Abstract
Tissue folding is a frequently observed phenomenon, from the cerebral cortex gyrification, to the gut villi formation and even the crocodile head scales development. Although its causes are not yet well understood, some hypotheses suggest that it is related to the physical properties of the tissue and its growth under mechanical constraints. In order to study the underlying mechanisms affecting tissue folding, experimental models are developed where epithelium monolayers are cultured inside hydrogel microcapsules. In this work, we use a 2D vertex model of circular cross-sections of cell monolayers to investigate how cell mechanical properties and proliferation affect the shape of in-silico growing tissues. We observe that increasing the cells’ contractility and the intercellular adhesion reduces tissue buckling. This is found to coincide with smaller and thicker cross-sections that are characterized by shorter relaxation times following cell division. Finally, we show that the smooth or folded morphology of the simulated monolayers also depends on the combination of the cell proliferation rate and the tissue size.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Tissue folding is a frequently observed phenomenon, from blastula gastrulation (Tamulonis et al. 2011; Rauzi et al. 2013; Polyakov et al. 2014), to the formation of brain convolutions (Tallinen et al. 2014, 2016; Mota and Herculano-Houzel 2015), gut villis (Simons 2013; Shyer et al. 2013), cyst (Bielmeier et al. 2016) and even crocodile head scales (Milinkovitch et al. 2013). However, the causes of tissue buckling and folding are not well understood. Some studies suggest that buckling results from cell mechanical properties. Bielmeier et al. (2016) demonstrated that buckling can emerge from interface contractility between differently fated cells, potentially leading to cyst formation. Tamulonis et al. (2011) showed that the blastula gastrulation can emerge from the endoderm and ectoderm being characterised by different apical cell constriction and intercellular adhesion. Štorgel et al. (2016) proposed a mechanical model explaining epithelial folds, based on intraepithelial stresses generated by differential tensions of apical, lateral and basal cell sides as well as on the elasticity of the basement membrane. Other hypotheses suggest that tissue folding is related to its growth under mechanical constraints. Tallinen et al. suggested that gyrification arises from mechanical instabilities driven by differential growth between the grey and the white matter (Tallinen et al. 2014, 2016). Shyer et al. (2013) explained that the formation of the gut villis is the consequence of the endoderm expansion under compressive stresses generated by sequential differentiation of distinct smooth muscle layers of the gut.
In order to support such hypotheses and investigate the mechanisms underlying tissue folding, numerical models were developed. Among them, one can find the vertex models that were used in the past decade to study the mechanics of confluent cell monolayers (Alt et al. 2017; Fletcher et al. 2013, 2014). Although 3D vertex models were recently proposed (Bielmeier et al. 2016; Monier et al. 2015; Misra et al. 2016), the 2D models are commonly used to describe both in-plane tissues as seen from the apical face for the study of phenomena like cell sorting (Aliee et al. 2012; Umetsu et al. 2014) or wound healing (Nagai and Honda 2009), and to describe transverse cross-sections of tissues for investigating phenomena that are three-dimensional in nature such as tissue folding (Štorgel et al. 2016) and gastrulation (Tamulonis et al. 2011; Rauzi et al. 2013; Polyakov et al. 2014).
Example of a confocal scan of an epithelium monolayer, which was cultured inside a hydrogel microcapsule, and that starts buckling. The cell membrane is coloured in green and the microcapsule in purple. Images from the Roux Lab, Department of Biochemistry, University of Geneva, Switzerland. (Color figure online)
Lately, experimental models, where cells are cultured inside hydrogel elastic microcapsules (Alessandri et al. 2013, 2016), are also used to investigate the effect of mechanics on cell monolayers development and buckling (see Fig. 1). In this work, we use a 2D vertex model to simulate a cross-section of a spherical cell monolayer. We investigate how cell mechanics combined with cell proliferation affect the morphology of the growing tissue. The remainder of the paper is organised as follows. Section 2 introduces our numerical vertex model. Results and discussion are presented in Sect. 3. In Sect. 3.1, we show how cell mechanical properties, namely cell contractility and intercellular adhesion, affect cell monolayer buckling. In Sect. 3.2, we analyse the consequence of increasing cell proliferation rate on the morphology of the simulated growing monolayer. Finally, we conclude in Sect. 4 by summarising the results and presenting the future work.
2 Numerical model
2.1 Cell monoloyer representation and dynamics
The model represents a cross-section along cell height of a spherical cell monolayer by a circular ring of cells. A cell is described by a quadrilateral, defined by four consecutive vertices interconnected by edges. The ring topology of the cell monolayer induces that each cell has two neighbours, and in this model two adjacent cells share a common edge (see Fig. 2 left). We differentiate the lateral edges of a cell, also referred as bulk edges in Merzouki et al. (2016), which are shared by the cell with its neighbours, and the apico-basal edges, which belong solely to the cell and are also designated as boundary edges. Similar representations of circular cross-sections of cell monolayers were used in Štorgel et al. (2016), Polyakov et al. (2014).
Numerical and experimental models of an epithelium monolayer. Left cell-monolayer cross-section, as represented by the 2D numerical vertex model, is a ring made up of quadrilateral cells. Each cell shares a common edge with its two neighbour cells. Shared edges at the junction of neighbour cells are coloured in yellow. Right confocal scan of an epithelium monolayer cultured inside an elastic alginate microcapsule coated with a matrigel layer that is used as an attachment substrate for cells. Cells’ membrane and the alginate microcapsule are coloured in purple and green, respectively. Image from the Roux Lab, Department of Biochemistry, University of Geneva, Switzerland. (Color figure online)
We note that no standard energy function exists in the literature for circular cross-sections of tissues (Štorgel et al. 2014, 2016). The energy function that we use in this work (see Eq. 1) is habitually used by 2D apical vertex models (Farhadifar et al. 2007; Merzouki et al. 2016), but it is also similar to that of Polyakov et al. (2014) for modeling tissue transverse cross-sections.
where \(A_{\alpha }\) is the area of a cell \(\alpha\), \(A_{\alpha }^0\) is its preferred area , \(L_{\alpha }\) is its perimeter, and \(L_{i,j}\) is the length of the edge \(e_{ij}\) connecting vertices \(v_i\) and \(v_j\). The first term of H represents the cell area elasticity. \(K_{a}\) being positive, this term is minimised when the area of the cell \(A_{\alpha }\) tends to its preferred area \(A_{\alpha }^0\). The second term of the energy represents the cell perimeter contractility, produced by the cytoskeleton. \(\Gamma _{\alpha }\) being positive, the minimisation of this term reduces the perimeter of the cell \(L_{\alpha }\) and makes the cell more round-shaped. Finally, the third term represents the adherence of cells. The line tension \(\Lambda _{i,j}\) along lateral edges \(e_{ij}\) is positive, which leads to edges that tend to be longer to minimise this third term of H, representing junctions between pairs of cells that have a tendency to adhere to each other. In contrast, we set \(\Lambda _{i,j} = 0\) along all the boundary edges \(e_{i,j}\).
Left counterclockwise ordering of cell vertices. Right representation of forces applied by one cell \(\alpha\) on its vertices. The forces \({\mathbf {F}}^{\mathbf{A}}\), coloured in green, are those related to the cell area elasticity. They make the cell area converge toward its preferred area \(A^0\). In red, \({\mathbf {F}}^{\mathbf{P}}\) are the forces related to the cell contractility. These forces tend to minimise the cell perimeter. Finally, in blue are the forces \({\mathbf {F}}^{\mathbf{L}}\) related to the adhesion of the cell \(\alpha\) to its neighbours. These forces increase the contact surface between two adjacent cells. (Color figure online)
Our 2D vertex model is similar to that of Farhadifar et al. (2007), in terms of its energy function (see Eq. 1). However, the fact of using the 2D vertex model to describe tissues’ cross-sections along cell height instead of in-plane tissues viewed from their apical surface, requires the implementation of different boundary conditions and a special care has to be taken for their management (see Sect. 2.3). Moreover, in contrast with Farhadifar et al. (2007) where the vertex movements are driven by the minimisation of the energy equation using the Conjugate Gradient Method, we use Newtonian dynamics in the same spirit as used in Tamulonis et al. (2011). This dynamics allows us to follow the physical time evolution of the tissue.
As we have done for the model dynamics in Merzouki et al. (2016), we derive the force \(\varvec{F_i}\) acting on each vertex \(v_i\) at a position \(\varvec{r_i} =<x_i,y_i>\), from the energy function H (see Eq. 1),
where \(<x_{\alpha _{i-1}},y_{\alpha _{i-1}}>\) and \(<x_{\alpha _{i+1}},y_{\alpha _{i+1}}>\) are the positions of the previous and next vertices, \(v_{\alpha _{i-1}}\) and \(v_{\alpha _{i+1}}\), of vertex \(v_i\) in cell \(\alpha\), when the vertices of \(\alpha\) are ordered counterclockwise (see Fig. 3 left). Figure 3 (right) displays the forces applied by one cell \(\alpha\) on its vertices, including the forces resulting from the cell area elasticity, its perimeter contractility and its adhesion to its adjacent cells.
A velocity-dependent friction is added to the force equation to dissipate energy and prevent the system to oscillate for ever. Newton mechanics are used to determine the acceleration \(\frac{d^2\varvec{r_i}}{dt^2}\) of the vertex \(v_i\),
where \(m_i\) is the mass of the vertex \(v_i\) and \(\eta _i\) is the damping parameter controlling the viscosity of the vertex’s movement.
To determine the position \(\varvec{r_i}\) of a vertex \(v_i\) at the time step \(t+\delta t\) based on its acceleration \(\frac{d^2\varvec{r_i}}{dt^2}\), the differential equation (Eq. 3) is solved using the Verlet integration method with a time step \(\delta t\),
and
2.2 Cell proliferation
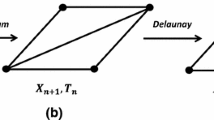
Similarly to Farhadifar et al. (2007), the process of cell proliferation starts by cells entering mitosis, growing in size and dividing. Cells are regularly picked at random to enter mitosis and each cell has a probability \(p_{mitosis}\) to be selected. A cell \(\alpha\) grows as a consequence of an increase of its preferred area \(A^0_{\alpha }\) by small increments (see Fig. 4 left). When a growing cell reaches a threshold area \(A^{thresh}\), which is set as the double of its size prior to entering mitosis, the cell is divided along its apical-basal axis (see Fig. 4 middle), creating two daughter cells (see Fig. 4 right). The cell proliferation is characterised by two parameters: (i) the probability of a cell to enter mitosis \(p_{mitosis}\) and (ii) its growth rate. The first one controls the average number of cells entering mitosis at the same time and the second one controls the speed at which the cells on mitosis grow.
Cell proliferation. Left cells grow in size during mitosis (in yellow), due to a progressive increase of their \(A^0_\alpha\). Middle the cells that reached their threshold size \(A^{thresh}_{\alpha }\), i.e. doubled their size preceding mitosis, are divided along their apico-basal axes (dashed line). Right cell division creates two daughter cells (in red). The daughter cells are assigned the same \(A^0\) as their parent prior to entering mitosis. (Color figure online)
2.3 Boundary management
Our model’s open boundary conditions, allowing us to simulate a circular cross-section of a spherical cell monolayer, with both bulk and boundary edges, requires additional care to be taken during simulations. Indeed, we saw from Eq. 2, that unless vertices are linked by an edge or belong to a common cell, they will not consider each other positions when being displaced according to their forces. Therefore, the displacement of a vertex may lead it to cross an edge and enter a cell it does not belong to, producing unrealistic cell overlapping (see Fig. 5 left). For this purpose, before moving a vertex, we check that this displacement does not cause such an unwanted event. In case it does, the movement of the vertex is corrected such that it stops at the edge it would have crossed in the absence of proper boundary management (see Fig. 5 right).
Boundary management to avoid cell overlapping. Left without boundary management, cells overlap. Yellow dashed lines highlight the regions of the cell monolayer that overlapped. Right with boundary management, cell overlapping is avoided. Green dashed lines highlight the regions of the cell monolayer where overlapping was avoided. (Color figure online)
3 Results and discussion
We focus in this work on investigating how our model parameters, related to cell mechanical properties and proliferation, control the buckling of the simulated tissues. In what follows, a unique type of cells compose the cell monolayer cross-section. All the cells are assigned with the same properties. The cells’ area elasticity K and the preferred area \(A^0\) are set to \(10^9\) N/m\(^3\) and \(70\times 10^{-12}\) m\(^2\), respectively. These values are similar to those experimentally measured and numerically estimated in Merzouki et al. (2016), Brückner and Janshoff (2015). The cell growth rate is set so that the value of \(A^0\) increases quasistatically.Footnote 1
3.1 Influence of cell mechanical properties on tissue buckling
In order to study the influence of the cell contractility and the intercellular adhesion on the tissue buckling, we perform a set of simulations where our model’s normalised parameters \(\bar{\Gamma }={\Gamma \over {K\cdot A^0}}\) and \(\bar{\Lambda }={\Lambda \over ({K\cdot A^0})^{3/2}}\) are varied. As described in Farhadifar et al. (2007), Merzouki et al. (2016), a high value of the normalised cell contractility parameter \(\bar{\Gamma }\) means that the cell contractile forces are high compared to those of the cell area elasticity. Similarly, a high value of the normalised adhesion parameter \(\bar{\Lambda }\) implies high intercellular adhesion forces compared to those resulting from the cell area elasticity. 50 simulations are performed for each couple of parameters \((\bar{\Gamma },\bar{\Lambda })\), with \(\bar{\Gamma }\) and \(\bar{\Lambda }\) ranging from 0.02 to 0.14 with a step of 0.02, and from 0 to 0.6 with a step of 0.1, respectively. Each simulation starts with a circular monolayer cross-section made up of 20 cells and ends after 100 cell divisions, with \(p_{mitosis} = 0.02\).
Diagrams showing how cell contractility \(\bar{\Gamma }\) and intercellular adhesion \(\bar{\Lambda }\) affect the tissue’s cross-section buckling. Results of simulations that start from circular cross-sections made up of 20 cells and end after 100 cells divisions, with \(p_{mitosis} = 0.02\). a Examples of resulting cell monolayers. The colour (from red to blue) of the tissue depends on the average cell aspect ratio (cell width divided by cell height), or the inverse of the relative thickness of the tissue. Red represents relatively thin tissues (\(\text {cell height} \sim \text {cell width}\)), while blue represents relatively thick tissues (\(\text {cell height} > \text {cell width}\)). b Buckling measure, \(B = {P^2\over A}\), where P is the circumference of the cross-section and A is the area delimited by the outer boundary edges of the cross-section. For a given area A, the circumference P and the buckling measure B are minimised when the cross-section is circular. In general, the smaller B, the more circular and smooth is the cross-section. Inversely, the higher B, the more folded is the cross-section. The quantitative results in b are averaged over 50 simulations for each \((\bar{\Gamma },\bar{\Lambda })\). The normalised standard deviation of the buckling measure \({\sigma \over \mu }[B] < 8\%\). Smaller contractility and intercellular adhesion yield more folded cell monolayers, while increased cell contractility and intercellular adhesion result in smoother tissues. (Color figure online)
Figure 6a presents one example of the resulting cross-sections obtained with different \((\bar{\Gamma },\bar{\Lambda })\). We note that no simulations were possible in the parameter regions 1 and 2. These regions correspond to cases where cells tend to have infinitely long lateral edges (region 1) or become infinitely small (region 2). The buckling of a simulated cross-section is quantified using the measure \(B = P^2/A\), where P is the circumference of the cross-section and A is the area delimited by its outer edges. For a given delimited area A, the circumference P of the cross-section is minimised (and therefore B is minimised) when the cross-section is circular. B increases for more folded cross-sections. The effect of the cell mechanical properties \((\bar{\Gamma },\bar{\Lambda })\) on the buckling measure B are presented in Fig. 6b. These quantitative results of tissue buckling, are averaged over the 50 simulations performed for each \((\bar{\Gamma },\bar{\Lambda })\). We find that higher cell contractility and intercellular adhesion yield more circular and smooth monolayer cross-sections. Inversely, the lowest cell contractility and intercellular adhesion result in the more folded tissues. The normalised standard deviation of the buckling measure \({\sigma \over \mu }[B]\) varies between 0.1 and \(8\%\) from the regions where the average buckling measure \(\mu [B]\) is low (high \(\bar{\Gamma }\) and \(\bar{\Lambda }\)) to the regions were the average buckling measure is large (low \(\bar{\Gamma }\) and \(\bar{\Lambda }\)).
Diagrams showing how cell contractility \(\bar{\Gamma }\) and intercellular adhesion \(\bar{\Lambda }\) affect the tissue’s cross-section thickness and circumference. These are the results of simulations that start from a circular cross-section made up of 20 cells and end after 100 cells divisions, with \(p_{mitosis} = 0.02\). a Average thickness of the cross-section T or average cell height. b Cross-section circumference P. Smaller contractility and intercellular adhesion yield larger and thinner cell monolayers, while increased cell contractility and intercellular adhesion result in smaller and thicker tissues. These measures are averaged over 50 simulations for each \((\bar{\Gamma },\bar{\Lambda })\). Normalised standard deviations of the cross-section thickness and circumference are \({\sigma \over \mu }[T] < 0.8\%\) and \({\sigma \over \mu }[P] < 1.4\%\)
Moreover, we measure the circumference and thickness of the tissue’s cross-sections. The average measures obtained with the varied cell contractility and intercellular adhesion are presented in Fig. 7. We see that \(\bar{\Gamma }\) and \(\bar{\Lambda }\) influence the size and thickness of the simulated monolayers. In general, increasing \(\bar{\Gamma }\) and \(\bar{\Lambda }\) lead to cross-sections with reduced circumference and increased thickness. We note that the tissue thickness and size vary very little over the 50 simulations performed with the same \((\bar{\Gamma },\bar{\Lambda })\); Their normalised standard deviations were \({{\sigma \over \mu }[T]}<0.8\%\) and \({{\sigma \over \mu }[P]}<1.4\%\). Indeed the tissue thickness and perimeter depend mainly on the cell mechanical properties and are not remarkably affected by the stochastic proliferation of cells. Therefore, we find that tissue folding coincides with larger and thinner tissues, while circular and smooth configurations are associated to smaller and thicker tissue cross-sections.
Finally, we also measure the time needed by the monolayer cross-section to relax (reach an equilibrium state of minimised energy) after one cell division, for the different \((\bar{\Gamma },\bar{\Lambda })\). The results are presented in Fig. 8a. We observe that smaller cell contractility and intercellular adhesion correspond to tissues that require longer times to relax after one cell division, while those characterised by higher cell contractility and intercellular adhesion relax faster. Therefore, buckling cross-sections coincide with longer relaxation time needed after cell division. We confirm this observation by plotting the average buckling measure B of the simulated tissues characterised by a given \((\bar{\Gamma },\bar{\Lambda })\) versus the relaxation time needed by these tissues to relax after one cell division (see Fig. 8b).
a Diagram showing how cell contractility \(\bar{\Gamma }\) and intercellular adhesion \(\bar{\Lambda }\) affect the time needed by the tissue’s cross-section to relax after one cell division. One cell division is performed on a cricular cross-section made up of 20 cells. Here, the cross-section is considered to be relaxed when the normalized standard deviation of the energy \({\sigma \over \mu }[E]\) over the last \(3\times 10^4\) iterations is smaller than \(10^{-6}\). b Scatter plot of the average tissue buckling versus the time needed to relax after one cell division. Smaller contractility and intercellular adhesion yield longer relaxation times after one cell division, which coincides with more folded tissues
3.2 Proliferation rate and tissue size
In order to study how the cell proliferation influences the tissue shape, we simulated cell monolayers made up initially of 20 cells that enter mitosis with a probability \(p_{mitosis} \in \{0.01, 0.02, 0.04, 0.08, 0.16, 0.32\}\). In what follows, cells were assigned \(\bar{\Gamma } = 0.04\) and \(\bar{\Lambda } = 0.3\).
Figure 9 displays the simulated growing tissues after 50, 100 and 200 cell divisions. We observe that the proliferation rate of cells affects the final morphology of cell monolayers. Tissues with the same number of cells may have a smooth or folded morphology, depending on whether they are the result of a low or high cell proliferation rate. Higher values of \(p_{mitosis}\), corresponding to larger proportions of cells growing and dividing at the same time, produce tissues that grow faster and are more folded. However, we see that even small proliferation rates end up buckling after a large number of cell divisions. In summary, although an infinite number of cell divisions seems to always lead the tissue to fold,Footnote 2 we note that the proliferation rate controls when buckling happens. The lower the proliferation rate, the later the tissue starts folding. For instance, at \(p_{mitosis} = 0.01\), the tissue buckles after 200 cell divisions, while at \(p_{mitosis} = 0.32\), it already starts buckling after 50 cell divisions.
Diagram showing how the probability of a cell to enter mitosis \(p_{mitosis}\) and the number of cell divisions control whether the tissue ends up smooth or folded. All these simulations started from a circular monolayer made up of 20 cells. A higher probability for a cell to enter mitosis enhances the occurrence of buckling after a given number of cell divisions. On the other hand, for any cell proliferation rate, an infinitely growing tissue always ends up buckling
In order to show that the above observations stay true whatever the initial size of the cell monolayer, we repeated these simulations starting now from larger tissues, made up of 60 cells. Figure 10 presents the obtained results. Similarly to earlier findings, a tissue of same size (ex. 110 cells, generated after 50 cell divisions) may end up either smooth (with \(p_{mitosis} = 0.01\)) or folded (as with \(p_{mitosis} = 0.32\)) depending on its cell proliferation rate. However, even the tissue proliferating at the lowest rate, \(p_{mitosis} = 0.01\), which stayed smooth after 50 cell divisions, ends up buckling after additional cell divisions (ex. from 100 cell divisions). Finally, when comparing two simulations (from Figs. 9, 10), where the same number of cell divisions are performed at the same proliferation rate but starting from distinct initial numbers of cells, we notice that the tissues that end with the more cells are those displaying the larger number of folds with the higher amplitudes.
Diagram showing how the probability of a cell to enter mitosis \(p_{mitosis}\) and the number of cell divisions control whether the tissue ends up smooth or folded. All these simulations started from a circular monolayer made up of 60 cells. A higher probability for a cell to enter mitosis enhances the occurrence of buckling after a given number of cell divisions. On the other hand, for any cell proliferation rate, an infinitely growing tissue always ends up buckling
4 Conclusion
In this work, we investigated how the parameters of our 2D vertex model of cell monolayer cross-sections, which are related to the mechanical properties and proliferation of cells, influence the buckling of the simulated growing tissues.
We found that increased cell contractility and intercellular adhesion lead to more circular and smoother tissues, and inversely that lower cell contractility and intercellular adhesion result in more folded monolayers. Furthermore, enhanced buckling was found to coincide with larger and thinner tissues, requiring longer time to relax after cell division, while smoother tissue cross-sections had smaller circumference, increased thickness, and were characterised by shorter relaxation times following cell division. Although our model simulates cross-sections of epithelial cell monolayers and that our current analysis is mainly qualitative, our first result relating tissue size and thickness through cell mechanics to folding, may remind us of the results of Mota and Herculano-Houzel (2015) who found that cortical folding scales across lissencephalic and gyrencephalic species as a function of the product of cortical surface area and the square root of the cortical thickness.
Moreover, we observed that besides cell mechanics, the cell proliferation rate also determines whether a growing tissue will be smooth or folded after a given number of cell divisions. The same number of cell divisions performed from the same simulated cell monolayer, can result in remarkably different morphologies depending on the rate at which cells proliferate. The higher the cell proliferation rate, the faster the tissue grows, the sooner it buckles and the more compact are its folds. However, our results showed that even at low proliferation rates, a tissue growing infinitely is expected to eventually buckle. A constant low proliferation rate may therefore postpone tissue folding or prevent it for a certain time but not indefinitely.
In general, we showed that in-silico growing cell monolayers end up folding despite the absence of external mechanical constraints, such as those potentially applied by an additional confining environment. Similarly, even though in the context of the grey matter folding, Tallinen et al. (2014) showed that compressive constraints of the skull or the meninges is not required for gyrification. Instead, the latter is a function of the relative cortical expansion and its relative thickness (compared with the brain size). Moreover, we showed that the folding of simulated growing cell monolayers can occur without the presence of different types of cells as in Tamulonis et al. (2011), Bielmeier et al. (2016), or differential tension between the boundary (apical and basal) edges of the cells, as in Štorgel et al. (2016), Höhn and Hallmann (2011). In our simulations, all the cells were identical and only characterised by their normalised overall contractility, adhesion to their neighbours and proliferation rate.
Although we did not know about the work of Drasdo (2000) during the writing of this paper, our obtained results using a 2D vertex model confirm his findings, when studiying the buckling of growing cell monolayers. He found that folding occurs when the bending rigidity is too small to compensate the cell proliferation. The monolayer’s bending rigidity can be related to the cell contractility and the intercellular adhesion in our vertex model. We showed that the larger the cell contractility and the intercellular adhesion, the more rigid, circular and smooth is the tissue. In this work, we also explicitely showed how our model parameters related to the cell mechanics affect the relaxation time of the tissue. We also evaluated how they affect the simulated tissue circumference and thickness, which are characteristic of folded tissues.
A future work will be to perform a more quantitative study on the relation between cell mechanical properties, tissue size and thickness, and monolayer buckling. Also, an interesting task will be to investigate the causes of our observations regarding the eventual buckling of infinitely growing tissues. Moreover, as our model allows a simple way to apply external mechanical constraints on the simulated cell monolayer, we will in the future study the impact of confinement on the development of cell monolayers. Finally, as the main limitation of the model presented in this work is its two-dimensionality, which misses the curvature of the tissue in the third dimension, we are implementing a vertex model that describes a cell monolayer evolving in a three-dimensional space to compare the obtained results.
Notes
\(A^0\) of a cell \(\alpha\) is increased each 3000 iterations by \(\Delta {A^0} = {0.1\cdot A_{\alpha }\text {(before mitosis)}}\), which is enough time for a cell to reach equilibrium between each \(A^0\) increment in our simulations, with a time step \(\delta t\) and a damping \(\eta\) parameters equal to \(10^{-1}\) sec and 1 sec\(^{-1}\). The choice of the cell growth rate corresponds to the implementation of a multi-scale simulation technique (combining the relaxation time scale in the order of minutes and the proliferation time scale in the order of 10–20 h), and does not correspond to biologically realistic growth times. We artificially accelerate the cell growth to speed up the execution of our simulations. However, the quasi-static growth of cells insures that we get the same results as those we would have obtained with a slower and more biologically realistic cell growth.
As long as the tissue does not have the time to return to its equilibrium state between two cell divisions.
References
Alessandri K, Sarangi BR, Gurchenkov VV, Sinha B, Kiessling TR, Fetler L, Rico F, Scheuring S, Lamaze C, Simon A, Geraldo S, Vignjević D, Domejean H, Rolland L, Funfak A, Bibette J, Bremond N, Nassoy P (2013) Cellular capsules as a tool for multicellular spheroid production and for investigating the mechanics of tumor progression in vitro. Proc Natl Acad Sci 110(37):14843. doi:10.1073/pnas.1309482110
Alessandri K, Feyeux M, Gurchenkov B, Delgado C, Trushko A, Krause KH, Vignjević D, Nassoy P, Roux A (2016) A 3D printed microfluidic device for production of functionalized hydrogel microcapsules for culture and differentiation of human neuronal stem cells (hNSC). Lab Chip 16(9):1593. doi:10.1039/C6LC00133E
Aliee M, Röper JC, Landsberg KP, Pentzold C, Widmann TJ, Jülicher F, Dahmann C (2012) Physical mechanisms shaping the Drosophila dorsoventral compartment boundary. Curr Biol 22(11):967. doi:10.1016/j.cub.2012.03.070
Alt S, Ganguly P, Salbreux G (2017) Vertex models: from cell mechanics to tissue morphogenesis. Philos Trans R Soc B Biol Sci 372(1720):20150520. doi:10.1098/rstb.2015.0520
Bielmeier C, Alt S, Weichselberger V, La Fortezza M, Harz H, Jülicher F, Salbreux G, Classen AK (2016) Interface contractility between differently fated cells drives cell elimination and cyst formation. Curr Biol 26(5):563. doi:10.1016/j.cub.2015.12.063
Mota B, Herculano-Houzel S (2015) Cortical folding scales universally with surface area and thickness, not number of neurons. Science 349(6243):74. doi:10.1126/science.aaa9101
Simons BD (2013) Getting your gut into shape. Science 342(6155):203. doi:10.1126/science.1245288
Farhadifar R, Röper JC, Aigouy B, Eaton S, Jülicher F (2007) The influence of cell mechanics, cell-cell interactions, and proliferation on epithelial packing. Curr Biol 17(24):2095. doi:10.1016/j.cub.2007.11.049
Fletcher AG, Osborne JM, Maini PK, Gavaghan DJ (2013) Implementing vertex dynamics models of cell populations in biology within a consistent computational framework. Prog Biophys Mol Biol 113(2):299. doi:10.1016/j.pbiomolbio.2013.09.003
Fletcher AG, Osterfield M, Baker RE, Shvartsman SY (2014) Vertex models of epithelial morphogenesis. Biophys J 106(11):2291. doi:10.1016/j.bpj.2013.11.4498
Štorgel N, Krajnc M, Mrak P, Štrus J, Ziherl P(2016) Quantitative morphology of epithelial folds. Biophys J 110(1):269. doi:10.1016/j.bpj.2015.11.024
Merzouki A, Malaspinas O, Chopard B (2016) The mechanical properties of a cell-based numerical model of epithelium. Soft Matter 12(21):4745. doi:10.1039/C6SM00106H
Milinkovitch MC, Manukyan L, Debry A, Di-Poï N, Martin S, Singh D, Lambert D, Zwicker M (2013) Crocodile head scales are not developmental units but emerge from physical cracking. Science 339(6115):78. doi:10.1126/science.1226265
Misra M, Audoly B, Kevrekidis IG, Shvartsman SY (2016) Shape transformations of epithelial shells. Biophys J 110(7):1670. doi:10.1016/j.bpj.2016.03.009
Monier B, Gettings M, Gay G, Mangeat T, Schott S, Guarner A, Suzanne M (2015) Apico-basal forces exerted by apoptotic cells drive epithelium folding. Nature 518(7538):245. doi:10.1038/nature14152
Mota B, Herculano-Houzel S (2015) Cortical folding scales universally with surface area and thickness, not number of neurons. Science 349(6243):74. doi:10.1126/science.aaa9101
Nagai T, Honda H (2009) Computer simulation of wound closure in epithelial tissues: cell basal-lamina adhesion. Phys Rev E 80(6):061903. doi:10.1103/PhysRevE.80.061903
Polyakov O, He B, Swan M, Shaevitz JW, Kaschube M, Wieschaus E (2014) Passive mechanical forces control cell-shape change during Drosophila ventral furrow formation. Biophys J 107(4):998. doi:10.1016/j.bpj.2014.07.013
Rauzi M, Hoevar Brezavek A, Ziherl P, Leptin M (2013) Physical models of mesoderm invagination in Drosophila embryo. Biophys J 105(1):3. doi:10.1016/j.bpj.2013.05.039
Shyer A, Talline T, Nerurkar N, Wei Z, Kim E, Kaplan D, Tabin C, Mahadevan L (2013) Villification: how the gut gets its villi. Science 342:212
Simons BD (2013) Getting your gut into shape. Science 342(6155):203. doi:10.1126/science.1245288
Štorgel N, Krajnc M, Mrak P, Štrus J, Ziherl P, (2016) Quantitative morphology of epithelial folds. Biophys J 110(1):269. doi:10.1016/j.bpj.2015.11.024
Tallinen T, Chung JY, Rousseau F, Girard N, Lefèvre J, Mahadevan L (2016) On the growth and form of cortical convolutions. Nat Phys 12(6):588. doi:10.1038/nphys3632
Brückner BR (1853) Janshoff A (2015), Elastic properties of epithelial cells probed by atomic force microscopy. Biochim Biophys Acta (BBA) Mol. Cell Res 11, Part B:3075. doi:10.1016/j.bbamcr.2015.07.010
Tamulonis C, Postma M, Marlow HQ, Magie CR, de Jong J, Kaandorp J (2011) A cell-based model of Nematostella vectensis gastrulation including bottle cell formation, invagination and zippering. Dev Biol 351(1):217. doi:10.1016/j.ydbio.2010.10.017
Umetsu D, Aigouy B, Aliee M, Sui L, Eaton S, Jülicher F, Dahmann C (2014) Local increases in mechanical tension shape compartment boundaries by biasing cell intercalations. Curr Biol 24(15):1798. doi:10.1016/j.cub.2014.06.052
Acknowledgements
We thank the SystemsX.ch initiative who supported this work (project EpiPhysX).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Merzouki, A., Malaspinas, O., Trushko, A. et al. Influence of cell mechanics and proliferation on the buckling of simulated tissues using a vertex model. Nat Comput 17, 511–519 (2018). https://doi.org/10.1007/s11047-017-9629-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11047-017-9629-y