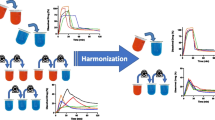
Abstract
For successful oral drug development, defining a bioequivalence (BE) safe space is critical for the identification of newer bioequivalent formulations or for setting of clinically relevant in vitro specifications to ensure drug product quality. By definition, the safe space delineates the dissolution profile boundaries or other drug product quality attributes, within which the drug product variants are anticipated to be bioequivalent. Defining a BE safe space with physiologically based biopharmaceutics model (PBBM) allows the establishment of mechanistic in vitro and in vivo relationships (IVIVR) to better understand absorption mechanism and critical bioavailability attributes (CBA). Detailed case studies on how to use PBBM to establish a BE safe space for both innovator and generic drugs are described. New case studies and literature examples demonstrate BE safe space applications such as how to set in vitro dissolution/particle size distribution (PSD) specifications, widen dissolution specification to supersede f2 tests, or application toward a scale-up and post-approval changes (SUPAC) biowaiver. A workflow for detailed PBBM set-up and common clinical study data requirements to establish the safe space and knowledge space are discussed. Approaches to model in vitro dissolution profiles i.e. the diffusion layer model (DLM), Takano and Johnson models or the fitted PSD and Weibull function are described with a decision tree. The conduct of parameter sensitivity analyses on kinetic dissolution parameters for safe space and virtual bioequivalence (VBE) modeling for innovator and generic drugs are shared. The necessity for biopredictive dissolution method development and challenges with PBBM development and acceptance criteria are described.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To support drug product quality and performance, the FDA has issued a draft guidance for the industry to advocate the use of oral biopharmaceutics modeling approaches in combination with in vivo bioavailability (BA)/bioequivalence (BE) studies [1]. The demonstration of BE, indicating that “rate and extent of drug absorption” are similar between a reference and a test formulation, is an important part of successful formulation development in both generic and innovator drug companies. Bioequivalence is needed for e.g. bridging a prototype formulation to a marketed formulation, or getting a new generic drug formulation approved [2, 3]. The consistency of clinical performance of drug products has to be ensured by appropriately setting in vitro specifications (e.g. in vitro dissolution, particle size distribution) [4] that are within the BE safe space.
A BE study is usually recommended for new drug products when pre- or post-approval changes lead to out of the specification results, or for generic drug products for direct comparison to a marketed reference listed drug (RLD) [2]. In vitro dissolution testing is a common method to compare batches or drug products. A similarity f2 test is recommended by the health authorities to demonstrate the similarity between dissolution profiles [2]. When the similarity factor f2 > 50, then similarity between two drug products/batches is demonstrated. However, f2 < 50 does not necessarily indicate non-BE [5,6,7] and f2 tests can be overdiscriminating [8]. The development of a BE safe space can offer the opportunity to confirm or expand the clinically relevant dissolution specification (CRDS) [1].
The BE safe space “defines the boundaries of in vitro specifications (dissolution or other relevant drug product quality attributes), within which drug product variants are anticipated to be bioequivalent to one another” [9]. The establishment of a safe space offers a mechanistic understanding of the impact of in vitro drug product properties on the in vivo pharmacokinetic (PK) performance and may help to reduce unexpected or suboptimal in vivo PK outcomes.
In generic and innovator product development [7, 10,11,12,13], establishing safe space offers multiple advantages and applications that can range from product development decisions to regulatory justifications. Selected case examples are shown in Table 1. Successful PBBM applications early in the product development help to identify CBAs (critical bioavailability attributes). CBAs are formulation attributes, such as particle size, dissolution rate, tablet compression force, which can be expected to “critically impact the bioavailability (absorption rate and extent) of a drug product” [28]. CBA is a newer term coined by regulatory agencies along with critical process parameters (CPP), critical material attributes (CMA), critical quality attributes (CQA) and critical formulation variables (CFV) [28]. Ideally CBA can be either of these attributes and if defined during the generic product development, it helps eventually to embed in vivo aspects into the product development. Defining the safe space for dissolutions and CBA’s can also help to justify the QC media dissolution specifications and helps to reject bio-inequivalent batches and to identify dissolution boundaries. Additionally, in cases where lower or higher strength biowaivers could not be achieved due to dissolution dissimilarity or due to lack of IVIVC, the safe space can serve as an effective tool to demonstrate BE. Apart from these applications, defining safe space can also aid to support biowaivers related to modification in the manufacturing process, composition changes and site transfers which can serve as backbone for biopharmaceutics risk assessments [29, 30].
PBBM may be useful to confirm or widen the safe space, sometimes superseding f2 similarity criteria (Table 1). For BCS class I/III drugs, BE is defined by in vitro dissolution, which sets the acceptable criteria as 80% dissolved in 30 min and 15 min for BCS I and III, respectively [31]. For some BCS I and III drugs, the dissolution BE criteria may be too stringent and not biopredictive. In such cases the justification of wider dissolution specifications may be pursued (Table 1). BCS II drugs, have low solubility in the physiology pH range and high permeability, for which absorption is a dissolution-limited process. Defining clinically relevant specification with PBBM for BCS II involves prediction of clinical exposure and examples are documented in the literature (Table 1) [12, 13]. For BCS IV drugs, with moderate solubility and permeability, PBBM may be used to show that absorption is more permeability rate controlled than by solubility/dissolution [7].
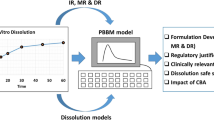
Mechanistic modeling or PBBM modeling both provide another perspective to link in vitro data and in vivo pharmacokinetics. PBBM modeling approaches have received increased attention in recent years, as demonstrated by an FDA draft guidance [9], publications [11] and workshops [28]. General PBBM applications are shown in Fig. 1 [10].
This review article lists selected case studies to highlight the development of PBBM BE safe space and its applications, from both innovators and generic sides. Detailed workflow of model development and technical discussion on the incorporation of dissolution profiles and virtual BE setup are reviewed. The case studies and examples are limited to oral administration.
Case Studies
General Considerations and Modeling Approaches
A general workflow of PBBM development is represented in detail, in Fig. 2. One application of PBBM is its use to supersede f2 results, wherein when f2 data demonstrate dissolution dissimilarity (f2<50); PBBM can be utilized to demonstrate that despite dissolution dissimilarity, clinical BE can be achieved (Fig. 3, case studies 2 & 4). The use of PBBM for safe space and knowledge space determinations in generic and innovator drug development is shown schematically in Fig. 4, also in case studies 1 & 3. To develop a PBBM, disposition parameters are needed to describe drug distribution and are independent of the route of administration. The disposition parameters are preferably generated from intravenous (IV) PK data, if not, oral solution PK with a low dose can be used. Physiochemical and biopharmaceutics parameters can be obtained from in vitro characterization or literature. Upon development of the initial PBBM model, a clinical observed PK profile will be used for model verification by comparing observed AUC and Cmax with predicted values. Parameter optimization and parameter sensitivity analysis can be conducted if necessary. The developed PBBM model will be challenged and validated against other independent clinical PK data from different sites or with different batches. If the model meets the validation criteria, it can be applied for its intended use.
PBPK modeling workflow with different dissolution model, including Johnson model [32]; Weibull function [33,34,35]; Takano model (z-factor) [36]; diffusion layer model (DLM) [37, 38]; p-psd model [13, 39] The Takano model figure was modified from [40]. The P-PSD figure was derived from [40]. Independent clinical data set should be used for model validation. * Innovator – datasets from Ph-I, II with different formulation variants. Generic – pilot BE data **Direct input of in vitro dissolution is not desirable due to the limited time points [41]
Illustration of a PBBM based BE safe space to supersede f2 similarity. PBBM indicates that test and reference are in the BE safe space (right side). This is in contrast to the f2 test, which had indicated non-similarity and non-BE (upper left). In this case PBBM indicates a wider dissolution BE safe space than the f2 similarity test, see also Case 4
Comparison of development of the BE safe space between innovator drugs and generic drugs [41]
For the case studies shown in this review, simulation was conducted using GastroPlus software v.9.8 (Simulations Plus, Inc. Lancaster, CA). Physicochemical and pharmacokinetic input parameters for all case studies are summarized in Table 2. The default optimized logD model SA/V 6.1 ASF model was considered in all the models, if not specified. Dissolution model selection follows the decision tree, shown in Fig. 2.
Case Study 1: PBBM to Widen Dissolution Specifications for a BCS II Drug Compound in Generic Drug Development
Biopharmaceutics Properties and Background
ZY-ABC is classified as a BCS class II compound with Log P of 2.5 and it is a weakly acidic compound with a pKa of 3.1. ZY-ABC has high permeability and low solubility at acidic pH. At pH greater than 6, the solubility is high. Upon oral administration, Tmax occurs between 1 and 4 hours. Following oral administration, pharmacokinetics are linear over the dose range of 7-14 mg and absorption is >90%. Food does not have a clinically relevant effect on the pharmacokinetics of ZY-ABC. The dissolution profiles of the test and reference formulations were generated experimentally in 1000 mL pH 6.8 phosphate buffer, using an USP apparatus II (Paddle) at 50 rpm.
The original dissolution specification of ZY-ABC Tablets was Q = 80% at 20 minutes, which had been recommended by a health authority. The objective of this PBBM modeling was to support the widening of clinically relevant dissolution specifications for ZY-ABC tablets to Q = 75% at 30 minutes.
Simulation Methodology
The modeling strategy consisted of three steps: model setup, model validation, and model application (Fig. 2). PKPlus® was used to obtain two-compartment disposition parameters from literature reported intravenous profile and exported to ACAT™ model. ‘Human physiology fasting’ was considered for the simulation because the fasting condition is more discriminative to reflect the impact of the dissolution differences on bioequivalence. The model was conducted using the z-factor model after fitting to the dissolution data at pH 6.8. Optimization was not done for any of the parameters. Additional drug records were produced in GastroPlus™; i.e. the exploratory test formulation with a 20% slower theoretical dissolution profile.
The PBBM model was developed with physicochemical, biopharmaceutics, and pharmacokinetics parameters, and it was validated against the clinical PK data of the pivotal test formulation batch. The developed model was then verified by comparing the predicted PK profiles with the observed PK profiles for IV formulation, the exploratory test formulations with a different release rate, and the reference formulation. The percentage of prediction error (%PE) was calculated for all the validations. The validated model was further applied to perform virtual bioequivalence simulations. An additional simulation was performed with the theoretical slower batch (i.e. 20% slower dissolution profile than the dissolution profile observed for the pivotal test batch). The population simulator was used to run a virtual bioequivalence trial between the test batch and a theoretical, 20% slower batch with a two-way crossover trial (n = 24 subjects). GastroPlus™ cross-over virtual trial feature was used where a population is first generated and then reloaded to run crossover studies with a new formulation. All the population parameters were kept as default values in GastroPlus™ during the population simulation because of the low intra-subject variability (ISCV) of the drug product observed in clinical studies. The mean and 90% confident interval (CI) for Cmax and AUC0-t from the virtual BE simulation were calculated. When all the BE parameters fell into the 90% CI BE criteria i.e. between 80% and 125%, the trial was classified as “pass”; otherwise, it was a “fail” [2].
Results and Discussion
The in vitro dissolution profiles are presented in Table 3. The results of parameter sensitivity analysis of z-factor on AUC and Cmax showed that z-factor does not impact AUC and Cmax due to the high solubility at high pH range (Fig. S2). The simulated mean PK profiles for the test formulation of ZY-ABC tablets is comparable with the observed mean PK profiles under fasting conditions, with the %PE of 2.6% and 10% for Cmax and AUC0-t, respectively (Table 4).
Validation of the model against the IV infusion profile, exploratory test formulation with a different release rate, and reference formulation by incorporating corresponding dissolution profiles and inputs, was shown in Table 5. The %PE values are well below the acceptance criteria (%PE < 15%).
The built model was further used to simulate virtual bioequivalence trial, (n = 24 subjects) and it indicated that drug product with a theoretical 20% slower dissolution profile was found to be bioequivalent to the actual test formulation, used in pivotal bioequivalence study (Table 5 and Fig. 5).
Case study 1, Virtual bioequivalence simulation and analysis with 90% confidence interval around the mean (90% CI) for 24 subjects between a virtual (theoretical) 20% slower formulation (test formulation: green) vs the exhibit test batch formulation (reference formulation: pink).The green highlighted box indicates bioequivalance between two batches
Conclusion
Oral biopharmaceutic modeling e.g. PBBM, provides increased confidence to understand the impact of dissolution profiles on absorption as compared with a mathematical method such as an IVIVC. Additionally, physiological variables such as pH, GI transit time, compartment volumes, and gastric emptying patterns, are the key to determining in vivo dissolution rate and absorption, which have been incorporated into the PBBM modeling.
A safe space was defined, and a clinically relevant dissolution specification for ZY-ABC was proposed based on the outcome of the virtual BE trials. It was demonstrated that the proposed Q = 75% dissolved within 30 minutes was well within a region of dissolution profiles where bioequivalence is anticipated.
Case Study 2 PBBM to Support Lower Strength Biowaiver in Case of f2 Mismatch (f2 Fails/BE Passes)
Biopharmaceutics Properties and Background
The DRL compound is a weakly acidic molecule formulated as an immediate-release tablet with five strengths: 40 mg, 80 mg, 120 mg, 160 mg, and 320 mg. The compound can be classified as a BCS III drug with Log D of −1.17 (pH 7) with acidic pKa of 5.6 and 3.7. The solubility is low in acidic condition (0.045 mg/mL at pH 1.2) and as pH increases, the solubility increases (0.333 mg/mL at pH 4.5 and 3.029 mg/mL at pH 6.8, all pH values are initial pH). After oral administration, the absorption was rapid with a Tmax of 2-4 h with linear pharmacokinetics. Dr. Reddy’s developed generic IR test formulation for all strengths, and as per regulatory guidance, a crossover bioequivalence study in the fasting study was conducted at higher strength 320 mg in healthy human subjects and bioequivalence was achieved between reference and generic formulations.
Based on the successful bioequivalence study on higher strength 320 mg, the waiver for 40 mg, 80 mg, 120 mg, 160 mg was requested. In order to obtain biowaiver for all lower strengths, dissolution tests in multi-media (pH 1.2, pH 4.5 and pH 6.8, 1000 mL, 50 rpm, USP II (Paddle) at 37 ± 0.5°C) were performed for all strengths as per the current EMA regulatory guidance [43]. The estimated similarity factor (f2) values were found >50 for all the batches, except for the lowest strength of test formulations, 40 mg, for which f2 was found to be 43 and less than 50 in pH 4.5 acetate buffer. Based on this f2 value, the regulatory agency denied the waiver for lower strength 40 mg, and hence the possibility of conducting a separate bioequivalence study for 40 mg study was anticipated. However, in order to obtain a waiver for the 40 mg bioequivalence study, physiologically based biopharmaceutics modeling (PBBM) was conducted to assess the impact of this lower f2 value on in vivo pharmacokinetics (PK) of the molecule.
Simulation Methodology
The PBBM was used to evaluate the impact of failing f2 on the bioequivalence. The modeling strategy included utilizing the pH-solubility profile generated in-house along with Log P and pKa values. The human effective permeability value was optimized to capture in vivo behavior and the default precipitation time was used. Plasma protein binding and blood to plasma ratio were obtained from literature and used in simulations. Elimination parameters were obtained by fitting literature-based intravenous pharmacokinetic profiles into the PKPlus module.
To simulate PK profiles, the rate of dissolution described by Z-factor as a function of pH was estimated by fitting multimedia in vitro dissolution profiles to built-in z-factor model in GastroPlus®. Overall, the PBBM model was developed using IV disposition parameters, physicochemical properties, and dissolution rate (Z-factor) vs pH data, and the model was validated against Cp-time profiles of pivotal test and reference formulations from clinical BE study, which was conducted by Dr. Reddy’s Laboratories in healthy volunteers in the fasting condition. The model validation was conducted with virtual population as well. Once the model was validated, the dissolution profiles of 40 mg strength were fitted with a Z-factor vs pH model and used as input for simulation of 320 mg strength. These simulated concentration-time profiles following oral administration using in vitro data of the test batch of 40 mg (given as input for 320 mg) were compared with clinical data of pivotal 320 mg test and reference formulations.
Results and Discussion
The PBBM model integrated DRL compound physicochemical properties with the use of the Z-factor dissolution model (multiple pH vs z-factor) method (Fig. 6) and a single compartmental model (top-down approach). The simulated plasma concentration-time profiles with virtual bioequivalence simulations of 68 subjects predicted the observed concentrations well (Fig. 7) with prediction errors lower than 10% for absolute PK parameters (i.e. Cmax and AUC) and for T/R ratio’s, which was considered acceptable from a model validation perspective (Table 6). The predicted T/R ratios correlated well with predicted T/R ratios of the pivotal study, and confidence intervals were within 80-125%, indicating the robustness of the PBBM model (Table 7).
Case study 2, Dissolution profiles of pivotal RLD and Test 320 mg products along with z-factor fitting at various pH conditions. The dissolution rate is highest at pH 6.8, moderate at pH 4.5 and slowest at pH 1.1 for both formulations. At pH 4.5 (middle profile), a dissolution mismatch with f2 < 50, was observed for 40mg against 320mg
Case study 2, Virtual BE simulations with 68 subjects (A) population simulation of pivotal RLD 320 mg (B) population simulation of pivotal test 320 mg. In Figure (A) and (B), individual PK profiles (pink dots) was compared with simulated mean exposure profiles (solid blue line), where simulations with a different level of confidence interval were presented as light blue line. (C) Virtual BE of pivotal RLD (pink: reference) vs test 320 mg (green: test). The probability contours are shown in thin blue lines in (A) and (B). Highlighted green box in the middle of each figure represents a bioequivalence comprising two batches at 90% confidence interval
The validated model was subsequently applied to 40 mg batch where dissolution mismatch was observed in pH 4.5 media. Virtual BE trial was then performed with the previously validated population using Z-factor vs pH data of lower strength test formulation batch of 40 mg as dissolution input. This represents the worst-case simulations wherein 40 mg dissolution data were used as input for simulating the highest strength 320 mg pharmacokinetics. These simulated concentration-time profiles following oral administration using in vitro data of the test batch of 40 mg were within BE limits (Cmax, AUC0-t, and AUC0-inf well within 80-125% range) of clinically observed mean PK data for the test batch of 320 mg.
Hence, PBBM simulations were successfully utilized to show that dissolution differences observed on the lower strength formulation at pH 4.5 were anticipated to have no impact on human PK.
Case Study 3 Safe Space Exploration for Etoricoxib Tablets [12]
Biopharmaceutics Properties and Background
Briefly, etoricoxib is a selective inhibitor of cyclooxygenase-2 (COX-2) which is completely absorbed in human, as indicated by a bioavailability of 100% under fasted conditions and the compound exhibited linear pharmacokinetics in a single- and multiple-dose study [44]. Etoricoxib is a weak base BCS Class 2 drug with pH-dependent solubility at physiological pH values. Etoricoxib solubility is >25 mg/mL at pH 2 and 0.05 at pH 6.9 [45].
Simulation Methodology
The original etoricoxib bioequivalence modeling is one of the earlier published PBBM applications where PBBM had been used successfully to predict bioequivalence for tablet batches when f2 multi-media dissolution comparisons had indicated non-similarity and non-bioequivalence, suggesting that f2 were non-biopredictive [45]. The modeling set-up for the base model had been described previously [45] and available clinical data from IV and PO administration were used [46]. Clinical data for a non-BE batch are not available.
The original etoricoxib model had used dissolution data directly as an input in the simulation [45]. More recently for the safe space building the mechanistic z-factor dissolution models were used which are described in detail in a previous article [36]. Etoricoxib PBBM input data are shown in Table 2 relisted from reference [45] . The use of dissolution z-factor model parameter sensitivity analyses at various pH-values has also been described for warfarin bioequivalence assessments using GastroPlus [46] and is suitable for IR formulations. For etoricoxib, IR formulations’ dissolution data were modeled in two steps. In the first step, data at pH 6.8 were fitted to the z-factor model. The z-factor was then subsequently varied in GastroPlus until the simulated Cmax was at least 20% lower, which was defined as the lower end of the safe space. The same steps were conducted for pH 2 media. Dissolution profiles were modeled and fitted in GastroPlus by choosing a stomach or small intestinal pH and volume which was mimicking the dissolution bath conditions. The stomach transit time was set to infinity to maintain a constant volume during the simulation as previously suggested [45]. The fitted z-factors were then used in the simulations. A surface pH correction was done, as etoricoxib is formulated as a weak base, and thus the surface pH, which controls dissolution is higher that the bulk pH at pH 2 [42].
Results and Discussion
Modeling results for predicted PK parameters derived in GastroPlus with a stepwise reduction of the z-factors in the stomach and the intestine are shown in Table 8, reproduced from [12]. The complete bioequivalence safe space plot for pH 2 and pH 6.8 media can be found in [12]. PBBM accurately described the observed pharmacokinetic data for etoricoxib [12]. The z-factor was overall similar and independent of the pH media used (0.003 vs. 0.001 mL/mg/s), consistent with the equation provided by Takano [36], which does not include pH or solubility as a variable in the definition of the z-factor. This PBBM approach, where a bioequivalence safe space can be explored and defined virtually by varying the z-factor is well suited for compounds where it is not practical or feasible to manufacture non-bioequivalent batches. Generating Cmax and AUC estimates as a function of the z-factor (Table 8), down to the edge of the dissolution (or z-factor) BE space (where Cmax is near 80% of the reference batch) has been used by previously [36].
Case Study 4: Establishment of Safe Space to Support a Biowaiver for Ribociclib (F2 Fails/BE Passes) [7]
Biopharmaceutics Properties and Background
Ribociclib is a weakly basic BCS 4 drug with highly pH-dependent solubility and low solubility within intestinal pH range (~0.8 mg/mL at pH 6.8) and moderate permeability [47]. Given these properties, permeability may rate-limit absorption. Ribociclib pharmacokinetics are not altered by food and there are no significant pH-mediated drug-drug interactions with acid reducing agents [47]. The evaluated formulations are with the same amount of API of ribociclib free base, and similar excipients with no impact on permeability [7].
A PBBM model was developed to confirm the bioequivalence between an early capsule formulation and a tablet (BE batch). Moreover, the bioequivalence between a tablet BE batch and a tablet commercial batch was evaluated. Finally, the tablet BE safe space using dissolution QC method was established.
Simulation Methodology
The GastroPlus PBBM model for ribociclib was previously reported [7]. The model strategy follows the PBBM development workflow, as shown in Fig. 2. In this study, four clinical studies were used for modeling development and validation. The model described the baseline PK data from the food effect study using capsule formulation in heathy volunteer, and then was validated against 1) FIH study using capsule formulation in patients, 2) BE study between capsule and tablet (BE batch) in healthy volunteer and 3) Phase 1b/2 study using capsule and tablet (commercial batch).
Dissolution data for the capsule, tablet (BE batches) and tablet (commercial batch) were available at pH 1, 2, 4.5 and pH 6.8. The f2 value for capsule and tablet (BE batch) was <50, indicating dissimilarity between these two formulations, as shown in Fig. 8. Three different dissolution models were evaluated as also shown schematically in Fig. 2: 1) using the ‘Johnson’ dissolution model to account for particle size; 2) fitting in vitro dissolution profiles at pH 2 to simulate dissolution in the stomach with a single Weibull function, 3) using the Takano z-factor model to fit dissolution profiles across pH 1, 2, 4.5, and 6.8 QC media.
Case study 4, based on f2 similarity test, capsule and tablet show an f2 mismatch, as f2 < 50, indicating non-similarity and non-BE (upper left graph). In vitro dissolution profiles are not considered similar according to f2 test. PBBM, using the ACAT model includes the moderate permeability of ribociclib. PBBM estimates that in vivo absorption is similar and correctly describes BE for capsule and tablet formulations (Replotted from [7]) Reprinted from Journal of Pharmaceutical Sciences, 111(1), Laisney, M., Heimbach, T., Mueller-Zsigmondy, M., Blumenstein, L., Costa, R. and Ji, Y., 2022. Physiologically Based Biopharmaceutics Modeling to Demonstrate Virtual Bioequivalence and Bioequivalence Safe-space for Ribociclib which has Permeation Rate-controlled Absorption. pp.274-284. Copyright (2022), with permission from Elsevier
Systemic PK parameters were derived “top-down” to fit the arithmetic mean PK profile from a food effect study using capsule in healthy volunteers using PKPlus and GastroPlus (Fig. S4). Virtual clinical trial simulations and VBE were conducted using Gastroplus (7). Each virtual BE assessment contained 250 subjects, including 25 subjects per trial and 10 trials.
Results and Discussion
The developed PBBM model has successfully described the observed clinical PK for four independent studies with an acceptable prediction errors (%PE) (<25%) for Cmax and AUC (Fig. S4) across studies. Three dissolution incorporation approaches have proved the absorption of ribociclib was dominated by permeability instead of dissolution rate, by comparing the in vivo dissolution and in vivo absorption plots and through parameter sensitivity analysis results (data not shown).
Capsule and tablet (BE batch) were predicted BE by the PBBM model, which was confirmed by the clinical BE study. Although the f2 value of dissolution profiles for the two formulations were smaller than 50, the PBBM model has superseded f2 testing and proved BE. Further, to establish the safe space, tablet BE batch was used as the reference batch, and simulations were run by changing “time-scale” factor of Weibull function, extrapolated from pH 2 dissolution profiles (QC method). A difference less than 20% between maximal and minimal AUC or Cmax was considered to denote a safe space following the principles of the IVIVC guidance [48]. VBE simulations with hypothetical dissolution profiles as inputs, demonstrate that the proposed dissolution specification (80% dissolved at 45 min at pH 2) fell within the safe-space where BE is anticipated (Figs. 9 and 10).
Case study 4, Example of a safe space plot. The ribociclib tablet BE safe space was defined based on in vitro dissolution in pH 2 media which was used as an input to a GastroPlus PBBM. The light green is the knowledge space, for which clinical data are available and which have been linked to biopredictive dissolution data. The light yellow area represents the extrapolated calculated BE safe space by simulation (Replotted from [7]). The light green line represents the edge of the calculated safe space. A clinically tested non-BE batch was not available, thus the knowledge space is limited to the light green area. Reprinted from Journal of Pharmaceutical Sciences, 111(1), Laisney, M., Heimbach, T., Mueller-Zsigmondy, M., Blumenstein, L., Costa, R. and Ji, Y., 2022. Physiologically Based Biopharmaceutics Modeling to Demonstrate Virtual Bioequivalence and Bioequivalence Safe-space for Ribociclib which has Permeation Rate-controlled Absorption. pp.274-284. Copyright (2022), with permission from Elsevier
Replotted from [7], permission received. Case study 4, Ribociclib, example of virtual BE simulations with 25 virtual subjects of a reference batch (green solid line) and test batch (pink solid line). The pink area and green area represent the simulated drug concentration with 90% confidence interval for test batch and reference batch, respectively. The light blue solid lines are the simulated drug concentration with confidence intervals ranging from 10 to 100%. The green highlighted box indicates a bioequivalance between the two batches. Reprinted from Journal of Pharmaceutical Sciences, 111(1), Laisney, M., Heimbach, T., Mueller-Zsigmondy, M., Blumenstein, L., Costa, R. and Ji, Y., 2022. Physiologically Based Biopharmaceutics Modeling to Demonstrate Virtual Bioequivalence and Bioequivalence Safe-space for Ribociclib which has Permeation Rate-controlled Absorption. pp.274-284. Copyright (2022), with permission from Elsevier
Discussion
Scientific applications of PBBM, also referred to as Physiologically based pharmacokinetic (PBPK) Modeling and Simulation in Biopharmaceutics, were recently reviewed by Wu et al. [29]. Health authorities’ interest in PBBM/PBPK and is underscored by the release of an FDA draft guidance on PBPK analyses for biopharmaceutic applications as discussed above [1]. Approximately twenty-five IQ companies have provided detailed feedbacks [19], requested clarification stating that PBPK could be used for study waivers under certain conditions. In general PBBM applications can be used at all stages of drug formulation development and broad applications have been found in generic and innovator drug companies [10] (Fig. 1).
Selected PBBM examples from the literature have been compiled in Table 1. Four detailed representative case studies from drug innovator and generic drug companies are shown in Table 2. These case studies show successful PBBM with %PE typically ≤25% and which were submitted and accepted for their intended application (Table 2). For dissolution profile data modeling, a key component of PBBM, several models have been used with success (Fig. 2, Table 1 and 2).
Common Challenges in PBBM Applications
Common challenges are highlighted in bold boxes in Fig. 2, and include appropriate input parameter selection, dissolution profile model choice, setting model acceptance criteria, and defining the intended purpose/applications, e.g. widening of safe space, superseding f2 etc.
PBBM Model Inputs & Parameter Optimization
PBBM modeling involves multiple inputs for API, formulation, physiology and pharmacokinetics. API inputs such as Log P, particle size, solubility (aqueous and biorelevant) can be obtained from in vitro experiments and can be directly used in the model. Formulation inputs such as dissolution can be obtained from in vitro experiments and other inputs such as hardness, disintegration time and precipitation time can be indirectly accounted by the dissolution. It is critical to use correctly measured parameters to inform modeling, however, some parameters, e.g. precipitation time, are not easy to obtained from in vitro testing, especially for weak bases and their salts form or amorphous compounds. A commonly used approach to estimate precipitation time is through transfer dissolution model, e.g. two-stage dissolution, gastrointestinal simulator (GIS) and biorelevant gastrointestinal transfer (BioGIT) [49,50,51]. The precipitation time should be further validated and optimized upon receiving clinical PK data. Most of the platforms come with default physiology settings and alterations of physiology is not usually recommended but can be done in few cases with sufficient explanation, e.g. the evaluation of pH-dependent DDI, the change of gastric transit time or the change of water content in the GI tract [13, 52]. For disposition parameters, intravenous data is preferred but in absence of that, an oral PK data with low dose corrected for bioavailability can be used as input. During the model development it may happen that with actual experimental in vitro data and PK data, the model is not able to capture plasma concentration profiles accurately due to variability or study-to-study difference between data used for model building and validation. In such cases, model parameters may be optimized “topdown”, but a mechanistic explanation is needed for justification. For parameter optimization, manual optimization or software based optimization can be utilized to fit the study data, and explanations need to be given. From the regulatory perspective, any optimization of model parameters requires a mechanistic explanation along with the submission of initial values, parameter range tested, the estimation method and optimization algorithm, as well as the in vitro and in vivo data used for optimization [1]. For such purpose, relevant parameter sensitivity analyses can be used to demonstrate that the optimized values represent the observed data.
Dissolution Profiles Modeling
In order to build a BE safe space, the first and foremost pre-requisite is to incorporate dissolution data into the model. Once the biopredictivity of the dissolution method has been established, dissolution profiles can be modeled using various approaches, as shown in Fig. 2. As such, there are different types of dissolution methods (biopredictive, biorelevant, bioindicative – please see the glossary, Table 9). For the purpose of PBBM, the dissolution methods should be biopredictive and bioindicative. Once such ability of dissolution method is confirmed, various dissolution models can be used to parameterize dissolution data.
The Z-factor model is a frequently used model to put dissolution data into the model for immediate release formulations. It was first defined by Takano et al. and assumes spherical particle shape and uniform particle size for dissolution and provides a dissolution factor with units of ml/mg/s [36]. The Z-factor is a function of diffusion coefficient, diffusion layer thickness, density and particle radius. It can be calculated under a certain dissolution condition. Theoretically, the z-factor is independent of the dissolution pH. However, there are circumstances where the z-factor may change with dissolution profiles determined at various pH value, e.g. for compounds with highly pH-dependent solubility or specific excipients, or the observation of coning in the dissolution or the compounds with wettability issue. Moreover, the surface pH could be different from the bulk pH in the dissolution media. Under such conditions, the dissolution kinetics may change. The dissolution profiles and z-factors can be incorporated as a function of pH into PBPK softwares such as Gastroplus. However, the z-factor assumes that the entire dose is available for dissolution immediately. Hence, the z-factor may not describe dissolution data accurately in cases of long disintegration time or incomplete dissolution where coning is observed. In such cases, appropriate correction factors are required to be incorporated [56]. Additionally, there are some challenges of using z-factor in terms of fitting. Ideally the z-factor is fitted for the entire dissolution profile, however, fitting initial points is also considered acceptable as they define the dissolution rate.
The Johnson model [32] and single or double Weibull models [17, 57] can be used to fit the dissolution data of modified release formulations. Weibull models can be used to generate virtual dissolution profiles for justifying dissolution specifications by adjusting mean dissolve time (MDT) and shape factor. For justifying dissolution specifications, sometimes it is required to generate virtual dissolution profiles, however there is no specific guidance on generating virtual dissolution profiles at lower and upper specifications. In such cases, using Weibull fitting by maintaining similar shape the virtual dissolution profiles can be generated. However, in cases where multiple specifications exist (e.g. for MR formulations where specifications are required to be established at least at 3 time points), it becomes a challenge to have virtual dissolution profile passing through all the specifications. This area is to be further explored.
Additionally, approaches such as product-particle size distribution (P-PSD) can also be used to incorporate dissolution data into the model [39]. Similar to z-factor model, the P-PSD model considers the in vitro dissolution conditions and drug substance solubility. The fitted P-PSD profile is able to reflect the dynamic change in the local conditions such as volume and media pH, while z-factor model assumes a consistent particle size in the dissolution. The mechanism of P-PSD model is treated the drug diffusion through a stagnant film layer [13]. The aspect of P-PSD was described by Pepin et al., wherein dissolution profiles are converted to a particle size distribution, which is then subsequently used as input into the model thereby establishing the specifications for particle size [13].
PBBM Virtual Trial Settings
Virtual Bioequivalence (VBE) trial simulations in PBPK softwares allow pharmaceutical scientists to assess the possibility of BE using virtual trials [17, 19]. Figure 2 shows schematically how modeled dissolution is integrated with a population simulator, such that VBE’s can be conducted for a chosen population. VBE trials account not only for differences between formulations, but account also for variability in the subject population. Intersubject variability can be included and default %CV values are sometimes used [17]. VBE uses a model to compare a hypothetical test versus reference formulations. VBE examples were included in case studies 1, 2 and 4.
Typically a new test formulation is compared to a marketed formulation (Generics, Innovators) or pivotal trial formulation (Innovators). Ideally the model should capture the observed variability of the reference population. It is recommended that the simulations would be conducted with an anticipated number of clinical trial subject. Virtual trial populations should mirror the clinical trial subjects [19].
While crossover trials are explored more with the simulation tools, other designs such as partial and full replicate are not fully incorporated into the available platforms. It is anticipated that the future versions of these platforms may include such complex study designs that can help to incorporate intra-subject coefficient of variation (ISCV) into the models [58, 59].
Acceptance Criteria
There are no formally accepted criteria to assess predictive performance of PBBM modeling [1, 29, 48, 60]. Notably, there have been comments from individual companies as well as the innovation and quality consortium (IQ consortium) on the proposed acceptance criteria, as the PBBM draft guidance recommended that the spirit of the IVIVC guidance criteria be followed, which specifies a 10% PE [1,48]. While a 10% PE is achievable and practical for carefully designed formulation crossover studies, for PBBM validation and model set-up, often independent clinical studies are required. Examples of such independent clinical studies are fasted clinical data from food effect studies or oral solution data, or clinical data from DDI study control arms lacking the modulator. Thus, when using independent studies a wider PE, e.g. “a maximal difference of 20% in the predicted Cmax and AUC as estimated by a PBPK model can be accepted (in line with IVIVC guidance)" was recommended [61].
For the interested reader, detailed comments for proposed acceptance criteria, as submitted by the IQ consortium, are shown in Supplement Table SI [60], but regulatory acceptance for a final version of the PBBM draft guidance [1] is uncertain. Based on the published studies on PBBM, commonly employed criteria include 1) to compare predicted and observed PK parameters e.g. AUC/Cmax and the difference are within 2-fold. The 2-fold criteria is well accepted in model use in drug-drug interaction [62]. 2) to calculate average fold error (AFE)/absolute average fold error (AAFE), AFE is compared to 1, and AAFE is set to below 2 [13]; 3) for risk assessment such as food effect assessment and pH-dependent DDI assessment, the ratio of predicted AUC and Cmax with or without food/acid-reducing agent is commonly used to compare with observed value, and deviation is within 25% [63]. The criteria will depend on the model use. In addition to the criteria, the agreement between predicted and observed PK profiles including the absorption phase and elimination phase is desirable as well. While such prediction errors and 2-folds limits hold good for exploratory analysis at discovery setting, they may not be acceptable for regulatory submissions. When such regulatory justifications are made with regard to dissolutions specification justification, biowaivers, f2 mismatch etc., a prediction error of 20% may be suitable. This limit is higher than the %PE as per IVIVC guidance. However, obtaining such low PE of 10% may be challenging for PBPK models as they involve physiological variability along with formulation variability. However, if the model is developed for internal decision making or for non-regulatory justification, the prediction error can be relaxed depending on the nature of molecule. For example, for highly variable molecules, BE limits may be extended for Cmax as per EMA guidance and AUC limits may be extended as per Canada guidance [43, 64]. In such cases the possibility of extending prediction errors acceptance criteria can be re-evaluated.
In addition to prediction errors, an obvious question comes with regard to the number of datasets for validation. The IVIVC guidance defines the number of validation datasets (internal/external) requirements however such guidance is currently not available for PBBM models [1, 9, 48]. In such cases, a typical external validation for PBBM models includes validation against literature datasets, other dosage forms or from other independent clinical studies. Internal validation includes model validation against the data used for building the model. In addition to single simulations, population simulations may provide additional validation. Since number of validation datasets is not defined, it is suggested to validate with maximum number of independent datasets.
Common Application: PBBM to Establish Dissolution Space, Superseding f2
One of most common applications of PBBM is to establish safe space for both innovator and generic drug development as indicated in Fig. 4. For innovator development, dissolution data of pivotal Ph-III lot or to be marketed formulation is used as reference to establish dissolution safe space. Whereas in generic development, both pivotal RLD and test formulations can be used to create dissolution safe space and to difference BE and non-BE lots. In both generic and innovator development, the developed safe space can be further utilized to identify & evaluate impact of CBA’s (CPP, CMA, CFV, CQA) in order to further refine or expand safe space. Traditionally the f2 similarity factor has been used as a metric for comparing dissolution profiles for various purposes such as biowaivers, site transfers, manufacturing process changes etc. However there are many limitations of utilizing f2 such as limited number of time points, percent relative standard deviation, and it doesn’t take the dissolution profiles shape into consideration as this is a model independent metric [65]. Other model dependent dissolution profiles comparison approaches such as single order, first order or Weibull based approaches are being utilized but the applications of such approaches are minimal. Regulatory agencies such as US FDA requires f2 calculations in quality control media only whereas other agencies such as EMA, China require dissolution profiles comparison in multimedia (pH 1.2, 4.5 and 6.8, with or without surfactant). In all of these cases, it is not thoroughly evaluated whether these media are biopredictive of in vivo performance. Additionally, failure of f2 in any of these comparisons may lead to regulatory rejections of biowaivers, site transfers or manufacturing process changes. In cases where in vivo absorption is controlled by permeability rather than dissolution (BCS Class III, IV), f2 may become overdiscriminating and non-biopredictive (Case study 4, ribociclib). Table 1 summarizes selected examples from generic companies and innovators of using PBBM to develop BE safe space, where some cases show that dissolution safe space supersedes f2 testing. With adequate model development, PBBM approaches can then be utilized to demonstrate that despite f2 failure (Fig. 3), bioequivalence can be achieved thus superseding the traditional f2 methodology. The case study 2 demonstrates this concept and such methodology can also makes many drug candidates eligible for BCS based biowaiver by extending the dissolution limits of BCS Class I (>85% in 30 min, rapid dissolution) and BCS Class III (>85% in 15 min very rapid dissolution) [31]. Moreover, if biopredictive media have been identified and proved during the initial product development using PBBM modeling, it can very well surpass the traditional f2 criteria and can avoid dissolution profiles similarity testing in multiple media’s thereby reducing the resources and saving time tremendously.
Additionally, there is a continued discussion on biopredictive media vs QC media in dissolution (Fig. 11). Traditionally QC media has been designed to understand formulation changes at manufacturing and plant level and is required to be discriminatory in nature towards formulation changes. But recently there has been a focus shift towards bio-predictivity of the QC media and its ability to reject non-BE batches. However as QC media is designed in such a way to only discriminate batches from formulation changes perspective it may not be bio-predictive. Hence it may be seen in future that while a QC method exists, a separate emphasis may be focused on having a separate media in the submission package as biopredictive media. Such trend can be seen for the future but if QC media can serve as both formulation discriminatory as well as bio-predictive to reject non-BE batches then that may be an ideal situation.
PBBM modeling provides mechanistic understanding of drug absorption and takes physiology impacts into consideration, which provides more flexibility and higher successful rate. A PBBM can be validated by observed data from multiple clinical studies and relevant in vitro data Recommended data to establish BE safe space is shown in Fig. 2 [41]. Once a solid model has been developed, a virtual BE study will be run in the software. Virtual BE allows to establish whether or not BE exists among batches by setting up a hypothetical population and including virtual or measured dissolution profiles.
There are published research papers and review papers of using PBBM to establish safe space, setting in vitro specification or to overcome f2 test criteria, selected case studies are shown in Table 1 [12]. The traditional f2 factor has limitations and is not always applicable for dissolution profile comparisons especially for cases where the % relative standard deviation (RSD) is high. In such cases alternative approaches such as f2 bootstrap, multivariate statistical analysis (model independent) and zero order or Weibull approaches (model dependent) can be used [66].
Outlook
PBPK modeling has found broad regulatory acceptance, particularly in designing drug-drug interaction studies, including DDI clinical study waivers [62]. PBPK for oral biopharmaceutics applications has gotten broader acceptance recently, including uses of PBBM for drug product quality [57]. The authors see PBBM as one of the key tools to establish a BE space, supplementing f2 criteria on a case by case basis. For formulation selection and bridging studies with innovator drugs, PBBM can be used select a formulation for Phase II or III trials, which is anticipated to show BE. In particular, for generic companies, PBBM has found broad applications, as study waivers or successfully chosen formulations are part of key drug development activities [10]. However, global regulatory acceptances are still not prevalent and PBBM is likely more used for company internal decision making, as opposed to seeking study waivers, or seeking specification settings.
Conclusion
With advances in science and technology, efforts are being made in the direction of in silico prediction of bioequivalence and clinical trials as part of model-integrated drug development (MIDD) [67, 68]. PBBM as part of MIDD is getting impetus and regulatory agencies encourage both innovator and generic companies to utilize such models in regulatory submissions. PBBM approaches have the possibility to reduce human trials, as healthy volunteers typically do not get a significant benefit from the studied drug in BE trials. Using PBBM modeling brings multiple perspectives into the clinical development by establishing clinically relevant dissolution specifications, bioequivalence safe space, establishing mechanistic IVIVC and identifying CBA’s. The case studies described in this article demonstrate potential of such tools in drug discovery and development for both innovator and generic companies. Along with case studies, regulatory perspectives, detailed workflow for model building, validation and application, approaches to input dissolution profiles and particle size were discussed. Critical discussion highlights a few challenging topics such as PBBM modeling for highly variable drugs, prediction errors, QC vs biopredictive dissolution media, virtual BE simulations, PBBM to supersede f2. This manuscript presents state of the art of PBBM modeling from both innovator and generic perspective and the authors feel that within next few years PBBM will be at the heart of innovator and generic drug product applications, driving product development.
Abbreviations
- ACAT:
-
Advanced compartmental absorption and transit
- AFE:
-
Average Fold Error
- API:
-
Active pharmaceutical ingredient
- AUC:
-
Area under the curve
- BA:
-
Bioavailability
- BCS:
-
Biopharmaceutics classification system
- BE:
-
Bioequivalence
- CBA:
-
Critical bioavailability attributes
- CFV:
-
Critical formulation variables
- CI:
-
Confidence interval
- CMA:
-
Critical material attributes
- Cmax :
-
Maximum concentration
- CPP:
-
Critical process parameters
- CQA:
-
Critical quality attributes
- CRDS:
-
Clinically relevant dissolution specifications
- EMA:
-
European Medicines Agency
- ISCV:
-
Intra subject variability
- IV:
-
Intravenous
- IVIVC:
-
In vitro in vivo correlation
- MR:
-
Modified release
- NCE:
-
New chemical entity
- PBBM:
-
Physiologically based biopharmaceutics modeling
- PBPK:
-
Physiologically based pharmacokinetic modeling
- PK:
-
Pharmacokinetics
- PO:
-
Per oral
- p-PSD:
-
product - particle size distribution
- PSA:
-
Parameter sensitivity analysis
- QbD:
-
Quality by design
- QC:
-
Quality control
- RLD:
-
Reference listed drug
- USFDA:
-
United States Food and Drug Administration
- VBE:
-
Virtual bioequivalence
References
FDA, US. The use of physiologically based pharmacokinetic analyses — biopharmaceutics applications for Oral drug product development, Manufacturing Changes, and Controls https://www.fda.gov/media/142500/download. 2020. Accessed July 1st, 2022.
FDA, US. Bioavailability and Bioequivalence Studies Submitted in NDAs or INDs — General Considerations https://www.fda.gov/media/88254/download. 2014. Accessed 1 July 2022.
FDA, US. Bioequivalence Studies With Pharmacokinetic Endpoints for Drugs Submitted Under an ANDA Guidance for Industry https://www.fda.gov/media/87219/download. 2021. Accessed 1 July 2022.
FDA, US. Dissolution testing of immediate release solid Oral dosage forms. 1997.
Moore J. Mathematical comparison of dissolution profiles. Pharm Technol. 1996;20:64–75.
Zhang Y, et al. DDSolver: an add-in program for modeling and comparison of drug dissolution profiles. AAPS J. 2010;12(3):263–71.
Laisney M, et al. Physiologically based biopharmaceutics modeling to demonstrate virtual bioequivalence and bioequivalence safe-space for Ribociclib which has permeation rate-controlled absorption. J Pharm Sci. 2022;111(1):274–84.
Tsong Y. Dissolution profile similarity factor, f2. 2020.
FDA, US. The use of physiologically based pharmacokinetic analyses — biopharmaceutics applications for Oral drug product development, Manufacturing Changes, and Controls 2020.
Yuvaneshwari K, et al. Applications of PBPK/PBBM modeling in generic product development: an industry perspective. J Drug Deliv Sci Technol. 2022;69:103152.
Wu F, et al. Biopharmaceutics applications of physiologically based pharmacokinetic absorption modeling and simulation in regulatory submissions to the U.S. Food and Drug Administration for new drugs. AAPS J. 2021;23(2):31.
Heimbach T, et al. Establishing the bioequivalence safe space for immediate-release Oral dosage forms using physiologically based biopharmaceutics modeling (PBBM): case studies. J Pharm Sci. 2021;110(12):3896–906.
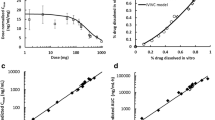
Pepin XJH, et al. Justification of drug product dissolution rate and drug substance particle size specifications based on absorption PBPK modeling for Lesinurad immediate release tablets. Mol Pharm. 2016;13(9):3256–69.
Zhang F, et al. In silico prediction of bioequivalence of isosorbide mononitrate tablets with different dissolution profiles using PBPK modeling and simulation. Eur J Pharm Sci. 2021;157:105618.
Paraiso RL, et al. The use of PBPK/PD to establish clinically relevant dissolution specifications for zolpidem immediate release tablets. Eur J Pharm Sci. 2020;155:105534.
Hsieh N-H, et al. A Bayesian population physiologically based pharmacokinetic absorption modeling approach to support generic drug development: application to bupropion hydrochloride oral dosage forms. J Pharmacokinet Pharmacodyn. 2021;48(6):893–908.
Macwan JS, et al. Application of physiologically based biopharmaceutics modeling to understand the impact of dissolution differences on in vivo performance of immediate release products: the case of bisoprolol. CPT Pharmacometrics Syst Pharmacol. 2021;10(6):622–32.
Bolger MB, et al. The irrelevance of in vitro dissolution in setting product specifications for drugs like dextromethorphan that are subject to lysosomal trapping. J Pharm Sci. 2019;108(1):268–78.
Loisios-Konstantinidis I, et al. Establishing virtual bioequivalence and clinically relevant specifications using in vitro biorelevant dissolution testing and physiologically-based population pharmacokinetic modeling. Case example: naproxen. Eur J Pharm Sci. 2020;143:105170.
Li X, et al. Justification of biowaiver and dissolution rate specifications for Piroxicam immediate release products based on physiologically based pharmacokinetic modeling: an in-depth analysis. Mol Pharm. 2019;16(9):3780–90.
Jaiswal S, et al. Development, validation and application of physiologically based biopharmaceutics model to justify the change in dissolution specifications for DRL ABC extended release tablets. Drug Dev Ind Pharm. 2021;47(5):778–89.
Miao L, et al. Using a physiologically based pharmacokinetic absorption model to establish dissolution bioequivalence safe space for oseltamivir in adult and pediatric populations. AAPS J. 2020;22(5):1–10.
FDA, US. Product quality review(s)-ORILISSA (elagolix) tablets. 2017.
Babiskin AH, Zhang X. Application of physiologically based absorption modeling for amphetamine salts drug products in generic drug evaluation. J Pharm Sci. 2015;104(9):3170–82.
Kato T, et al. Establishment of a clinically relevant specification for dissolution testing using physiologically based pharmacokinetic (PBPK) modeling approaches. Eur J Pharm Biopharm. 2020;151:45–52.
Jereb R, et al. In vitro–in vivo relationship and bioequivalence prediction for modified-release capsules based on a PBPK absorption model. AAPS PharmSciTech. 2020;21(1):1–11.
Holmstock N. Development of canagliflozin: mechanistic absorption modeling during late-stage formulation and process optimization. 2017.
Raines K. PBPK Biopharmaceutics guidance and progress on risk assessment, in regulatory utility of mechanistic modeling to support alternative bioequivalence approaches workshop 2021.
Wu F, et al. Biopharmaceutics applications of physiologically based pharmacokinetic absorption modeling and simulation in regulatory submissions to the US food and drug administration for new drugs. AAPS J. 2021;23(2):1–14.
Anand O. Clinically relevant dissolution specifications: a biopharmaceutics’ risk based approach: an FDA perspective. 2021.
FDA, US. M9 Biopharmaceutics Classification SystemBased Biowaivers 2021.
Noyes AA, Whitney WR. The rate of solution of solid substances in their own solutions. J Am Chem Soc. 1897;19(12):930–4.
Dokoumetzidis A, Papadopoulou V, Macheras P. Analysis of dissolution data using modified versions of Noyes–Whitney equation and the weibull function. Pharm Res. 2006;23(2):256–61.
Papadopoulou V, et al. On the use of the Weibull function for the discernment of drug release mechanisms. Int J Pharm. 2006;309(1):44–50.
Langenbucher F. Letters to the editor: linearization of dissolution rate curves by the Weibull distribution. J Pharm Pharmacol. 1972;24(12):979–81.
Takano R, et al. Rate-limiting steps of oral absorption for poorly water-soluble drugs in dogs; prediction from a miniscale dissolution test and a physiologically-based computer simulation. Pharm Res. 2008;25(10):2334–44.
Wang J, Flanagan DR. General solution for diffusion-controlled dissolution of spherical particles. 1. Theory. J Pharm Sci. 1999;88(7):731–8.
Chirumamilla SK, et al. Mechanistic PBPK modelling to predict the advantage of the salt form of a drug when dosed with acid reducing agents. Pharmaceutics. 2021;13(8):1169.
Okazaki A, Mano T, Sugano K. Theoretical dissolution model of poly-disperse drug particles in biorelevant media. J Pharm Sci. 2008;97(5):1843–52.
Pepin XJH. In silico PBPK modelling in support of drug product dissolution and drug substance particle size specifications. M-CERSI, 2017.
Suarez-Sharp S. Utilization of PBBM/PBPK models for building a safe space and regulatory applications in support of drug product quality. 2020.
Pepin XJH, et al. Bridging in vitro dissolution and in vivo exposure for acalabrutinib. Part I. mechanistic modelling of drug product dissolution to derive a P-PSD for PBPK model input. Eur J Pharm Biopharm. 2019;142:421–34.
EMA. Guideline on the Investigation of Bioequivalence 2010.
Agrawal NGB, et al. Single- and multiple-dose pharmacokinetics of etoricoxib, a selective inhibitor of cyclooxygenase-2, in man. J Clin Pharmacol. 2003;43(3):268–76.
Mitra A, Kesisoglou F, Dogterom P. Application of absorption modeling to predict bioequivalence outcome of two batches of etoricoxib tablets. AAPS PharmSciTech. 2015;16(1):76–84.
Zhang X, et al. Integrating in vitro, modeling, and in vivo approaches to investigate warfarin bioequivalence. CPT: Pharmacometrics Syst Pharmacol. 2017;6(8):523–31.
Samant TS, et al. Ribociclib bioavailability is not affected by gastric pH changes or food intake: In silico and clinical evaluations. Clin Pharmacol Ther. 2018;104(2):374–83.
FDA, US. Extended release oral dosage forms: development, evaluation, and application of in vitro/in vivo correlations. 1997.
Butler J, et al. In vitro models for the prediction of in vivo performance of oral dosage forms: recent progress from partnership through the IMI OrBiTo collaboration. Eur J Pharm Biopharm. 2019;136:70–83.
Tsume Y, et al. In vitro dissolution methodology, mini-gastrointestinal simulator (mGIS), predicts better in vivo dissolution of a weak base drug, dasatinib. Eur J Pharm Sci. 2015;76:203–12.
Mann J, et al. Validation of dissolution testing with biorelevant media: an OrBiTo study. Mol Pharm. 2017;14(12):4192–201.
Li M, et al. Understanding in vivo dissolution of immediate release (IR) solid Oral drug products containing weak acid BCS class 2 (BCS class 2a) drugs. AAPS J. 2021;23(6):1–13.
FDA, US. Retention of Bioavailability and Bioequivalence Testing Samples. https://www.fda.gov/science-research/clinical-trials-and-human-subject-protection/retention-bioavailability-and-bioequivalence-testing-samples. 1993. Accessed 1 July 2022.
Zhang X, et al. Mechanistic oral absorption modeling and simulation for formulation development and bioequivalence evaluation: report of an FDA public workshop. CPT: Pharmacometrics Syst Pharmacol. 2017;6(8):492–5.
Sharp SS. Part 2 Utilization of PBBM PBPK Models for Building a Safe Space with Dr Sandra Suarez Sharp. Simulations Plus, Inc. 2020. https://www.youtube.com/watch?v=kJCC_3I-7eY. Accessed 5 July 2022.
Hofsäss MA, Dressman J. Suitability of the z-factor for dissolution simulation of solid Oral dosage forms: potential pitfalls and refinements. J Pharm Sci. 2020;109(9):2735–45.
Aburub A, et al. An IQ consortium perspective on connecting dissolution methods to in vivo performance: analysis of an industrial database and case studies to propose a workflow. AAPS J. 2022;24(3):49.
Bego M, et al. Proof of concept in assignment of within-subject variability during virtual bioequivalence studies: propagation of intra-subject variation in gastrointestinal physiology using physiologically based pharmacokinetic modeling. AAPS J. 2022;24(1):21.
Mitra A. PBBM workshop 2019: "virtual bioequivalence: model development, verification and applications". 2019.
(IQ) IC. Comment from IQ for the use of physiologically based pharmacokinetic analyses — biopharmaceutics applications for Oral drug product development, Manufacturing Changes, and Controls https://www.regulations.gov/docket/FDA-2020-D-1517/comments 2020. Accessed 1 July 2022.
Inc MC. Comment from IQ for the use of physiologically based pharmacokinetic analyses — biopharmaceutics applications for Oral drug product development, Manufacturing Changes, and Controls https://www.regulations.gov/comment/FDA-2020-D-1517-0005 2020. Accessed 1 July 2022.
Shebley M, et al. Physiologically based pharmacokinetic model qualification and reporting procedures for regulatory submissions: a consortium perspective. Clin Pharmacol Ther. 2018;104(1):88–110.
Dong Z, et al. Application of physiologically-based pharmacokinetic modeling to predict gastric pH-dependent drug–drug interactions for weak base drugs. CPT Pharmacometrics Syst Pharmacol. 2020;9(8):456–65.
Canada H. Conduct and Analysis of Comparative Bioavailability Studies. https://www.canada.ca/content/dam/hc-sc/documents/services/drugs-health-products/drug-products/applications-submissions/guidance-documents/bioavailability-bioequivalence/conduct-analysis-comparative.pdf. 2012. Accessed 1 July 2022.
Abend AM, et al. Current approaches for dissolution similarity assessment, requirements, and global expectations. AAPS J. 2022;24(3):50.
Kollipara S, et al. Simplified model-dependent and model-independent approaches for dissolution profile comparison for Oral products: regulatory perspective for generic product development. AAPS PharmSciTech. 2022;23(1):53.
Impact Story. Modeling Tools Could Modernize Generic Drug Development, https://www.fda.gov/drugs/regulatory-science-action/impact-story-modeling-tools-could-modernize-generic-drug-development. 2022 [cited 2022 April 8th, 2022]. Accessed 1 July 2022.
Kuemmel C, et al. Consideration of a credibility assessment framework in model-informed drug development: potential application to physiologically-based pharmacokinetic modeling and simulation. CPT Pharmacometrics Syst Pharmacol. 2020;9(1):21–8.
Funding
This review article was sponsored by Merck Sharp & Dohme Corp., a subsidiary of Merck & Co., Inc., Kenilworth, NJ, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
DW and TH are employees of Merck Sharp & Dohme Corp., a subsidiary of Merck & Co., Inc., Kenilworth, NJ, USA and may hold equity stock in Merck & Co., Inc., Kenilworth, NJ, USA. MS and AS are employees of Zydus Lifesciences Limited (formerly known as Cadila Healthcare Limited), Ahmedabad, India. SK and TA are employees of Dr. Reddy’s Laboratories Ltd. and report no conflicts interest. The authors alone are responsible for the content and writing of this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 563 kb)
Rights and permissions
About this article
Cite this article
Wu, D., Sanghavi, M., Kollipara, S. et al. Physiologically Based Pharmacokinetics Modeling in Biopharmaceutics: Case Studies for Establishing the Bioequivalence Safe Space for Innovator and Generic Drugs. Pharm Res 40, 337–357 (2023). https://doi.org/10.1007/s11095-022-03319-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-022-03319-6