass NTSR to herbicides inhibiting acetyl-CoA carboxylase (ACCase), the second herbicide class most used against rye-grass, were recently identified using a RNA-Seq approach (Gaines et al. 2014). However, NTSR genetic bases remain largely unknown in rye-grasses. Here, we considered rye-grass NTSR to herbicide inhibiting acetolactate synthase (ALS), the herbicide class most used against rye-grass. RNA-Seq allowed establishing a transcriptomic resource that was used to check whether (1) gene regulation was at the basis of herbicide response, and (2) whether particularities in gene regulation could be related to differences in herbicide sensitivity in rye-grass. Comparison of the transcriptomic responses of plants with NTSR and of herbicide-sensitive plants over a time-course experiment provided insight into key biological processes and molecular functions involved in rye-grass response to ALS inhibitors, and identified contigs that can be used as NTSR markers.
Materials and methods
Plant material and sample collection
Seeds were collected in 2008 in one French field where rye-grass control using ALS inhibitors failed. Seedlings were checked for the absence of ALS alleles conferring herbicide resistance by genotyping as described (Délye et al. 2009). Twenty genotyped seedlings were grown in individual 2L-pots in a glasshouse at 22/18 °C day/night with 14-h photoperiod. At the eighteen tiller stage, they were subjected to vegetative propagation: all individual tillers were separated and transplanted into individual pots. For each plant, this yielded 18 clones at the 3–4 leaf growth stage, at which ALS-inhibiting herbicide application is recommended.
A time-course experiment including seven time-points was conducted: untreated (UT), two, six, 12, 24, 36 and 48 h after treatment (HAT). Two clones (biological replicates) intended for RNA extraction were used per plant at each time-point (i.e., 14 clones per plant in total). The four remaining clones of each plant were not sampled for RNA extraction. Two clones sprayed with herbicide (phenotype control) and two clones sprayed with water (water-sprayed control) together with the clones used for the time-course experiment served to characterise plant phenotype (resistant or sensitive) 4 weeks after treatment. Two herbicide-sprayed and two water-sprayed clones from a reference herbicide-sensitive plant were included in the spraying experiment to check herbicide application efficacy. The commercial herbicide formulation Abak that contains pyroxsulam (7.5 % w/w; Dow AgroScience, Saint Quentin en Yvelines, France), an ALS inhibitor frequently sprayed against rye-grasses, was used in the time-course experiment at its recommended field rate (18.75 g pyroxsulam ha−1) with an adjuvant enhancing herbicide penetration into leaf tissues (Actirob B; Bayer CropScience, Lyon, France, 1 L ha−1). Herbicide application was as described (Petit et al. 2012).
ALS inhibitors essentially act in the meristems (Zhou et al. 2007) that are at the leaf basis in grasses (Briske 1991). One sample collected for RNA extraction thus included the basis of the above-ground part of the two clones of one given plant in one given modality that was cut, immediately frozen in liquid nitrogen and stored at −80 °C until phenotype assessment had been performed 4 weeks after herbicide application using the clones intended for this purpose. Three resistant plants and three sensitive plants were selected for RNA-Seq. Forty-two samples, each consisting of two clones for each of the six selected plants and seven time-points, were thus used for subsequent analyses.
RNA extraction and Illumina sequencing
Total RNA was extracted from the 42 samples using RNeasy plant mini kit (Qiagen, Courtaboeuf, France) following manufacturer’s instructions. Potential genomic DNA contaminations were removed using the RNase-Free DNase Set (Qiagen). RNA quality was checked on a 2100 Bioanalyser (Agilent, Waldbroon, Germany). Our aim was to identify as many transcripts present in each phenotype at each time-point as possible. For this purpose and because sequencing 42 samples was not affordable, the 42 RNA samples were pooled prior to sequencing. One pooled sample was an equimolar mixture of total RNA extracted from the six biological replicates per phenotype at a given time-point (two clones for each of the three resistant or sensitive plants at each time-point). Fourteen pooled samples were thus sequenced. This design allowed to investigate the average transcriptomic pattern of one resistant pool and one sensitive pool each consisting of three plants with two biological replicates per plant over the seven time-points.
RNA-Seq was performed by Fasteris (Plan-Les-Ouates, Switzerland). One µg RNA from each of the 14 pooled samples was processed using TruSeq RNA Sample Prep kit (Illumina). Transcripts were purified and fragmented by zinc breaking using mRNA-SEQ kit (Illumina). Non-normalised, double-stranded cDNA libraries were prepared with random primers and RNaseH and subjected to 15 PCR cycles. Fragments with 160–240 base pair inserts were recovered after agarose gel electrophoresis. Each cDNA library was indexed with a unique 6-base identifier and sequenced in two independent runs following manufacturer’s instructions to generate paired-end, 100-base reads. In each sequencing run, two lanes with seven libraries per lane were used on a HiSeq 2000 (Illumina). Sequence data was extracted using the CASAVA 1.8.1 pipeline (Illumina) and subjected to quality control using an indexed PhiX reference sequence on each call lane to estimate sequencing error. Identifier sequences and contaminant sequences were removed and the quality reads were subsequently attributed to the corresponding pooled samples, yielding one set of quality reads per experimental modality.
De novo transcriptome assembly and annotation
All quality reads obtained for all 14 experimental modalities were pooled and used as starting material for de novo transcriptome assembly. As the pooled samples sequenced were obtained from plants issued from seeds collected in a field, nucleotide variation among plants was expected to complicate transcriptome assembly. Preliminary assembly tests with available software and protocols generated highly fragmented assemblies. A custom iterative procedure was thus designed to handle heterozygosity and sequencing errors. The first step implemented iterative Velvet (Zerbino and Birney 2008) runs with stringent parameters (k-mer ranging from 61 to 85, step = 4, max_divergence = 0.01). Reads were considered stranded (-strand_specific parameter) even if the actual data were not. At each iteration, the contig sequences assembled at the previous iteration were integrated as long reads (-long). This highly time-consuming procedure generated a set of accurate long reads. This data was very similar to Sanger EST sequence data. An iterative pipeline integrating an nrcl-like program and the cap3 assembler (Huang and Madan 1999) was developed. Each pipeline iteration combined an nrcl run removing redundancy by including short redundant contigs into longer contigs and a cap3 assembly merging contigs. At each iteration, less and less stringent parameters were used, because each iteration added constraints to the subsequent one. The thresholds used for nrcl and cap3 were an Identity percentage ranging from 99 to 93 %, an overhang length of 5 and 40 nucleotides, respectively, and a high-scoring segment pair length of 100 and 40 nucleotides, respectively.
The assembly quality was manually checked on a set of contigs expected to correspond to single-copy genes by checking that a single consensus contig sequence was assembled for each gene (not shown). Putative peptide-coding sequences were sought in all assembled contigs using FrameDP (Gouzy et al. 2009) and annotated using InterProScan (Quevillon et al. 2005). Annotations included Gene Ontology (GO) terms, Enzyme Commission (EC) codes and Pfam domains. Similarity searches for peptide annotation were also conducted against TAIR10, Brachypodium distachyon proteome, Swiss-Prot and TrEMBL using the BLASTp algorithm with an E value <10–5. Additional Pfam domain assignation was performed using the Pfam-A 27.0 database and the hmmsearch program (Finn et al. 2011).
Contig expression analysis
All sequence reads corresponding to each pooled sample were mapped against the assembled contigs using Glint (Faraut and Courcelle unpublished; available at http://lipm-bioinfo.toulouse.inra.fr/download/glint/) configured to keep only the best scoring pairs without gaps in the alignment (maximum number of mismatches = 5, minimum score = 24, length of each read aligned ≥50 %). Redundancy was expected in the assembly. Thus, each read in each experimental modality could be mapped to different contigs, so that differential expression analysis was not biased by redundancy. The total number of reads mapped per contig was computed for each pooled sample and normalised using the RPKM method (Mortazavi et al. 2008).
RT-qPCR validation of RNA-Seq expression patterns
Primers targeting 19 contigs showing RNA-Seq expression patterns stable or variable among modalities (Table S1) were designed using Primer3Plus (Untergasser et al. 2007). Amplification specificity and qPCR efficacy were checked as described (Duhoux and Délye 2013). The expression level of each contig was measured in each of the 42 original RNA samples used to create the 14 pooled samples subjected to RNA-Seq. cDNA synthesis was performed in duplicate (technical replicates) from 5 µg total RNA using the Masterscript RT-PCR System (5PRIME, Hamburg, Germany). qPCR using a StepOnePlus Real-Time PCR System (Applied Biosystems, Foster City, USA) and contig expression level normalization using three validated rye-grass reference genes were as described (Duhoux and Délye 2013). Correlation between contig expression level obtained by RNA-Seq and average contig expression level measured by RT-qPCR over the three original RNA samples used to create each pooled sample was checked using Pearson’s product-moment coefficient correlation.
Identification of differentially expressed contigs and GO term enrichment
Because of the absence of biological replicates in our experimental design, no statistic test could be performed to identify contigs with differences in expression among modalities. Thus, to identify differentially expressed contigs, we used R DESeq package (Anders and Huber 2010) with the following, stringent criteria implemented: a difference in expression > threefold between two modalities, a contig length ≥400 nucleotides and a RPKM value ≥0.5 in at least one modality. The TopGO package was subsequently implemented with the elim method (Alexa et al. 2006) to identify GO terms significantly enriched in contigs differentially regulated among modalities. The transcriptomic herbicide response of each phenotype (resistant or sensitive) over time was investigated via pairwise comparisons between each treated modality and the corresponding UT modality. To identify phenotype-associated differences in herbicide response, pairwise comparisons were performed between phenotypes at each time point.
Candidate NTSR contig validation
Contigs with a possible role in NTSR were identified after GO enrichment analyses and/or based on the analysis of individual expression patterns of contigs assigned to Pfam families involved in NTSR. Differential expression was first confirmed by measuring the expression level of these contigs in each of the 42 individual samples using RT-qPCR (Table S2). For further validation, the expression level of all contigs up-regulated in each resistant plant compared to each sensitive plant at each time-point was measured in untreated clones of 212 additional plants originating from 15 geographically different French field populations (Table 1). For each of these plants, the absence of mutant ALS alleles conferring herbicide resistance had been checked and the phenotype characterised beforehand using the commercial herbicide formulations Abak (pyroxsulam) and Archipel WG (Bayer CropScience, Lyon, France) that is a mixture of two ALS inhibitors (iodosulfuron + mesosulfuron, 3 % w/w each). Briefly, each plant was split into eight clones. Two clones intended for RNA extraction were collected as before just prior to treatment. Abak or Archipel were each applied on two clones per plant at their recommended field rates (18.75 g pyroxsulam ha−1 or 7.5 g iodosulfuron + 7.5 g mesosulfuron ha−1, respectively) with adjuvant (Actirob B, 1 L ha−1). The last two clones were water-treated controls. Four weeks after herbicide application, plants which treated clones were killed, markedly affected but survived or unaffected or moderately affected but healthily growing were rated sensitive, moderately resistant or resistant, respectively.
Results
LOLbase, a rye-grass transcriptome resource
A range of phenotypes were observed 4 weeks after treatment among the 20 plants sprayed. The three plants showing the most resistant phenotypes (phenotype control most similar to the water-sprayed control) and the three plants showing the most sensitive phenotypes (phenotype control most rapidly killed) were selected for RNA-Seq. RNA integrity numbers measured prior to sequencing ranged from 8.6 to 9.8, indicating good RNA quality. After filtering, sequencing the 14 pooled samples yielded 323,833,502 quality 100-base reads (>32 × 109 nucleotides) that were pooled for de novo transcriptome assembly. Reads are available at the NCBI Sequence Read Archive (http://ncbi.nlm.nih.gov/sra, accessions SRR1141056 to SRR1141083).
147,829 unique sequences with sizes ranging from 145 to 13,078 nucleotides were assembled de novo. Assembled contigs shorter than 400 nucleotides or encoding predicted peptides shorter than 133 amino-acids that probably corresponded to truncated transcripts were poorly annotated (Table 2, Fig. S1). They were considered as assembly waste and filtered out. The proportion of contigs with at least one mapped read was similar in all modalities (Fig. S2). Previous studies proposed that contigs with low RPKM values were assembly errors and that a twofold average sequencing coverage was an acceptable threshold to consider a contig as representing a potentially expressed transcript (Zhao et al. 2010; Burke and Strand 2012). Herein, this threshold corresponded to a RPKM value of 0.5. Accordingly, all sequences with RPKM values <0.5 in all modalities were discarded from subsequent analyses. This yielded a rye-grass transcriptomic resource containing 92,381 contigs (N50 = 771 nucleotides) predicted to encode 81,563 potential peptides (Table 2). In the following, the name “LOLbase” will refer to these contigs and their predicted peptides. Sequences are available at https://iant.toulouse.inra.fr/Lolium?download=1.
Expression data validation using RT-qPCR
LOLbase expression data were validated by RT-qPCR measurement of the expression level of 19 randomly selected contigs in each of the 42 original individual RNA samples. The expression level of the 19 contigs showed variation among individual plants, including plants with the same phenotype at the same time-point (Figs. 1, S3). Yet, the average contig expression levels computed for the three resistant plants or for the three sensitive plants at each time-point matched the corresponding RNA-Seq expression data, with Pearson’s product-moment correlation coefficient >0.90 for all contigs except LOLSS028240 (0.77) LOLSS003821 (0.42) and LOLSS001274 (0.47) (Figs. 1, S3). These relatively low values were due to the moderate and stable expression level of these contigs over all modalities.
RT-qPCR validation of the RNA-Seq contig expression patterns of five of the 19 contigs used for RNAseq expression data validation (see Supplemental Fig. S3 for the other transcripts). The relative expression values were computed using the UT experimental modality with the highest expression level as a reference. White bars, relative expression values computed from the RPKM counts in LOLbase; black bars, relative expression values measured by RT-qPCR; r values: Pearson’s product-moment correlation coefficient values
Transcriptional response to the ALS inhibitor pyroxsulam common to sensitive plants and plants with NTSR (treated vs. UT)
23,343 and 38,420 contigs were respectively up-regulated or down-regulated (>threefold difference in RPKM value) in the resistant and/or in the sensitive phenotype in at least one herbicide-treated modality compared to the respective UT modalities, of which 8,284 and 10,600 were respectively up-regulated or down-regulated in both phenotypes in at least one herbicide-treated modality. The numbers of contigs regulated at a given time-point after herbicide application or starting from one time-point after herbicide application in each phenotype are shown in Figs. 2 or S4, respectively.
Number of contigs regulated or with a stable expression after herbicide application compared to UT. A Number of contigs up-regulated, down-regulated or with a stable expression in the resistant phenotype (left) or in the sensitive phenotype (right) in each herbicide-treated modality compared to the corresponding UT modality. B Total number of contigs up-regulated, down-regulated or with a stable expression in the resistant phenotype (left) or in the sensitive phenotype (right) compared to the corresponding UT modality
In the following, we only considered enriched terms with a p value <10−10 in at least one modality (Table S3). Twelve molecular functions or four biological processes were enriched in up-regulated contigs in at least one herbicide-treated modality (Table S3). Eleven molecular functions and three biological processes were common to both phenotypes. They included terms involved in gene expression regulation, amino-acid metabolism (e.g., leucine biosynthetic process) or potentially herbicide response (i.e., monooxygenase activity, transferase activity-transferring hexosyl groups, drug transmembrane transport/multidrug transport and FMN binding). Five molecular functions, six biological processes or seven cellular components were enriched in down-regulated contigs in at least one herbicide-treated modality (Table S3). Four molecular functions, five biological processes and the six cellular components were common to both phenotypes. They included terms involved in photosynthesis or gene expression regulation.
The terms drug transmembrane transport (molecular function) and multidrug transport (biological process) included the same set of contigs, as was the case for 2-isopropylmalate synthase activity (molecular function) and leucine biosynthetic process (biological process), and for drug transmembrane transport (molecular function) and multidrug transport (biological process). Thus, 12 and 15 non-redundant terms were enriched in up-regulated or down-regulated contigs after pyroxsulam application, respectively, in both phenotypes. The herbicide-regulated contigs assigned to these terms in each phenotype included contigs with a similar regulation in both phenotypes that could belong to the general herbicide response and contigs with a phenotype-specific regulation. The proportion of contigs potentially involved in pyroxsulam general response (up- or down-regulated in both phenotypes) varied greatly among terms, ranging from 100 % (leucine biosynthesis process and photosystem I reaction center) to 24 % (nucleus) (Fig. 3). Terms involved in protein or amino-acid biosynthesis (leucine biosynthesis process, aminoacyl-tRNA ligase activity, arginine-tRNA ligase) or photosynthesis had a high proportion of herbicide-regulated contigs common to both phenotypes (85–00 %) (Fig. 3).
Phenotype-specific transcriptional response to the ALS inhibitor pyroxsulam (resistant vs. sensitive)
The numbers of contigs up-regulated in one phenotype at a given time-point or starting from one time-point are shown in Figs. 4 or S5, respectively. 9,583 and 10,805 contigs were respectively up-regulated UT in the resistant or in the sensitive phenotype (Fig. 4). In the following, we only considered terms significantly enriched with a p value <10−5 (Table S4). Three molecular functions, three biological processes or one cellular component were enriched UT in the resistant phenotype. They included terms involved in gene regulation or ATP metabolism. Three molecular functions, one biological processes or two cellular components were enriched in the sensitive phenotype, most of which were involved in protein biosynthesis or catabolism (Table S4).
21,001 and 27,209 contigs were up-regulated in the resistant or in the sensitive phenotype, respectively, in a least one herbicide-treated modality, of which 1,981 and 1,749 were respectively up-regulated in the resistant or in the sensitive phenotype over the whole time-course (Fig. 4). Twenty-one molecular functions, 13 biological processes or four cellular components were significantly enriched in up-regulated contigs in at least one herbicide-treated modality (Table S4). Among these, 12 molecular functions, seven biological processes or two cellular components were only enriched in up-regulated contigs in the resistant phenotype, including terms involved in gene expression regulation, stress response or potentially herbicide response (transferase activity-transferring acyl groups other than amino-acyl groups). Five molecular functions, two biological processes or two cellular components were only enriched in up-regulated contigs in the sensitive phenotype. They included terms involved in oxidative stress response or protein catabolism (Table S4). Among the terms enriched in contigs differentially regulated between phenotypes, monooxygenase activity, transferase activity-transferring hexosyl groups, ATP binding, iron ion binding, electron carrier activity, chitinase activity/chitin catabolic process, protein amino acid phosphorylation, response to water, microtubule motor activity/microtubule-based movement, nucleosome and nucleus were also identified when comparing phenotype response to pyroxsulam (Fig. 3).
Expression patterns of contigs assigned to Pfam families involved in herbicide response
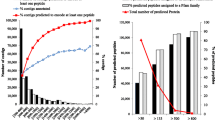
Pfam families known to be involved in herbicide response and NTSR are cytochromes P450, glutathione-S-transferases, glycosyltransferases and ABC transporters (Yuan et al. 2007; Délye 2013). Annotation identified 3,498 potential peptides assigned to these families in LOLbase, for which a considerable variation in the expression patterns was observed between phenotypes, over the time-course, and with the contig (Figs. 5, S6). The number of contigs up-regulated in one modality varied from 28 (ABC transporters) to 127 (UDP-glycosyltransferases) in the resistant phenotype, and from 55 (ABC transporters) to 204 (cytochromes P450) in the sensitive phenotype. Overall, more contigs assigned to these Pfam families were regulated by pyroxsulam application in the sensitive phenotype than in the resistant phenotype (Figs. 5, S6). Five, seven, six, four and three contigs annotated as cytochromes P450, UDP-glycosyltransferases, glycosyltransferases, glutathione-S-transferases or ABC transporters, respectively, were up-regulated at least threefold in the resistant phenotype over the whole time-course. Eight contigs could be considered as relevant candidates for NTSR. The 17 remaining contigs had low RPKM values (<0.5) in most modalities, or an expression level sharply decreasing after herbicide application (not shown).
Differences in the regulation of contigs assigned to Pfam families known to be involved in herbicide response and NTSR. P450, cytochromes P450; UDPGT, UDP-glycosyltransferases; ABC_tran, ABC transporters, GST, glutathione-S-transferases; Glyco_transf, glycosyltransferases. R resistant phenotype; S sensitive phenotype
The glutathione-S-transferase LrGSTF1 plays a significant role in rye-grass NTSR to ACCase inhibitors, an herbicide group distinct from ALS inhibitors (Cummins et al. 2013). Nine additional rye-grass contigs annotated as cytochromes P450, glutathione-S-transferases, monooxygenases or glycosyl-transferases were also proposed to have a link with NTSR to ACCase inhibitors (Gaines et al. 2014). A BlastN search in LOLbase identified a predicted glutathione-S-transferase fully encompassing LrGSTF1 coding sequence (identity = 94.7, E value = 2.8 × 10−168), and homologs of the nine other NTSR candidates (identities between 92.5 and 99.1 %, E-values between 2.1 × 10−56 and 0.0). LOLbase RNA-Seq expression patterns of these 10 contigs showed a constant increase with the time after herbicide application in both phenotypes, and their difference in expression between phenotypes was never above three-fold (not shown). These contigs could therefore not be considered as contigs potentially involved in NTSR to ALS inhibitors in our study.
Candidate NTSR contigs identification
To avoid possible confusing effects due to the commercial herbicide formulation, candidate NTSR contigs were selected from LOLbase on the basis of an up-regulation > threefold in the resistant phenotype at each time-point, including UT. Affiliation of the predicted peptide to a GO term enriched in up-regulated contigs exclusively in the resistant phenotype and/or to a Pfam family involved in NTSR was an additional selection criterion. Thirty candidate NTSR contigs were identified. Annotation identified six cytochromes P450, six glutathione-S-transferases, one glycosyltransferase, one zinc-finger protein, one carbonic anhydrase, two uromethyl-transferases and four DUF proteins. The nine remaining contigs could not be assigned an annotation.
As our RNA-Seq was conducted on pooled samples with no true biological replicates, RNA-Seq yielded the average expression of each contig over the three plants contained in each pooled sample. RT-qPCR based validation of LOLbase expression data revealed variation in contig expression among individual plants in each pooled sample (Figs. 1, S3). Accordingly, contigs up-regulated in each resistant plant compared to each sensitive plant at each time-course point were sought by measuring the expression level of the 30 contigs in all 42 individual samples used to generate the 14 pooled samples subjected to RNA-Seq. Four of the 30 contigs met this criterion. Their expression level was subsequently measured using RT-qPCR in 212 additional, untreated plants not containing herbicide resistant, mutant ALS and with characterised sensitivity to pyroxsulam and (iodosulfuron + mesosulfuron): 123 plants were resistant or moderately resistant to both herbicides, 56 sensitive to both herbicides and 33 resistant or moderately resistant to one herbicide (Table 1). All four contigs displayed a significantly higher expression level in the resistant plants (Figs. 6, S7). They putatively encoded cytochromes P450 from families 72A or 81B (referred to hereafter as CYP72A and CYP81B1), one glycosyltransferase (GTA) and one glutathione-S-transferase (GSTA).
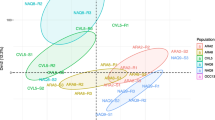
Principal component analysis of the expression levels of the four candidate contigs in 212 rye-grass plants with characterised sensitivity to pyroxsulam and (iodosulfuron + mesosulfuron). Expression levels were measured using RT-qPCR. Red dots, plants moderately or highly resistant to both herbicides; orange dots plants moderately or highly resistant to one herbicide and sensitive to the other; green dots plants sensitive to both herbicides. Principal component axes 1 and 2 explained 59.5 and 21.0 % of the variation, respectively
Cytochromes P450, glutathione-S-transferases and glycosyltransferases can be involved in NTSR via enhanced expression (Délye 2013). There was a 2,217-, 1,768-, 676-, and 125,032-fold difference in expression between the highest and lowest relative expression levels observed for CYP72A, CYP81B1, GTA and GSTA, respectively (Fig. S8). Variation was higher among plants than among populations. Expression of each of these four contigs was almost undetectable in a few plants, and very high in a few others (Fig. S8). For all these plants, both biological replicates showed the same extreme expression level, and an expression level of the reference genes similar to those of the other plants analysed. No plant displayed an extreme expression level for more than one of the four contigs.
Although some sensitive plants had expression levels higher than some resistant plants for a given contig, the highest expression levels for each contig were always observed in resistant plants (Fig. S8). Considering each herbicide separately, the nine, 23, one and three plants most transcribing CYP72A, CYP81B1, GTA and/or GSTA, respectively, were all moderately resistant or resistant to pyroxsulam. The 45, 23, eleven and ten plants most transcribing CYP72A, CYP81B1, GTA and/or GSTA, respectively, were all moderately resistant or resistant to (iodosulfuron + mesosulfuron). Plants sensitive to one herbicide could be resistant or moderately resistant to the other, thus not being “true” sensitive plants. Considering sensitivity to both herbicides improved phenotype discrimination, with the 47, 23, ten and 26 plants most transcribing CYP72A, CYP81B1, GTA and GSTA, respectively, being moderately resistant or resistant to at least one herbicide (Fig. S8).
NTSR is considered to evolve by accumulation of genes in a plant (Délye 2013; Délye et al. 2013). Considering the expression levels of all four contigs together should thus improve phenotype discrimination. Principal component analysis was implemented to check the hypothesis that expression data from the four contigs in the 212 plants could be a predictor for NTSR. A clear separation of many, but not all, plants resistant to at least one herbicide from sensitive plants was obtained (Fig. 7). Separation was driven by higher expression of several of the four contigs in plants with NTSR. From this finding, we computed a cumulated relative expression level (CREL) value for each plant as the sum of the relative expression levels of each contig in this plant divided by the highest corresponding relative expression level observed among the plants studied (Figs. 8, S9). Considering each herbicide separately, the 11 and 28 plants with the highest CREL were all moderately resistant or resistant to pyroxsulam or to (iodosulfuron + mesosulfuron), respectively. Considering both herbicides improved phenotype discrimination: the 47 plants with the highest CREL value (≥0.85) were moderately resistant or resistant to at least one herbicide (Figs. 8, S9).
Plant phenotypes sorted according to the cumulated relative expression level (CREL) value. CREL values and expression levels of the four candidate NTSR contigs are given in a heatmap. The average CREL value computed over the 212 plants analysd or the average contig expression value computed from expression values measured using RT-qPCR in the 212 plants analysed were used as a reference to establish the shade gradient in the corresponding column. Red, blue or white shades indicates CREL or contig expression values higher, lower or not different from the average value. Plant sensitivity to pyroxsulam and (iodosulfuron + mesosulfuron): orange highly resistant (R); grey moderately resistant (r); blue sensitive (S). Plant sensitivity to both herbicides: red moderately or highly resistant to both herbicides; yellow moderately or highly resistant to one herbicide and sensitive to the other; green sensitive to both herbicides
Discussion
LOLbase, a reliable rye-grass transcriptome resource
Our aim was to establish a resource enabling to study transcriptomic patterns in herbicide-resistant and herbicide-sensitive rye-grass plants in the absence of herbicide and at the early stages of herbicide response using experimental conditions as similar as possible to realistic field conditions. Herbicide damage to plants starts occurring 3–8 h after herbicide application (reviewed in Délye 2013). To be efficient, NTSR must be implemented before herbicide damage is irreversible, and must be upheld long enough to allow resistant plants to recover (Délye 2013). Accordingly, RNA-Seq data was obtained from a time course experiment ranging from UT until 48HAT. A commercial pyroxsulam formulation applied in the field was used together with its recommended adjuvant in our experiments, because NTSR is selected for in field populations by recurrent applications of not only herbicide molecules, but also associated formulations and adjuvants. The Illumina technology that is considered most adequate for de novo transcriptome assembly without a reference sequence (Ward et al. 2012) was used for RNA-Seq. LOLbase contains 91,928 contigs potentially representing active transcripts and predicted to encode 81,563 peptides, of which 46,451 could be assigned an annotation. LOLbase was established from the aerial part of young plants at a vegetative growth stage, and is thus not expected to be a comprehensive transcriptomic resource. Indeed, 185,833 contigs obtained by an RNA-Seq approach from leaf, root, stem and flowers from the perennial Lolium perenne were recently made available in GenBank (Farrell et al. 2014; accession GAYX00000000). As a comparison, the genomes of the five grass species fully sequenced (rice, sorghum, barley, maize and the weed Brachypodium distachyon) contain 25,532–63,540 genes. Rye-grass being a diploid species, the number of contigs in LOLbase suggests redundancy among the assembled contigs, i.e., more than one assembled contig may represent a same unigene and/or several contigs may represent different segments of the same unigene. LOLbase contig N50 size value was 771 nucleotides (Table 2), similar to those obtained for recent de novo plant transcriptome assemblies based on Illumina reads (Fan et al. 2013; Xu et al. 2013) or on a combination of Illumina and 454 reads (Yang et al. 2013), but shorter than the average length of protein-encoding genes in grasses (1,000–1,300 nucleotides; Carels and Bernardi 2000). LOLbase data compares favourably to a previous partial rye-grass transcriptome resource obtained from similar plant material using 454 sequencing (Gaines et al. 2014). The assembly statistics of this previous assembly (N50 = 1,150 nucleotides, average contig size = 1,049 nucleotides) were similar to those of LOLbase (Table 1), but a lower number of contigs were assembled [19,623 and 12,450 contigs >100 nucleotides and >500 nucleotides, respectively, vs. 92,381 contigs >400 nucleotides in LOLbase (Table 1)]. While redundancy was present in both assemblies, LOLbase representation of rye-grass transcriptome is most likely more comprehensive, due to the higher sequencing depth in our work (>32 × 109 nucleotides sequenced) compared to Gaines et al. (2014) (>405 × 106 nucleotides sequenced).
LOLbase contigs were assembled from RNA-Seq data obtained from three resistant and three sensitive plants from a field population, i.e., not genetically homogenised. Thus, nucleotide variation among the plants sequenced likely contributed to the redundancy observed in LOLbase. Considering these limitations, LOLbase annotation identified a total of 3,498 potential peptides assigned to Pfam families potentially involved in NTSR (cytochromes P450, glutathione-S-transferases, glycosyltransferases and/or transporters) (Yuan et al. 2007; Délye 2013), thereby providing a strong basis for further NTSR investigation.
Rye-grass response to pyroxsulam
The effect of ALS inhibitors had essentially been studied on broadleaved plants. It was previously summarised (Zhou et al. 2007) and further elucidated in recent works (Das et al. 2010; Orcaray et al. 2011; Zulet et al. 2013). Briefly, ALS inhibition is followed by growth arrest and subsequent slow plant death. ALS inhibitor application causes a rapid activation of specific stress response pathways, including xenobiotic metabolism (Manabe et al. 2007; Das et al. 2010). The biosynthesis of branched amino-acids (BCAAs) is interrupted, causing a decrease in the cell contents in free BCAAs followed by a rapid (12–24 h after herbicide application) overall increase in free amino-acid contents, including BCAAs, due to amino-acid remobilisation from proteins via enhanced degradation and to reduced protein synthesis (Trenkamp et al. 2009; Orcaray et al. 2011; Zulet et al. 2013). Side-effects of ALS inhibitors also include an increase in the leaf cell carbohydrate contents that triggers alternative respiratory pathways (Orcaray et al. 2011; Zulet et al. 2013) and modifications of central energy pathways in response to oxidative damage (Das et al. 2010).
LOLbase allowed insight into a grass response to ALS inhibitors. Part of rye-grass transcriptional response to pyroxsulam application was shared by resistant and sensitive plants: pyroxsulam caused a rapid (from 2HAT on) up-regulation of contigs involved in stress response, metabolism or detoxification that preceded an up-regulation of contigs assigned to BCAA biosynthesis or central energy pathways and a down-regulation of contigs assigned to photosynthesis (Table S4). This was consistent with plant growth arrest and with the expected early effects of ALS inhibition. Later effects of ALS inhibition such as enhanced protein degradation or carbohydrate accumulation were not detected in our experiment, probably because our time-course ended at 48HAT, while these effects were observed starting from 72HAT (Orcaray et al. 2010; Zabalza et al. 2013; Zulet et al. 2013).
In a previous transcriptome-wide survey, A. thaliana plants carrying an herbicide-resistant ALS displayed no transcriptional changes after application of an ALS-inhibiting herbicide, while their sensitive counterparts showed extensive transcriptional changes (Manabe et al. 2007). This is in contrast to our data, which suggest that rye-grass plants expressing NTSR to pyroxsulam underwent a stress exerted by ALS inhibitor application, with part of their transcriptional response to pyroxsulam action similar to that observed in the sensitive plants (Tables S4, S6; Figs. 3, 5). Our data are consistent with plants resistant because of NTSR being affected by the herbicide action but ultimately overcoming it, as previously proposed (Délye 2013).
Considering phenotype-specific response to pyroxsulam, herbicide-induced oxidative stress response seemed more marked in the sensitive phenotype (Table S7), which could reflect an herbicide stress stronger than in the resistant phenotype. General analysis of contigs differentially expressed between the resistant and the sensitive phenotypes did not allow identification of processes or functions that could directly explain NTSR. More contigs assigned to Pfam families involved in herbicide response were regulated by herbicide application in the sensitive phenotype than in the resistant phenotype (Fig. 5). This could reflect a random herbicide response of the sensitive phenotype, while the resistant phenotype response would be more focussed and more efficient.
From all the foregoing, it is clear that despite some redundancy and the genetic heterogeneity of the plant material used for RNA-Seq, LOLbase is a reliable resource to investigate rye-grass transcriptomic response and NTSR to ALS-inhibiting herbicides.
Candidate NTSR contigs
Transcriptomic data obtained from each pooled sample reflected the average variation in gene expression induced by biological variation among individuals (three plants per phenotype) and by the environment (two clones per plant). Yet, the pooled samples used for RNA-Seq did not contain distinct biological replicates, as is generally recommended to identify contigs differentially expressed among experimental modalities (e.g., Egan et al. 2012; Ward et al. 2012). Our experimental design was thus not optimal to identify contigs specifically up-regulated in resistant plants, especially because of variation in expression level among the individual samples constituting each pooled sample (Figs. 1, S3). However, rye-grass transcriptomic response to ALS inhibitor obtained from LOLbase expression data was fully consistent with the literature. This encouraged us to seek potential NTSR determinants using LOLbase expression data despite the limitations inherent to our experimental design. We considered contigs with a constitutive up-regulation in the resistant plants that was maintained during the early phases of the transcriptional response to pyroxsulam because NTSR is expected to be most effective in avoiding irremediable physiological damage if constitutive, and target contigs to be used as a target for NTSR detection would be most useful if constitutively differentially expressed in resistant plants. Only contigs which were confirmed by RT-qPCR to be up-regulated in each resistant plant compared to each sensitive plant at each time-point were retained as candidate NTSR contigs.
The four candidate contigs identified herein potentially encoded peptides with functions consistent with herbicide degradation: two cytochromes P450 (CYP72A and CYP81B1), one glycosyltransferase (GTA) and one glutathione-S-transferase (GSTA). Some cytochromes P450 in families 81B1 and 72A were directly involved in herbicide degradation (Werck-Reichhart et al. 2000). Rice CYP72A31 and barnyardgrass (Echinochloa phyllopogon) CYP72A254 that respectively shared 62.4 and 68.4 % amino-acid identity with CYP72A were very recently demonstrated to confer NTSR to ALS-inhibiting herbicides (Iwakami et al. 2014; Saika et al. 2014). CYP72A and CYP81B1 are thus promising NTSR candidates in rye-grass. Induction of glycosyltransferases like GTA triggered by the application of ALS-inhibiting herbicides was observed in A. thaliana (e.g., Manabe et al. 2007), although it was not clear whether these enzymes detoxified the herbicide or were part of a broader response. Glutathione-S-transferases like GSTA can catalyse the conjugation of an herbicide to glutathione, or act as peroxidases protecting the cell from oxidative damage that can derive from herbicide action. Although both roles of glutathione-S-transferases were demonstrated in NTSR to ACCase inhibitors (Cummins et al. 1999, 2009, 2013), no data are currently available regarding glutathione-S-transferase-mediated degradation of ALS-inhibiting herbicides.
The CREL of the four candidate NTSR contigs allowed to clearly identify many plants that were resistant to at least one of the ALS inhibitors studied (plants with CREL value ≥0.85). Considering plant sensitivity to both herbicides improved the reliability of the diagnosis (Figs. 8, S9), suggesting the candidate contigs would play a role in or be markers for NTSR to both herbicides. Despite pyroxsulam (a triazolopyrimidine) and (iodosulfuron + mesosulfuron) (sulfonylureas) belonging to dissimilar chemical families, 123 out of the 156 plants resistant to one herbicide were also resistant to the other. Thus, some mechanisms conferring NTSR to one herbicide likely also conferred resistance to the other. This is consistent with previous data supporting the hypothesis that evolution of NTSR to one herbicide decreases sensitivity to other herbicides, whatever their cellular target or chemical family, thereby facilitating further evolution of NTSR to a broader range of herbicides (Délye et al. 2013). In this hypothesis, purely chemical grass weed control is ultimately doomed, and integrated weed management must be implemented (Délye et al. 2013).
In summary, we identified four candidate NTSR contigs. Their expression levels varied among plants: some resistant plants showed a low level of expression for all four transcripts, and no resistant plant displayed a high level of expression for all contigs. This suggests that not all plants studied were resistant because of the same NTSR mechanisms and that NTSR determinants remain to be identified. This is in accordance with previous data showing that mechanisms controlling NTSR can vary among rye-grass populations and even among rye-grass plants (Busi et al. 2011, 2013), or can be herbicide-induced (Délye 2013).
Our data support the hypothesis that herbicide response and NTSR are driven by differential expression of a set of genes. Despite the limitations inherent to our experimental design and the genetic heterogeneity of the plant material studied, four contigs were identified that enabled reliable identification of many, but not all, NTSR plants in 15 different rye-grass populations on the basis of their expression levels. These four contigs are clearly potentially useful for developing molecular assays to help detecting NTSR in rye-grass populations. As gene expression regulation also involves post-transcriptional steps, the possible direct role in NTSR of these four contigs must be further investigated by confirming the increase in transcription causes an increase in the corresponding protein amount. The possible degrading activity against pyroxsulam and (iodosulfuron + mesosulfuron) of the proteins predicted to be encoded by the four contigs also needs to be assessed. Also, other NTSR determinants clearly exist in rye-grass that need to be identified. Additional RNA-Seq experiments using LOLbase and an appropriate experimental design including biological replicates are clearly a relevant way towards this purpose.
Herbicide degradation is a complex process involving the coordinated actions of detoxifying enzymes and regulatory genes (reviewed in Délye 2013). To fully understand NTSR, identifying the underlying genes and unravelling their interactions is a first step. This will enable developing DNA- or RNA-based NTSR diagnosis tools, and possibly contemplating neutralising NTSR using emerging technologies such as RNA interference (e.g., Li et al. 2013). In a second step, the evolutionary dynamics of NTSR selection needs to be elucidated so that effective integrated weed management strategies tailored to delay NTSR evolution and/or disadvantage NTSR plants in the field are implemented (Délye et al. 2013). Identifying NTSR genes seems near at hand, and LOLbase will clearly be of great help for this purpose.
References
Alexa A, Rahnenfuhrer J, Lengauer T (2006) Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 22:1600–1607
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106
Baker HG (1974) The evolution of weeds. Annu Rev Ecol Syst 5:1–24
Beckie HJ, Tardif FJ (2012) Herbicide cross resistance in weeds. Crop Prot 35:15–28
Briske DD (1991) Developmental morphology and physiology of grasses. In: Heitschmidt RK, Stuth JW (eds) Grazing management: an ecological perspective. Timber Press, Portland, pp 85–108
Burke GR, Strand MR (2012) Deep sequencing identifies viral and wasp genes with potential roles in replication of Microplitis demolitor bracovirus. J Virol 86:3293–3306
Busi R, Vila-Aiub MM, Powles SB (2011) Genetic control of a cytochrome P450 metabolism-based herbicide resistance mechanism in Lolium rigidum. Heredity 106:817–824
Busi R, Neve P, Powles S (2013) Evolved polygenic herbicide resistance in Lolium rigidum by low-dose herbicide selection within standing genetic variation. Evol Appl 6:231–242
Carels N, Bernardi G (2000) Two classes of genes in plants. Genetics 154:1819–1825
Charmet G, Balfourier F, Chatard V (1996) Taxonomic relationships and interspecific hybridization in the genus Lolium (grasses). Genet Resour Crop Evol 43:319–327
Cummins I, Cole DJ, Edwards R (1999) A role for glutathione transferases functioning as glutathione peroxidases in resistance to multiple herbicides in black-grass. Plant J 18:285–292
Cummins I, Bryant DN, Edwards R (2009) Safener responsiveness and multiple herbicide resistance in the weed black-grass (Alopecurus myosuroides). Plant Biotechnol J 7:807–820
Cummins I, Wortley DJ, Sabbadin F, He ZS, Coxon CR, Straker HE, Sellars JD, Knight K, Edwards L, Hughes D et al (2013) Key role for a glutathione transferase in multiple-herbicide resistance in grass weeds. Proc Natl Acad Sci USA 110:5812–5817
Das M, Reichman JR, Haberer G, Welzl G, Aceituno FF, Mader MT, Watrud LS, Pfleeger TG, Gutierrez RA, Schaffner AR, Olszyk DM (2010) A composite transcriptional signature differentiates responses towards closely related herbicides in Arabidopsis thaliana and Brassica napus. Plant Mol Biol 72:545–556
Délye C (2013) Unravelling the genetic bases of non-target-site-based resistance (NTSR) to herbicides: a major challenge for weed science in the forthcoming decade. Pest Manag Sci 69:176–187
Délye C, Boucansaud K, Pernin F, Le Corre V (2009) Variation in the gene encoding acetolactate-synthase in Lolium species and proactive detection of mutant, herbicide-resistant alleles. Weed Res 49:326–336
Délye C, Jasieniuk M, Le Corre V (2013) Deciphering the evolution of herbicide resistance in weeds. Trends Genet 29:649–658
Duhoux A, Délye C (2013) Reference genes to study herbicide stress response in Lolium sp.: up-regulation of P450 genes in plants resistant to acetolactate-synthase inhibitors. PLoS ONE 8:e63576
Egan AN, Schlueter J, Spooner DM (2012) Applications of next-generation sequencing in plant biology. Am J Bot 99:175–185
Fan HK, Xiao Y, Yang YD, Xia W, Mason AS, Xia ZH, Qiao F, Zhao SL, Tang HR (2013) RNA-Seq analysis of Cocos nucifera: transcriptome sequencing and de novo assembly for subsequent functional genomics approaches. PLoS ONE 8:e59997
Farrell JD, Byrne S, Paina C, Asp T (2014) De novo assembly of the perennial ryegrass transcriptome using an RNA-Seq strategy. PLoS ONE 9:e103567
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39:W29–W37
Gaines TA, Lorentz L, Figge A, Herrmann J, Maiwald F, Ott MC, Han H, Busi R, Yu Q, Powles SB, Beffa R (2014) RNA-Seq transcriptome analysis to identify genes involved in metabolism-based diclofop resistance in Lolium rigidum. Plant J. doi:10.1111/tpj.12514
Gouzy J, Carrère S, Schiex T (2009) FrameDP: sensitive peptide detection on noisy matured sequences. Bioinformatics 25:670–671
Heap I (2014) International survey of herbicide-resistant weeds. [WWW document]. http://www.weedscience.org. Accessed 31 July 2014
Huang XQ, Madan A (1999) CAP3: a DNA sequence assembly program. Genome Res 9:868–877
Humphreys J, O’Kiely P (2006) Amount and quality of grass harvested for first-cut silage for differing spring-grazing frequencies of two mixtures of perennial ryegrass cultivars with contrasting heading date. Grass Forage Sci 61:77–88
Iwakami S, Uchino A, Kataoka Y, Shibaike H, Watanabe H, Inamura T (2014) Cytochrome P450 genes induced by bispyribac-sodium treatment in a multiple-herbicide-resistant biotype of Echinochloa phyllopogon. Pest Manag Sci 70:549–558
Li JY, Mu JY, Bai JT, Fu FY, Zou TT, An FY, Zhang J, Jing HW, Wang Q, Li Z, Yang S, Zuo J (2013) PARAQUAT RESISTANT1, a Golgi-localized putative transporter protein, is involved in intracellular transport of paraquat. Plant Physiol 162:470–483
Manabe Y, Tinker N, Colville A, Miki B (2007) CSR1, the sole target of imidazolinone herbicide in Arabidopsis thaliana. Plant Cell Physiol 48:1340–1358
Martin JA, Wang Z (2011) Next-generation transcriptome assembly. Nat Rev Genet 12:671–682
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Oerke EC (2006) Crop losses to pests. J Agr Sci 144:31–43
Orcaray L, Igal M, Marino D, Zabalza A, Royuela M (2010) The possible role of quinate in the mode of action of glyphosate and acetolactate synthase inhibitors. Pest Manag Sci 66:262–269
Orcaray L, Igal M, Zabalza A, Royuela M (2011) Role of exogenously supplied ferulic and p-coumaric acids in mimicking the mode of action of acetolactate synthase inhibiting herbicides. J Agric Food Chem 59:10162–10168
Ozsolak F, Milos PM (2011) RNA sequencing: advances, challenges and opportunities. Nat Rev Genet 12:87–98
Petit C, Pernin F, Heydel JM, Délye C (2012) Validation of a set of reference genes to study response to herbicide stress in grasses. BMC Res Notes 5:18
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:W116–W120
Saika H, Horita J, Taguchi-Shiobara F, Nonaka S, Nishizawa-Yokoi A, Iwakami S, Hori K, Matsumoto T, Tanaka T, Itoh T, Yano M, Kaku K, Shimizu T, Toki S (2014) A novel rice cytochrome P450 gene, CYP72A31, confers tolerance to acetolactate synthase-inhibiting herbicides in rice and Arabidopsis. Plant Physiol. doi:10.1104/pp.113.231266
Trenkamp S, Eckes P, Busch M, Fernie AR (2009) Temporally resolved GC-MS-based metabolic profiling of herbicide treated plants treated reveals that changes in polar primary metabolites alone can distinguish herbicides of differing mode of action. Metabolomics 5:277–291
Untergasser A, Nijveen H, Rao X, Bisseling T, Geurts R, Leunissen JAM (2007) Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res 35:W71–W74
Vigueira CC, Olsen KM, Caicedo AL (2013) The red queen in the corn: agricultural weeds as models of rapid adaptive evolution. Heredity 110:303–311
Ward JA, Ponnala L, Weber CA (2012) Strategies for transcriptome analysis in nonmodel plants. Am J Bot 99:267–276
Werck-Reichhart D, Hehn A, Didierjean L (2000) Cytochromes P450 for engineering herbicide tolerance. Trends Plant Sci 5:116–123
Xu P, Liu Z, Fan X, Gao J, Zhang X, Zhang X, Shen X (2013) De novo transcriptome sequencing and comparative analysis of differentially expressed genes in Gossypium aridum under salt stress. Gene 525:26–34
Yang X, Yu XY, Li YF (2013) De novo assembly and characterization of the barnyardgrass (Echinochloa crus-galli) transcriptome using next-generation pyrosequencing. PLoS ONE 8:e69168
Yuan JS, Tranel PJ, Stewart CN Jr (2007) Non-target-site herbicide resistance: a family business. Trends Plant Sci 12:6–13
Zabalza A, Zulet A, Gil-Monreal M, Igal M, Royuela M (2013) Branched-chain amino acid biosynthesis inhibitors: herbicide efficacy is associated with an induced carbon-nitrogen imbalance. J Plant Physiol 170:814–821
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18:821–829
Zhao J, Ohsumi TK, Kung JT, Ogawa Y, Grau DJ, Sarma K, Song JJ, Kingston RE, Borowsky M, Lee JT (2010) Genome-wide identification of polycomb-associated RNAs by RIP-seq. Mol Cell 40:939–953
Zhou Q, Liu W, Zhang Y, Liu KK (2007) Action mechanisms of acetolactate synthase-inhibiting herbicides. Pestic Biochem Physiol 89:89–96
Zulet A, Gil-Monreal M, Villamor JG, Zabalza A, van der Hoorn RAL, Royuela M (2013) Proteolytic pathways induced by herbicides that inhibit amino acid biosynthesis. PLoS ONE 8:e73847
Acknowledgments
We thank Dr Valérie Le Corre (INRA Dijon, France) for assistance in statistical analyses, and Genotoul—Génopôle Toulouse Midi-Pyrénées for allowing access to computational facilities.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Duhoux, A., Carrère, S., Gouzy, J. et al. RNA-Seq analysis of rye-grass transcriptomic response to an herbicide inhibiting acetolactate-synthase identifies transcripts linked to non-target-site-based resistance. Plant Mol Biol 87, 473–487 (2015). https://doi.org/10.1007/s11103-015-0292-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0292-3