Abstract
The TNF receptor superfamily member Fn14 is overexpressed by many solid tumor types, including glioblastoma (GBM), the most common and lethal form of adult brain cancer. GBM is notable for a highly infiltrative growth pattern and several groups have reported that high Fn14 expression levels can increase tumor cell invasiveness. We reported previously that the mesenchymal and proneural GBM transcriptomic subtypes expressed the highest and lowest levels of Fn14 mRNA, respectively. Given the recent histopathological re-classification of human gliomas by the World Health Organization based on isocitrate dehydrogenase 1 (IDH1) gene mutation status, we extended this work by comparing Fn14 gene expression in IDH1 wild-type (WT) and mutant (R132H) gliomas and in cell lines engineered to overexpress the IDH1 R132H enzyme. We found that both low-grade and high-grade (i.e., GBM) IDH1 R132H gliomas exhibit low Fn14 mRNA and protein levels compared to IDH1 WT gliomas. Forced overexpression of the IDH1 R132H protein in glioma cells reduced Fn14 expression, while treatment of IDH1 R132H-overexpressing cells with the IDH1 R132H inhibitor AGI-5198 or the DNA demethylating agent 5-aza-2′-deoxycytidine increased Fn14 expression. These results support a role for Fn14 in the more aggressive and invasive phenotype associated with IDH1 WT tumors and indicate that the low levels of Fn14 gene expression noted in IDH1 R132H mutant gliomas may be due to epigenetic regulation via changes in DNA methylation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The cytokine TNF-like weak inducer of apoptosis (TWEAK) and its cell surface receptor fibroblast growth factor-inducible 14 (Fn14) are minimally expressed in normal, uninjured tissues; however, increased expression of the TWEAK and/or Fn14 genes has been detected in numerous solid tumor types [1,2,3,4,5]. In the setting of glioblastoma (GBM), the most prevalent and lethal form of adult brain cancer [6, 7], elevated Fn14 expression is associated with poor patient survival [8]. Four major transcriptional subtypes of GBM—classical, mesenchymal, neural and proneural—have been identified [9,10,11] and we reported previously that Fn14 mRNA was expressed at the highest levels in the more invasive and aggressive mesenchymal subtype tumors [4]. Glioma cell invasion into normal brain parenchyma is a hallmark of GBM pathophysiology and generally leads to tumor recurrence [6, 7]. Interestingly, glioma cells located in the invasive rim express Fn14 at higher levels than those cells residing in the tumor core [8, 12], and TWEAK:Fn14 engagement as well as Fn14 overexpression can trigger glioma cell invasion in vitro and in vivo [4, 8, 12, 13].
Recently, the histopathological classification of GBM has been modified based on genetic mutations in the isocitrate dehydrogenase (IDH) gene [14, 15]. IDH mutations were first identified in GBM tumors in 2008 during an analysis of 20,661 protein-coding genes in 22 GBM samples [16]. This work laid the foundation for more extensive genomic analyses, which identified mutations in the IDH1 or IDH2 isoforms in 60–80% of grade II and III gliomas [17]. These mutations are also found in 80% of secondary GBMs, which are tumors that have developed from low grade gliomas (LGGs), but only 3–7% of primary GBMs (i.e. those that have arisen spontaneously) [18]. The most commonly identified mutation is the IDH1 R132H gain-of-function mutation, which catalyzes the NADPH dependent reduction of α-ketoglutarate (α-KG) to produce the R enantiomer of 2-hydroxyglutarate (2-HG) [19]. In turn, 2-HG inhibits α-KG-dependent dioxygenases, with resultant downstream effects that mediate tumor cell interactions with the environment, collagen modification, responses to hypoxia, and immune evasion [18, 20,21,22,23]. IDH1 mutant GBMs are more likely to involve the frontal lobes, demonstrate less contrast enhancement, and produce less peritumoral changes on MRI [24]. Patients with IDH1 mutant tumors are, on average, younger (33 vs. 53 years) and survive significantly longer than those with IDH1 WT tumors [16,17,18, 25, 26]. Gliomas with the IDH1 mutation are found to have a specific cellular phenotype, characterized by global DNA hypermethylation at CpG islands [18, 27]. Evidence suggests that these IDH mutation-related epigenetic changes impact a variety of biological processes, including metabolic pathways and transcriptional programs linked to tumor growth and development [18].
Although the TWEAK-Fn14 axis and the IDH1 mutation have both been shown to have important effects on glioma cell biology and patient outcome, the inter-relationship between these patho-biological systems has not been studied. Recent work has shown that forced overexpression of the IDH1 R132H protein in glioma cells reduces migration [28] and invasion [28, 29] in vitro. Given the role that Fn14 may play in promoting glioma cell invasion [4, 8, 12, 13], we investigated Fn14 gene expression levels in IDH1 WT and R132H mutant low-grade gliomas (LGGs) and GBM tumors as well as IDH1 R132H-overexpressing glioma cell lines. We report that resected gliomas and glioma cell lines carrying an IDH1 R132H mutation express Fn14 at relatively low levels compared to their IDH1 WT counterparts, supporting a possible link between low Fn14 levels, reduced cell invasion, and improved patient prognosis associated with the IDH1 mutation.
Materials and methods
Normal (non-neoplastic) brain and glioma mRNA expression analysis
TWEAK (queried as TNFSF12) and Fn14 (queried as TNFRSF12A) mRNA expression data corresponding to 10 normal brain and 548 GBM specimens were downloaded from the TCGA data portal (Level 3 data) [30]. In addition, Fn14 mRNA expression data from 286 LGGs and 240 GBMs whose IDH1 mutational status was available were downloaded from cBioPortal [31]. Among the LGG dataset, 65 IDH1 WT tumors and 221 IDH1 mutant tumors were identified, while among the GBM dataset, 227 IDH1 WT and 13 IDH1 mutant tumors were identified (12 containing the R132H point mutation, and 1 with an R132G mutation). Additionally, GBM subtype data (including samples with unknown IDH1 mutational status) were downloaded from the UCSC Cancer Genomics Browser [32]. Among this dataset, 144 classical, 155 mesenchymal, 87 neural, and 138 proneural tumors were identified. All 13 IDH1 mutant GBM samples were of the proneural subtype; these were analyzed as a distinct subset relative to the 125 proneural tumors with either WT (n = 45) or unknown (n = 80) IDH1 status. Gene expression data is presented as normalized z-scores, which reflect the number of standard deviations away from the mean of expression in the reference population. In this context, the reference population refers to either all tumors that are diploid for the gene in question, or matched normal brain tissue.
Immunohistochemical analysis of Fn14 expression in glioma patient samples
Formalin-fixed, paraffin-embedded (FFPE) gliomas were obtained from both the University of Maryland School of Medicine Department of Pathology and the Dignity Health/St. Joseph’s Hospital and Medical Center Department of Neuropathology (Phoenix, AZ). Samples were categorized into 4 groups: (1) IDH1 WT LGG, (2) IDH1 mutant LGG, (3) IDH1 WT GBM and (4) IDH1 mutant GBM. The IDH1 mutational status was assessed during each patient’s clinical workup via immunohistochemistry with an IDH1 R132H-specific antibody (Dianova GmbH, Hamburg, Germany), and was identified via chart review. Six IDH1 WT LGGs, five IDH1 mutant LGGs, five IDH1 WT GBMs, and three IDH1 mutant GBM samples were retrieved. Institutional Research Board (IRB) approval was obtained prior to collecting the archived tissue.
The FFPE glioma samples were immunostained using the UltraVision Quanto (HRP) detection system (Thermo Fischer Scientific; Waltham, MA). After routine deparaffinization with a series of xylene and alcohols, antigen retrieval was performed using 90% formic acid. Slides were then rinsed with distilled water and wash buffer. Endogenous peroxidase activity was blocked with hydrogen peroxide solution (TA-125-HP, Thermo Fisher Scientific) for 10 min prior to incubation with a rabbit anti-TWEAKR (Fn14) monoclonal antibody (ab109365, Abcam; Cambridge, MA) at 1:200 for 60 min at room temperature. The primary antibody signal was developed with Quanto detection reagents and DAB chromogen as per manufacturer’s instructions. Finally, the sections were counterstained with hematoxylin, rinsed, and mounted in Cytoseal XYL (Thermo Fischer Scientific). The immunostained slides were scanned using the Aperio CS2 digital pathology scanner (Leica Biosystems Inc; Buffalo Grove, IL).
Cell culture and western blot analysis
U138 and LN18 human glioma cell lines stably transfected with either vector or IDH1 R132H plasmid DNA [33] were obtained from Dr. Craig Horbinski (Northwestern University, Evanston, IL) and grown at 37 °C in a humidified incubator (95% air, 5% CO2) in DMEM supplemented with 10% fetal bovine serum and 1% penicillin–streptomycin (1000 units/l). For the AGI-5198 treatment experiment, LN18 IDH1 R132H cells were either treated with vehicle (DMSO) or AGI-5198 (1 µM; MedChem Express; Monmouth Junction, NJ) for 1, 2 or 3 days. For the 5-aza-2`-deoxycytidine (5-aza-dC) treatment experiment, LN18 IDH1 R132H cells were either treated with vehicle (DMSO) or three concentrations of 5-aza-dC (Millipore Sigma; St. Louis, MO) for 3 days. Cells were harvested by scraping, snap frozen, and lysed in HNTG buffer (20 mM HEPES, 150 mM NaCl, 1.5 mM MgCl2, 10% glycerol, and 1% Triton X-100) supplemented with a protease inhibitor cocktail (Millipore Sigma) and two phosphatase inhibitor cocktails (Millipore Sigma). The protein concentration of each lysate was determined by BCA protein assay (Pierce; Rockford, IL). Equal amounts of protein were subjected to SDS-PAGE (Life Technologies; Carlsbad, CA) and electrotransferred to PVDF membranes (Thermo Fischer Scientific). Immunoblotting was performed as previously described [34] using the following primary antibodies: IDH1 R132H (Dianova GmbH; Hamburg, Germany), Fn14 (Cell Signaling Technology (CST); Danvers, MA), PAR-4 (CST), MLH1 (CST), and GAPDH (CST). Densitometric analysis of the Western blot data was performed using Image J software and all Fn14 expression values were normalized to GAPDH values.
Statistical analysis
A statistical analysis was performed using a two-tailed Student’s t test or an analysis of variance (ANOVA) with a post-hoc Tukey’s honest significance difference (HSD) test, as appropriate. All tests were performed using JMP Pro 12 (SAS Institute Inc., Cary, NC) and R 3.2.2 (R Foundation for Statistical Computing, Vienna, Austria) software. p values < 0.05 were considered significant.
Results
Fn14 mRNA, but not TWEAK mRNA, is highly expressed in GBM
We first examined both TWEAK and Fn14 mRNA expression levels in 10 normal brain specimens and 548 GBM specimens by interrogating the TCGA GBM dataset. We found that Fn14 mRNA levels, but not TWEAK mRNA levels, were significantly elevated in GBM specimens (Fig. 1).
Comparison of TWEAK and Fn14 mRNA expression levels in normal brain (NB) versus GBM. a TWEAK and b Fn14 mRNA expression data in 10 NB and 548 GBM specimens were downloaded from the TCGA data portal and converted to z-scores. Patients used in this analysis were limited to the provisional TCGA dataset with gene expression availability for those two genes. Patient/sample sets for each expression cluster were plotted as box and whisker plots for each gene. The whiskers of the plot map the maximum and minimum z-score for each expression cluster. The bar and dotted bar in each box represent the median and mean value, respectively, for the expression z-score of each group. The top and bottom of each box represents the 25th and 75th percentile, respectively, of the expression z-score values for each group. TWEAK expression was not significantly different (NS, p = 0.419) while Fn14 expression was significantly different (p < 0.0001) between NB and GBM as determined using Student’s t test (two-tailed)
Fn14 mRNA is expressed at relatively low levels in the proneural, IDH1 mutant GBM subtype relative to the other molecular subtypes
We recently reported that the highest level of Fn14 mRNA expression occurred in mesenchymal subtype tumors [4]. However, this prior analysis utilized a relatively small GBM subtype sample size and did not distinguish between IDH1 WT and R132H mutant tumors. Therefore, we conducted additional Fn14 expression profiling studies using GBM subtype data obtained from the UCSC Cancer Genomics Browser. We identified 144 classical, 155 mesenchymal, 87 neural, and 138 proneural tumors. We also identified 13 IDH1 mutant GBM samples in the proneural group, which were analyzed as a distinct subset relative to the 125 proneural tumors with either WT (n = 45) or unknown (n = 80) IDH1 gene status.
We found that the Fn14 transcript was expressed at the highest levels in the classical and mesenchymal GBM subtypes (Fig. 2). The neural subtype demonstrated significantly lower Fn14 expression than tumors of the classical (p = 0.0013) and mesenchymal subtypes (p < 0.0001). Fn14 mRNA levels were even lower in GBMs of the proneural subtype compared to the neural subtype (p = 0.0022). Similarly, IDH1 mutant GBMs demonstrated lower Fn14 expression than GBMs of the classical (p < 0.0001), mesenchymal (p < 0.0001), and neural subtypes (p = 0.0016). Fn14 mRNA in the 13 proneural, IDH1 mutant GBMs showed a trend toward lower expression when compared to the other 125 proneural GBM subtype tumors, but this difference was not statistically significant (p = 0.2234).
Analysis of Fn14 mRNA expression levels in GBM subtypes. a Fn14 mRNA expression data in 524 GBM specimens were downloaded as z-scores from cBioPortal. The z-scores were computed by cBioPortal relative to the expression distribution of each gene in tumors that are diploid for this gene. Samples within each subtype (classical n = 144, mesenchymal n = 155, neural n = 87, proneural n = 125, IDH1 mutant = 13) were plotted as box and whisker plots. b Table showing p values for significance of mean across GBM subtypes for Fn14 expression (ANOVA followed by the Tukey–Kramer HSD test)
Fn14 mRNA is expressed at relatively low levels in IDH1 mutant LGGs and GBMs
Since IDH1 mutations are most frequently found in secondary GBMs (defined as those that develop from LGGs) [35], we next examined the relative levels of Fn14 expression in IDH1 WT and mutant LGGs. Of the 286 LGGs with sequencing data in the TCGA database, 221 samples (77.3%) exhibited an IDH1 mutation. Fn14 mRNA expression was significantly lower in LGGs with mutant IDH1 relative to those with WT IDH1 (p < 0.0001) (Fig. 3a). Within the GBM dataset, 524 samples were identified: the IDH1 mutational status of 284 samples (54.2%) was unknown, 227 (43.3%) were IDH1 WT, and 13 (2.5%) exhibited an IDH1 mutation. GBM tumors with the IDH1 mutation had, on average, significantly lower Fn14 mRNA levels than those with WT IDH1 (p < 0.0001) (Fig. 3b).
Analysis of Fn14 mRNA expression levels in IDH1 WT versus IDH1 mutant LGGs and GBMs. a Fn14 mRNA expression data from 286 LGGs whose IDH1 mutational status was available were downloaded from the cBioPortal. LGG samples (IDH1 WT n = 65, IDH1 mutant n = 221) were plotted as box and whisker plots. Fn14 expression levels were significantly lower in IDH1 mutant samples relative to IDH1 WT LGG samples (p < 0.0001) using Student’s t test (two-tailed). b Fn14 mRNA expression data from 240 GBMs were downloaded and analyzed as above. GBM samples (IDH1 WT n = 227, IDH1 mutant n = 13) were plotted as box and whisker plots. Fn14 expression levels differed significantly between the two groups (p < 0.0001) using Student’s t test (two-tailed)
Fn14 protein is also expressed at low levels in IDH1 mutant gliomas
As our Fn14 gene expression profiling analysis identified relatively low Fn14 mRNA levels in IDH1 mutant low and high-grade gliomas, we next studied whether this relationship was also present at the protein level in 19 resected glioma specimens. Immunohistochemistry revealed moderate Fn14 levels in IDH1 WT LGGs, low Fn14 levels in IDH1 mutant LGGs, moderate-to-high Fn14 levels in IDH1 WT GBMs, and perhaps most strikingly, low-to-moderate Fn14 levels in IDH1 mutant GBM samples. Representative immunohistochemistry images for these four tumor types are shown in Fig. 4.
Ectopic expression of IDH1 R132H protein in glioma cell lines decreases Fn14 expression and AGI-5198 or 5-aza-dC treatment of IDH1 R132H-overexpressing cells increases Fn14 expression
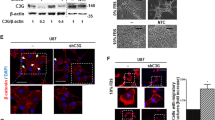
We next investigated whether IDH1 R132H overexpression in human glioma cells altered Fn14 levels using U138 and LN18 cells engineered to overexpress this protein [33]. Control, vector-transfected cells and the IDH1 R132H cells were grown under similar conditions, harvested and Fn14 expression was assayed by Western blot analysis. We found that Fn14 levels were reduced in the IDH1 R132H-overexpressing cells compared to their corresponding vector control cell lines, with the greatest difference, an ~ 7.0-fold decrease, noted in the LN18 cells (Fig. 5a). To confirm that IDH1 R132H enzymatic activity was responsible for the observed reduction in Fn14 levels, we treated LN18 IDH1 R132H cells with the selective IDH1 R132H inhibitor AGI-5198 [36] for various lengths of time. Cells were harvested and Fn14 expression was assayed by Western blot analysis. We also blotted for prostate apoptosis response-4 (PAR-4) expression as a positive control for a protein regulated by IDH1 R132H activity [33]. Drug treatment led to a transient increase in Fn14 levels, with the maximal increase in Fn14 expression (~ 2.2-fold) detected after 2 days of drug treatment (Fig. 5b).
Analysis of Fn14 protein expression in IDH1 R132H-overexpressing glioma cells and the effect of the IDH R132H inhibitor AGI-5198 or the demethylation agent 5-aza-dC on Fn14 levels. a Stably-transfected U138 and LN18 glioma cell lines were harvested and IDH1 R132H, Fn14 and GAPDH levels were evaluated by Western blot analysis. b LN18 IDH1 R132H cells were either treated with vehicle (DMSO) or 1 µM AG-5198 for the indicated number of days. Cells were harvested and Fn14, PAR-4 and GAPDH levels were evaluated by Western blot analysis. c LN18 IDH1 R132H cells were either treated with vehicle (DMSO) or three doses of 5-aza-dC for 3 days. Cells were harvested and Fn14, MLH1 and GAPDH levels were evaluated by Western blot analysis
It is well established that the IDH1 R132H metabolic product 2-HG promotes DNA hypermethylation [17, 22, 27]. Furthermore, it has been reported that the human Fn14 promoter contains a CpG island close to the transcription start site [37]. Therefore, to test the hypothesis that DNA methylation contributes to the IDH1 R132H-mediated decrease in Fn14 protein expression, we treated LN18 IDH1 R132H cells with several concentrations of the DNA methyltransferase inhibitor 5-aza-dC [38]. Cells were harvested after 3 days of treatment and Fn14 expression was assayed by Western blot analysis. We also blotted for MutL Homolog 1 (MLH1) expression as a positive control for a protein whose expression is regulated by DNA methylation [39]. We found that drug treatment increased Fn14 levels, with the maximal increase in Fn14 expression (~ 2.3-fold) detected using 5-aza-dC at a concentration of 10 µg/ml (Fig. 5c).
Discussion
There is increasing recognition of a significant role for the TWEAK receptor Fn14 in the molecular pathogenesis of GBM [4]. Prior studies have revealed major contributions related to tumor cell migration, invasion, and chemotherapy resistance [8, 12, 13, 40,41,42,43]. Here, we first examined TWEAK and Fn14 gene expression levels in normal brain and GBM samples. We found that TWEAK mRNA levels were not elevated in GBM compared to normal brain tissue, consistent with our prior study evaluating TWEAK gene expression in many fewer samples (one normal brain and 10 GBM tumors) by real-time quantitative PCR [43]. In contrast, Fn14 was significantly overexpressed in GBM, consistent with our earlier reports using smaller sample sizes and either quantitative PCR [43] or analysis of human GBM gene expression databases [4, 8]. In our prior report, we found that GBM tumors of the mesenchymal subtype exhibited a statistically significant higher level of Fn14 expression when compared to the classical, neural, and proneural subtypes [4]. Of note, a recent report indicates that the neural subtype is not a tumor-intrinsic transcriptional subtype in IDH WT GBMs [44]. Here, we evaluated ~ 3X more tumor specimens and found that the Fn14 expression level was similar in the classical and mesenchymal subtypes although there was a trend toward higher expression in the mesenchymal tumors. The Fn14 expression level in both the classical and mesenchymal subtypes was significantly higher than the other three subtypes examined.
We then focused on the analysis of Fn14 gene expression levels in IDH1 WT and mutant (R132H) gliomas. As mentioned above, the IDH mutation in gliomas has considerable prognostic implications, where mutated LGG patients live ~ 2.3X longer [45] and mutated GBM patients live ~ 2.0–3.5X longer [17, 45] than IDH1 WT patients. Emerging evidence suggests that these tumors co-opt numerous different biological processes compared to their WT counterparts [18, 20,21,22,23]. One of these differentially adapted processes is tumor cell invasion. IDH mutated tumors have more discrete imaging patterns, intraoperative borders, and histological margins [46, 47]. We report here that expression of the pro-invasive receptor Fn14 is lower in GBM tumors carrying the IDH1 gene mutation compared to those with an IDH1 WT gene. In the first analysis, where we examined Fn14 mRNA levels in the five GBM subtypes using data from the UCSC Cancer Genomics Browser (Fig. 2), proneural, IDH1 mutant GBMs showed a trend toward lower expression when compared to the other proneural GBM subtype tumors, but this difference was not statistically significant (p = 0.2234). This most likely reflects the fact that this proneural group contained 80 tumors of unknown IDH1 gene status, and if some of these were IDH1 mutant tumors, then we predict that the difference between the two groups would be masked. In any case, in the second analysis where we examined Fn14 mRNA expression in IDH1 WT and mutant GBMs using data from the cBioPortal (Fig. 3), the difference between Fn14 mRNA levels in IDH1 WT and mutant GBM tumors was highly significant (p < 0.0001). Of note, in this analysis we found a 5.4% incidence of the IDH1 mutation in GBMs and we did not classify GBMs as primary or secondary. The difference between Fn14 mRNA levels in IDH1 WT and mutant LGGs was also highly significant (p < 0.0001) (Fig. 3). We identified a 77.3% incidence of the IDH1 mutation in LGGs, which agrees with the incidence cited by prior studies [18, 26].
We then examined Fn14 gene expression levels in cell lines engineered to overexpress the IDH1 R132H enzyme. We found that ectopic IDH1 R132H overexpression in two glioma cell lines decreased Fn14 levels. This finding, in combination with the prior data linking elevated Fn14 signaling with glioma cell invasive capacity [4, 8, 13], suggests that there may be a link between the IDH1 gene mutation, low Fn14 levels, and decreased tumor invasiveness. This would be consistent with recent studies demonstrating that IDH1 R132H overexpression in human glioma cells decreases migration, invasion, and matrix metalloproteinase expression in vitro [28, 29]. Of note, Baldock et al. [47] used mathematical modeling to study the invasiveness of contrast-enhancing gliomas on MR imaging, and found no difference in the net rate of invasion or velocity of radial expansion in IDH1 WT and IDH1 mutant tumors. However, their analysis may have been limited by a reliance on MR imaging rather than histology to detect glioma cell invasion.
We found that when IDH1 R132H-overexpressing LN18 glioma cells were treated with an IDH1 R132H-specific small molecule inhibitor (AGI-5198), Fn14 expression was up-regulated. This result confirms that IDH1 R132 enzymatic activity, not some other potential undefined genetic or metabolic alteration in the cell line, was triggering the observed decrease in Fn14 levels. Since it is well established that the IDH1 R132H metabolic product 2-HG promotes DNA hypermethylation [22, 27] and it has been reported that the Fn14 promoter contains a CpG island close to the transcription start site [37], we also investigated whether DNA methylation might be responsible for IDH1 R132-mediated Fn14 down-regulation. IDH1 R132H-expressing cells were treated with 5-aza-dC, an analogue of cytidine that is incorporated into DNA and RNA. This compound inhibits DNA methyltransferase leading to a reduction in DNA methylation [38]. We observed that Fn14 expression was up-regulated after 5-aza-dC treatment, but the effect was not strictly dose-dependent, consistent with a previous report examining 5-aza-dC activity on antigen presentation in breast cancer cells [48]. Our results suggest that the low level of Fn14 protein expression noted in both IDH1 mutant-overexpressing cells and IDH1 mutant tumors may be due, at least in part, to transcriptional repression by an epigenetic mechanism.
Given the recent efforts to develop TWEAK- or Fn14-targeted therapeutic agents for the treatment of cancer patients [2, 3, 5], including GBM patients [4, 41, 42, 49], our findings also suggest that the IDH1 mutation status might serve as an important eligibility criterion in GBM clinical trials involving Fn14-targeted agents. Furthermore, our results support a role for Fn14 in the more aggressive phenotype demonstrated by IDH1 WT tumors, consistent with our previous reports showing that Fn14 expression levels positively correlate with glioma grade and poor GBM patient survival [8]. Additional work will be needed to further elucidate the connections between the IDH1 R132H mutation and Fn14 gene expression, Fn14 signaling and glioma cell invasion.
References
Winkles JA (2008) The TWEAK-Fn14 cytokine-receptor axis: discovery, biology and therapeutic targeting. Nat Rev Drug Discov 7:411–425
Culp PA, Choi D, Zhang Y, Yin J, Seto P, Ybarra SE, Su M, Sho M, Steinle R, Wong MH, Evangelista F, Grove J, Cardenas M, James M, Hsi ED, Chao DT, Powers DB, Ramakrishnan V, Dubridge R (2010) Antibodies to TWEAK receptor inhibit human tumor growth through dual mechanisms. Clin Cancer Res 16:497–508
Lassen UN, Meulendijks D, Siu LL, Karanikas V, Mau-Sorensen M, Schellens JH, Jonker DJ, Hansen AR, Simcox ME, Schostack KJ, Bottino D, Zhong H, Roessler M, Vega-Harring SM, Jarutat T, Geho D, Wang K, DeMario M, Goss GD (2015) A phase I monotherapy study of RG7212, a first-in-class monoclonal antibody targeting TWEAK signaling in patients with advanced cancers. Clin Cancer Res 21:258–266
Perez JG, Tran NL, Rosenblum MG, Schneider CS, Connolly NP, Kim AJ, Woodworth GF, Winkles JA (2016) The TWEAK receptor Fn14 is a potential cell surface portal for targeted delivery of glioblastoma therapeutics. Oncogene 35:2145–2155
Cheng E, Armstrong CL, Galisteo R, Winkles JA (2013) TWEAK/Fn14 axis-targeted therapeutics: moving basic science discoveries to the clinic. Front Immunol 4:473
Wen PY, Kesari S (2008) Malignant gliomas in adults. N Engl J Med 359:492–507
Ricard D, Idbaih A, Ducray F, Lahutte M, Hoang-Xuan K, Delattre JY (2012) Primary brain tumours in adults. Lancet 379:1984–1996
Tran NL, McDonough WS, Savitch BA, Fortin SP, Winkles JA, Symons M, Nakada M, Cunliffe HE, Hostetter G, Hoelzinger DB, Rennert JL, Michaelson JS, Burkly LC, Lipinski CA, Loftus JC, Mariani L, Berens ME (2006) Increased fibroblast growth factor-inducible 14 expression levels promote glioma cell invasion via Rac1 and nuclear factor-kappaB and correlate with poor patient outcome. Cancer Res 66:9535–9542
Phillips HS, Kharbanda S, Chen R, Forrest WF, Soriano RH, Wu TD, Misra A, Nigro JM, Colman H, Soroceanu L, Williams PM, Modrusan Z, Feuerstein BG, Aldape K (2006) Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 9:157–173
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O’Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN, Cancer Genome Atlas Research Network (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98–110
Brennan C, Momota H, Hambardzumyan D, Ozawa T, Tandon A, Pedraza A, Holland E (2009) Glioblastoma subclasses can be defined by activity among signal transduction pathways and associated genomic alterations. PLoS ONE 4:e7752
Fortin SP, Ennis MJ, Savitch BA, Carpentieri D, McDonough WS, Winkles JA, Loftus JC, Kingsley C, Hostetter G, Tran NL (2009) Tumor necrosis factor-like weak inducer of apoptosis stimulation of glioma cell survival is dependent on Akt2 function. Mol Cancer Res 7:1871–1881
Cherry EM, Lee DW, Jung JU, Sitcheran R (2015) Tumor necrosis factor-like weak inducer of apoptosis (TWEAK) promotes glioma cell invasion through induction of NF-kappaB-inducing kinase (NIK) and noncanonical NF-kappaB signaling. Mol Cancer 14:9
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Reifenberger G, Wirsching HG, Knobbe-Thomsen CB, Weller M (2017) Advances in the molecular genetics of gliomas—implications for classification and therapy. Nat Rev Clin Oncol 14:434–452
Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P, Mankoo P, Carter H, Siu IM, Gallia GL, Olivi A, McLendon R, Rasheed BA, Keir S, Nikolskaya T, Nikolsky Y, Busam DA, Tekleab H, Diaz LA Jr, Hartigan J, Smith DR, Strausberg RL, Marie SK, Shinjo SM, Yan H, Riggins GJ, Bigner DD, Karchin R, Papadopoulos N, Parmigiani G, Vogelstein B, Velculescu VE, Kinzler KW (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321:1807–1812
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins GJ, Friedman H, Friedman A, Reardon D, Herndon J, Kinzler KW, Velculescu VE, Vogelstein B, Bigner DD (2009) IDH1 and IDH2 mutations in gliomas. New Engl J Med 360:765–773
Dunn GP, Andronesi OC, Cahill DP (2013) From genomics to the clinic: biological and translational insights of mutant IDH1/2 in glioma. Neurosurg Focus 34:E2
Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, Marks KM, Prins RM, Ward PS, Yen KE, Liau LM, Rabinowitz JD, Cantley LC, Thompson CB, Vander Heiden MG, Su SM (2009) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462:739–744
Clark O, Yen K, Mellinghoff IK (2016) Molecular pathways: isocitrate dehydrogenase mutations in cancer. Clin Cancer Res 22:1837–1842
Amankulor NM, Kim Y, Arora S, Kargl J, Szulzewsky F, Hanke M, Margineantu DH, Rao A, Bolouri H, Delrow J, Hockenbery D, Houghton AM, Holland EC (2017) Mutant IDH1 regulates the tumor-associated immune system in gliomas. Genes Dev 31:774–786
Noushmehr H, Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RG, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H, Neuvial P, Cope LM, Buckley J, Herman JG, Baylin SB, Laird PW, Aldape K, Cancer Genome Atlas Research Network (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17:510–522
Dang L, Su SM (2017) Isocitrate dehydrogenase mutation and (R)-2-hydroxyglutarate: from basic discovery to therapeutics development. Annu Rev Biochem 86:305–331
Lai A, Kharbanda S, Pope WB, Tran A, Solis OE, Peale F, Forrest WF, Pujara K, Carrillo JA, Pandita A, Ellingson BM, Bowers CW, Soriano RH, Schmidt NO, Mohan S, Yong WH, Seshagiri S, Modrusan Z, Jiang Z, Aldape KD, Mischel PS, Liau LM, Escovedo CJ, Chen W, Nghiemphu PL, James CD, Prados MD, Westphal M, Lamszus K, Cloughesy T, Phillips HS (2011) Evidence for sequenced molecular evolution of IDH1 mutant glioblastoma from a distinct cell of origin. J Clin Oncol 29:4482–4490
Hartmann C, Hentschel B, Wick W, Capper D, Felsberg J, Simon M, Westphal M, Schackert G, Meyermann R, Pietsch T, Reifenberger G, Weller M, Loeffler M, von Deimling A (2010) Patients with IDH1 wild type anaplastic astrocytomas exhibit worse prognosis than IDH1-mutated glioblastomas, and IDH1 mutation status accounts for the unfavorable prognostic effect of higher age: implications for classification of gliomas. Acta Neuropathol 120:707–718
Dunn GP, Rinne ML, Wykosky J, Genovese G, Quayle SN, Dunn IF, Agarwalla PK, Chheda MG, Campos B, Wang A, Brennan C, Ligon KL, Furnari F, Cavenee WK, Depinho RA, Chin L, Hahn WC (2012) Emerging insights into the molecular and cellular basis of glioblastoma. Genes Dev 26:756–784
Turcan S, Rohle D, Goenka A, Walsh LA, Fang F, Yilmaz E, Campos C, Fabius AW, Lu C, Ward PS, Thompson CB, Kaufman A, Guryanova O, Levine R, Heguy A, Viale A, Morris LG, Huse JT, Mellinghoff IK, Chan TA (2012) IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483:479–483
Bralten LB, Kloosterhof NK, Balvers R, Sacchetti A, Lapre L, Lamfers M, Leenstra S, de Jonge H, Kros JM, Jansen EE, Struys EA, Jakobs C, Salomons GS, Diks SH, Peppelenbosch M, Kremer A, Hoogenraad CC, Smitt PA, French PJ (2011) IDH1 R132H decreases proliferation of glioma cell lines in vitro and in vivo. Ann Neurol 69:455–463
Cui D, Ren J, Shi J, Feng L, Wang K, Zeng T, Jin Y, Gao L (2016) R132H mutation in IDH1 gene reduces proliferation, cell survival and invasion of human glioma by downregulating Wnt/beta-catenin signaling. Int J Biochem Cell Biol 73:72–81
The Cancer Genome Atlas (TCGA) Research Network (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455:1061–1068
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:pl1
Cline MS, Craft B, Swatloski T, Goldman M, Ma S, Haussler D, Zhu J (2013) Exploring TCGA pan-cancer data at the UCSC cancer genomics browser. Sci Rep 3:2652
Liu Y, Gilbert MR, Kyprianou N, Rangnekar VM, Horbinski C (2014) The tumor suppressor prostate apoptosis response-4 (Par-4) is regulated by mutant IDH1 and kills glioma stem cells. Acta Neuropathol 128:723–732
Armstrong CL, Galisteo R, Brown SA, Winkles JA (2016) TWEAK activation of the non-canonical NF-kappaB signaling pathway differentially regulates melanoma and prostate cancer cell invasion. Oncotarget 7:81474–81492
Ichimura K, Pearson DM, Kocialkowski S, Backlund LM, Chan R, Jones DT, Collins VP (2009) IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas. Neuro Oncol 11:341–347
Rohle D, Popovici-Muller J, Palaskas N, Turcan S, Grommes C, Campos C, Tsoi J, Clark O, Oldrini B, Komisopoulou E, Kunii K, Pedraza A, Schalm S, Silverman L, Miller A, Wang F, Yang H, Chen Y, Kernytsky A, Rosenblum MK, Liu W, Biller SA, Su SM, Brennan CW, Chan TA, Graeber TG, Yen KE, Mellinghoff IK (2013) An inhibitor of mutant IDH1 delays growth and promotes differentiation of glioma cells. Science 340:626–630
Tajrishi MM, Shin J, Hetman M, Kumar A (2014) DNA methyltransferase 3a and mitogen-activated protein kinase signaling regulate the expression of fibroblast growth factor-inducible 14 (Fn14) during denervation-induced skeletal muscle atrophy. J Biol Chem 289:19985–19999
Gnyszka A, Jastrzebski Z, Flis S (2013) DNA methyltransferase inhibitors and their emerging role in epigenetic therapy of cancer. Anticancer Res 33:2989–2996
Herman JG, Umar A, Polyak K, Graff JR, Ahuja N, Issa JJ, Markowitz S, Willson JKV, Hamilton SR, Kinzler KW, Kane MF, Kolodner RD, Vogelstein B, Kunkel TA, Baylin SB (1998) Incidence and functional consequences of hMLH1 promoter hypermethylation in colorectal carcinoma. Proc Nat Acad Sci USA 95:6870–6875
Ensign SP, Roos A, Mathews IT, Dhruv HD, Tuncali S, Sarkaria JN, Symons MH, Loftus JC, Berens ME, Tran NL (2016) SGEF is regulated via TWEAK/Fn14/NF-kappaB signaling and promotes survival by modulation of the DNA repair response to temozolomide. Mol Cancer Res 14:302–312
Roos A, Dhruv HD, Mathews IT, Inge LJ, Tuncali S, Hartman LK, Chow D, Millard N, Yin HH, Kloss J, Loftus JC, Winkles JA, Berens ME, Tran NL (2017) Identification of aurintricarboxylic acid as a selective inhibitor of the TWEAK-Fn14 signaling pathway in glioblastoma cells. Oncotarget 8:12234–12246
Dhruv H, Loftus JC, Narang P, Petit JL, Fameree M, Burton J, Tchegho G, Chow D, Yin H, Al-Abed Y, Berens ME, Tran NL, Meurice N (2013) Structural basis and targeting of the interaction between fibroblast growth factor-inducible 14 and tumor necrosis factor-like weak inducer of apoptosis. J Biol Chem 288:32261–32276
Tran NL, McDonough WS, Donohue PJ, Winkles JA, Berens TJ, Ross KR, Hoelzinger DB, Beaudry C, Coons SW, Berens ME (2003) The human Fn14 receptor gene is up-regulated in migrating glioma cells in vitro and overexpressed in advanced glial tumors. Am J Pathol 162:1313–1321
Wang Q, Hu B, Hu X, Kim H, Squatrito M, Scarpace L, deCarvalho AC, Lyu S, Li P, Li Y, Barthel F, Cho HJ, Lin YH, Satani N, Martinez-Ledesma E, Zheng S, Chang E, Sauve CG, Olar A, Lan ZD, Finocchiaro G, Phillips JJ, Berger MS, Gabrusiewicz KR, Wang G, Eskilsson E, Hu J, Mikkelsen T, DePinho RA, Muller F, Heimberger AB, Sulman EP, Nam DH, Verhaak RGW (2017) Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32:42–56
Sanson M, Marie Y, Paris S, Idbaih A, Laffaire J, Ducray F, El Hallani S, Boisselier B, Mokhtari K, Hoang-Xuan K, Delattre JY (2009) Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J Clin Oncol 27:4150–4154
Kawaguchi T, Sonoda Y, Shibahara I, Saito R, Kanamori M, Kumabe T, Tominaga T (2016) Impact of gross total resection in patients with WHO grade III glioma harboring the IDH 1/2 mutation without the 1p/19q co-deletion. J Neurooncol 129:505–514
Baldock AL, Yagle K, Born DE, Ahn S, Trister AD, Neal M, Johnston SK, Bridge CA, Basanta D, Scott J, Malone H, Sonabend AM, Canoll P, Mrugala MM, Rockhill JK, Rockne RC, Swanson KR (2014) Invasion and proliferation kinetics in enhancing gliomas predict IDH1 mutation status. Neuro Oncol 16:779–786
Klar AS, Gopinadh J, Wadle A, Renner C (2015) Treatment with 5-aza-2′-deoxycytidine induces expression of NY-ESO-1 and facilitates cytotoxic T lymphocyte-mediated tumor cell killing. PLoS ONE 10:e0139221
Schneider CS, Perez JG, Cheng E, Zhang C, Mastorakos P, Hanes J, Winkles JA, Woodworth GF, Kim AJ (2015) Minimizing the non-specific binding of nanoparticles to the brain enables active targeting of Fn14-positive glioblastoma cells. Biomaterials 42:42–51
Acknowledgements
We thank Dr. Craig Horbinski for providing the glioma cell lines. We also thank Ashley Cellini and Kimberly Tuttle for their assistance with obtaining tissue slides from the University of Maryland Greenebaum Cancer Center’s Pathology and Biorepository Shared Service. This work was supported in part by National Institutes of Health Grant K08 NS09043 (G.F.W.), American Cancer Society Research Scholar Grant 128970-RSG-16-012-01-CDD (G.F.W.), and The Ben and Catherine Ivy Foundation (N.L.T.). J.G.D. was supported by the NIH T32 Training Grant CA154274 and an NIGMS Initiative for Maximizing Student Development Grant (R25 GM55036).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
All procedures performed in studies involving tissue from human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Given that this tissue was obtained from a tumor bank retrospectively, the institutional review board determined that formal consent is not required.
Rights and permissions
About this article
Cite this article
Hersh, D.S., Peng, S., Dancy, J.G. et al. Differential expression of the TWEAK receptor Fn14 in IDH1 wild-type and mutant gliomas. J Neurooncol 138, 241–250 (2018). https://doi.org/10.1007/s11060-018-2799-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-018-2799-3