Abstract
Background
In Bangladesh, only a fraction of prostate cancer patients are diagnosed annually due to lack of symptom awareness and screening challenges, resulting in high mortality. Aiming to improve screening methods, we evaluated X-ray cross-complementing gene 1 (XRCC1) Arg194Gln and Xeroderma pigmentosum group D (XPD) Lys751Gln polymorphisms to determine their relevance as potential markers for predicting prostate cancer risk, severity and clinical parameters in Bangladeshi population.
Methods and results
This study included 132 prostate cancer patients and 135 healthy controls. Genotype analysis was done from blood samples by the PCR-RFLP method. The XRCC1 Trp/Trp genotype was associated with prostate cancer (ORadj = 5.51; 95% CI = 1.13–26.78; p-value = 0.03) compared to Arg/Arg genotype. No significant association was found between the XPD variants and prostate cancer risk. The XRCC1 Trp/Trp genotype increased prostate cancer risk in smokers and non-smokers but was statistically non-significant. In individuals without a family history of cancer, the XRCC1 Trp/Trp genotype had a non-significant 4.64-fold higher risk (ORadj=4.64; 95% CI = 0.88–24.36; p-value = 0.07), while the XPD Gln/Gln had a 2.66-fold non-significant higher risk (ORadj=2.66; 95% CI = 0.88–8.10; p-value = 0.09). The XRCC1 Trp/Trp variant was associated with hematuria risk, higher mean serum creatinine, and mean prostate-specific antigen (PSA) levels in prostate cancer patients. The XPD Gln/Gln variant was only associated with higher mean serum creatinine levels.
Conclusion
Our findings suggest that XRCC1 screening may be used as a biomarker for prostate cancer to improve early diagnosis in Bangladesh.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prostate cancer (PCa) remains the second most prevalent carcinoma and the fifth leading cause of male cancer death [1]. Current therapeutic approaches for high-risk non-metastatic PCa patients include the combination of abiraterone with prednisolone and androgen deprivation therapy (ADT) [2]. In the metastatic setting, enzalutamide and abiraterone should not be combined for those starting long-term ADT, as recently reported by Attard et al. (2023) [2]. Despite the excellent outcomes in the trials, there remains concerns regarding the risks of harm from continuous treatment to progression [2]. Thus, novel therapeutic approaches are needed. These advances in therapy also underscore the importance of early detection and proper screening, as available treatment options can significantly improve survival rates if the disease is diagnosed early. The occurrence of PCa in developing countries is much lower, yet survival rates are low [3, 4]. Developing countries account for a significant proportion of PCa incidence and mortality globally [5]. According to the Global Cancer Statistics 2020, PCa ranked second in countries with higher Human Development Index (HDI) following lung cancer and first in countries with lower HDI countries among men [1]. In Bangladesh, out of 1.3–1.5 million PCa patients, only 0.2 million are diagnosed annually due to a lack of symptoms, awareness, and screening techniques [6]. Additionally, most deaths occur due to late diagnosis. While PCa is screened on prostate-specific antigen (PSA) and digital examination, there remains an unmet need for an effective PCa screening method to reduce cancer-related mortality in Bangladesh.
Genetic variations are increasingly implicated in prostate carcinogenesis and its use in PCa prediction and prognosis [7,8,9]. Abnormalities in DNA repair genes, either germline or somatic, are detected in 19% of primary PCa cases and nearly 23% in advanced castration-resistant PCa cases [10]. For instance, pathogenic variants of BRCA1 have been linked to high Gleason score (aggressive tumor grades), distant metastases, and high chances of recurrence [10]. Polymorphisms of genes involved in DNA repair may alter protein function and an individual’s capacity to repair damaged DNA, thus modulating susceptibility to cancer [11]. Base excision repair (BER) is a DNA repair pathway that precisely repairs non-bulky adducts, strand breaks, and endogenous base damage [12]. The X-ray Cross Complementing 1 (XRCC1) protein is an essential enzyme in the BER pathway encoded by the XRCC1 gene on chromosome 19q13.2-13.3 [13, 14]. The protein recognizes and binds to single-strand breaks (SSBs) by complexing with DNA ligase III at its COOH terminus, DNA polymerase β at its NH2 terminus domain, human AP endonuclease (APE1), polynucleotide kinase, and poly(ADP-ribose) polymerase at the repair site [13]. One polymorphism in codon 194 in exon 6 of the XRCC1 gene (at position 26,304 on exon 6, base C to T, amino acid Arg to Trp, rs1799782) resulting in an arginine-to-tryptophan transition has been linked to cancer susceptibility in case-control studies of numerous cancers such as breast cancer [15], lung cancer [16], thyroid [17] and bladder cancer [18]. Data in PCa are much more conflicting across different races [19,20,21,22].
Another DNA repair pathway, nucleotide excision repair (NER), corrects various DNA lesions, including chemically induced bulky adducts, cross-links, and pyrimidine dimers, by excising the short single-stranded DNA segment containing the lesion and then repairing it through DNA synthesis and ligation [23]. Xeroderma Pigmentosum Complementary group D (XPD) is an ATP-dependent 5′-3′ helicase involved in NER, which interestingly, like XRCC1, also maps to chromosome 19q13.3 [24]. The XPD protein, an essential subunit of the transcription factor IIH (TFIIH) complex, unwinds DNA in damaged regions [25]. An XPD variant at position 751 in exon 23 (at position 35,931 on exon 23, base A to C, rs13181) causing a lysine-to-glutamine transition has been associated with non-small cell lung cancer [26], breast cancer [27], bladder cancer [28], and colorectal cancer [29]. However, several studies failed to identify an increased PCa risk with XPD Lys751Gln polymorphism, namely in the Indian population [30], the Taiwanese population [31], and the South Australian population [32]. The precise role of XRCC1 Arg194Trp and XPD Lys751Gln as genetic polymorphisms in PCa risk among men of Bangladesh remains largely unknown.
In this study, we evaluated the associations between these genetic polymorphisms and the risk of PCa in the Bangladeshi population. We also considered their smoking history and family history of cancer. In addition, studies on the Single Nucleotide Polymorphisms (SNPs) association with biochemical parameters and clinical features of PCa patients are relatively scarce. Therefore, we also explored the SNPs’ association with the patient’s clinical features, including PSA levels, serum creatinine levels, tumor staging, histopathological grading and digital rectal examination (DRE) findings.
Materials and methods
Subject selection criteria
A total of 132 PCa patients with histologically diagnosed PCa were recruited from the Department of Urology of Bangladesh Institute of Research and Rehabilitation in Diabetes, Endocrine, and Metabolic Disorders (BIRDEM) General Hospital, Bangabandhu Sheikh Mujib Medical University (BSMMU) and Dhaka Medical College Hospital (DMCH), Dhaka, Bangladesh. The control group comprised 135 healthy subjects who were unrelated to the cases from the same geographic area. Participants with a history of other chronic diseases and cancer were excluded from the study.
Data collection by interview
With informed consent, participant details including age, smoking history, and family history of chronic diseases were obtained using a structured questionnaire. Clinical data were recorded in the presence of the attending physician. Present and former smokers were considered smokers, whereas individuals who had never smoked were classified as non-smokers. The tumor grade was classified as either low grade (Gleason’s score < 7) or moderate to the high-grade tumor (Gleason’s score ≥ 7). The cancer stage was categorized into two groups: Localized or Organ-confined, which included T1a-c/T2a-b N0 M0, and Locally advanced or metastatic, which had T3a-b/T4/N1/M1, according to TNM classification. The study was conducted following the approval from the Ethical Review Committees of the Faculty of Biological Sciences, University of Dhaka (Ref no. 206/Biol. Scs).
Sample Collection and DNA extraction
About three milliliters (3.0 mL) of venous blood was drawn from each participant by a phlebotomist. The drawn blood was immediately transferred to EDTA (1.20 mg/mL) tubes. Genomic DNA was extracted from the whole blood samples using FavorPrep™ Blood Genomic DNA Extraction Mini Kit according to the manufacturer’s protocol.
Genotyping
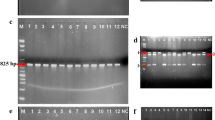
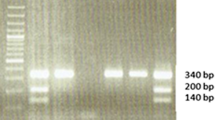
The XRCC1 codon 194 and XPD 751 genotypes were determined using the PCR-RFLP technique. Briefly, two separate PCR assays were used to amplify the XRCC1 codon 194 and XPD codon 751 polymorphisms using primer sequences (Table 1) and PCR conditions following previously published papers [33,34,35]. Restriction enzymes PvuII for XRCC1 codon 194 and PstI for XPD codon 751 polymorphisms (New England Biolabs, USA) were used for restriction fragment analyses (Supplementary Figure S1 and S2).
Statistical analysis
Statistical analysis was performed using IBM SPSS Statistics (v20.0; IBM Corp). Using logistic regression models, the relative associations were determined by calculating the odds ratio (OR) with 95% confidence intervals (CIs) and level of significance (p). Graph Pad Prism (v 8.4.2) was used to run the t-tests and Fisher’s exact tests. p-value < 0.05 was taken as the significance level.
Results
Study subjects
Table 2 lists the baseline characteristics and the clinical parameters of the study subjects. The patients age ranged from 49 to 85, with a mean age of 67 years. Controls were between the ages of 47 and 88 years, with a mean age of 65 years. More than half (56.06%) of the cancer patients were between the ages of 60–70, while the least proportion (15.91%) of patients was observed in the < 60-year group. Smoking was more common among the cases compared to the controls (60.61% vs. 38.52%, p-value < 0.001). Additionally, the cases had a higher prevalence of family history of cancer than the controls (16.67% vs. 3.7%, p-value < 0.05).
Most of the patients had low-grade (Gleason’s score < 7) tumors (59.09%) and localized organ-confined tumors (63.64%). Above 60% of patients had UTIs, while only 27.27% had hematuria. Most patients had BMI in the normal range of 18.5–24.9 kg (83.33%) and average serum creatinine levels (0.7–1.3 mg/dl) (68.94%). However, majority (82.58%) had elevated PSA values (> 10 ng/ml).
Frequency distribution of XRCC1 and XPD polymorphism and PCa risk
The XRCC1 homozygous mutant type (Trp/Trp) genotype increased PCa risk 5.51-fold (ORadj = 5.51; 95% CI = 1.13–26.78; p-value = 0.03) compared to homozygous wild type (Arg/Arg) genotype (Table 3). Heterozygous mutant (Arg/Trp) risk was not associated with PCa (ORadj = 1.21; 95% CI = 0.73–1.98; p-value = 0.46). The recessive model analysis was used to assess the effect of the minor allele (Trp allele) on the association of PCa risk. The recessive model, i.e., carrying two copies of the XRCC1 194 codon Trp allele, was positively associated by 4.88 times with the risk of PCa (ORadj=4.88; 95% CI = 1.24–22.62.12; p-value = 0.03).
In the analysis of XPD genotypes, the homozygous mutant variants (Gln/Gln) of XPD had 2.25 times the risk of PCa compared to the wild type variant, however, the association was non-significant (ORadj=2.25; 95% CI = 0.81–6.28; p-value = 0.12). Similarly, carrying two copies of the XPD codon 751 Gln allele was non-significantly associated (2.5-folds) with the risk of PCa (ORadj=2.55; 95% CI = 1.01–6.67; p-value = 0.07) as observed in the recessive model analysis.
Combined genotypic effects of XRCC1 and XPD polymorphisms on PCa Risk
Pairwise joint associations of XRCC1 Arg194Trp and XPD Lys751Gln genotypes with PCa risk were all non-significant (Table 4, Supplementary Table S1). The combined effect of XRCC1 Trp/Trp and XPD Lys/Lys has the highest risk of PCa (8.23 folds) but was statistically insignificant (OR = 8.23; 95% CI = 1.33–95.83; p-value = 0.06). No healthy control was homozygous mutant in both genes, so its combined effect could not be compared.
XRCC1 and XPD polymorphism on risk of PCa based on Smoking Status and Family History of Cancer
Smokers with XRCC1 Trp/Trp genotype had a non-significant 5.45 times increased PCa risk (ORadj=5.45; 95% CI = 0.63–47.39; p-value = 0.13) and XPD Gln/Gln genotype had a non-significant 3-fold increased PCa risk (ORadj=3.04; 95% CI = 0.59–15.40; p-value = 0.18) (Table 5). Surprisingly, none of the variants of XRCC1 or XPD were significantly associated with PCa risk in individuals with a family history of cancer. In those without a family history of cancer, the XRCC1 Trp/Trp genotype had a 4.64-fold higher risk of PCa (ORadj=4.64; 95% CI = 0.88–24.36; p-value = 0.07), while the XPD Gln/Gln had a 2.66-fold higher risk (ORadj=2.66; 95% CI = 0.88–8.10; p-value = 0.09). Both were statistically insignificant (p-value > 0.05).
Association of XRCC1 and XPD polymorphism with clinical parameters of patients
XRCC1 Arg194Trp or XPD Lys751Gln polymorphism was not significantly associated with tumor grade, tumor stage, DRE results, or UTIs (p-value > 0.05) (Table 6). However, the XRRC1 Trp/Trp genotype doubled the likelihood of high-grade tumors (Gleason score ≥ 7). Conversely, the XPD Lys/Gln and Gln/Gln genotypes increased the risk of having locally advanced or metastatic tumors by 3-fold but did not reach statistical significance. The XRCC1 Trp/Trp genotype increased hematuria risk by 12.57 times (p-value > 0.05). The homozygous mutant genotypes of XRCC1 and XPD increased the risk of higher serum creatine levels (Table 6). Having the mutant homozygous genotype of XRCC1 also increased the risk of having higher PSA levels in patients.
Mean serum creatinine levels (1.41 mg/dl vs. 1.19 mg/dl) and mean PSA levels (73.2 ng/ml vs. 44.2 ng/ml) were higher in patients with XRCC1 Trp/Trp genotype (p-value < 0.05) (Fig. 1). Accordingly, the homozygous mutant Gln/Gln genotype of the XPD gene was also significantly associated with higher creatinine levels in PCa patients (1.34 mg/dl vs. 1.17 mg/dl) (p-value = 0.04) (Fig. 2).
Discussion
Predictive factors can assist clinicians in tailoring treatments for individual patients. Recently, liquid biopsy has emerged as a promising alternative to invasive tissue biopsies, offering minimal invasiveness for prostate cancer diagnoses. Liquid biopsy is used to evaluate several biomarkers in body fluids, including circulating tumor cells (CTCs), extracellular vesicles (EVs), circulating tumor DNA (ctDNA) and RNA (ctRNA), as well as serum biomarkers like miRNA, AR-Vs, bone metabolism, neuroendocrine, and metabolic markers [36, 37]. However, the high cost, the discrepancies in definition, isolation methods and standardization pose challenges to validating these tests and being approved for clinical use. Given the ongoing difficulties in developing liquid biopsy biomarkers for clinical use, SNPs in DNA repair genes emerge as a promising area for developing noninvasive biomarkers with prognostic significance [36]. Identifying gene polymorphisms in PCa-associated pathways could improve early diagnosis, provide selective chemoprevention, and better understand the complexity of biological pathways involved in prostate carcinogenesis [38]. Our present study was a molecular population-based case-control study examining whether the XRCC1 codon 194 and XPD 751 polymorphisms influence the risk of PCa in the Bangladeshi population.
The XRCC1 Arg194Trp polymorphism resides in the linker region separating the NH2-terminal domain and the BRCT1 (BRCA1 C-terminus) domain, directly impacting enzymatic function [39]. However, the polymorphism does not cause a complete loss of protein function, as evidenced by the lethal condition observed in mice during embryonic development when XRCC1 is inactivated [40]. In our study, we found that PCa patients with XRCC1 codon 194 Trp/Trp genotype were 5.51 times at higher risk of PCa compared with controls than those carrying the Arg/Arg genotype. Consistent with our result, the genotype 194Trp/Trp was found to be associated with an increased risk of PCa in the Chinese population [19]. In a Japanese population, however, the 194 Arg/Trp genotype instead of the Trp/Trp genotype was significantly more prevalent in PCa patients than in controls [20]. The same study also discovered that having only one Trp allele (Arg/Trp and Trp/Trp genotypes) doubled the chance of developing prostate cancer. Their findings conflict with our dominant model analysis, which indicated that the genotypes Arg/Trp + Trp/Trp did not significantly increase the risk of PCa. However, the risk of PCa was raised when both copies of the Trp allele were present in comparison to the presence of Arg/Arg and Arg/Trp genotype as suggested by the recessive model (Table 3). Their findings may not apply to us as only patients with a family history of PCa only were included in their study. Contradictory data also exist where codon 194 did not show significant results with PCa risk [21, 22]. The lack of association of the studies may be attributed to variances in ethnicity, geographic and environmental factors. While it is possible that the 194 Arg/Trp or Trp/Trp genotype led to reduces DNA repair capacity and increased tumor progression, the extent to which this polymorphism impairs the DNA repair capacity is uncertain. Alternatively, Rahman & Zein et al. discovered that the wild-type Arg/Arg variation had a larger rise in mean sister chromatid exchange (SCE), a sign of gene damage, in response to the genotoxic chemical than the Arg/Trp varian [41]. These data imply a possible genomic instability associated with the wild-type variant and not with the mutant variants. Due to the lack of participants carrying the rare homozygous mutant Trp/Trp genotype, its effect on SCE could not be studied. Even so, the association between SCP and Arg/Arg genotype was observed in a small study population in their study and did not reach statistical significance. Wang and colleagues also reported significantly more chromosomal breaks in the wild-type Arg/Arg when compared to the homozygous/heterozygous variant group [42], suggesting the wild-type Arg allele, rather than the mutant Trp allele, may be more likely to increase cancer risk in the presence of certain environmental exposures. Since nearly all studies use the wild-type variant as the reference when comparing genotype associations, more functional studies evaluating the DNA repair capacity of XRCC1 codon 194 polymorphism are required to reach a consensus. The disparity in findings could be explained by the likelihood that other cancer SNPs of susceptibility genes in linkage disequilibrium with XRCC1 were present that contribute to higher cancer risk. Moreover, the influence of the polymorphism on DNA repair ability may differ based on the type and degree of DNA damaging insults. It is also possible that some of these outcomes are due to chance.
XPD, which maps close to XRCC1 on chromosome 19, is a probable cancer susceptibility gene in linkage disequilibrium with XRCC1. Therefore, the polymorphism in codon 751 affects the COOH-terminal domain of the XPD protein, an essential domain for TFIIH function. Mutations in the XPD gene may affect its interactions with other subunits, reducing XPD DNA helicase activity in TFIIH and defects in NER [43, 44]. However, it has been speculated that TFIIH transcriptional activity is relatively tolerant of amino acid alterations in the XPD protein. The 751 residue lies near the C-terminal end of the polypeptide, well past the point where disease-associated mutations have been observed [45]. Accordingly, we observed no statistically significant association between the XPD codon 751 genotype and PCa risk. A positive association was observed for XPD codon 751 Gln/Gln with PCa risk compared to wild type variant, but the link did not reach statistical significance (Table 3). Our findings align with some previous studies of XPD codon 751 polymorphism and PCa conducted in other countries [30, 31]. We also investigated the combined effect of the two SNPs that may have a collective impact on DNA repair outcomes (Table 4), but the results are not statistically significant. The lack of statistically significant association may be due to the selection of DNA repair genes of two different pathways. No healthy control was homozygous mutant in both genes, so its combined effect could not be compared.
Tobacco smoke is a rich source of various potent carcinogens and Reactive Oxygen Species (ROS), including Polycyclic Aromatic Hydrocarbon (PAHs), aromatic amines, and N-nitroso compounds which can produce DNA bulky adducts, base damage, and SSBs and Double Strand Breaks (DSBs) [46, 47]. The formation of PAH-DNA adducts in the prostate caused by cigarette smoke exposure has been demonstrated to differ between races, implying that genetic differences in the DNA repair genes contribute to individual susceptibility to cancer risk [47, 48]. The mutant homozygous Trp/Trp variant of XRCC1 and Gln/Gln variant of XPD was positively associated with PCa in smokers in this study, although not statistically significant (Table 5). While previous studies have also shown similar findings about XPD as our study [30], another study showed an association of XRCC1 Arg/Trp and Trp/Trp genotype with PCa risk in smokers [19], suggesting a strong gene-environment interaction in PCa susceptibility. In the same study, the XRCC1 Arg/Trp and Trp/Trp genotypes were found to raise PCa risk in individuals with no family history of cancer. Our study observed a statistically insignificant association between homozygous mutant genotypes of both genes with PCa in individuals without a family history of cancer. Inconsistencies in results across epidemiological studies could be explained by the varying distributions of genetic and environmental factors in the study cohort, which may influence the effects of any particular genetic variant.
Finally, we explored the association of the SNPs with clinical features and biochemical parameters with genotypes of PCa patients. We found that these polymorphisms did not affect tumor grade, tumor aggressiveness, DRE results, or UTIs (p-value > 0.05) (Table 6). Only the XRCC1 Trp/Trp genotype significantly increased hematuria risk by 12.6 times (p-value < 0.05). The XRCC1 Trp/Trp genotype was also significantly associated with higher serum creatinine levels and PSA compared to patients having the wild type (Arg/Arg) genotype (Fig. 1). The effect on serum creatine levels was also seen in patients with XPD Gln/Gln genotype (Fig. 2). Serum creatinine level is an independent predictor of high-risk PCa prognosis; low and high serum creatinine levels have a significantly higher prognostic risk of PCa [49]. Most patients (68.94%) included in our study had serum creatinine levels within the normal range of 0.7–1.3 mg/dL. Survival rates were not evaluated in our study, and the prognostic risk was, therefore, outside the scope of our investigation. Although the association of XRCC1 and XPD polymorphisms with PCa risk has been studied, it is unclear how SNPs affect clinical factors such as tumor stage, tumor grade, creatinine, and PSA levels. Hematuria and high creatinine levels are indicators of compromised kidney function. The association of XRCC1 homozygous wild-type Trp/Trp genotype with increased hematuria risk and serum creatinine levels suggests that patients with this genotype are particularly at risk of kidney function impairment. Vacher and colleagues [50] found that renal insufficiency is highly prevalent in PCa patients even when serum creatinine levels are normal. Even though most of the patients in our study had normal serum creatinine levels, the renal function of patients with the Trp/Trp genotype and PCa should be evaluated.
One of the merits of our study is that our control group included both hospital-based and non-hospital-based participants, eliminating the possibility of a selection bias. Moreover, by excluding tribal or other ethnic groups from the study groups, we achieved a homogeneous genetic background of the subject. We did, however, assess the reproducibility of results by repeating genotype assays on 10% of the samples chosen at random, and the replicates showed 100% concordance.
Conclusion
Our study suggested a prospective connection between genetic polymorphisms and increased PCa susceptibility. We found that the Trp/Trp variant at codon 194 of XRCC1 is associated with the PCA risk; however, XPD Lys751Gln gene polymorphism did not show a statistically significant association with an increased risk of developing PCa in the Bangladeshi population. XRCC1 Trp/Trp variant appears to be linked to hematuria risk, higher mean serum creatinine, and mean serum PSA levels. At the same time, XPD Gln/Gln is associated with only higher mean serum creatinine levels in PCa patients. Additional studies using a bigger sample size are required to explore the relationship of these polymorphisms with serum creatinine and PSA levels.
Data availability
No datasets were generated or analysed during the current study.
Abbreviations
- APEI:
-
Apurinic/apyrimidinic Endonuclease
- BER:
-
Base excision repair
- CI:
-
Confidence Interval
- DRE:
-
Digital Rectal Examination
- DSB:
-
Double Strand Breaks
- HDI:
-
Human Development Index
- NER:
-
Nucleotide Excision Repair
- OR:
-
Odds Ratio
- PAH:
-
Polycyclic Aromatic Hydrocarbon
- PCa:
-
Prostate Cancer
- PSA:
-
Prostate-Specific Antigen
- ROS:
-
Reactive Oxygen Species
- SCE:
-
Sister Chromatid Exchange
- SNP:
-
Single Nucleotide Polymorphisms
- SSB:
-
Single-Strand Breaks
- TFIIH:
-
Transcription factor II
- UTI:
-
Urinary Tract Infection
- XPD:
-
Xeroderma pigmentosum group D
- XRCC1:
-
X-ray cross-complementing gene 1
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global Cancer statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Attard G, Murphy L, Clarke NW, Sachdeva A, Jones C, Hoyle A et al (2023) Abiraterone acetate plus prednisolone with or without enzalutamide for patients with metastatic prostate cancer starting androgen deprivation therapy: final results from two randomised phase 3 trials of the STAMPEDE platform protocol. Lancet Oncol 24(5):443–456. https://doi.org/10.1016/s1470-2045(23)00148-1
Hassanipour S, Delam H, Arab-Zozani M, Abdzadeh E, Hosseini SA, Nikbakht HA et al (2020) Survival rate of prostate Cancer in Asian countries: a systematic review and Meta-analysis. Ann Glob Health 86(1):2. https://doi.org/10.5334/aogh.2607
Center MM, Jemal A, Lortet-Tieulent J, Ward E, Ferlay J, Brawley O et al (2012) International variation in prostate cancer incidence and mortality rates. Eur Urol 61(6):1079–1092. https://doi.org/10.1016/j.eururo.2012.02.054
Sharma R (2019) The burden of prostate cancer is associated with human development index: evidence from 87 countries, 1990–2016. Epma j 10(2):137–152. https://doi.org/10.1007/s13167-019-00169-y
Imtiaz H, Afroz S, Hossain MA, Bellah SF, Rahman MM, Kadir MS et al (2019) Genetic polymorphisms in CDH1 and Exo1 genes elevate the prostate cancer risk in Bangladeshi population. Tumour Biol 41(3):1010428319830837. https://doi.org/10.1177/1010428319830837
Abida W, Cyrta J, Heller G, Prandi D, Armenia J, Coleman I et al (2019) Genomic correlates of clinical outcome in advanced prostate cancer. Proc Natl Acad Sci U S A 116(23):11428–11436. https://doi.org/10.1073/pnas.1902651116
Li-Sheng Chen S, Ching-Yuan Fann J, Sipeky C, Yang TK, Yueh-Hsia Chiu S, Ming-Fang Yen A et al (2019) Risk prediction of prostate Cancer with single nucleotide polymorphisms and prostate specific Antigen. J Urol 201(3):486–495. https://doi.org/10.1016/j.juro.2018.10.015
Chen WS, Aggarwal R, Zhang L, Zhao SG, Thomas GV, Beer TM et al (2019) Genomic drivers of poor prognosis and Enzalutamide Resistance in Metastatic Castration-resistant prostate Cancer. Eur Urol 76(5):562–571. https://doi.org/10.1016/j.eururo.2019.03.020
Boussios S, Rassy E, Shah S, Ioannidou E, Sheriff M, Pavlidis N (2021) Aberrations of DNA repair pathways in prostate cancer: a cornerstone of precision oncology. Expert Opin Ther Targets 25(5):329–333. https://doi.org/10.1080/14728222.2021.1951226
Jiang M, Jia K, Wang L, Li W, Chen B, Liu Y et al (2020) Alterations of DNA damage repair in cancer: from mechanisms to applications. Ann Transl Med 8(24):1685. https://doi.org/10.21037/atm-20-2920
Nemec AA, Wallace SS, Sweasy JB (2010) Variant base excision repair proteins: contributors to genomic instability. Semin Cancer Biol 20(5):320–328. https://doi.org/10.1016/j.semcancer.2010.10.010
Caldecott KW (2019) XRCC1 protein; form and function. DNA Repair (Amst) 81:102664. https://doi.org/10.1016/j.dnarep.2019.102664
Mohrenweiser HW, Carrano AV, Fertitta A, Perry B, Thompson LH, Tucker JD et al (1989) Refined mapping of the three DNA repair genes, ERCC1, ERCC2, and XRCC1, on human chromosome 19. Cytogenet Cell Genet 52(1–2):11–14. https://doi.org/10.1159/000132829
Jalali C, Ghaderi B, Amini S, Abdi M, Roshani D (2016) Association of XRCC1 Trp194 allele with risk of breast cancer, and Ki67 protein status in breast tumor tissues. Saudi Med J 37(6):624–630. https://doi.org/10.15537/Smj.2016.6.13540
Chen S, Tang D, Xue K, Xu L, Ma G, Hsu Y et al (2002) DNA repair gene XRCC1 and XPD polymorphisms and risk of lung cancer in a Chinese population. Carcinogenesis 23(8):1321–1325. https://doi.org/10.1093/carcin/23.8.1321
Wang X, Zhang K, Liu X, Liu B, Wang Z (2015) Association between XRCC1 and XRCC3 gene polymorphisms and risk of thyroid cancer. Int J Clin Exp Pathol 8(3):3160–3167
Stern MC, Umbach DM, van Gils CH, Lunn RM, Taylor JA (2001) DNA repair gene XRCC1 polymorphisms, smoking, and bladder cancer risk. Cancer Epidemiol Biomarkers Prev 10(2):125–131
Zhu H, Jiu T, Wang D (2015) Impact of polymorphisms of the DNA repair gene XRCC1 and their role in the risk of prostate cancer. Pak J Med Sci 31(2):290–294. https://doi.org/10.12669/pjms.312.6653
HAMANO T, MATSUI H, OHTAKE N (2008) Polymorphisms of DNA repair genes, XRCC1 and XRCC3, and susceptibility to familial prostate cancer in a Japanese population. Asia-Pac J Clin Oncol 4(1):21–26. https://doi.org/10.1111/j.1743-7563.2008.00140.x
Langsenlehner T, Renner W, Gerger A, Hofmann G, Thurner EM, Kapp KS et al (2011) Association between single nucleotide polymorphisms in the gene for XRCC1 and radiation-induced late toxicity in prostate cancer patients. Radiother Oncol 98(3):387–393. https://doi.org/10.1016/j.radonc.2011.01.021
Huang SP, Huang CY, Wang JS, Liu CC, Pu YS, Yu HJ et al (2007) Prognostic significance of p53 and X-ray repair cross-complementing group 1 polymorphisms on prostate-specific antigen recurrence in prostate cancer post radical prostatectomy. Clin Cancer Res 13(22 Pt 1):6632–6638. https://doi.org/10.1158/1078-0432.Ccr-07-1437
Petruseva IO, Evdokimov AN, Lavrik OI (2014) Molecular mechanism of global genome nucleotide excision repair. Acta Naturae 6(1):23–34
Sameer AS, Nissar S (2018) XPD-The Lynchpin of NER: molecule, Gene, polymorphisms, and role in colorectal carcinogenesis. Front Mol Biosci 5:23. https://doi.org/10.3389/fmolb.2018.00023
Beck BD, Hah DS, Lee SH (2008) XPB and XPD between transcription and DNA repair. Adv Exp Med Biol 637:39–46. https://doi.org/10.1007/978-0-387-09599-8_5
Du Y, He Y, Mei Z, Qian L, Shi J, Jie Z (2016) Association between genetic polymorphisms in XPD and XRCC1 genes and risks of non-small cell lung cancer in East Chinese Han population. Clin Respir J 10(3):311–317. https://doi.org/10.1111/crj.12218
Samson M, Singh SS, Rama R, Sridevi V, Rajkumar T (2011) XPD Lys751Gln increases the risk of breast cancer. Oncol Lett 2(1):155–159. https://doi.org/10.3892/ol.2010.220
Gao W, Romkes M, Zhong S, Nukui T, Persad RA, Smith PJ et al (2010) Genetic polymorphisms in the DNA repair genes XPD and XRCC1, p53 gene mutations and bladder cancer risk. Oncol Rep 24(1):257–262. https://doi.org/10.3892/or_00000854
Gan Y, Li XR, Chen DJ, Wu JH (2012) Association between polymorphisms of XRCC1 Arg399Gln and XPD Lys751Gln genes and prognosis of colorectal cancer in a Chinese population. Asian Pac J Cancer Prev 13(11):5721–5724. https://doi.org/10.7314/apjcp.2012.13.11.5721
Sobti RC, Berhane N, Melese S, Mahdi SA, Gupta L, Thakur H et al (2012) Impact of XPD gene polymorphism on risk of prostate cancer on north Indian population. Mol Cell Biochem 362(1–2):263–268. https://doi.org/10.1007/s11010-011-1152-3
Bau DT, Wu HC, Chiu CF, Lin CC, Hsu CM, Wang CL et al (2007) Association of XPD polymorphisms with prostate cancer in Taiwanese patients. Anticancer Res 27(4c):2893–2896
Dhillon VS, Yeoh E, Fenech M (2011) DNA repair gene polymorphisms and prostate cancer risk in South Australia–results of a pilot study. Urol Oncol 29(6):641–646. https://doi.org/10.1016/j.urolonc.2009.08.013
Chacko P, Rajan B, Joseph T, Mathew BS, Pillai MR (2005) Polymorphisms in DNA repair gene XRCC1 and increased genetic susceptibility to breast cancer. Breast Cancer Res Treat 89(1):15–21. https://doi.org/10.1007/s10549-004-1004-x
Mitra AK, Singh N, Garg VK, Chaturvedi R, Sharma M, Rath SK (2009) Statistically significant association of the single nucleotide polymorphism (SNP) rs13181 (ERCC2) with predisposition to squamous cell carcinomas of the Head and Neck (SCCHN) and breast cancer in the north Indian population. J Exp Clin Cancer Res 28(1):104. https://doi.org/10.1186/1756-9966-28-104
Nairuz T, Bushra YU, Kabir Y (2021) Effect of XPD and TP53 gene polymorphisms on the risk of platinum-based Chemotherapy Induced Toxicity in Bangladeshi Lung Cancer patients. Asian Pac J Cancer Prev 22(12):3809–3815. https://doi.org/10.31557/apjcp.2021.22.12.3809
Crocetto F, Russo G, Di Zazzo E, Pisapia P, Mirto BF, Palmieri A et al (2022) Liquid biopsy in prostate Cancer Management-Current challenges and Future perspectives. Cancers (Basel) 14(13). https://doi.org/10.3390/cancers14133272
Saxby H, Mikropoulos C, Boussios S (2020) An update on the prognostic and predictive serum biomarkers in metastatic prostate Cancer. Diagnostics (Basel) 10(8). https://doi.org/10.3390/diagnostics10080549
Salvi S, Conteduca V, Gurioli G, Calistri D, Casadio V, De Giorgi U (2016) Impact of candidate genetic polymorphisms in prostate Cancer: an overview. Mol Diagn Ther 20(1):1–12. https://doi.org/10.1007/s40291-015-0169-9
Ceja Galvez HR, Salazar Flores J, Torres Sanchez ED, Rojas Bravo D, Reyna Villela MZ, Reyes Uribe E (2021) Genetic profile for the detection of susceptibility to poisoning by exposure to pesticides. Ann Agric Environ Med 28(2):208–213. https://doi.org/10.26444/aaem/136362
Tebbs RS, Flannery ML, Meneses JJ, Hartmann A, Tucker JD, Thompson LH et al (1999) Requirement for the Xrcc1 DNA base excision repair gene during early mouse development. Dev Biol 208(2):513–529. https://doi.org/10.1006/dbio.1999.9232
Abdel-Rahman SZ, El-Zein RA (2000) The 399Gln polymorphism in the DNA repair gene XRCC1 modulates the genotoxic response induced in human lymphocytes by the tobacco-specific nitrosamine NNK. Cancer Lett 159(1):63–71. https://doi.org/10.1016/s0304-3835(00)00532-2
Wang Y, Spitz MR, Zhu Y, Dong Q, Shete S, Wu X (2003) From genotype to phenotype: correlating XRCC1 polymorphisms with mutagen sensitivity. DNA Repair (Amst) 2(8):901–908. https://doi.org/10.1016/s1568-7864(03)00085-5
Coin F, Marinoni JC, Rodolfo C, Fribourg S, Pedrini AM, Egly JM (1998) Mutations in the XPD helicase gene result in XP and TTD phenotypes, preventing interaction between XPD and the p44 subunit of TFIIH. Nat Genet 20(2):184–188. https://doi.org/10.1038/2491
Coin F, Bergmann E, Tremeau-Bravard A, Egly JM (1999) Mutations in XPB and XPD helicases found in xeroderma pigmentosum patients impair the transcription function of TFIIH. Embo j 18(5):1357–1366. https://doi.org/10.1093/emboj/18.5.1357
Clarkson SG, Wood RD (2005) Polymorphisms in the human XPD (ERCC2) gene, DNA repair capacity and cancer susceptibility: an appraisal. DNA Repair (Amst) 4(10):1068–1074. https://doi.org/10.1016/j.dnarep.2005.07.001
Barnes JL, Zubair M, John K, Poirier MC, Martin FL (2018) Carcinogens and DNA damage. Biochem Soc Trans 46(5):1213–1224. https://doi.org/10.1042/bst20180519
Nair J, Ohshima H, Nair UJ, Bartsch H (1996) Endogenous formation of nitrosamines and oxidative DNA-damaging agents in tobacco users. Crit Rev Toxicol 26(2):149–161. https://doi.org/10.3109/10408449609017928
Nock NL, Tang D, Rundle A, Neslund-Dudas C, Savera AT, Bock CH et al (2007) Associations between smoking, polymorphisms in polycyclic aromatic hydrocarbon (PAH) metabolism and conjugation genes and PAH-DNA adducts in prostate tumors differ by race. Cancer Epidemiol Biomarkers Prev 16(6):1236–1245. https://doi.org/10.1158/1055-9965.Epi-06-0736
Gu X, Wu J, Liu X, Hong Y, Wu Y, Tian Y (2022) Role of serum creatinine levels in prognostic risk stratification of prostate Cancer patients. Med Sci Monit 28:e937100. https://doi.org/10.12659/msm.937100
Launay-Vacher V, Ayllon J, Janus N, Spano JP, Ray-Coquard I, Gligorov J et al (2009) Drug management of prostate cancer: prevalence and consequences of renal insufficiency. Clin Genitourin Cancer 7(3):E83–E89. https://doi.org/10.3816/CGC.2009.n.029
Acknowledgements
The Authors acknowledge the support of the physicians and nurses of Bangladesh Institute of Research and Rehabilitation in Diabetes, Endocrine and Metabolic Disorders (BIRDEM) General Hospital, Bangabandhu Sheikh Mujib Medical University (BSMMU), and Dhaka Medical College Hospital (DMCH), Dhaka, Bangladesh for their technical assistance during blood collection and patient counseling. We also thank all the study subjects for participating in this study. The authors thank Dr. AyatunNesa, Department of Laboratory Medicine, BIRDEM General Hospital, Dhaka, Bangladesh. for collecting patient samples. The authors are also grateful to all the Institutional Ethical Review Committees members of the Department of Biochemistry and Molecular Biology, University of Dhaka, Bangladesh, for approving the study protocol. This research did not receive any funding.
Funding
This research did not receive any funding.
Author information
Authors and Affiliations
Contributions
Y.K. and M.H. conceptualized the research idea and provided overall supervision and project administration throughout the study. N.A. designed the methodology, curated the data, engaged in the investigation process, and contributed to data visualization. M.A.I. refined the methodology, secured essential resources, conducted formal analysis and validated the results. N.A. wrote the original draft. All authors reviewed, edited and revised the final draft.
Corresponding author
Ethics declarations
Informed consent and ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by The Ethical Review Committees of the Department of Biochemistry and Molecular Biology, University of Dhaka, Bangladesh.
Consent to participate
Informed consent was obtained from all participants before the sample and data collection.
Consent to publish
The authors affirm that human research participants provided informed consent for publication of their clinical data.
Competing interests
The authors declare that they have no conflicting interests. All authors declare they have no financial interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ahmed, N., Islam, M.A., Hossain, M.M. et al. XRCC1 and XPD polymorphisms: clinical outcomes and risk of prostate cancer in Bangladeshi population. Mol Biol Rep 51, 893 (2024). https://doi.org/10.1007/s11033-024-09707-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09707-y