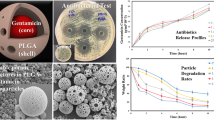
Abstract
Several strategies for the delivery and release of drugs have been studied, among them the use of polymeric nanoparticles of PLGA (poly lactic-co-glycolic acid). These nanoparticles (NPs) have been shown to be promising for the controlled and effective release of drugs due to their biodegradability and biocompatibility. Regarding this, the aim of this study is to synthesize and characterize PLGA polymer nanoparticles associated with antimicrobials in order to reduce adverse reactions and to have a more effective local delivery. Empty PLGA nanoparticles and conjugated to vancomycin and meropenem antibiotics were synthesized by the double emulsification-solvent evaporation technique. Then, they were characterized by the analysis of the mean particle diameter, dynamic light scattering (DLS), Fourier transform infrared (FTIR) vibrational spectroscopy, contact angle measurement, and atomic force microscopy (AFM) and scanning electron microscopy (SEM). The DLS analysis of the NPs obtained showed approximate sizes of 263.5 nm (NPs-PLGA), 239.3 nm (NPs-VAN) and 284.2 nm (NPs-MER), and monodispersivity. FTIR results and contact angle measurements suggest encapsulation of antibiotics to NPs. The morphology evaluated through AFM and SEM indicates homogeneous, uniform, and spherical distribution of NPs and also apparently smooth. The antibacterial action of PLGA nanoparticles carried with antibiotics was effective when using concentrations of 5–2.5 mg/mL (PLGA-VAN) versus Staphylococcus aureus and 10–2.5 mg/mL (PLGA-MER) to Pseudomonas aeruginosa. The release kinetics of the antibiotics revealed a release profile of 43.9% at the end of 24 h and 96% at the end of 96 h for PLGA-VAN formulation and 8.25% and 16% at the end of 24 h and 96 h, for PLGA-MER, respectively. This study supports the potential application of PLGA particles containing antibiotics as facilitators of drug delivery and release effectively.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introdução
Poly lactic-co-glycolic acid (PLGA) is widely used for the development of nanoparticles (NPs) because of their biocompatibility and biodegradability and has been approved as a component in various drug delivery formulations (Kim et al. 2014; Danhier et al. 2012). The PLGA nanoparticles provide high versatility when used in the transport of therapeutic compounds, with effective and controlled release at the site of action (Duncan and Gaspar 2011), which improve the antimicrobial potential (SILVA et al. 2014), besides promoting focalization of the active compound at the target site and leading to reduction of the side effects, which makes this copolymer a viable option in current chemotherapy. A number of hydrophilic and hydrophobic antimicrobials are administered using NPs as carriers (Günday Türeli et al. 2017; Parisi et al. 2017; Venkatesh et al. 2015), which have presented satisfactory results against gram-positive and -negative microorganisms. Such formulations are based on the dispersion of preformed polymers and polymerization from polymer monomers and ionic gelling (Mohanraj and Chen 2006). The morphology, diameter, and polydispersity index (PDI) of NPs are features that directly affect biodistribution in vivo (Zaidi et al. 2017). Also, nanoparticles with diameters of up to 200 nm are considered to be more effective because they are not immediately detected and opsonized by macrophages, as well as promoting efficient retention and penetration into injured tissues (Yameen et al. 2014). In addition, antimicrobial-loaded NPs can enter the cells through endocytosis, releasing the drug to counteract intracellular infections caused by microorganisms.
Bacterial infections have generated major problems in public health. Despite of wide availability of antibiotics, new difficulties in the treatment of infections have arisen, mainly due to the emergence of increasingly resistant microorganisms (World Health Organization (WHO) 2017; Parisi et al. 2017). Thus, formulations of nanoparticles with antibiotics may help to combat new resistance mechanisms, increasing the effectiveness of the antibiotic and reducing the adverse effects of the drugs. The World Health Organization (WHO) (2017) classified some bacterial species as priority for the development of new forms of treatment, among them: Staphylococcus aureus and Pseudomonas aeruginosa. Therefore, the present study reports the development of antimicrobial-loaded NPs in two formulations, one containing vancomycin and the other with meropenem, in order to develop and characterize PLGA polymer nanoparticles containing antimicrobials for the future treatment of biofilms produced by pathogenic microorganisms, since increasing resistance to antibiotics added to biofilm formation is a worldwide challenge.
Material and methods
Obtaining and yielding of nanoparticles
PLGA nanoparticles were obtained by double emulsification/solvent evaporation (Jelvehgari et al. 2010). A solution A (organic phase) was prepared with 0.1 g of 50:50 PLGA (Purasorb® PDLG 5002) and 4 mL of dichloromethane (DCM) (Sigma-Aldrich®, St. Louis, USA). Additionally, solutions with the antibiotics vancomycin (VAN) and meropenem (MER) (5 mg/mL) were added to this organic phase by dripping during ice bath production for 5 min. A solution B (aqueous phase) composed of 0.04 g of polyvinyl alcohol (PVA) (Mw 9000–10,000, 80% hydrolyzed, Sigma-Aldrich®, St. Louis, USA) and 20 mL of ultrapure Milli-Q water heated to 65 °C, placed under magnetic stirring for total dissolution of surfactant. Then, solution A was added to solution B by dripping with the aid of a microsyringe. A final solution obtained was emulsified in an ice bath sonicator (37 Khz, 240 W-S10H-Elma ®) for 20 min and subjected to magnetic stirring for 18 h to evaporate the solvent. Subsequently, the material was centrifuged for 15 min at 5 °C at 234×g to remove impurities, and its supernatant was collected and again centrifuged for 20 min at 5 °C at 29,756×g to be deposited as nanoparticles. After obtaining, the NPs were lyophilized. The yield of the nanoparticles was calculated from the following equation:
*ATB-antibiotic.
Analysis of mean hydrodynamic diameter and polydispersity index
The mean hydrodynamic diameter was determined by dynamic light scattering (DLS) using Delsa™ Nano C Particle Analyzer (Beckman Coulter, Brea, CA, EUA.). The nanoparticles were resuspended in 1 mL of ultrapure water at a concentration of 10 mg/mL and submitted to ultrasound for 5 min (Hill et al. 2013). The analysis was repeated on days 15, 30, 45, 60 for stability check.
Fourier transform infrared vibrational spectroscopy
The chemical groups of the antibiotics, vancomycin (VAN), and meropenem (MER) were determined using the vibrations of the functional groups obtained by Fourier-transform infrared (FTIR) spectroscopy (Shimadzu IRAffinity-1, Columbia, MA) with a resolution of 2 cm−1. Measurement of the absorption spectra of the samples was performed in the infrared region (4000 to 400 cm−1) using KBr.
Contact angle measurement
Glass substrates (1 × 1-cm glass cover) containing the samples were submitted for contact angle measurements (Teclis Tracker, IT Concept) to verify surface modification. The measurements were performed at five random points of each substrate prepared in this work. Then, 5 μL distilled and deionized water was dripped onto the substrate and the contact angle between the water and the surface of the substrate was determined.
Microscopy
Atomic force microscopy (AFM) and scanning electron microscopy (SEM) performed morphological analysis, diameter, and distribution. In this last technique, the scanning electron microscope with STEM (scanning transmission electron microscopy) double beam detector was used (FEI Helios 660 FEG). The samples were prepared by dispersing the solution in copper-coated water of the holey carbon type. AFM analysis was performed according to the methodology of Georgiev et al. (2013), where the material was dried for 12 h in the absence of light. Topographic measurements were performed in dynamic mode with AFM (SPM-9700 Shimadzu, Kyoto, Japan). A cantilever having a radius of curvature of less than 10 nm was used and scans were performed at 150 Hz frequency and a spring constant of 5 N/m provided by Budget Sensors.
Efficiency of antibiotic encapsulation
The efficiency of antibiotic encapsulation (VAN and MER) was performed by indirect method, which relates to the amount of free drug after the synthesis of NPs, containing the antibiotics, with the amount of drug added at the beginning of the process. The quantification of each compound was determined spectrophotometrically 280 nm (VAN) and 300 nm (MER) (Shimadzu UV-1601 spectrophotometer, Columbia, MA) and compared with the calibration curve. The efficiency and encapsulation were calculated according to the equation below (Moreira Gomes et al. 2011).
Antimicrobial evaluation of Nps
The antimicrobial evaluation of NPs was performed using the following bacterial strains: hospital-associated (HA) MRSA Staphylococcus aureus and multidrug-resistant hospital-associated (MDR-HA) Pseudomonas aeruginosa, S. aureus ATCC 43300, and P. aeruginosa ATCC 27853, through analysis of the minimum inhibitory concentration (MIC) by the micro-dilution method in broth, according to the protocol of the Clinical and Laboratory Standards Institute (CLSI 2017). After primary incubation of the microorganisms in blood agar at 37 °C for 24 h, bacterial suspension was performed in saline solution equivalent to McFarland’s Scale 0.5 (approximately 1 × 108 CFU/mL), followed by dilution of 1:20 in saline solution. In a 96-well polyethylene micro-plate, 100 μL of concentrated Muxer Hinton Agar 2× was added. Subsequently, 100 μL of the NPs containing the antibiotics was inserted, followed by serial dilution in the subsequent wells. Ten microliters of the inoculum suspension was added and incubated at 35 ± 2 °C for 24 h. To read the results, 20 μL of 0.5% triphenyl tetrazolium chloride (TTC) was added, with reincubation of the plates in a bacteriological oven at 35 ± 2 °C for 2 h to show the wells with bacterial growth, marked in red. To determine the minimum bactericidal concentration (MBC), 20 μL of the non-growth wells was withdrawn for inoculation in Muller Hinton agar médium and followed by incubation for 24 h at 35 °C.
Kinetics of antibiotic release
Release kinetics were performed according to Nandakumar et al. (2013) with adaptations, using dialysis bags with about 25 cm2 of area. NPs of 10 mg/mL were placed into a dialysis bag, previously hydrated and properly closed. The material was dipped into a falcon tube containing 30-mL phosphate-buffered saline (PBS), pH 7.4, at a temperature of 37 °C and shaking at 150 rpm. Aliquots of 3 mL were collected at time intervals of 0 h, 2 h, 24 h, 48 h, 72 h, and 96 h with medium replacement. Then, the quantificaton was performed spectrophotometrically at 280 nm (VAN) and 305 nm (MER) (Shimadzu UV-1601 spectrophotometer, Columbia, MA). From the obtained concentrations, the graph with the release profile of the NPs was elaborated, calculating the release kinetics, through the more linear range of the presented profile.
Statistics
The results presented were expressed as mean ± standard deviation (SD) in triplicate. Significant differences were calculated using Wilcoxon and the differences between the groups were used Mann-Whitney test. Values of p < 0.05 were considered significant.
Results and discussion
Obtaining, yielding, and characterization of NPs
The synthesis of the nanoparticles was evidenced by the analysis through DLS, which showed NPs with DHM between 263.5 ± 2.6 nm (NPs PLGA), 239.3 ± 1.53 nm (NPs PLGA-VAN) e 284.2 ± 1.85 nm (NPs PLGA-MER) and PDI of 0.1 suggesting a monodispersivity of the particles obtained. The yield analysis and encapsulation efficiency showed an income of 78.3 to 90.2% and 28.3 to 74.79% (Table 1) encapsulation. Regardless of the encapsulation efficiency, the formulations obtained presented satisfactory DHM and PDI when compared to the literature. Posadowska et al. (2015) obtained NPs encapsulated with gentamicin, ranging in size from 307 ± 8 to 391 ± 23 nm. Studies using vancomycin and meropenem carried to PLGA demonstrated variable encapsulation results between 12.7 and 78.6% vancomycin (Lotfipour et al. 2013; Zakeri-Milani et al. 2013) and 82% for meropenem (Nandakumar et al. 2013). However, several variables are responsible for a difference in EE%, such as structural differences of the molecules and chemical interaction with the polymer.
Studies report a relationship between the hydrophilic feature of the substance to be encapsulated by the emulsion/evaporation method with the EE% rate, and there may be a decrease of this encapsulation (Rao and Geckeler 2011). The stability of the formulations was evaluated at 0, 15, 30, 45, and 60 days, using dynamic light scattering (DLS) and PDI, with no significant changes (p < 0.05) (Fig. 1).
The presence of the antibiotics in the NPs was evaluated by Fourier transform infrared (FTIR) vibrational spectroscopy and by contact angle measurements. The FTIR spectra of PLGA, vancomycin, and meropenem as well as NPs-MER and NPs-VAN are shown in Fig. 2a, b and represent spectra obtained from the IR studies, which correspond to 4000 cm to 500 cm−1, respectively. Regarding vancomycin, the absorption peak at 3220.42 cm−1 is attributed to the hydroxyl terminal group and 1636.30 cm−1 is due to the stretching of the C=O bond. The band at 1224.43 cm−1 is related to the asymmetric COC elongation. Other bands characteristic of the vancomycin spectrum appear at 1576.66, 1489.07–1399.62, 1142.43–1054.84 cm−1 corresponding to C=C, C–O–C, and C–N–H vibrations, respectively. The vancomycin CH2 vibration band is present at 1483.48 cm−1. While the FTIR analysis of the meropenem shows stretching bands of the N–H group at 3554.02 cm−1, stretching bands of the O–H group at 3367.65 cm−1, stretching bands of the methyl group C–H at 2959.51 cm−1, 2929.69 cm−1, and 2940.84 cm−1, elongation CO in COOH and pyrrolidine ring at 1735.08 cm−1, stretching bands of C–O of the hydroxyethyl group at 1140.57 cm−1 and flexion OH in COOH at 663.47 cm−1 (Cielecka-Piontek et al. 2013; Abdelkader et al. 2017).
The poly lactic-co-glycolic acid has bands corresponding to the stretching of the aliphatic CH (2918.50 cm−1), the CO ester (1746.44 cm−1), the carbonyl CO stretch (827.47 cm−1) and to the (C–O) ether group was observed in 1082.79 cm−1, (Nandakumar et al. 2013; Javed et al. 2015), which was also evidenced in figure. The presence of vancomycin is confirmed by stretching of CH (2922.23 cm−1), CONH (1744.39 cm−1), OH flexion in COOH (846.10 cm−1), and stretching of the CO of the hydroxyethyl group (1082.79, 1166.67 cm−1). The presence of meropenem in PLGA is confirmed by the presence of stretching of CH (2983.73 cm−1), CONH 1742.53, OH flexion in COOH (836.78 cm−1) and stretching of the C–O of the hydroxyethyl group (1082.79 and 1168.52 cm−1) suggesting encapsulation in the NPs. These results converge with those presented by the contact angle measurements of the formulations, where PLGA-VAN (38.9°) and PLGA-MER (42.7°) reduced when compared to the empty PLGA formulation (57.6°), suggesting the presence of the antibiotics in the NPs, due to the more hydrophilic behavior of the encapsulated antibiotics.
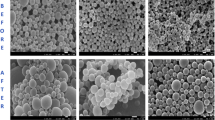
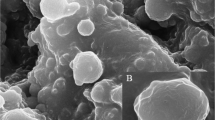
Microscopic analysis (SEM and AFM) (Figs. 3, 4, and 5) indicates homogeneous, uniform and spherical, and apparently smooth distribution of NPs, with sizes presented for SEM at 105 ± 13.5 nm (PLGA), 81.9 ± 21,5 nm (PLGA-VAN), and 172.9 ± 39.2 nm (PLGA-MER), presenting difference when compared with results obtained through DLS. This fact is related to the presence of aggregates of particles in the suspension measured by DLS, occurring the reading of larger particles, being the average diameter measured in the DLS greater in comparison with microscopic techniques (Tomaszewska et al. 2013). The findings in AFM do not show morphological differences and surface area between the formulations obtained. Additionally, the diameters and morphology are considered satisfactory for use; therefore, they will probably not be detected and opsonized by cells, providing a greater deposit and penetration in the injured tissues (Yameen et al. 2014).
Evaluation of the antimicrobial activity of NPs
The evaluation of the antimicrobial activity of the NPs carried with antibiotics in comparison to the free drugs was performed by MIC (Table 2). The strains of S. aureus MRSA and P. aeruginosa presented resistance profile to the antibiotic models (vancomycin and meropenem) used in their free form (CLSI 2017). Differently from the strains S. aureus ATCC 43300 (resistant to methicillin) and P. aeruginosa ATCC 27853, which were considered sensitive to the respective antibiotic models, as predicted. In relation to the action of PLGA NPs carried with antibiotics, an effective action was observed when using concentrations 5–2.5 mg/mL (PLGA-VAN) against S. aureus MRSA and 10–2.5 mg/mL (PLGA-MER) against P. aeruginosa.
The results of the MICs against the strains tested with antibiotics carried to PLGA NPs were higher when compared to the free antibiotics. This fact may be related to the later release of the antibiotics in the medium, allowing an initial growth of the microorganisms.
Although MICs of NP PLGA-VAN and PLGA-MER have shown higher values in vitro, controlled and sustained release generates advantages such as therapeutic efficacy, decreased systemic side effects, and reduced frequency of drug administration. Other studies have also demonstrated MICs for drugs carried at higher NPs when compared to drugs in their free form (Sabaeifard et al. 2016; Lotfipour et al. 2014). The MIC was only tested with pure PLGA; however, there was no inhibition of bacterial growth.
In vitro release studies
In order to study the release kinetics model of the obtained formulations (Table 3), different mathematical models (zero-order, first-order, Higuchi, and Kors-peppas models) were used to study the in vitro release profile.
After comparing the models, it is possible to notice that the Higuchi and Kors-peppas models presented higher coefficients of linear correlation in both formulations, demonstrating that they were better adjusted to the release kinetics of NPs containing vancomycin and meropenem. These results suggest a combined release by diffusion and erosion of the polymer matrix (Paul 2011).
The accumulated percentage of drug release was plotted as a function of time. The release kinetics of the antibiotics revealed that NPGA PLGA-VAN presented 43.9% at the end of 24 h and 96% at the end of 96 h and the PLGA-MER formulation at 8.25% and 16% at the end of 24 and 96 h, respectively (Fig. 6).
Reliable kinetics of in vitro release is an essential step since knowing its behavior, it becomes possible to establish future correlations with in vivo conditions (Modi and Anderson 2013). The in vitro release profile may also provide information on the dosage form, which allows a rational approach in the development of pharmaceuticals (D’Souza 2014).
Conclusion
NP-PLGA formulations, PLGA-VAN NPs and PLGA-MER NPs, were successfully obtained by double emulsification-solvent evaporation (A/O/A), presenting satisfactory yield. The NPs presented homogeneous, uniform and spherical, and apparently smooth distribution. The release profile for PLGA-VAN formulation was 43.9% at the end of 24 h and 96% at the end of 96 h. The PLGA-MER formulation presented a release of 8.25% and 16% at the end of 24 h and 96 h, respectively. The results presented suggest the possibility of formulations reaching free microorganisms and biofilms. This study supports the potential application of PLGA particles containing antibiotics as facilitators of drug delivery and release effectively.
References
Abdelkader A, El-Mokhtar MA, Abdelkader O, Hamad MA, Elsabahy M, El-Gazayerly ON (2017) Ultrahigh antibacterial efficacy of meropenem-loaded chitosan nanoparticles in a septic animal model. Carbohydr Polym 174:1041–1050. https://doi.org/10.1016/j.carbpol.2017.07.030
Cielecka-Piontek J, Paczkowska M, Lewandowska K, Barszcz B, Zalewski P, Garbacki P (2013) Solid-state stability study of meropenem – solutions based on spectrophotometric analysis. Chem Cent J 7:98. https://doi.org/10.1186/1752-153X-7-98
Clinical and Laboratory Standards Institute (2017) M100-S27. Performance standards for antimicrobial susceptibility testing: 27th informational supplement. CLSI, Wayne
D’Souza S (2014) A review of in vitro drug release test methods for Nano-sized dosage forms. Adv Pharm 2014:1–12. https://doi.org/10.1155/2014/304757
Danhier F, Ansorena E, Silva JM, Coco R, Le Breton A, Préat V (2012) PLGA-based nanoparticles: an overview of biomedical applications. J Control Release 161(2):505–522. https://doi.org/10.1016/j.jconrel.2012.01.043
Duncan D, Gaspar R (2011) Nanomedicine(s) under the microscope. Mol Pharm 8:2101–2141. https://doi.org/10.1021/mp200394t
Georgiev AG, Johansen J, Ramanathan VD, Sere YY, Beh CT, Menon AK (2013) Arv1 regulates PM and ER membrane structure and homeostasis but is dispensable for intracellular sterol transport. Traffic 14(8):912–921
Günday Türeli N, Torge A, Juntke J, Schwarz BC, Schneider-Daum N, Türeli AE, Lehr CM, Schneider M (2017) Ciprofloxacin-loaded PLGA nanoparticles against cystic fibrosis P. aeruginosa lung infections. Eur J Pharm Biopharm 117:363–371. https://doi.org/10.1016/j.ejpb.2017.04.032
Hill LE, Taylor MT, Gomes C (2013) Antimicrobial efficacy of poly(DL-lactideco-glycolide) (PLGA) nanoparticles with entrapped cinnamon bark extract against listeria monocytogenes and Salmonella typhimurium. J Food Sci 78(4):626–632. https://doi.org/10.1111/1750-3841.12069
Javed KR, Ahmad M, Ali S et al (2015) Comparison of Doxorubicin Anticancer Drug Loading on Different Metal Oxide Nanoparticles. Medicine 94(11):e617. https://doi.org/10.1097/MD.0000000000000617
Jelvehgari M, Barar J, Valizadeh H, Heidari N (2010) Preparation and evaluation of poly (s-caprolactone) nanoparticles-in- microparticles by W/O/W emulsion method. Iran J Basic Med Sci 13(3):85–96. https://doi.org/10.22038/ijbms.2010.5090
Kim JK, Kim HJ, Chung J-Y, Lee JH, Young SB, Kim YH (2014) Natural and synthetic biomaterials for controlled drug delivery. Arch Pharm Res 37:60–68. https://doi.org/10.1007/s12272-013-0280-6
Lotfipour F, Abdollahi S, Jelvehgari M, Valizadeh H, Hassan M, Milani M (2014) Study of antimicrobial effects of vancomycin loaded PLGA nanoparticles against enterococcus clinical isolates. Drug Res (Stuttg) 64(7):348–352. https://doi.org/10.1055/s-0033-1358747
Modi S, Anderson BD (2013) Determination of drug release kinetics from nanoparticles: overcoming pitfalls of the dynamic Dialysis method. Mol Pharm 10(8):3076–3089. https://doi.org/10.1021/mp400154a
Mohanraj VJ, Chen Y (2006) Nanoparticles – a review. Trop J Pharm Res 5(1):561–573. https://doi.org/10.4314/tjpr.v5i1.14634
Moreira Gomes C, Moreira RG, Castell-Perez E (2011) Poly (DL-lactide-coglycolide) (PLGA) nanoparticles with entrapped transcinnamaldehyde and eugenol for antimicrobial delivery applications. J Food Sci 76:16–24. https://doi.org/10.1111/j.1750-3841.2010.01985.x
Nandakumar V, Geetha V, Chittaranjan S, Doble M (2013) High glycolic poly (DL lactic co glycolic acid) nanoparticles for controlled release of meropenem. Biomed Pharmacother 67:431–436. https://doi.org/10.1016/j.biopha.2013.02.004
Parisi OI, Scrivano L, Sinicropi MS, Puoci F (2017) Polymeric nanoparticle constructs as devices for antibacterial therapy. Curr Opin Pharmacol 36:72–77. https://doi.org/10.1016/j.coph.2017.08.004
Paul DR (2011) Elaborations on the Higuchi model for drug delivery. Int J Pharm 418:13–17. https://doi.org/10.1016/j.ijpharm.2010.10.037
Posadowska U, Brzychczy-Włoch M, Pamuła E (2015) Gentamicin loaded PLGA nanoparticles as local drug delivery system for the osteomyelitis treatment. Acta Bioeng Biomech 17(3):41–48. https://doi.org/10.1007/s10856-015-5604-2
Rao JP, Geckeler KE (2011) Polymer nanoparticles: preparation techniques and size-control parameters. Prog Polym Sci 36:887–913. https://doi.org/10.1016/j.progpolymsci.2011.01.001
Sabaeifard P, Abdi-Ali A, Soudi MR, Gamazo C, Irache JM (2016) Amikacin loaded PLGA nanoparticles against Pseudomonas aeruginosa. Eur J Pharm Sci 93:392–398. https://doi.org/10.1016/j.ejps.2016.08.049
Silva ISM, Santos RFP, Melo TVC, Silva AJC, Sarmento PA, Lúcio IML, Campesatto EA, Padilha FF, Conserva LM, Bastos MLA (2014) In vitro biological potential of Guanxuma-of-horn [Sebastiania corniculata (Vahl) Mull. Arg.] in infection control. J Chem Pharm Res 6(4):663–669
Tomaszewska E, Soliwoda K, Kadziola K, Tkacz-Szczesna B, Celichowski G, Cichomski M, Szmaja W, Grobelny J (2013) Detection limits of DLS and UV-vis spectroscopy in characterization of Polydisperse nanoparticles colloids. J Nanomater 2013:1–10. https://doi.org/10.1155/2013/313081
Venkatesh DN, Baskaran M, Karri VVSR, Mannemala SS, Radhakrishna K, Goti S (2015) Fabrication and in vivo evaluation of Nelfinavir loaded PLGA nanoparticles for enhancing oral bioavailability and therapeutic effect. Saudi Pharm J 23(6):667–674. https://doi.org/10.1016/j.jsps.2015.02.021
World Health Organization (WHO) (2017) Global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics. IOP Publishing PhysicsWeb. http://www.who.int/medicines/publications/global-priority-list-antibiotic-resistant-bacteria/en/. Accessed 10 December 2017
Yameen B, Choi WI, Vilos C, Swami A, Shi J, Farokhzad OC (2014) Insight into nanoparticle cellular uptake and intracellular targeting. J Control Release 190:485–499. https://doi.org/10.1016/j.jconrel.2014.06.038
Zaidi S, Misba L, Khan AU (2017) Nano-therapeutics: a revolution in infection control in post antibiotic era. Nanomedicine 13(7):2281–2301. https://doi.org/10.1016/j.nano.2017.06.015
Zakeri-Milani P, Loveymi BD, Jelvehgari M, Valizadeh H (2013) The characteristics and improved intestinal permeability of vancomycin PLGA-nanoparticles as colloidal drug delivery system. Colloids Surf B: Biointerfaces 103:174–181. https://doi.org/10.1016/j.colsurfb.2012.10.021
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
Rights and permissions
About this article
Cite this article
Gaspar, L.M.d.A.C., Dórea, A.C.S., Droppa-Almeida, D. et al. Development and characterization of PLGA nanoparticles containing antibiotics. J Nanopart Res 20, 289 (2018). https://doi.org/10.1007/s11051-018-4387-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11051-018-4387-z