Abstract
MicroRNAs (miRNAs) may act as prognostic biomarkers in a variety of cancers. The aim of this study was to identify and evaluate a prognostic miRNA signature in patients with lower-grade gliomas (LGGs). miRNA expression profiles and clinical data of patients with LGGs from the Chinese Glioma Genome Atlas (CGGA; the training cohort) and The Cancer Genome Atlas (TCGA; the validation cohort) were analyzed, and the least absolute shrinkage and selection operator Cox regression model was used to identify the miRNA signature, which was combined with clinical prognostic factors to develop an individualized survival prediction model. Gene ontology analysis and Kyoto Encyclopedia of Genes and Genomes pathway analysis were conducted to reveal the biological implications of the signature. We identified a four-miRNA signature that stratified patients in the training cohort into low- or high-risk groups according to overall survival time, a finding that was verified in the validation cohort. Multivariate Cox regression analysis indicated that the four-miRNA signature was an independent prognostic biomarker, and a nomogram combining this miRNA signature with clinicopathological and molecular factors showed high prognostic accuracy for individualized survival prediction in both TCGA (C-index = 0.83) and CGGA (C-index = 0.68) cohorts. Functional annotation indicated that the major biological processes of this prognostic miRNA signature were enriched in cell cycle and DNA repair-associated biological processes. Our findings indicated that our newly discovered four-miRNA signature may be an effective independent prognostic factor for the prediction of patients with LGGs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lower-grade gliomas (LGGs) are a group of heterogeneous neuroepithelial tumors comprising diffuse low-grade and intermediate-grade gliomas [World Health Organization (WHO) grades II and III] [1, 2]. The clinical course of the disease spans a broad spectrum, highlighting the need to stratify patients into distinct subgroups with more uniform clinical outcomes. According to the 2016 WHO brain tumor classification [3], a number of genetic and molecular abnormalities, including IDH mutation and 1p/19q codeletion, have been progressively incorporated as supportive markers to facilitate the comprehensive assessment of patients with LGGs and as an integral part of LGG subclassification [4, 5]; these features have been shown to distinguish biologically and clinically distinct subtypes. Nevertheless, a deep understanding of the biological differences between individual patients remains distant, and new prognostic biomarkers are required to better determine clinical outcome and devise patient-tailored treatment.
Recently, the discovery of microRNAs (miRNAs) has led to new discoveries for both the diagnosis and prognosis of various tumors [6, 7]. miRNAs are short non-coding RNAs consisting of approximately 22 nucleotides that play an effective tumor-regulatory function by modulating multiple targets at the posttranscriptional level. Ample evidence has indicated that abnormal expression of miRNAs in glioma is closely associated with multiple biological processes including cellular proliferation, angiogenesis, apoptosis, metastasis, and invasion [8,9,10]. However, whether a miRNA signature derived from one or more miRNAs can predict clinical outcomes in patients with LGGs is unknown.
In the present study, we used available miRNA data from 100 patients with LGGs from the Chinese Glioma Genome Atlas (CGGA) to identify a unique miRNA signature as a putative prognostic biomarker, and further validated its predictive properties using data from a different cohort of 420 patients from The Cancer Genome Atlas (TCGA) database. Furthermore, we developed and validated a miRNA-based predictive model that integrated our newly discovered four-miRNA signature with traditional clinicopathological risk factors for patient-tailored survival prediction in patients with LGGs.
Methods
Patient selection
We used 520 patients with miRNA expression data and corresponding clinical information in this study, of which 100 samples were downloaded from the CGGA database (http://www.cgga.org.cn) as the training set, and 420 samples were obtained from TCGA (cancergenome.nih.gov) as the validation set. Selection criteria for both cohorts were as follows: (a) histopathologically confirmed LGGs according to WHO classification; (b) high-quality miRNA expression data available; and (c) no history of radiation therapy, chemotherapy, or corticosteroid therapy before surgery. Patients with incomplete prognostic information were excluded. Overall survival (OS) was calculated from the date of initial diagnosis until death or last follow-up examination. All patients in the CGGA provided written informed consent to participate and the privacy of patients was strictly protected. The study protocol was approved by the ethics committee of Beijing Tiantan Hospital.
miRNA expression, mRNA expression, and biomarker detection
For the training cohort, miRNA expression data was generated using the human v2.0 miRNA Expression BeadChip (Illumina, Inc., San Diego, CA, USA) with 1146 miRNAs covering 97% of the miRBase 12.0 database [11], mRNA expression was generated by the Agilent Whole Human Genome Array platform [12], and mutations in IDH were detected by pyrosequencing [13], all of which have been described in detail in our previous study. For the validation cohort, miRNA expression data, level 3 RNA sequencing data (RSEM normalized) and molecular data (IDH mutation, 1p/19q co-deletion, and ATRX mutation status) were obtained from TCGA database.
Feature selection and risk score construction
Evidence of an association between differentially expressed miRNAs and patient OS was determined using a univariate Cox proportional regression model, and P values of < 0.05 were considered statistically significant. Because it is suitable for regression analysis of high-dimensional data, the least absolute shrinkage and selection operator (LASSO) method was used to identify and select the most useful predictive features. The following risk score formula was used to calculate for each patient by applying a linear combination of selected features that were weighted by their respective coefficients (Coef) from LASSO:
Identical β values were applied to the validation cohort.
Prediction of survival outcome using a miRNA-based risk score
Patients with assigned risk scores in the training and validation cohorts were classified into the high-risk or low-risk group using the median as the cutoff point. The OS rates of patients in the high-risk and low-risk groups were first assessed in the training dataset and then validated in the validation dataset using the Kaplan–Meier method. The log-rank test was used to determine survival differences between groups. Similarly, differences in the survival curves for each identified significant feature were evaluated. The survival impact of the four-miRNA signature within each LGGs subtype was also investigated. Associations between risk score and clinical characteristics were assessed with Chi square or Fisher’s exact tests. Univariate and multivariate Cox regression analysis was used to evaluate whether the four-miRNA signature was an independent prognostic factor.
Individualized prediction model construction
Nomograms for individualized prediction of patient outcomes were generated based on the results of the multivariate analysis to predict 1-, 2-, 3-, and 5-year OS with the rms package in R [14]. To minimize information loss, a backward stepdown selection process was performed to recruit the independent prognostic factors into the final nomogram model conforming to the Akaike information criterion [15]. Harrell’s concordance-index (C-index) and calibration curves were used to assess predictive performance and discriminative ability of the nomograms [16]. Furthermore, the prognostic nomogram was validated in an independent external cohort.
Bioinformatics analysis
Pearson correlation analysis, which was performed using the R programming language (cran.r-project.org), was used to identify genes associated with a miRNA-based risk score, identified based on a P value of < 0.05 and a Pearson correlation coefficient of > 0.3. Furthermore, we performed gene ontology analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways analysis to identify the biological mechanisms of the miRNA-based signature and the pathways in which they may be involved using DAVID (david.ncifcrf.gov). False discovery rates (FDRs) were used to address multiple comparisons. The biological process network was visualized using Cytoscape.
Results
Identification of prognostic miRNAs and association with OS
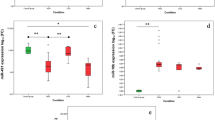
Using univariate Cox proportional regression analysis, we identified 133 protective miRNAs and 128 oncogenic miRNAs that were significantly associated with patient OS (Fig. 1a). Next, we used a LASSO Cox regression model to identify the most informative putatively prognostic miRNAs, and identified four miRNAs (miR-590-3p, miR-10b, miR-196a, and miR-15b-3p) with non-zero regression coefficients (Fig. 1b, c). All four miRNAs showed high expression (using median expression as a cutoff) and were associated with shorter patient survival, findings which were verified in the validation cohort (Fig. 2a–h).
Kaplan–Meier plot of OS in patients according to the expression of four miRNAs in patients with LGGs. a, c, e, g Each of the four identified miRNAs divides patients into groups with significantly different prognosis in the training dataset. b, d, f, h Expression of the four identified miRNAs retained prognostic significance for patients in the validation set
Development of a miRNA risk score and association with OS
A risk score formula was constructed based on the individual expression levels of the four miRNAs and their respective coefficients as follows: \({\text{Risk score}}=\left( {{\text{expressio}}{{\text{n}}_{{\text{miR-59}}0{\text{-3p}}}} \times \left( {{\text{6}}.{\text{84}} \times {\text{1}}{0^{ - 0{\text{5}}}}} \right)} \right)+\left( {{\text{expressio}}{{\text{n}}_{{\text{miR-1}}0{\text{b}}}} \times \left( {{\text{4}}.{\text{18}} \times {\text{1}}{0^{ - 0{\text{5}}}}} \right)} \right)+\left( {{\text{expressio}}{{\text{n}}_{{\text{miR-196a}}}} \times \left( {{\text{4}}.{\text{11}} \times {\text{1}}{0^{ - 0{\text{5}}}}} \right)} \right)+\left( {{\text{expressio}}{{\text{n}}_{{\text{miR-15b-3p}}}} \times \left( {{\text{9}}.{\text{43}} \times {\text{1}}{0^{ - 0{\text{6}}}}} \right)} \right)\). The dichotomized risk score (using the median risk score as a cutoff) enabled us to segment patients into high- (n = 50) and low-risk (n = 50) groups [hazard ratio (HR) = 0.1213, 95% confidence interval (CI) 0.060–0.287] in the training cohort (Fig. 3a). To confirm that the four-miRNA signature had a similar prognostic value in different populations, we then applied it to predict OS in an independent validation cohort using the median risk score as a cutoff. We found that the dichotomized risk score could also stratify patients from the validation cohort into high- (n = 210) and low-risk groups (n = 210) (HR = 0.3892, 95% CI 0.256–0.593, Fig. 3d). When stratifying patients by IDH mutation status, the four-miRNA signature remained a significant prognostic factor in both cohorts (Fig. 3b, c, e, f). In addition, this signature allowed patients within each LGG subtype to be further stratified (Fig. S1).
Kaplan–Meier plot of OS in patients according to the four-miRNA signature risk score. The dichotomized risk score allowed the segmentation of patients into high- and low-risk groups in both the a training and d validation cohorts. When stratifying by IDH mutation status, the four-miRNA signature remained a significant prognostic factor in both the b, c training and e, f validation cohorts
Associations between the four-miRNA signature with clinicopathological variables and patient outcome
Next, we investigated whether there were associations between the four-miRNA signature and widely accepted prognostic factors in patients with LGGs. We found that the risk score was significantly associated with several known prognostic factors (age, sex, WHO grade, histology, IDH status, and 1p/19q status) in the large TCGA dataset (Table 1).
We then performed univariate and multivariate Cox regression analysis to ascertain whether the four-miRNA signature could be an independent predictor for patients with LGGs. Based on the results from univariate analysis, we found that the risk score (P < 0.001), age (P = 0.039), WHO grade (P < 0.001), and IDH status (P = 0.042) were significantly associated with survival in the training cohort. Similar results were found in the validation cohort (Table 2). After multivariable adjustment using the above factors, the risk score remained a powerful and independent factor (P < 0.001) in both the training and validation cohort (Table 2).
Establishment and validation of the individualized prediction models
Using a backward stepwise method based on the smallest Akaike information criterion, prognostic nomograms that integrated independent prognostic parameters (WHO grade, age at diagnosis, IDH status, and risk score) were constructed. In TCGA cohort, we found that the C-index for the prediction nomogram was 0.83, whereas the C-index of the nomogram for predicting OS was 0.68 in the CGGA cohort. The calibration curve also demonstrated excellent agreement between prediction and observation in the probabilities of 1-, 2-, 3-, and 5-year OS in both cohorts (Fig. 4).
Significant functions and pathway enrichment analysis
Paired miRNA and mRNA expression data of 93 and 420 samples representing the training and validation data sets, respectively, were eligible for subsequent analysis. In the training cohort, we identified 3012 genes that were significantly associated with different risk score groups, including 1487 that were up-regulated in high-risk patients and 1525 that were downregulated in low-risk patients (Fig. 5a). We then used gene ontology analysis and network analysis to explore the underlying biological functions of genes associated with high-risk patients, and found that the major biological processes were enriched in cell cycle and modification, nucleosome assembly, RNA processing, respiratory electron transport, translational elongation, and DNA repair (Fig. 5b, c). KEGG pathway enrichment analysis showed that cell cycle, DNA replication, and mismatch repair were the main associated pathways (Fig. 5d). Similar four-miRNA signature-related biological processes and signal pathways, which are widely thought to play important roles in tumor proliferation, were observed in the validation dataset (Fig. S2).
Significant functions and pathway enrichment analysis of the four-miRNA signature in the training dataset. a Heat map of differentially expressed genes between high- and low-risk four-miRNA signature groups from 93 samples of LGGs. b Network analysis and c gene ontology analysis showing that the score from a high-risk group was associated with certain biological processes, such as cell cycle and nucleosome assembly. d KEGG pathway enrichment analysis showing that patient classification as high-risk was associated with several specific pathways
Discussion
To elucidate the biological mechanisms underlying LGGs, previous studies have analyzed DNA mutations, RNA expression, DNA copy number, and DNA methylation data [1, 2, 5, 17, 18]. However, information at the post-transcriptional level would provide additional information to improve our understanding of the biological processes underlying LGGs. In the present study, we performed an analysis of post-transcriptional data and identified a four-miRNA signature for prediction of OS in patients with LGGs. This unique signature successfully and robustly stratified patients into high-risk and low-risk groups in both the training (CGGA) and validation (TCGA) cohorts, and the four-miRNA-based nomogram provided an effective approach for individual survival estimation. Furthermore, in silico biological analyses identified potential functional roles of the miRNAs that comprise this prognostic signature in processes implicated in tumorigenesis.
Using a LASSO regression model, we identified four miRNAs (miR-590-3p, miR-10b, miR-196a, and miR-15b-3p) whose expression profiles were significantly associated with patient prognosis and showed that some of these miRNAs play important roles in glioma pathogenesis and progression. For instance, increased expression of miR-10b has been shown to be significantly associated with glioma grade progression [19], and in vitro studies have demonstrated that overexpression of miR-10b promotes invasion, migration, extracellular matrix remodeling, and tumor progression [20,21,22]. In addition, functional analysis revealed that miR-10b mediates tumor progression through the cell cycle and increased hypoxia [23, 24]. Similarly, miR-196a plays a role in the progression of malignancy in patients with glioma and is correlated with OS in patients with glioblastoma [25]. Previous studies of potential regulatory mechanisms demonstrated that overexpression of miR-196a may contribute to cell proliferation and the suppression of apoptosis by activation of NF-kB in glioma cells [26]. Hence, we proposed that the prognostic value of our identified miRNAs may be derived from their important roles in regulating the initiation and progression of LGGs. Furthermore, using LASSO modeling, we integrated multiple miRNAs into a single signature that may better reflect the complex nature and biological processes that underlie LGGs compared with that of a single miRNA. Although TCGA follow-up times were longer than those in the CGGA database, our four-miRNA signature remained a significant predictor in both the training and validation cohorts, which demonstrate that the signature is robust and reliable.
IDH mutations and the 1p/19q co-deletion are particularly notable molecular alterations that occur in the very early stage of gliomagenesis and are considered key prognostic factors for patients with LGGs according to a new WHO classification [1, 17, 27]. Although recent studies have shown that molecular-based stratification (i.e., groups with IDH1 or IDH2 mutations and the 1p/19q co-deletion, IDH1 or IDH2 mutations and no 1p/19q co-deletion, and IDH1/IDH2 wildtype) can be used to categorize patients in clinically and etiologically similar groups [1, 5], genetic characterization alone may be insufficient to comprehensively delineate or define tumor behaviors or mechanisms. In fact, previous studies have identified and validated age, WHO grade, seizure, and other prognostic factors [28,29,30]; however in this study, we generated a miRNA-based nomogram integrating molecular markers as well as clinical and genomic data that yielded a more comprehensive and individual prognostic prediction for patients with LGGs. The key benefits of this model are that it provides a complementary perspective about a single tumor and develops an individual scoring system for patients.
Our study had several potential clinical applications; specifically, our identified miRNA signature may serve as a novel biomarker for prognostic response and prediction towards existing adjuvant treatments for LGGs. Although maximal safe resection combined with adjuvant radiotherapy and chemotherapy is the recommended treatment for patients with LGG and any poor prognostic feature [31], the clinical outcome remains extremely variable with the same treatment regimen. Therefore, this finding indicates that efforts should be focused on the identification of subgroups of patients who are more likely to benefit from adjuvant treatments and thus enables greater personalized care. Houillier et al. demonstrated that patients with IDH-mutated gliomas have a significantly increased response to the oral alkylating agent temozolomide [32]. Baumert et al. [33] performed an EORTC clinical trial and concluded that patients with IDH1/IDH2 mutations and 1p/19q non-codeletion benefitted from radiotherapy more than chemotherapy. Despite the importance of these findings, integrating miRNA data as part of a molecular-based stratification approach may provide new clues in the identification of candidate therapeutic targets in patients with LGGs or enable the provision of better personalized medicine for certain patient subgroups. In a previous study, several miRNAs were examined as potential biomarkers in response to temozolomide treatment in patients with glioblastoma [11]. Therefore, suitable therapies combining more tolerable targeted drugs as an adjunct could be selected for patients with different risk scores, which will improve the efficacy of systemic control.
We identified a few limitations to our study. First, although our study has a large sample size containing independent training and validation sets to robustly demonstrate the putative prognostic value of our miRNA-signature in patients with LGGs, prospective studies are warranted to more fully assess whether these miRNAs are clinically valuable in patient prognosis. Second, the inclusion of additional known variables implicated in LGGs, such as the Karnofsky Performance Score, tumor location, neuroimaging data, and other genomic characteristics are needed to better determine the prognostic contribution of our miRNA signature with existing measures. Third, the absence of a 1p/19q codeletion study in the training cohort is also a limitation of this study. Fourth, our study was retrospective in nature, and therefore, we cannot exclude the possibility that variable treatment regimens with surgery, radiation, and chemotherapy may have had confounding effects on survival outcomes in the patients in our cohorts. Thus, the predictive value of this newly discovered miRNA signature requires further characterization and validation in a separate dataset with a uniform treatment regimen.
In conclusion, we identified and reliably validated a four-miRNA signature associated with OS in two cohorts of patients with LGGs and developed a miRNA-based nomogram for favorable individual prognostic assessment. Further understanding of the biological processes that underlie these miRNAs will provide new insights into the pathogenesis of LGGs and will be an important step towards improved decision making in personalized clinical management of patients with gliomas.
References
Cancer Genome Atlas Research Network, Brat DJ, Verhaak RG, Aldape KD, Yung WK, Salama SR, Cooper LA, Rheinbay E, Miller CR, Vitucci M, Morozova O, Robertson AG, Noushmehr H, Laird PW, Cherniack AD, Akbani R, Huse JT, Ciriello G, Poisson LM, Barnholtz-Sloan JS, Berger MS, Brennan C, Colen RR, Colman H, Flanders AE, Giannini C, Grifford M, Iavarone A, Jain R, Joseph I, Kim J, Kasaian K, Mikkelsen T, Murray BA, O’Neill BP, Pachter L, Parsons DW, Sougnez C, Sulman EP, Vandenberg SR, Van Meir EG, von Deimling A, Zhang H, Crain D, Lau K, Mallery D, Morris S, Paulauskis J, Penny R, Shelton T, Sherman M, Yena P, Black A, Bowen J, Dicostanzo K, Gastier-Foster J, Leraas KM, Lichtenberg TM, Pierson CR, Ramirez NC, Taylor C, Weaver S, Wise L, Zmuda E, Davidsen T, Demchok JA, Eley G, Ferguson ML, Hutter CM, Mills Shaw KR, Ozenberger BA, Sheth M, Sofia HJ, Tarnuzzer R, Wang Z, Yang L, Zenklusen JC, Ayala B, Baboud J, Chudamani S, Jensen MA, Liu J, Pihl T, Raman R, Wan Y, Wu Y, Ally A, Auman JT, Balasundaram M, Balu S, Baylin SB, Beroukhim R, Bootwalla MS, Bowlby R, Bristow CA, Brooks D, Butterfield Y, Carlsen R, Carter S, Chin L, Chu A, Chuah E, Cibulskis K, Clarke A, Coetzee SG, Dhalla N, Fennell T, Fisher S, Gabriel S, Getz G, Gibbs R, Guin R, Hadjipanayis A, Hayes DN, Hinoue T, Hoadley K, Holt RA, Hoyle AP, Jefferys SR, Jones S, Jones CD, Kucherlapati R, Lai PH, Lander E, Lee S, Lichtenstein L, Ma Y, Maglinte DT, Mahadeshwar HS, Marra MA, Mayo M, Meng S, Meyerson ML, Mieczkowski PA, Moore RA, Mose LE, Mungall AJ, Pantazi A, Parfenov M, Park PJ, Parker JS, Perou CM, Protopopov A, Ren X, Roach J, Sabedot TS, Schein J, Schumacher SE, Seidman JG, Seth S, Shen H, Simons JV, Sipahimalani P, Soloway MG, Song X, Sun H, Tabak B, Tam A, Tan D, Tang J, Thiessen N, Triche T, Jr., Van Den Berg DJ, Veluvolu U, Waring S, Weisenberger DJ, Wilkerson MD, Wong T, Wu J, Xi L, Xu AW, Yang L, Zack TI, Zhang J, Aksoy BA, Arachchi H, Benz C, Bernard B, Carlin D, Cho J, DiCara D, Frazer S, Fuller GN, Gao J, Gehlenborg N, Haussler D, Heiman DI, Iype L, Jacobsen A, Ju Z, Katzman S, Kim H, Knijnenburg T, Kreisberg RB, Lawrence MS, Lee W, Leinonen K, Lin P, Ling S, Liu W, Liu Y, Liu Y, Lu Y, Mills G, Ng S, Noble MS, Paull E, Rao A, Reynolds S, Saksena G, Sanborn Z, Sander C, Schultz N, Senbabaoglu Y, Shen R, Shmulevich I, Sinha R, Stuart J, Sumer SO, Sun Y, Tasman N, Taylor BS, Voet D, Weinhold N, Weinstein JN, Yang D, Yoshihara K, Zheng S, Zhang W, Zou L, Abel T, Sadeghi S, Cohen ML, Eschbacher J, Hattab EM, Raghunathan A, Schniederjan MJ, Aziz D, Barnett G, Barrett W, Bigner DD, Boice L, Brewer C, Calatozzolo C, Campos B, Carlotti CG, Jr., Chan TA, Cuppini L, Curley E, Cuzzubbo S, Devine K, DiMeco F, Duell R, Elder JB, Fehrenbach A, Finocchiaro G, Friedman W, Fulop J, Gardner J, Hermes B, Herold-Mende C, Jungk C, Kendler A, Lehman NL, Lipp E, Liu O, Mandt R, McGraw M, McLendon R, McPherson C, Neder L, Nguyen P, Noss A, Nunziata R, Ostrom QT, Palmer C, Perin A, Pollo B, Potapov A, Potapova O, Rathmell WK, Rotin D, Scarpace L, Schilero C, Senecal K, Shimmel K, Shurkhay V, Sifri S, Singh R, Sloan AE, Smolenski K, Staugaitis SM, Steele R, Thorne L, Tirapelli DP, Unterberg A, Vallurupalli M, Wang Y, Warnick R, Williams F, Wolinsky Y, Bell S, Rosenberg M, Stewart C, Huang F, Grimsby JL, Radenbaugh AJ, Zhang J (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372:2481–2498. https://doi.org/10.1056/NEJMoa1402121
Ceccarelli M, Barthel FP, Malta TM, Sabedot TS, Salama SR, Murray BA, Morozova O, Newton Y, Radenbaugh A, Pagnotta SM, Anjum S, Wang J, Manyam G, Zoppoli P, Ling S, Rao AA, Grifford M, Cherniack AD, Zhang H, Poisson L, Carlotti CG Jr, Tirapelli DP, Rao A, Mikkelsen T, Lau CC, Yung WK, Rabadan R, Huse J, Brat DJ, Lehman NL, Barnholtz-Sloan JS, Zheng S, Hess K, Rao G, Meyerson M, Beroukhim R, Cooper L, Akbani R, Wrensch M, Haussler D, Aldape KD, Laird PW, Gutmann DH, TCGA Research Network, Noushmehr H, Iavarone A, Verhaak RG (2016) Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164:550–563. https://doi.org/10.1016/j.cell.2015.12.028
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820. https://doi.org/10.1007/s00401-016-1545-1
Weller M, Weber RG, Willscher E, Riehmer V, Hentschel B, Kreuz M, Felsberg J, Beyer U, Loffler-Wirth H, Kaulich K, Steinbach JP, Hartmann C, Gramatzki D, Schramm J, Westphal M, Schackert G, Simon M, Martens T, Bostrom J, Hagel C, Sabel M, Krex D, Tonn JC, Wick W, Noell S, Schlegel U, Radlwimmer B, Pietsch T, Loeffler M, von Deimling A, Binder H, Reifenberger G (2015) Molecular classification of diffuse cerebral WHO grade II/III gliomas using genome- and transcriptome-wide profiling improves stratification of prognostically distinct patient groups. Acta Neuropathol 129:679–693. https://doi.org/10.1007/s00401-015-1409-0
Suzuki H, Aoki K, Chiba K, Sato Y, Shiozawa Y, Shiraishi Y, Shimamura T, Niida A, Motomura K, Ohka F, Yamamoto T, Tanahashi K, Ranjit M, Wakabayashi T, Yoshizato T, Kataoka K, Yoshida K, Nagata Y, Sato-Otsubo A, Tanaka H, Sanada M, Kondo Y, Nakamura H, Mizoguchi M, Abe T, Muragaki Y, Watanabe R, Ito I, Miyano S, Natsume A, Ogawa S (2015) Mutational landscape and clonal architecture in grade II and III gliomas. Nat Genet 47:458–468. https://doi.org/10.1038/ng.3273
Liu N, Chen NY, Cui RX, Li WF, Li Y, Wei RR, Zhang MY, Sun Y, Huang BJ, Chen M, He QM, Jiang N, Chen L, Cho WC, Yun JP, Zeng J, Liu LZ, Li L, Guo Y, Wang HY, Ma J (2012) Prognostic value of a microRNA signature in nasopharyngeal carcinoma: a microRNA expression analysis. Lancet Oncol 13:633–641. https://doi.org/10.1016/S1470-2045(12)70102-X
Zhang JX, Song W, Chen ZH, Wei JH, Liao YJ, Lei J, Hu M, Chen GZ, Liao B, Lu J, Zhao HW, Chen W, He YL, Wang HY, Xie D, Luo JH (2013) Prognostic and predictive value of a microRNA signature in stage II colon cancer: a microRNA expression analysis. Lancet Oncol 14:1295–1306. https://doi.org/10.1016/S1470-2045(13)70491-1
Pang JC, Kwok WK, Chen Z, Ng HK (2009) Oncogenic role of microRNAs in brain tumors. Acta Neuropathol 117:599–611. https://doi.org/10.1007/s00401-009-0525-0
Kouri FM, Ritner C, Stegh AH (2015) miRNA-182 and the regulation of the glioblastoma phenotype - toward miRNA-based precision therapeutics. Cell Cycle 14:3794–3800. https://doi.org/10.1080/15384101.2015.1093711
Ames HM, Yuan M, Vizcaino MA, Yu W, Rodriguez FJ (2017) MicroRNA profiling of low-grade glial and glioneuronal tumors shows an independent role for cluster 14q32.31 member miR-487b. Mod Pathol 30:204–216. https://doi.org/10.1038/modpathol.2016.177
Zhang W, Zhang J, Yan W, You G, Bao Z, Li S, Kang C, Jiang C, You Y, Zhang Y, Chen CC, Song SW, Jiang T (2013) Whole-genome microRNA expression profiling identifies a 5-microRNA signature as a prognostic biomarker in Chinese patients with primary glioblastoma multiforme. Cancer 119:814–824. https://doi.org/10.1002/cncr.27826
Bao ZS, Chen HM, Yang MY, Zhang CB, Yu K, Ye WL, Hu BQ, Yan W, Zhang W, Akers J, Ramakrishnan V, Li J, Carter B, Liu YW, Hu HM, Wang Z, Li MY, Yao K, Qiu XG, Kang CS, You YP, Fan XL, Song WS, Li RQ, Su XD, Chen CC, Jiang T (2014) RNA-seq of 272 gliomas revealed a novel, recurrent PTPRZ1-MET fusion transcript in secondary glioblastomas. Genome Res 24:1765–1773. https://doi.org/10.1101/gr.165126.113
Yang P, Cai J, Yan W, Zhang W, Wang Y, Chen B, Li G, Li S, Wu C, Yao K, Li W, Peng X, You Y, Chen L, Jiang C, Qiu X, Jiang T, CGGA Project (2016) Classification based on mutations of TERT promoter and IDH characterizes subtypes in grade II/III gliomas. Neuro-oncology. https://doi.org/10.1093/neuonc/now021
Feng LH, Dong H, Lau WY, Yu H, Zhu YY, Zhao Y, Lin YX, Chen J, Wu MC, Cong WM (2017) Novel microvascular invasion-based prognostic nomograms to predict survival outcomes in patients after R0 resection for hepatocellular carcinoma. J Cancer Res Clin Oncol 143:293–303. https://doi.org/10.1007/s00432-016-2286-1
Li J, Zhou J, Yang PH, Xia Y, Shi YH, Wu D, Lv G, Zheng W, Wang K, Wan XY, Lau WY, Wu MC, Fan J, Shen F (2016) Nomograms for survival prediction in patients undergoing liver resection for hepatitis B virus related early stage hepatocellular carcinoma. Eur J Cancer 62:86–95. https://doi.org/10.1016/j.ejca.2016.04.011
Wang Y, Li J, Xia Y, Gong R, Wang K, Yan Z, Wan X, Liu G, Wu D, Shi L, Lau W, Wu M, Shen F (2013) Prognostic nomogram for intrahepatic cholangiocarcinoma after partial hepatectomy. J Clin Oncol 31:1188–1195. https://doi.org/10.1200/JCO.2012.41.5984
Eckel-Passow JE, Lachance DH, Molinaro AM, Walsh KM, Decker PA, Sicotte H, Pekmezci M, Rice T, Kosel ML, Smirnov IV, Sarkar G, Caron AA, Kollmeyer TM, Praska CE, Chada AR, Halder C, Hansen HM, McCoy LS, Bracci PM, Marshall R, Zheng S, Reis GF, Pico AR, O’Neill BP, Buckner JC, Giannini C, Huse JT, Perry A, Tihan T, Berger MS, Chang SM, Prados MD, Wiemels J, Wiencke JK, Wrensch MR, Jenkins RB (2015) Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N Engl J Med 372:2499–2508. https://doi.org/10.1056/NEJMoa1407279
Johnson BE, Mazor T, Hong C, Barnes M, Aihara K, McLean CY, Fouse SD, Yamamoto S, Ueda H, Tatsuno K, Asthana S, Jalbert LE, Nelson SJ, Bollen AW, Gustafson WC, Charron E, Weiss WA, Smirnov IV, Song JS, Olshen AB, Cha S, Zhao Y, Moore RA, Mungall AJ, Jones SJ, Hirst M, Marra MA, Saito N, Aburatani H, Mukasa A, Berger MS, Chang SM, Taylor BS, Costello JF (2014) Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343:189–193. https://doi.org/10.1126/science.1239947
Visani M, de Biase D, Marucci G, Cerasoli S, Nigrisoli E, Bacchi Reggiani ML, Albani F, Baruzzi A, Pession A, PERNO Study Group (2014) Expression of 19 microRNAs in glioblastoma and comparison with other brain neoplasia of grades I-III. Mol Oncol 8:417–430 https://doi.org/10.1016/j.molonc.2013.12.010
Ji Y, Wei Y, Wang J, Gong K, Zhang Y, Zuo H (2015) Correlation of microRNA-10b upregulation and poor prognosis in human gliomas. Tumour Biol 36:6249–6254. https://doi.org/10.1007/s13277-015-3310-9
Gabriely G, Yi M, Narayan RS, Niers JM, Wurdinger T, Imitola J, Ligon KL, Kesari S, Esau C, Stephens RM, Tannous BA, Krichevsky AM (2011) Human glioma growth is controlled by microRNA-10b. Cancer Res 71:3563–3572. https://doi.org/10.1158/0008-5472.CAN-10-3568
El Fatimy R, Subramanian S, Uhlmann EJ, Krichevsky AM (2017) Genome editing reveals glioblastoma addiction to microRNA-10b. Mol Ther 25:368–378. https://doi.org/10.1016/j.ymthe.2016.11.004
Guessous F, Alvarado-Velez M, Marcinkiewicz L, Zhang Y, Kim J, Heister S, Kefas B, Godlewski J, Schiff D, Purow B, Abounader R (2013) Oncogenic effects of miR-10b in glioblastoma stem cells. J Neuro-oncol 112:153–163. https://doi.org/10.1007/s11060-013-1047-0
Cheng W, Ren X, Zhang C, Han S, Wu A (2017) Expression and prognostic value of microRNAs in lower-grade glioma depends on IDH1/2 status. J Neuro-oncol 132:207–218. https://doi.org/10.1007/s11060-016-2368-6
Guan Y, Mizoguchi M, Yoshimoto K, Hata N, Shono T, Suzuki SO, Araki Y, Kuga D, Nakamizo A, Amano T, Ma X, Hayashi K, Sasaki T (2010) MiRNA-196 is upregulated in glioblastoma but not in anaplastic astrocytoma and has prognostic significance. Clin Cancer Res 16:4289–4297. https://doi.org/10.1158/1078-0432.CCR-10-0207
Yang G, Han D, Chen X, Zhang D, Wang L, Shi C, Zhang W, Li C, Chen X, Liu H, Zhang D, Kang J, Peng F, Liu Z, Qi J, Gao X, Ai J, Shi C, Zhao S (2014) MiR-196a exerts its oncogenic effect in glioblastoma multiforme by inhibition of IkappaBalpha both in vitro and in vivo. Neuro-oncology 16:652–661. https://doi.org/10.1093/neuonc/not307
Flavahan WA, Drier Y, Liau BB, Gillespie SM, Venteicher AS, Stemmer-Rachamimov AO, Suva ML, Bernstein BE (2016) Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529:110–114. https://doi.org/10.1038/nature16490
Pignatti F, van den Bent M, Curran D, Debruyne C, Sylvester R, Therasse P, Afra D, Cornu P, Bolla M, Vecht C, Karim AB, European Organization for Research, Treatment of Cancer Brain Tumor Cooperative Group, European Organization for Research, Treatment of Cancer Radiotherapy Cooperative Group (2002) Prognostic factors for survival in adult patients with cerebral low-grade glioma. J Clin Oncol 20:2076–2084. https://doi.org/10.1200/JCO.2002.08.121
Schiff D, Brown PD, Giannini C (2007) Outcome in adult low-grade glioma: the impact of prognostic factors and treatment. Neurology 69:1366–1373. https://doi.org/10.1212/01.wnl.0000277271.47601.a1
Forst DA, Nahed BV, Loeffler JS, Batchelor TT (2014) Low-grade gliomas. Oncologist 19:403–413. https://doi.org/10.1634/theoncologist.2013-0345
Buckner JC, Shaw EG, Pugh SL, Chakravarti A, Gilbert MR, Barger GR, Coons S, Ricci P, Bullard D, Brown PD, Stelzer K, Brachman D, Suh JH, Schultz CJ, Bahary JP, Fisher BJ, Kim H, Murtha AD, Bell EH, Won M, Mehta MP, Curran WJ Jr (2016) Radiation plus procarbazine, CCNU, and vincristine in low-grade glioma. N Engl J Med 374:1344–1355. https://doi.org/10.1056/NEJMoa1500925
Houillier C, Wang X, Kaloshi G, Mokhtari K, Guillevin R, Laffaire J, Paris S, Boisselier B, Idbaih A, Laigle-Donadey F, Hoang-Xuan K, Sanson M, Delattre JY (2010) IDH1 or IDH2 mutations predict longer survival and response to temozolomide in low-grade gliomas. Neurology 75:1560–1566. https://doi.org/10.1212/WNL.0b013e3181f96282
Baumert BG, Hegi ME, van den Bent MJ, von Deimling A, Gorlia T, Hoang-Xuan K, Brandes AA, Kantor G, Taphoorn MJ, Hassel MB, Hartmann C, Ryan G, Capper D, Kros JM, Kurscheid S, Wick W, Enting R, Reni M, Thiessen B, Dhermain F, Bromberg JE, Feuvret L, Reijneveld JC, Chinot O, Gijtenbeek JM, Rossiter JP, Dif N, Balana C, Bravo-Marques J, Clement PM, Marosi C, Tzuk-Shina T, Nordal RA, Rees J, Lacombe D, Mason WP, Stupp R (2016) Temozolomide chemotherapy versus radiotherapy in high-risk low-grade glioma (EORTC 22033–26033): a randomised, open-label, phase 3 intergroup study. Lancet Oncol 17:1521–1532. https://doi.org/10.1016/S1470-2045(16)30313-8
Acknowledgements
The authors conducting this work represent the Chinese Glioma Cooperative Group (CGCG).
Funding
This work was supported by Beijing Postdoctoral Research Foundation (2016ZZ-37).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they haves no competing interests associated with this work.
Ethical approval
The glioma tissue collection was approved by the Institutional Review Board of Beijing Tiantan Hospital affiliated to Capital Medical University.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11060_2017_2704_MOESM1_ESM.tif
Fig S1. The prognostic value of the four-miRNA signature in the three molecular subgroups of LGGs. (A) Scatter plot demonstrating that the risk score was distributed differently in the three molecular subgroups. The signature enabled the segmentation of patients into high- and low-risk groups in the (B) IDHMUT +1p/19q codel, (C) IDHMUT +1p/19q non-codel, and (D) IDHWT subgroups (TIF 1460 KB)
11060_2017_2704_MOESM2_ESM.tif
Fig S2. Figure showing significant functions and pathway enrichment analysis of the four-miRNA signature in the validation dataset. (A) Heat map of differentially expressed genes between high- and low-risk four-miRNA signature groups from 420 LGG samples. (B) Network analysis and (C) gene ontology analysis showing that high-risk was associated with specific biological processes. (D) Pathway enrichment analysis showing that the high-risk group was associated with several KEGG pathways, including cell cycle and DNA replication (TIF 57539 KB)
Rights and permissions
About this article
Cite this article
Qian, Z., Li, Y., Fan, X. et al. Prognostic value of a microRNA signature as a novel biomarker in patients with lower-grade gliomas. J Neurooncol 137, 127–137 (2018). https://doi.org/10.1007/s11060-017-2704-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-017-2704-5