Abstract
Purpose
Somatic mutations of the isocitrate dehydrogenase 1 (IDH1) gene, mostly substituting Arg132 with histidine, are associated with better patient survival, but glioma recurrence and progression are nearly inevitable, resulting in disproportionate morbidity and mortality. Our previous studies demonstrated that in contrast to hemizygous IDH1R132H (loss of wild-type allele), heterozygous IDH1R132H is intrinsically glioma suppressive but its suppression of three-dimensional (3D) growth is negated by extracellular glutamate and reducing equivalent. This study sought to understand the importance of 3D culture in IDH1R132H biology and the underlying mechanism of the glutamate effect.
Methods
RNA sequencing data of IDH1R132H-heterozygous and IDH1R132H-hemizygous glioma cells cultured under two-dimensional (2D) and 3D conditions were subjected to unsupervised hierarchal clustering and gene set enrichment analysis. IDH1R132H-heterozygous and IDH1R132H-hemizygous tumor growth were compared in subcutaneous and intracranial transplantations. Short-hairpin RNA against glutamate dehydrogenase 2 gene (GLUD2) expression was employed to determine the effects of glutamate and the mutant IDH1 inhibitor AGI-5198 on redox potential in IDH1R132H-heterozygous cells.
Results
In contrast to IDH1R132H-heterozygous cells, 3D-cultured but not 2D-cultured IDH1R132H-hemizygous cells were clustered with more malignant gliomas, possessed the glioblastoma mesenchymal signature, and exhibited aggressive tumor growth. Although both extracellular glutamate and AGI-5198 stimulated redox potential for 3D growth of IDH1R132H-heterozygous cells, GLUD2 expression was required for glutamate, but not AGI-5198, stimulation.
Conclusion
3D culture is more relevant to IDH1R132H glioma biology. The importance of redox homeostasis in IDH1R132H glioma suggests that metabolic pathway(s) can be explored for therapeutic targeting, whereas IDH1R132H inhibitors may have counterproductive consequences in patient treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Malignant gliomas represent 81% of primary brain malignancy and cause significant mortality and morbidity [1]. Glioblastoma—World Health Organization (WHO) grade IV, the most common and advanced form of glioma—has a 5-year survival of only 5.5%, and the inevitable recurrence and progression of WHO grade II and III (lower-grade) gliomas also contribute to the disproportionately high mortality and morbidity [2]. As such, there is an unmet need to improve the current treatment strategy.
Somatic IDH1 missense mutations occur in > 70% of the lower-grade gliomas and secondary glioblastomas, substituting arginine 132 with histidine at a frequency of 92% among gliomas with mutations [3,4,5,6]. IDH1 is a cytosolic enzyme that produces 2-oxoglutarate and NADPH, which are further converted by the IDH1R132H neomorphic activity to D-2-hydroxyglutarate (D-2HG) [7]. High levels of D-2HG induce hypermethylation of lysine residues in histones and CpG islands in DNA through inhibition of histone demethylases and 5-methylcytosine hydroxylases [8], blocking cell differentiation and establishing the glioma-CpG island methylator phenotype (G-CIMP), respectively [9, 10].
Although IDH1R132H is believed to be oncogenic, this theory is predicated primarily on the findings of exogenous IDH1R132H expression, which have yet to be corroborated by endogenous, heterozygous IDH1R132H [11]. The use of exogenous IDH1R132H in reference to wild-type IDH1 has been virtually the norm because heterozygous IDH1R132H is scarcely preserved in experimental systems [12,13,14,15]. By exploring IDH1R132H-hemizygous BT142 mut/ − glioma cells [14, 16], we demonstrated that restoration of IDH1R132H heterozygosity (by a wild-type IDH1 transgene) restores D-2HG production and suppresses anchorage-independent three-dimensional (3D) spheroid growth [17]. Conversely, selection against IDH1R132H heterozygosity or exogenous IDH1R132H transgene occurs during 3D growth in vitro and in vivo but not during anchorage-dependent two-dimensional (2D) adherent growth [17, 18]. The antagonism between IDH1R132H heterozygosity and 3D growth indicates that IDH1R132H is tumor suppressive, as supported by the lack of gliomagenesis in heterozygous Idh1R132H mice [19,20,21]. Furthermore, IDH1R132H not only reduced glioma incidence and extended survival in Trp53-deficient background [21,22,23,24] but also obliterated gliomagenesis in Trp53-proficient background [18].
It should be noted, however, that IDH1R132H tumor-suppressive activity is undermined by events including loss of IDH1R132H heterozygosity, inactivation of tumor-suppressor genes, and abundant extracellular metabolites [11]. In particular, IDH1R132H suppression of 3D growth can be negated by glutamate and the reducing agent N-acetyl cysteine (NAC) [17, 18]. IDH1R132H gliomas depend on upregulation of hominoid-specific GLUD2 (glutamate dehydrogenase 2) to alleviate metabolic stress [23, 25,26,27], but the underlying mechanism remains unclear. We sought to determine the importance of 3D culture and the mechanism of glutamate effect in IDH1R132H biology.
Methods
Cell culture and spheroid quantification
The anaplastic oligoastrocytoma BT142 mut/ − cells, which showed no TP53 mutation but inconclusive chromosome 1p/19q codeletion [16], were purchased from ATCC (Manassas, VA). IDH1R132H heterozygosity was restored by reintroduction of a transgene expressing YFP–IDH1 to yield IDH1R132H-heterozygous BT142 mut/IDH1 in reference to IDH1R132H-hemizygous BT142 mut/YFP* [17]. A firefly luciferase gene in the lentiviral vector pLenti6.3/TO/luc [28] was transduced into the cells. Conditions for 2D and 3D cultures were described previously [17]. For quantification of neurosphere growth, spheroids were dissociated with StemPro Accutase (ThermoFischer, Waltham, MA) and quantified in triplicate using Bright-Glo Luciferase Assay (Promega, Madison, WI) or CellTiter-Glo Luminescent Cell Viability Assay (Promega) according to the manufacturers’ recommendations.
Chemical treatment
Adherent cells were treated with 3 µM AGI-5198 (Sigma-Aldrich, St. Louis, MO) for 3 or 5 days in reference to vehicle control. Culture medium was replaced every other day to maintain the concentrations for 5 days before the cells were seeded at a density of 5 × 104 per well in a 48-well plate for spheroid growth, with continued dosing every three days. Generally, spheroid growth was photographed 7 days after seeding and terminated for further analyses. For animal studies, 150 mg/kg AGI-5198 [29] was administered orally daily. Spheroid cultures involving the addition of sodium glutamate, NAC, and oxaloacetate (Sigma-Aldrich) were performed as described previously [17, 18] with additional dosing every other day.
Gene expression
Total RNA was extracted and converted to cDNA as described [17]. Quantitative PCR was performed in quadruplicate with LightScanner Master Mix (BioFire Diagnostics, Salt Lake City, UT) using CFX96 Touch Real-Time PCR Detection System (Bio-Rad Laboratories, Hercules, CA). The primer sets are listed in Supplementary Table 1. Annealing temperature was set at 63 ºC for 45 cycles. Quantitation cycle (Cq) values were obtained through CFX Manager Software (Bio-Rad) and normalized by the Cq values of reference genes RPL30, YWHAZ, and UBC. Western blotting was performed as described [17, 28] with1:2000 anti-GLUD (Invitrogen, Carlsbad, CA).
RNA sequencing and analysis
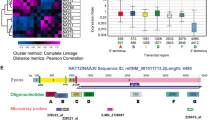
Total RNA was extracted as above from IDH1R132H-hemizygous and IDH1R132H-heterozygous BT142 cells in three biological replicates. The RNA library was prepared using Illumina Truseq Stranded mRNA Library Preparation Kit (San Diego, CA) with polyA selection and sequenced with HiSeq 50 Cycle Single-Read Sequencing (Illumina). The reads were aligned to the reference human genome hg38; H_sapiens_Dec_2013, GRCh38. USeq [30] was used to generate log2-fold change ratios for each gene. Gene set enrichment analysis (GSEA) was performed using the java GSEA desktop application (Broad Institute, Cambridge, MA) [31] for comparison between IDH1R132H-hemizygous and IDH1R132H-heterozygous cells. The MSigDB gene sets (Broad Institute) hallmark, KEGG, and Verhaak_Glioblastoma [32] were used. Top-10 gene sets were obtained based on highest normalized enrichment score (NES) in concomitance with most significant false discovery rate (FDR) and nominal p-value. Both FDRs and nominal p-values were presented in ± log10 conforming to the direction of NES. Unsupervised hierarchical clustering was performed using the hciR package in RStudio. Heatmaps of top-40 genes and specified gene sets were generated according to adjusted p-values of the top genes. The TCGA (The Cancer Genome Atlas) Lower-Grade Glioma (TCGA-LGG) data set was downloaded using TCGAbiolinks and joined with the above RNA-sequencing data. After correction of batch effect, a heatmap of sample distances was generated to reveal the distribution of IDH-wildtype and IDH-mutant gliomas as well as 2D- and 3D-cultured IDH1R132H-heterozygous and IDH1R132H-hemizygous cells.
Xenograft mouse models
Tumor transplantation experiments and procedures were approved by the University of Utah Institutional Animal Care and Use Committee and were performed as described [18, 28]. Non-obese diabetic/severe-combined immunodeficient IL-2Rg-null mice of both sexes were used at the age of 6–10 weeks. Transplantations required 2 × 106 BT142 cells for subcutaneous injections and 2 × 104 cells for intracranial injections. Bioluminescent imaging with inhalant isoflurane was performed essentially as described [18, 28]. Bioluminescent intensity as a surrogate of tumor volume was quantitatively analyzed using LivingImage software (Xenogen, Alameda, CA).
Tumors were harvested 6–8 weeks after injection for formalin fixation and paraffin embedding. Histological assessment and immunohistochemistry analysis were as previously described [18]. Primary antibodies were diluted as follows: 1:10,000 anti-5hmC (Invitrogen), 1:25 anti-Nestin (Invitrogen), and 1:500 anti-GLUD (Invitrogen). To confirm IDH1R132H heterozygosity in the resultant tumor, genomic DNA was extracted from paraffin-embedded intracranial tumors using DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany). PCR amplification and DNA sequencing were performed to distinguish YFP* from YFP‑IDH1 transgene [17].
GLUD2 knockdown
A GLUD2-specific small-hairpin (sh) RNA targeting the open reading frame (5′-CATGTCAGCTATGGCCGTT-3′) was expressed in the SMARTvector Inducible Lentiviral System (Dharmacon, Lafayette, CO). Transduced BT142 mut/IDH1 cells were enriched by fluorescence-activated cell sorting and selected with 1 µg/ml puromycin for 7 days. GLUD2 shRNA expression was induced by doxycycline at 1 µg/ml for in-vitro studies and 2 mg/ml in the drinking water for in vivo studies. GLUD2 knockdown was confirmed at the RNA and protein levels for ≥ 48 h after the induction or with ethanol control.
Redox assays
Intracellular reduced (GSH) and oxidized (GSSG) glutathione and NADP+ and NADPH concentrations were determined using the GSH/GSSG-Glo and NADP/NADPH-Glo assays (Promega), respectively, according to the manufacturer’s recommendations. The GSH/GSSG-Glo Assay was performed with 1 × 104 cells per condition. For NADP/NADPH-Glo assay, 5 × 103 cells per condition were used for acid and base treatments. Both assays were performed in triplicate.
Results
3D culture distinguishes IDH1R132H-hemizygous from IDH1R132H-heterozygous cells in gene expression profile
To understand the impact of 2D and 3D cultures on gene expression of IDH1R132H glioma cells, we performed RNA sequencing of IDH1R132H-hemizygous and IDH1R132H-heterozygous BT142 cells [17]. Unsupervised hierarchical clustering revealed that 3D-cultured IDH1R132H-heterozygous cells belonged to a specific cluster, whereas 2D-cultured IDH1R132H-heterozygous cells clustered with 2D- and 3D-cultured IDH1R132H-hemizygous cells (Fig. 1a), a finding in agreement with the differential growth of IDH1R132H-hemizygous and IDH1R132H-heterozygous cells in 3D, but not 2D, culture [17]. GSEA confirmed profound differences in the top-10 gene sets between 3D and 2D cultures. Notably, Oxidative_Phosphorylation was most significantly enriched in 3D-cultured IDH1R132H-hemizygous cells with both KEGG and Hallmark gene sets (Fig. 1b,c; Supplementary Fig. 1a), in agreement with the reports that malignant gliomas rely on oxidative phosphorylation for aggressive growth [33] and hemizygous IDH1R132H promotes 3D growth [17]. By contrast, Oxidative_Phosphorylation in 2D-cultured IDH1R132H-hemizygous cells was neither significant in the KEGG gene sets nor among the top 10 of Hallmark gene sets (Fig. 1b; Supplementary Fig. 1b, c). As expected, the significantly enriched genes in Oxidative_Phosphorylation were upregulated in IDH1R132H-hemizygous cells but downregulated in IDH1R132H-heterozygous cells (Fig. 1d). Furthermore, 3D and 2D cultures gave rise to divergent directions of enrichment between IDH1R132H-hemizygous and IDH1R132H-heterozygous cells among gene sets shared in the top-10 lists, including KEGG Lysosome and Hallmark E2F_Targets and G2M_Checkpoint (Fig. 1b; Supplementary Fig. 1).
A distinct gene expression profile from 3D-cultured IDH1R132H-heterozygous cells. a Unsupervised hierarchical clustering of RNA-sequencing data from 2D- and 3D-cultured IDH1R132H-heterozygous (het) and IDH1R132H-hemizygous (hem) BT142 cells revealing a unique cluster of 3D-cultured IDH1R132H-heterozygous cells distinct from the rest. Each column represents a single sample. Top-20 upregulated (red) and downregulated (blue) genes are indicated. b Differential enrichment of KEGG gene sets between 3D and 2D cultures. The top-10 gene sets based on the most significant false-discovery rates (FDR) and nominal p-values were plotted with the normalized enrichment scores (NES) and the log10-transformed nominal p-values and FDR. Orange dashed lines indicate the cutoff for FDR and nominal p-values at 0.05, and asterisks indicate the gene sets shared between 3D and 2D cultures (see Supplementary Fig. 1). c, d Oxidative phosphorylation gene set enrichment plot (c) and hierarchical clustering (d) of 3D-cultured IDH1R132H glioma cells
Previous transcriptomic studies indicated that 3D culture of various cancer cell types is closer to in-vivo tumor growth than 2D culture [34, 35]. To test the relevance of 3D culture to glioma biology, we incorporated the clustering analysis the TCGA-LGG data set consisting of 168 cases of IDH-mutant with 1p/19q codeletion, 246 cases of IDH-mutant without codeletion, and 93 cases of IDH-wildtype (Fig. 2). Hierarchical clustering gave rise to two major clusters: Cluster 1 composed essentially of IDH-mutant gliomas with or without codeletion and Cluster 2 containing both IDH-wildtype and IDH-mutant gliomas mostly without codeletion (Supplementary Table 2). In contrast to the congregation of 2D-cultured IDH1R132H cells, 3D-culture IDH1R132H-hemizygous cells belonged to the more malignant Cluster 2 whereas IDH1R132H-heterozygous cells were in the less malignant Cluster 1. Taken together, these results support the notion that 3D culture improves cell-line model systems for cancer research [34], particularly in the investigation of IDH1R132H biology.
3D culture distinguishes IDH1R132H-heterozygous cells from IDH1R132H-hemizygous cells. Unsupervised hierarchical clustering was performed by incorporating the above RNA-sequencing result with the TCGA-LGG data set consisting of IDH-mutant and 1p/19q codeleted, IDH-mutant (without codeletion), and IDH-wildtype gliomas. Enlarged views underscore that 3D culture distinguishes IDH1R132H-heterozygous (het) cells and IDH1R132H-hemizygous (hem) in the clustering
Tumorigenicity of IDH1R132H-heterozygous glioma cells is context dependent
IDH-mutant gliomas exhibit G-CIMP and belong to the proneural subtype [36], whereas recurrent gliomas are associated with decreased DNA methylation, mesenchymal transformation, and enrichment of the tumor-initiating marker gene CD44 [37, 38]. Interestingly, we observed significant enrichment of the Verhaak_GBM_Mesenchymal gene set in 3D-, but not 2D-, cultured IDH1R132H-hemizygous cells (Fig. 3a). The vast majority of genes in Verhaak_GBM_Mesenchymal were upregulated in the IDH1R132H-hemizygous cells, including CTSC (cathepsin C), ALDH3B1 (aldehyde dehydrogenase 3 family member B1), and CHI3L1 (chitinase-3-like 1, aka YKL-40) (Fig. 3b). Furthermore, quantitative PCR analysis of CD44 mRNA levels revealed a marked increase in IDH1R132H-hemizygous BT142 [39]. Together, these results suggest that loss of IDH1R132H heterozygosity is associated with mesenchymal transition.
Loss of IDH1R132H heterozygosity drives glioma progression. a Enrichment of glioblastoma mesenchymal gene set in 3D-, but not 2D-, cultured IDH1R132H-hemizygous BT142 cells. b Hierarchical clustering of 3D-cultured cells using the Verhaak_GBM_Mesenchymal gene set. c Histological examination of intracranial tumor growth derived from IDH1R132H-hemizygous and IDH1R132H-heterozygous BT142. Micrographs of hematoxylin‑eosin (H‑E) and immunohistochemical staining are presented with 20 × objectives and specified antibodies. Scale bar: 200 μm
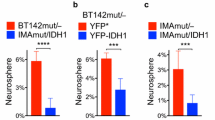
Next, we tested whether IDH1R132H-hemizygous BT142 cells were more tumorigenic. Subcutaneous xenograft studies showed robust tumor growth of IDH1R132H-hemizygous cells; however, IDH1R132H heterozygosity obliterated tumorigenesis (Supplementary Fig. 2a–c), which supports our previous findings that heterozygous IDH1R132H suppresses anchorage-independent growth and is intrinsically tumor suppressive [17, 18]. By contrast, in an orthotopic model, IDH1R132H-heterozygous cells exhibited tumor growth, albeit much smaller than that of IDH1R132H-hemizygous cells (Supplementary Fig. 2d, e). Histological examination confirmed pronounced reduction of cellularity and Ki67 staining in IDH1R132H-heterozygous tumors even though IDH1R132H staining remained strong but heterogeneous (Fig. 3c). However, selection against the IDH1 transgene was indicated by the YFP marker gene, which was much less expressed in IDH1R132H-heterozygous cells than IDH1R132H-hemizygous cells [17]. These results not only support the notion that the glutamate-rich cerebral environment is conducive to the growth of IDH1R132H glioma [18] but also indicate loss of IDH1R132H heterozygosity as a mechanism of glioma progression [40, 41].
Glutamate boosts redox homeostasis for 3D growth of IDH1R132H-heterozygous cells
IDH-mutant gliomas depend on glutamate to alleviate metabolic stress through GLUD2 upregulation [23, 25,26,27]. We asked first whether GLUD2 is required for glutamate stimulation of 3D growth by transducing IDH1R132H-heterozygous cells with a doxycycline-inducible GLUD2 shRNA. The addition of doxycycline resulted in marked reduction of GLUD2 expression at mRNA and protein levels (Fig. 4a,b). As expected, GLUD2-depleted cells were no longer responsive to glutamate but remained highly responsive to NAC, with a sevenfold increase in 3D growth (Fig. 4c; Supplementary Fig. 3a). In light of the critical role of increased redox potential in 3D growth [17, 42], this finding suggested a role for glutamate in redox homeostasis. Indeed, glutamate treatment of IDH1R132H-heterozygous cells increased the reduced to oxidized glutathione ratio (GSH/GSSG) by twofold, resulting from a 12% increase of GSH concomitant with a 47% decrease in GSSG, a finding similar to the treatment with NAC and oxaloacetate (Supplementary Fig. 3b), which are known to increase redox potential [17, 43]. Furthermore, glutamate treatment also significantly increased NADPH/NADP+ ratio by mainly decreasing NADP+ concentration (Supplementary Fig. 3c). GLUD2 depletion, however, completely blocked the effect of glutamate but not of NAC (Fig. 4d,e). Taken together, these results indicate that glutamate boosts redox homeostasis through GLUD2 to increase GSH/GSSG and NADPH/NADP+ ratios.
Glutamate depends on GLUD2 to boost redox homeostasis of IDH1R132H-heterozygous cells for 3D growth. Doxycycline (+ Dox) induction of GLUD2 shRNA markedly reduced GLUD2 expression at mRNA (a) and protein (b) levels in IDH1R132H-heterozygous BT142 cells. cGLUD2 shRNA (Dox) obliterated glutamate (+ Glu) but not N-acetyl cysteine (+ NAC) stimulation of 3D growth. Results are plotted in log2 ratios of treated over untreated (+ / −). Similarly, GLUD2 shRNA blocked glutamate effects to increase GSH/GSSG (d) and NADPH/NADP+ (e) ratios (n = 6) in reference to untreated cells (Ctr). One-way ANOVA was used in reference to the untreated
AGI-5198 enhances redox potential of IDH1R132H-heterozygous glioma cells independent of the glutamate pathway
Although the IDH1R132H inhibitor AGI-5198 is potently effective in diminishing D-2HG levels in vivo, its effect on tumor growth in preclinical studies remains controversial [11]. We observed that AGI-5198 stimulated 3D growth of IDH1R132H-heterozygous BT142 by twofold irrespective of GLUD2 status (Fig. 5a), suggesting a GLUD2-independent mechanism of AGI-5198. The stimulatory effect of AGI-5198 on 3D growth prompted us to ask whether AGI-5198 could induce tumor growth in the presence of GLUD2 shRNA. Two weeks after intracranial transplantation, injected mice were divided randomly into group 1 treated with doxycycline only and group 2 treated with doxycycline and AGI-5198. Despite the initially equivalent bioluminescent signals in the two groups, group 1 exhibited no tumor development—a steady decline in bioluminescent signal, whereas group 2 exhibited increasing tumor growth as a function of time (Fig. 5c). Tumor incidence was corroborated by direct imaging of autopsied brains (Fig. 5d) and confirmed by histological examination (data not shown). In support of preclinical studies showing shortened survival in intracranial tumor models treated with IDH1R132H inhibitor [44], our results indicated that AGI-5198 promotes glioma growth independent of the glutamate pathway.
AGI-5198 promotes 3D growth of IDH1R132H-heterozygous cells by boosting redox potential independent of the glutamate/GLUD2 pathway. a AGI-5198 treatment increased 3D growth of IDH1R132H-heterozygous BT142 irrespective of doxycycline induction. RLU, relative luciferase units. b A diagram depicts how extracellular glutamate (Glu) and AGI-5198 independently negate IDH1R132H tumor-suppressive activity by enhancing redox homeostasis in glioma. AGI-5198 induced intracerebral tumor growth, as indicated by bioluminescent imaging of live animals (c) and autopsied brains (d), of IDH1R132H-heterozygous BT142 in the presence of doxycycline induction. The starting time of both treatments is indicated. AGI-5198 treatment resulted in increased GSH/GSSG (e) and NAPDH/NADP+ (f) ratios independent of doxycycline induction
AGI-5198 has been shown to reduce radiosensitivity by restoring NADPH levels [45]. Although GLUD2 depletion increased GSSG concentration by 22%, AGI-5198 treatment elevated GSH/GSSG ratios similarly in GLUD2-proficient and -deficient cells by increasing GSH concentrations and decreasing GSSG concentrations (Fig. 5e). Whereas NADPH concentration decreased modestly in GLUD2-proficient cells with glutamate treatment (Supplementary Fig. 3c), AGI-5198 treatment of GLUD2-deficient cells yielded a modest NADPH concentration increase and pronounced NADP+ concentration decrease, consequently a lesser increase in NADPH/NADP+ ratio than AGI-5198–treated GLUD2-proficient cells (Fig. 5f). Together with the requirement of GLUD2 for glutamate stimulation of redox potential, these results indicate that AGI-5198 independently enhances redox potential to promote glioma growth of IDH1R132H-heterozygous cells.
Discussion
Choosing the proper IDH1R132H model systems is critical to the understanding of IDH1R132H glioma biology. Studies with endogenous model systems have demonstrated that heterozygous, but not hemizygous, IDH1R132H is tumor suppressive, in accordance with its association with better survival and the association of hemizygous IDH1R132H with malignant progression. In contrast, studies with exogenous expression tend to support the oncogenic theory where selection against IDH1R132H heterozygosity during 3D growth has essentially been overlooked [11]. We provide evidence that 3D, but not 2D, culture distinguished IDH1R132H-hemizygous cells from IDH1R132H-heterozygous cells (Fig. 1). Specifically, 3D-cultured IDH1R132H-heterozygous cells belonged to the cluster of lower malignancy, whereas 3D-cultured IDH1R132H-hemizygous cells fell into the cluster of higher malignancy; in contrast, 2D cultures gave rise to clustering together of these two cell types (Fig. 2). Furthermore, the glioblastoma mesenchymal gene set was exclusively enriched in 3D-cultured IDH1R132H-hemizygous cells (Fig. 3), consistent with the report that glioblastoma heterogeneity and molecular signatures are better represented by 3D than 2D culture [35]. The mesenchymal characterization of IDH1R132H-hemizygous cells is consistent with aggressive tumor growth (Fig. 3), less D-2HG production [17], and IDH1 and/or IDH1R132H copy-number alteration in glioma progression [40, 41]. Thus, 3D culture is more relevant to IDH1R132H glioma biology.
Glutamate metabolism has been recognized for its importance in IDH1R132H glioma growth. GLUD2 upregulation promotes lipid synthesis by increasing 2-oxoglutarate and citrate concentrations through the TCA cycle [25]. Furthermore, GLUD2 upregulation stimulates amino acid uptake for glutamate-dependent TCA cycle anaplerosis and glutathione recycling [23]. Likewise, upregulation of GLUD1 and GLUD2 in IDH1R132H gliomas is believed to provide sufficient NADPH and 2-oxoglutarate for TCA cycle anaplerosis [26, 27]. In keeping with our previous observation that glutamate, similar to NAC, stimulated 3D growth of IDH1R132H gliomagenic cells [18], this study indicates that the glutamate effect is to increase GSH/GSSG and NADPH/NADP+ ratios in IDH1R132H-heterozygous cells (Fig. 4), which is consistent with the metabolic changes of depleted glutathione and glutamate in IDH1R132H cells [46]. It should be noted, however, that the increased NADPH/NADP+ ratio arose from marked reduction of NADP+ concentrations without any increase of NADPH concentrations, supporting the role of GLUD2 in glutathione recycling [23]. Importantly, results from the rescue of GLUD2-depleted cells by NAC but not glutamate strongly indicate that glutamate stimulation of redox potential is sufficient to promote 3D growth of IDH1R132H-heterozygous cells (Fig. 5b). Although the mechanism underlying the glutamate effect on redox homeostasis requires further investigation, GLUD2 has been proposed as a therapeutic target of IDH1R132H glioma [23, 27].
AGI-5198 was reported initially to inhibit subcutaneous IDH1R132H glioma growth [29], but further studies failed to reproduce the finding despite near-complete inhibition of D-2HG [47]. In contrast, AGI-5198 has been shown to decrease radiosensitivity and cisplatin killing of IDH1R132H cancer cells by reducing reactive oxygen species [45, 48]. Furthermore, treatment of glioma patients with a mutant-selective IDH1 inhibitor revealed a trend of inverse correlation between D-2HG and GSH levels [49]. Results from this study showed that AGI-5198 increased redox potential and promoted 3D growth of IDH1R132H-heterozygous cells (Fig. 5), which maintain a low level of reducing equivalent associated with limited 3D growth [17]. Although the mechanism underlying redox regulation in IDH1R132H glioma cells requires further investigations, our results indicate that the effect of AGI-5198 is independent of the glutamate/GLUD2 pathway but converging on boosting redox homeostasis (Fig. 5b). Together with the result that AGI-5198 revived GLUD2-deficient tumor growth (Fig. 5c, d), these results not only provide an explanation for the disappointing and even detrimental outcomes of preclinical studies with IDH1 inhibitors [44, 47, 50] but also indicate the need to explore additional therapeutic targets including glutathione metabolism [51,52,53].
The evidence that AGI-5198 promotes glioma growth by boosting redox homeostasis lend credence to our original hypothesis that IDH mutations in glioma are beneficial [54]. Although the field of glioma research is still debating whether IDH1R132H is oncogenic or tumor suppressive, results from this study warn of counterproductive outcomes from the treatment of glioma patients with IDH1R132H inhibitors [11]. Our finding that glutamate and AGI-5198 negate IDH1R132H tumor-suppressive activity via independently boosting redox potential suggests a metabolic vulnerability for therapeutic targeting.
References
Ostrom QT, Gittleman H, Xu J et al (2016) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2009–2013. Neuro Oncol 18:v1–v75. https://doi.org/10.1093/neuonc/now207
Wen PY, Kesari S (2008) Malignant gliomas in adults. N Engl J Med 359:492–507. https://doi.org/10.1056/NEJMra0708126
Parsons DW, Jones S, Zhang X et al (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321:1807–1812. https://doi.org/10.1126/science.1164382
Balss J, Meyer J, Mueller W et al (2008) Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol 116:597–602. https://doi.org/10.1007/s00401-008-0455-2
Yan H, Parsons DW, Jin G et al (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360:765–773. https://doi.org/10.1056/NEJMoa0808710
Pusch S, Schweizer L, Beck A-C et al (2014) D-2-Hydroxyglutarate producing neo-enzymatic activity inversely correlates with frequency of the type of isocitrate dehydrogenase 1 mutations found in glioma. Acta Neuropathol Commun 2:19. https://doi.org/10.1186/2051-5960-2-19
Dang L, White DW, Gross S et al (2009) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462:739–744. https://doi.org/10.1038/nature08617
Xu W, Yang H, Liu Y et al (2011) Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19:17–30. https://doi.org/10.1016/j.ccr.2010.12.014
Turcan S, Rohle D, Goenka A et al (2012) IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483:479–483. https://doi.org/10.1038/nature10866
Lu C, Ward PS, Kapoor GS et al (2012) IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483:474–478. https://doi.org/10.1038/nature10860
Huang LE (2019) Friend or foe-IDH1 mutations in glioma 10 years on. Carcinogenesis 11:1299–1307. https://doi.org/10.1093/carcin/bgz134
Piaskowski S, Bienkowski M, Stoczynska-Fidelus E et al (2011) Glioma cells showing IDH1 mutation cannot be propagated in standard cell culture conditions. Br J Cancer 104:968–970. https://doi.org/10.1038/bjc.2011.27
Borodovsky A, Salmasi V, Turcan S et al (2013) 5-azacytidine reduces methylation, promotes differentiation and induces tumor regression in a patient-derived IDH1 mutant glioma xenograft. Oncotarget 4:1737–1747. https://doi.org/10.18632/oncotarget.1408
Luchman HA, Chesnelong C, Cairncross JG, Weiss S (2013) Spontaneous loss of heterozygosity leading to homozygous R132H in a patient-derived IDH1 mutant cell line. Neuro Oncol 15:979–980. https://doi.org/10.1093/neuonc/not064
Chesnelong C, Chaumeil MM, Blough MD et al (2014) Lactate dehydrogenase A silencing in IDH mutant gliomas. Neuro Oncol 16:686–695. https://doi.org/10.1093/neuonc/not243
Luchman HA, Stechishin OD, Dang NH et al (2012) An in vivo patient-derived model of endogenous IDH1-mutant glioma. Neuro Oncol 14:184–191. https://doi.org/10.1093/neuonc/nor207
Tiburcio PDB, Xiao B, Berg S et al (2018) Functional requirement of a wild-type allele for mutant IDH1 to suppress anchorage-independent growth through redox homeostasis. Acta Neuropathol 135:285–298. https://doi.org/10.1007/s00401-017-1800-0
Tiburcio PDB, Xiao B, Chai Y et al (2018) IDH1R132H is intrinsically tumor-suppressive but functionally attenuated by the glutamate-rich cerebral environment. Oncotarget 9:35100–35113. https://doi.org/10.18632/oncotarget.26203
Sasaki M, Knobbe CB, Itsumi M et al (2012) D-2-hydroxyglutarate produced by mutant IDH1 perturbs collagen maturation and basement membrane function. Genes Dev 26:2038–2049. https://doi.org/10.1101/gad.198200.112
Bardella C, Al-Dalahmah O, Krell D et al (2016) Expression of Idh1R132H in the murine subventricular zone stem cell niche recapitulates features of early gliomagenesis. Cancer Cell 30:578–594. https://doi.org/10.1016/j.ccell.2016.08.017
Pirozzi CJ, Carpenter AB, Waitkus MS et al (2017) Mutant IDH1 disrupts the mouse subventricular zone and alters brain tumor progression. Mol Cancer Res 15:507–520. https://doi.org/10.1158/1541-7786.MCR-16-0485
Amankulor NM, Kim Y, Arora S et al (2017) Mutant IDH1 regulates the tumor-associated immune system in gliomas. Genes Dev 31:774–786. https://doi.org/10.1101/gad.294991.116
Waitkus MS, Pirozzi CJ, Moure CJ et al (2018) Adaptive evolution of the GDH2 allosteric domain promotes gliomagenesis by resolving IDH1R132H-induced metabolic liabilities. Cancer Res 78:36–50. https://doi.org/10.1158/0008-5472.CAN-17-1352
Núñez FJ, Mendez FM, Kadiyala P et al (2019) IDH1-R132H acts as a tumor suppressor in glioma via epigenetic up-regulation of the DNA damage response. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aaq1427
Chen R, Nishimura MC, Kharbanda S et al (2014) Hominoid-specific enzyme GLUD2 promotes growth of IDH1R132H glioma. Proc Natl Acad Sci USA 111:14217–14222. https://doi.org/10.1073/pnas.1409653111
Khurshed M, Molenaar RJ, Lenting K et al (2017) In silico gene expression analysis reveals glycolysis and acetate anaplerosis in IDH1 wild-type glioma and lactate and glutamate anaplerosis in IDH1-mutated glioma. Oncotarget 8:49165–49177. https://doi.org/10.18632/oncotarget.17106
Lenting K, Khurshed M, Peeters TH et al (2019) Isocitrate dehydrogenase 1-mutated human gliomas depend on lactate and glutamate to alleviate metabolic stress. FASEB J 33:557–571. https://doi.org/10.1096/fj.201800907RR
Choi H, Gillespie DL, Berg S et al (2015) Intermittent induction of HIF-1α produces lasting effects on malignant progression independent of its continued expression. PLoS ONE 10:e0125125. https://doi.org/10.1371/journal.pone.0125125
Rohle D, Popovici-Muller J, Palaskas N et al (2013) An inhibitor of mutant IDH1 delays growth and promotes differentiation of glioma cells. Science 340:626–630. https://doi.org/10.1126/science.1236062
Nix DA, Courdy SJ, Boucher KM (2008) Empirical methods for controlling false positives and estimating confidence in ChIP-Seq peaks. BMC Bioinform 9:523. https://doi.org/10.1186/1471-2105-9-523
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Verhaak RGW, Hoadley KA, Purdom E et al (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98–110. https://doi.org/10.1016/j.ccr.2009.12.020
Marin-Valencia I, Yang C, Mashimo T et al (2012) Analysis of tumor metabolism reveals mitochondrial glucose oxidation in genetically diverse human glioblastomas in the mouse brain in vivo. Cell Metab 15:827–837. https://doi.org/10.1016/j.cmet.2012.05.001
Birgersdotter A, Sandberg R, Ernberg I (2005) Gene expression perturbation in vitro–a growing case for three-dimensional (3D) culture systems. Semin Cancer Biol 15:405–412. https://doi.org/10.1016/j.semcancer.2005.06.009
Smith SJ, Wilson M, Ward JH et al (2012) Recapitulation of tumor heterogeneity and molecular signatures in a 3D brain cancer model with decreased sensitivity to histone deacetylase inhibition. PLoS ONE 7:e52335. https://doi.org/10.1371/journal.pone.0052335
Noushmehr H, Weisenberger DJ, Diefes K et al (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17:510–522. https://doi.org/10.1016/j.ccr.2010.03.017
Bhat KPL, Balasubramaniyan V, Vaillant B et al (2013) Mesenchymal differentiation mediated by NF-κB promotes radiation resistance in glioblastoma. Cancer Cell 24:331–346. https://doi.org/10.1016/j.ccr.2013.08.001
de Souza CF, Sabedot TS, Malta TM et al (2018) A distinct DNA methylation shift in a subset of glioma CpG island methylator phenotypes during tumor recurrence. Cell Rep 23:637–651. https://doi.org/10.1016/j.celrep.2018.03.107
Tiburcio PDB, Locke MC, Bhaskara S et al Association of gene upregulation with DNA hypomethylation and better outcome in IDH-mutant glioma. J Neurosurg (manuscript in revision)
Jin G, Reitman ZJ, Duncan CG et al (2013) Disruption of wild-type IDH1 suppresses D-2-hydroxyglutarate production in IDH1-mutated gliomas. Cancer Res 73:496–501. https://doi.org/10.1158/0008-5472.CAN-12-2852
Mazor T, Chesnelong C, Pankov A et al (2017) Clonal expansion and epigenetic reprogramming following deletion or amplification of mutantIDH1. Proc Natl Acad Sci USA 114:10743–10748. https://doi.org/10.1073/pnas.1708914114
Jiang L, Shestov AA, Swain P et al (2016) Reductive carboxylation supports redox homeostasis during anchorage-independent growth. Nature 532:255–258. https://doi.org/10.1038/nature17393
Son J, Lyssiotis CA, Ying H et al (2013) Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496:101–105. https://doi.org/10.1038/nature12040
Tateishi K, Wakimoto H, Iafrate AJ et al (2015) Extreme vulnerability of IDH1 mutant cancers to NAD+ depletion. Cancer Cell 28:773–784. https://doi.org/10.1016/j.ccell.2015.11.006
Molenaar RJ, Botman D, Smits MA et al (2015) Radioprotection of IDH1-mutated cancer cells by the IDH1-mutant inhibitor AGI-5198. Cancer Res 75:4790–4802. https://doi.org/10.1158/0008-5472.CAN-14-3603
Reitman ZJ, Jin G, Karoly ED et al (2011) Profiling the effects of isocitrate dehydrogenase 1 and 2 mutations on the cellular metabolome. Proc Natl Acad Sci USA 108:3270–3275. https://doi.org/10.1073/pnas.1019393108
Turcan S, Fabius AWM, Borodovsky A et al (2013) Efficient induction of differentiation and growth inhibition in IDH1 mutant glioma cells by the DNMT inhibitor decitabine. Oncotarget 4:1729–1736. https://doi.org/10.18632/oncotarget.1412
Khurshed M, Aarnoudse N, Hulsbos R et al (2018) IDH1-mutant cancer cells are sensitive to cisplatin and an IDH1-mutant inhibitor counteracts this sensitivity. FASEB J 32:6344–6352. https://doi.org/10.1096/fj.201800547R
Andronesi OC, Arrillaga-Romany IC, Ly KI et al (2018) Pharmacodynamics of mutant-IDH1 inhibitors in glioma patients probed by in vivo 3D MRS imaging of 2-hydroxyglutarate. Nat Commun 9:1474–1479. https://doi.org/10.1038/s41467-018-03905-6
Kopinja J, Sevilla RS, Levitan D et al (2017) A brain penetrant mutant IDH1 inhibitor provides In vivo survival benefit. Sci Rep 7:13853. https://doi.org/10.1038/s41598-017-14065-w
Waitkus MS, Diplas BH, Yan H (2018) Biological role and therapeutic potential of IDH mutations in cancer. Cancer Cell 34:186–195. https://doi.org/10.1016/j.ccell.2018.04.011
Liu Y, Lu Y, Celiku O et al (2019) Targeting IDH1-mutated malignancies with NRF2 blockade. J Natl Cancer Inst. https://doi.org/10.1093/jnci/djy230
Tang X, Fu X, Liu Y et al (2019) Blockade of glutathione metabolism in IDH1-mutated glioma. Mol Cancer Ther. https://doi.org/10.1158/1535-7163.MCT-19-0103
Huang LE, Cohen AL, Colman H et al (2017) IGFBP2 expression predicts IDH-mutant glioma patient survival. Oncotarget 8:191–202. https://doi.org/10.18632/oncotarget.13329
Acknowledgements
Research reported in this publication utilized the High-Throughput Genomics and Bioinformatic Analysis Shared Resource and the Biorepository and Molecular Pathology Shared Resource at Huntsman Cancer Institute at the University of Utah. This work was supported in part by funds in conjunction with grant P30 CA042014 from the National Cancer Institute of the National Institutes of Health. This work was supported in part by the National Institute of Neurological Disorders and Stroke R21NS108065 Grant. The authors thank Chris Stubben for providing consultation for the analysis of RNA sequencing data, Luming Zhou and Carl T. Wittwer for the assistance in quantitative PCR analysis, Laura Roberts for technical assistance, and Kristin Kraus for editorial assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tiburcio, P.D.B., Gillespie, D.L., Jensen, R.L. et al. Extracellular glutamate and IDH1R132H inhibitor promote glioma growth by boosting redox potential. J Neurooncol 146, 427–437 (2020). https://doi.org/10.1007/s11060-019-03359-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-019-03359-w