Abstract
Background
Bacterial leaf blight (BLB) is one of the major biotic stress in rice cultivation. Management techniques, such as the development of BLB-resistant cultivars, are required to lessen the severity of the disease attack and yield losses. Pratikshya was selected in the present investigation as the recipient parent, as it is one of the popular high-yielding rice varieties of Odisha, India, which is having excellent grain as well as cooking quality. However, Pratikshya is highly susceptible to BLB which is prevalent in Eastern Indian region.
Methods and results
Three major BLB resistance genes xa5, xa13, and Xa21 from the donor source Swarna MAS (CR Dhan 800) were attempted to introduce into Pratikshya through a marker-assisted backcross breeding program. Those markers closely linked to the target genes were employed for foreground selection in the segregating generations till BC2F3. In each backcross generation, progenies containing all three targeted resistance genes and phenotypically more similar to the recipient parent, Pratikshya were selected and backcrossed. Screening of 1,598 plants of the BC2F2 population was conducted against BLB using Xoo inoculum and 35 resistant plants similar to Pratikshya were carried forward to the next generation. In the BC2F3 generation, 31 plants were found to possess all the three resistance genes. For background selection of plants carrying resistance genes 45 polymorphic SSR markers were employed. Evaluation of the pyramided lines at BC2F4 generation exhibited that, most pyramided lines were similar to Pratikshya in terms of morphological features and yield parameters, and some lines were superior to the recurrent parent in terms of morphological features and yield parameters.
Conclusion
The three-gene pyramided lines showed a high level of resistance to BLB infection and are anticipated to offer a significant yield advantage over the recipient parent Pratikshya. The pyramided lines can further be used for multi-location trial, so as to be released as a variety or can be used as a potential donor for BLB resistance genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa L.) is one of the most important cereals, serving as a principal source of nutrition to half of the global population, and particularly in India, it accounts a major portion of total food grain production [1]. India must produce 120 million tonnes of rice by 2030 to maintain self-sufficiency and to fulfil future food demands, which must be accomplished with limited land, water, labour, and using minimum amount of chemicals (i.e., fertilizers and pesticides etc.), as well as a continuous fight against developing disease, pests and the possible harmful consequences of climate change [1, 2]. Biotic stresses like disease and insect pests cause major yield loss in rice varieties across the globe [3]. Among many biotic stresses that affect rice production, bacterial leaf blight (BLB) is one of the most dreadful disease, causing yield reduction upto 74–81% depending on weather, location, and rice cultivar used for cultivation [4]. In both tropical and temperate rice-growing zones, the gram-negative bacterium Xanthomonas oryzae pv. oryzae (Xoo) causes bacterial leaf blight (BLB) disease [5]. During the maximum tillering stage, the BLB infection occurs, resulting in water-soaked lesions on the leaves that steadily enlarge and finally cause the rice plant to wilt [5]. Chemical pesticides and antibiotic sprays do not work against BLB [6], however antibiotics are utilized to control bacterial disease such as Pseudomonas spp., and Xanthomonas campestris [7, 8]. Therefore, host plant resistance represents the most practical and cost-effective strategy for disease management [9,10,11,12,13]. Till date, 47 genes have been identified, that provide resistance from BLB, of which many of them have been incorporated into high-yielding, popular rice cultivars across the globe [14,15,16]. The deployment of resistant cultivars with a single major resistance gene has been proven significant due to the development of novel strain of Xoo due to mutation [17].
Hence, the development of varieties with several resistance genes offer a viable alternative for broad-spectrum, long-term resistance against BLB in rice [18]. To achieve the goal, multiple resistance genes can be stacked or pyramided into the elite genetic background as a breeding strategy [19]. Meanwhile, marker-assisted selection (MAS) is a more effective and simpler approach for gene introgression than conventional breeding [19, 20]. The utilisation of MAS holds significant potential in assisting plant breeders to accomplish their objectives. However, its influence on the development of plant varieties has been limited [20]. In order to fully exploit the potential of MAS, it is crucial to establish a higher level of integration between MAS and breeding programmes. Additionally, it is important to understand the existing barriers and develop suitable methods to overcome them. The utilisation of the benefits of MAS in comparison to traditional breeding methods has the potential to significantly influence the improvement of cereal crops [20, 21].
Therefore, the present experiment was conducted to transfer valuable genes in rice variety Pratikshya for BLB resistance using MAS with the objectives: validation of parental lines for BLB resistance genes by using molecular markers, development of F1 and the backcross populations and marker-assisted selection of lines possessing BLB resistance genes xa5, xa13, and Xa21 in the populations F1, BC1F1, BC2F1, BC2F3 (foreground selection), phenotypic screening of segregating population (BC2F2) for BLB resistance through artificial inoculation, marker-assisted selection of lines possessing background of recurrent parent Pratikshya in the population BC2F3 (background selection), and evaluation of pyramided lines for yield and agro-morphological characters in BC2F4 population.
Materials and methods
Plant materials
Pratikshya is a popular high-yielding rice variety of Odisha, India with good grain and cooking quality released by Odisha University of Agriculture and Technology (OUAT), Bhubaneswar, which is suitable for late sown rainfed medium land, moderately resistance to brown spot, sheath rot, sheath blight, leaf folder, white backed plant hopper, gall midge, stem borer, but highly susceptible to BLB. The donor parent for BLB resistance employed in the current crossing program was Swarna MAS (CR Dhan 800) released by ICAR-National Rice Research Institute (ICAR-NRRI), Cuttack, India that carries three BLB resistance genes, i.e., xa5, xa13, Xa21 in the genetic background of Swarna variety [22].
Hybridization and marker-assisted selection
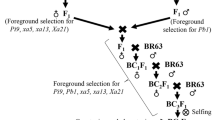
A step-by-step marker-assisted backcross breeding approach was implemented as shown in Fig. 1 for the effective introgression of xa5, xa13, and Xa21 genes into Pratikshya. In Kharif season, 2017, F1 seeds were successfully generated by crossing recipient parent Pratikshya with donor parent Swarna MAS. In the shallow pots located within the net house, F1 seeds were planted in Rabi season 2017. True hybridity was checked in the F1 generation plants using the three SSR markers (Table 1) linked to the xa5, xa13, and Xa21 genes. The selected true F1 plants were hybridized with the recurrent parent, Pratikshya, during Rabi season 2017 to produce BC1F1 seeds. The BC1F1 plants were then subjected to both foreground selection in Kharif season 2018 using MAS and phenotypic selection for agronomic similarity with Pratikshya.
Backcrossed seeds from selected BC1F1 plants were grown in the next generation as BC2F1 plants in Rabi season 2018. Selected plants from BC2F1 carrying the xa5, xa13, and Xa21 genes were allowed to self-pollinate and then carried forward to BC2F2 generation during Kharif season, 2019. Resistant plants from BC2F2 generation were selected by phenotypic screening using clip inoculation technique of Xoo inoculum (Xanthomonas oryzae pv. oryzae brought from Crop Protection Division, National Rice Research Institute, Cuttack, India) by following a standard method [23], and by assessing the similarity of agronomic characteristics to Pratikshya. During Rabi season, 2019, plants from the BC2F3 generation were used for foreground selection and selected plants bearing all three resistance genes were subjected to background selection with markers showing parental polymorphism, and phenotypic selection for agronomic similarity with Pratikshya. Plants having all three resistance genes and good recipient parent genome recovery percentage were carried forward to BC2F4 generation during Kharif season, 2020 for agronomic trials.
Molecular characterization and SSR markers analysis
Total genomic DNA was isolated by the modified CTAB method of DNA extraction for rice [24], and quantified on Nanodrop - Spectrophotometer (NANODROP 2000c), by diluting in 1X TE buffer to a final concentration of around 50 ng/µl. The PCR reaction mixture for foreground selection of xa5, xa13, and Xa21 consisted of 50 ng of genomic DNA, 10 mM of each primer, 10 mM of each dNTP, 10x PCR buffer, 3U Taq polymerase in a volume of 10 µl. The amplified products were subjected to electrophoresis on a 1.5% agarose gel for xa13 prom primer (for xa13 gene), pTA248 primer (for Xa21 gene), 3% for RM122 primer (for xa5 gene) (Table 1), and visualized on a gel documentation system (Bio-Rad Laboratories Inc., USA). To confirm the presence of the target genes, foreground selection was conducted till BC2F3 generation. However, 45 polymorphic SSR markers after the parental survey (Table 2) were employed for background selection to detect parental genome recovery in the pyramided lines [25].
The SSR bands from the gel photograph of BC2F3 generation were manually scored as 1 (if band present) and 0 (if band absent) to obtain the binary data for molecular analysis. Missing data were verified twice by repeating the genotyping process. The genetic distance between pyramided lines was calculated through the construction of the distance matrix using Jaccard’s similarity coefficient from the binary data to deduce the genetic relationships between the pyramided lines and two parents, by constructing a dendrogram by following sequential UPGMA (unweighted pair group method with arithmetic mean) using software package TASSEL 5.0 [26].
Screening for bacterial blight resistance and morphological characterization
During Kharif season, 2017, field screening for BLB resistance was conducted by utilizing the inoculum of Xanthomonas oryzae pv. oryzae. Each Plant artificially inoculated by the leaf clip inoculation method, in which, the top leaves of each plant were clipped at the maximum tillering stage and then inoculated [23]. The symptoms started to develop five to six days after the inoculation, and the observation period was between 14 and 21 days after inoculation. The plants with a score of 1 were regarded as resistant, those with a score of 3 as moderately resistants, those with a score of 7 as susceptible, and a score of 9 as highly susceptible [27]. The observations were recorded for percentage of diseased leaf area (DLA) followed by a standard evaluation system [27]. However, a multi-location trail was conducted at Rice Research Station, OUAT, Bhubaneswar, India (GPS co-ordinates: latitude NL 20° 15′ 55″, longitude of EL 85° 48′ 33″), and at agricultural farm, OUAT, Bhubaneswar, India (GPS co-ordinates: latitude of 20° 16’ 09.3’’ N, longitude of 85° 47’ 29.0’’ E), for proper validation of the screening.
Thirty-days-old seedlings of BC2F4 generation carrying BLB resistance genes were transplanted along with the donor and recipient parents in the main field of Rice research station, OUAT, Bhubaneswar with 20 cm × 15 cm spacing. The crop was successfully grown using conventional agronomic practices in Kharif season, 2020. The phenotypic trait observations were recorded for 10 plants in three replications and the replicated data were used to calculate mean, coefficient of variation (CV), and critical difference (CD). The observations were recorded for yield component characters viz., days to 50% flowering, plant height (cm), total number of tillers per plant, number of productive tillers per plant, panicle length (cm), filled grains/panicle, 1000-grain weight (g), and seed yield per plant (g). Data analyses were carried out with the use of the software, Grapes version 1.0.0 [28].
Results
Molecular validation of parental lines
The resistant parent (Swarna MAS) and susceptible parent (Pratikshya) were validated for the presence of BLB resistance genes xa5, xa13, and Xa21 with help of gene-linked markers RM122, xa13 prom, and pTA248, respectively. Genomic DNA from both parents was amplified using the above-mentioned three SSR markers and then parental polymorphism was revealed from gel electrophoresis. The resistant allele of xa5 showed a band at 200 bp using the RM122 marker. The resistant allele of the xa13 gene generated a fragment at 500 bp, whereas the resistant allele of the Xa21 gene was found in the resistant parent Swarna MAS at 1000 bp (Fig. 2).
Pyramiding of bacterial leaf blight resistance genes
During each generation from F1 to BC2F3 (except for BC2F2, due to more number of population and use of clip inoculation technique), foreground selection was conducted to select plants having all three resistance genes (positive plants) and only those plants were advanced to the next generation. The hybridity of F1 plants was assessed using molecular markers, and it was determined that out of a total of 26 F1 plants, 18 exhibited characteristics consistent with being true F1 plants. True F1 plants were backcrossed with the recipient parent Pratikshya to generate BC1F1 seeds. In the BC1F1 generation, 117 plants were grown and out of these, 58 plants were identified to possess all three resistance genes. These 58 BC1F1 plants positive for resistance genes were backcrossed with recurrent parent Pratikshya. Of the 170 BC2F1 plants grown, 40 were detected to possess three-resistance genes. Thus only these 40 BC2F1 plants were allowed to self-pollinate and advanced to BC2F2 generation (Fig. 3).
In BC2F2 generation phenotypic screening procedures were followed instead of the use of molecular screening procedures to identify resistant plants. Selected 35 BC2F2 plants were grown as a total of 35 lines in BC2F3 generation and plants from those lines homozygous for three resistance genes combinations were identified (Fig. 2). Out of the 35 lines only thirty-one plants were found to possess all the three genes and hence were subjected to background selection. The background selection of these 31 BC2F3 plants with forty-five polymorphic SSR markers showed genome recovery of Pratikshya in the range of 64.44–93.33%. Out of 31 plants, 5 were showed genome recovery from 91.11 to 93.33% (Table 3; Fig. 4). Thereafter, thirty-one BC2F3 plants were allowed to self-pollinate to obtain plants for BC2F4 population.
Background selection of plants from BC2F3generationwith different polymorphic SSR markers. The amplified fragments with respect to marker RM26885 is present in Pratikshya, plant no. 2, 3, 4, 5, 7, 8, 9, 10, 11, 12, 13, 14, 17, 18, 19, 20, 21, 22, 23, 24, 28, 29, 30, 31. The amplified fragments with respect to marker RM27180 is present in Pratikshya, plant no, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 17, 18, 19, 21, 22, 23, 27, 28, 29, 31.The amplified fragments with respect to marker RM27446 is present in Pratikshya, plant no 1, 2, 3, 4, 5, 6, 7, 8, 9, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31. Lane 1- 100 bp ladder, Lane 2- Pratikshya (P). However, the best improved line no. 19 - OR 2772-19 gave a seed yield/plant of 65.5 g
Genetic similarity of the pyramided lines with the recipient parent using SSR markers
The dendrogram was constructed from SSR data grouped the 31 three-gene pyramided lines along with both parents into two major clusters with Swarna MAS in cluster I and the remaining 31 pyramided lines, including Pratikshya, in cluster II along with the similarity matrix of the pyramided lines (Fig. 5; Table 4). Cluster II was further divided into two sub-groups, cluster II-A and cluster II-B (Table 5). Cluster II-A was further subcategorized into two sub-groups, cluster II-Aa, and cluster II-Ab. Cluster II-Aa consists of pyramided lines OR 2772-4, OR 2772-28, OR 2772-73, OR 2772-31, OR 2772-2, OR 2772-20, OR 2772-15, OR 2772-74, OR 2772-77, OR 2772-92, OR 2772-1, OR 2772-17, OR 2772-32, OR 2772-90, OR 2772-16, OR 2772-3, OR 2772-19, OR 2772-21, OR 2772-27 and OR 2772-30, OR 2772-33, OR 2772-54, OR 2772-64, OR 2772-51, OR 2772-65, OR 2772-62, OR 2772-10, and OR 2772-93, while Cluster II-Ab contains OR 2772-13 and OR 2772-18. Cluster II-B consists of OR 2772-76.
Phenotypic screening of the BC2F2 against bacterial leaf blight
The recipient parent Pratikshya showed a disease score of 7 indicating a greater level of susceptibility to BLB. Swarna MAS displayed a high level of resistance to the infection with a disease score of 0. The disease score of BC2F2 plants ranged from 0 to 1 showing resistance to the disease. Out of all the plants of the BC2F2 population (1598 plants), 1412 were observed to be resistant and 186 were susceptible, across the multi-location trail.
Evaluation of pyramided lines for agro-morphological traits in BC2F4 generation
The mean values of eight agronomic traits viz., days to 50% flowering, plant height (cm), number of tillers per plant, number of productive tillers per plant, panicle length (cm), number of filled grains per panicle, seed yield per plant (g) and 1000 grain weight (g) are presented in Table 6. From the statistical analysis, it was noticed that a significant difference was observed between the pyramided lines and both of the parents for all the characters except for total tillers, though, the difference was very less. The pyramided line OR 2772-19 displayed the highest seed yield per plant of 65.5 g. The average seed yield per plant of pyramided lines in the field was found to be 42.37 g, and the yield is varying due to the change in their genotyping composition after introgression.
The recurrent parent, Pratikshya, recorded a mean seed yield of 44 g/plant, while the donor parent (Swarna MAS) recorded 49.4 g/plant. Among pyramided lines, ten lines attained more seed yield per plant than Pratikshya. For panicle length, among the pyramided lines, OR 2772-21 had the longest panicle length (34.7 cm) followed by OR 2772-20 (28.9 cm). The parent Pratikshya displayed a panicle length of 24.5 cm. Out of thirty-one pyramided lines, nineteen lines produced longer panicles, while ten lines produced smaller panicles than Pratikshya. The highest plant height of 109.6 cm was observed in OR 2772-76 followed by 107 cm in OR 2772-27, while the shortest plant height of 88.3 cm was observed in OR 2772-13. Majority of the pyramided lines attained more grain weight than the Pratikshya, while OR 2772-92 showed a similar grain weight with the recurrent parent. From these observations, it was concluded that agro-morphological features play a significant part in the yield of the plant.
Discussion
Pratikshya (Parentage: Swarna × IR64) is late-maturing, popular high-yielding rice variety in Eastern India [25]. It can be harvested in 146 days, and has a high degree of adaptability, and it creates opportunities for cropping system intensification in the coastal zone of India by earlier establishment of Rabi season crops [29]. However, the main limitation of Pratikshya is its susceptibility to BLB, which significantly reduces yield, despite the fact that it has potential for high yield with a desirable plant type. BLB acts as a biotic constraint in the decline of productivity in South-East Asian countries. The extent of severity of 10-20% annual reduction in rice production worldwide caused by BLB in rice demands the development of effective management strategies [30]. However, in order to prevent the spread of BLB, it has been found that host plant resistance is the most efficient environmental friendly method [31]. The incorporation of resistance genes in combination remains as an effective strategy for managing BLB and providing durable resistance [32]. In contrast to conventional phenotypic evaluation, marker-assisted backcross breeding permits for the selection of recessive alleles, selection of desirable plants at the seedling stage before the formation of a visible phenotype, and the pyramiding of important traits into a single genetic background. The application of molecular markers allow for the genetic categorization of progeny at each generation and speeds up the selection process [33].
Pyramiding multiple resistance genes with potential characteristics into a single genotype through MAS can improve the efficiency of generating new crop varieties exhibiting disease resistance, as well as other desirable traits, however, it is crucial for the maintenance of yield stability in the rice variety Pratikshya [34]. Meanwhile, in several studies, this approach has been successfully employed to introduce resistance genes into the elite genetic background [11, 30, 31, 34]. The BLB resistance genes xa5, xa13, and Xa21 have been successfully incorporated into different genetic backgrounds of rice [11, 35].
Hence, these three resistance alleles covering three chromosome regions were selected to transfer into the popular variety Pratikshya in the present investigation. Out of 46 identified BLB resistance genes, three major R genes xa5, xa13, and Xa21 have been used in the current study. However, previous research has successfully cloned and described all three of these resistances, i.e., Xa21 resistance gene is dominant and has been cloned with a high level of resistance from many Xoo strains [36]. The gene xa5 is in the recessive gene category and codes an alternative form of transcription factor cIIa [37]. The xa13 is a recessive gene that originated because of a mutation in the promoter region of a gene which is homolog to nodulin MtN3 [38].
The pyramided lines carrying these three genes in the present study exhibited higher degrees of resistance than both of the parents. This aligns with the study of some scientists elucidating the contribution of synergistic action of resistance genes in achieving increased levels of resistance [39, 40]. These three R genes are resistant to all the prevalent races of BLB pathogen in the coastal region of East India. However, previously many workers have used these three genes for the transfer of BLB resistance to many traditional varieties of Odisha, India, as well as in other parts of the country [22, 41].
Meanwhile, due to the masking effect of dominant genes over recessive genes, combining them at the same time might be a challenge via phenotypic selection. In such cases, molecular markers for both recessive and dominant resistance genes can assist in the identification and selection of desirable plants with multiple resistance genes [41, 42]. However, pyramided lines with the highest recovery of the recipient parent genome were found with the help of marker-assisted background selection, in the present study. In the BC2F3 generation, the highest genome recovery rate was 91.11–93.33% in five pyramided lines. The low recovery of the background of recurrent parent observed in a few lines can be attributed to linkage drag, referring to the reduction in fitness in cultivars due to deleterious genes introduced along with the beneficial gene during backcrossing [42]. It is possible that the functional portion of the genome is not recovered since SSR markers are often used to target non-coding and heterochromatic areas of chromosomes [43].
The phenotypic selection was also used in each generation to identify plants that are similar to Pratikshya in terms of agronomic characteristics, but also better from it. However, the functionally expressed area of the genome is an indirect target of phenotypic selection and helps to accelerate the recovery of the recurrent parent phenotype [41, 42]. Meanwhile, the linkage drag is kept to a minimum in this study by using genetically related parents in the crossing program. Swarna MAS (CR Dhan 800) is a derivative of the variety Swarna, which is a highly adaptable variety. Previous researchers concluded that using a highly adaptable variety as a donor parent results in better performance and less linkage drag than using a wild species or landrace as a donor parent [42]. However, the parentage of the recipient parent is Swarna and IR64, and the shared ancestry of Pratikshya with donor parent Swarna MAS speeds up the recovery of the recipient parent genome [42, 43].
Yield and agro-morphological parameters from thirty-one pyramided lines, Pratikshya and Swarna MAS showed that the pyramided lines have the outstanding yielding ability of recipient parent and resistance to BLB in the present study. The analysis of variance for yield and agro-morphological traits displayed that the mean for all characters were at par with Pratikshya in most of the lines, many lines performed to a greater degree in terms of seed yield per plant, and the majority of the lines were close to the recurrent parent, in the present study. However, various studies have revealed that the high seed yield of some pyramided lines may be attributable to the transmission of yield features from the donor source to the recipient parent [30, 34]. Meanwhile, in the background of Pratikshya, pyramided lines had no antagonistic effect due to the introgression of resistance genes. High degrees of resistance to BLB, the lack of a penalty on yield, and good recovery of recurrent parent genome point towards a successful selection approach at both the molecular and phenotypic levels in the preset study. However, some scientists also found similar results from their study [11, 22, 34].
Due to the inadequacy of a single resistance gene in providing resistance against different pathogen races of BLB in Odisha, India, a wide range of resistance against BLB is critical. However, some scientists, reached a similar conclusion, observing a broad spectrum of resistance against bacterial leaf blight when multiple genes were introgressed into an elite genetic background rather than a single gene [44]. However, in the present study, pyramiding three resistance genes in a single genetic background was found to be an effective barrier against prevalent pathogen races of BLB in Odisha, India conditions, and expected to provide durable resistance.
Conclusion
Marker-assisted foreground and background selection along with phenotypic selection was found to be the best way to breed resistant varieties, making the breeding program much more effective. The current research concludes that, the marker-assisted backcross breeding program was successful in introducing three BLB resistance genes (xa5, xa13, and Xa21) into the high-yielding rice variety Pratikshya. The resulting pyramided lines exhibited a high level of resistance to BLB infection and were similar to Pratikshya in terms of morphological features and yield parameters. This study suggests that, the three-gene pyramided lines could offer a significant yield advantage over the recipient parent Pratikshya. The pyramided lines can further be used for testing at multilocation, so as to be released as a variety or can be used as a potential donor for BLB resistance genes. The present study also suggests the deployment of multiple resistance genes can be more effective than the use of a single resistance gene. The BLB pyramided lines are proven to be successful in providing long-lasting and durable resistance in the genetic background of Pratikshya, which is an accomplishment for developing BLB-resistant pyramided lines bearing three resistance genes.
Data Availability
All data generated or analysed during this study are included in this published article.
References
Rahman H, Dakshinamurthi V, Ramasamy S, Manickam S, Ashok Kumar Kaliyaperumal AK, Raha S, Panneerselvam N, Ramanathan V, Nallathambi J, Sabarippan R, Raveendran M (2018) Introgression of Submergence Tolerance into CO 43, a Popular Rice Variety of India, through marker-assisted Backcross breeding. Czech J Genet Plant Breed. 54:101–108
Khush GS (2005) What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol. 59:1–6
Muduli L, Dash M, Das Mohapatra S, Mohapatra KK, Nayak HS, Bastia DN, Pradhan B, Tripathy SK, Jena RC, Pradhan SK (2023) Phenotypic and genotypic assessment of elite rice varieties for brown plant hopper (Nilaparvata lugens Stål.) Resistance. Cereal Res Commun. 1–13
Srinivasan B, Gnanamanickam S (2005) Identification of a new source of resistance in wild rice, Oryza rufipogon to bacterial blight of rice caused by indian strains of Xanthomonas oryzae pv. Oryzae. Curr Sci. 88:1229–1231
Mew TW (2003) Current status and future prospects of research on bacterial blight of rice. Annu Rev Phytopathol. 25:359–382
Lee K, Rasabandith S, Angeles E, Khush G (2003) Inheritance of resistance to bacterial blight in 21 cultivars of rice. Phytopathol. 93:147–152
McManus PS, Stockwell VO, Sundin GW, Jones AL (2002) Antibiotic use in plant agriculture. Annu Rev Phytopathol. 40:443–465. https://doi.org/10.1146/annurev.phyto.40.120301
Ahmad N, Joji RM, Shahid M (2023) Evolution and implementation of one health to control the dissemination of antibiotic-resistant bacteria and resistance genes: a review. Front Cell Infect Microbiol. 12:1065796. https://doi.org/10.3389/fcimb.2022.1065796
Khush GS, Mackill DJ, Sidhu GS (1989) Breeding rice for resistance to bacterial blight. Bacterial blight of rice. IRRI, pp 207–217
Xie LH, Lin QY, Wu ZJ, Zhou ZJ, Duan YP (1994) Diagnosis, detection and control of rice virus disease in China. J Fujian Agric Univ. 23:280–285
Arunakumari K, Durgarani CV, Satturu V, Sarikonda KR, Chittoor PD, Vutukuri B, Laha GS, Nelli AP, Gattu S, Jamal M, Prasadbabu A (2016) Marker-assisted pyramiding of genes conferring resistance against bacterial blight and blast diseases into indian rice variety MTU1010. Rice Sci. 23:306–316
He D, Zhan J, Xie L (2016) Problems, challenges and future of plant disease management from an ecological point of view. J Integr Agric. 15:705–715
Pradhan M, Bastia DN (2022) Validation of linked markers of bacterial leaf blight resistance genes in rice variety of Odisha (Oryza sativa). J Pharm Innov. 11:446–448
Das G, Rao G (2015) Molecular marker assisted gene stacking for biotic and abiotic stress resistance genes in an elite rice cultivar. Front Plant Sci. 6
Chen S, Wang C, Yang J, Chen B, Wang W, Su J (2020) Identification of the novel bacterial blight resistance gene Xa46(t) by mapping and expression analysis of rice mutant H120. Sci Rep. 10:14642
Chen L, Yin F, Zhang D, Xiao S, Zhong Q, Wang B, Cheng Z (2022) Unveiling a novel source of resistance to bacterial blight in medicinal wild rice, Oryza officinalis. Life. 12:827. https://doi.org/10.3390/life12060827
Jiang N, Yan J, Liang Y (2020) Resistance genes and their interactions with bacterial Blight/Leaf Streak Pathogens (Xanthomonas oryzae) in Rice (Oryza sativa L.) - an updated review. Rice. 13:3. https://doi.org/10.1186/s12284-019-0358-y
Singh S, Sidhu J, Huang N, Vikal Y, Li Z, Brar D, Dhaliwal H, Khush G (2001) Pyramiding three bacterial blight resistance genes (xa5, xa13 and Xa21) using marker-assisted selection into indica rice cultivar PR106. Theor Appl Genet. 102:1011–1015
Rajpurohit D, Kumar R, Kumar M, Paul P, Awasthi A, Basha PO, Puri A, Jhang T, Singh K, Dhaliwal HS (2010) Pyramiding of two bacterial blight resistance and a semi dwarfing gene in type 3 basmati using marker-assisted selection. Euphytica. 178:111–126
Collard BC, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos Trans R Soc Lond B Biol Sci. 363(1491):557–572. https://doi.org/10.1098/rstb.2007.2170
Balachiranjeevi CH, Bhaskar NS, Abhilash V, Akanksha S, Viraktamath BC, Madhav MS, Hariprasad AS, Laha GS, Prasad MS, Balachandran SM, Neeraja CN, Kumar MS, Senguttvel P, Kemparaju KB, Bhadana VP, Ram T, Harika G, Swamy HKM, Hajira SK, Yugandhar A, Pranathi K, Anila M, Rekha G, Kousik BVN, Kumar TD, Swapnil RK, Archana G, Sundaram RM (2015) Marker-assisted introgression of bacterial blight and blast resistance into DRR17B, an elite, fine-grain type maintainer line of rice. Mol Breed. 35:151
Mohapatra S, Panda AK, Bastia AK, Mukherjee AK, Sanghamitra P, Meher J, Mohanty SP, Pradhan SK (2021) Development of Submergence-Tolerant, bacterial Blight-Resistant, and high-yielding Near Isogenic Lines of Popular Variety, ‘Swarna’ through marker-assisted breeding Approach. Front Plant Sci. 12:672618
Kauffman HE, Reddy APK, Hsien SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Disease Rep. 57:537–554
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus. 12:13–15
Jena PP, Bharathkumar S, Reddy JN, Mohapatra T (2015) Introgression of Sub1 locus into highly Preferred Rice Cultivars (Pooja and Pratikshya) in Eastern Region of India for Submergence Tolerance through marker assisted Backcrossing. Adv Biores. 6:45–53
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: Software for association mapping of complex traits in diverse samples. J Bioinform 23:2633–2635
IRRI (2013) Standard evaluation system for rice. International Rice Research Institute, Philippines
Gopinath PP, Parsad R, Joseph B, Adarsh VS (2020) GRAPES: General Rshiny Based Analysis Platform Empowered by Statistics. Available at: https://www.kaugrapes.com/home. version 1.0.0. Accessed on: 1st May 2022
Sarangi, S. K., Maji, B., Mahanta, K. K., Digar, S., Burman, D., Mandal, S., … Bell,R. W. (2019). Alternate kharif rice crop establishment methods and medium duration varieties to enable cropping system intensification in coastal saline region. Journal of Indian Society of Coastal Agricultural Research. 37(2), 115–122. Available at: https://krishi.icar.gov.in/jspui/handle/123456789/49625
Duy PN, Lan DT, Pham Thu H, Thi Thu HP, Nguyen Thanh H, Pham NP, Auguy F, Bui Thi Thu H, Manh TB, Cunnac S, Pham XH (2021) Improved bacterial leaf blight disease resistance in the major elite vietnamese rice cultivar TBR225 via editing of the OsSWEET14 promoter. PLoS ONE. 16(9):e0255470. https://doi.org/10.1371/journal.pone.0255470. PMID: 34499670; PMCID: PMC8428762
Dokku P, Das KM, Rao GJN (2013) Pyramiding of four resistance genes of bacterial blight in Tapaswini, an elite rice cultivar, through marker-assisted selection. Euphytica. 192:87–96
Hajira SK, Yugander A, Balachiranjeevi CH, Pranathi K, Anila M, Mahadevaswamy HK (2014) Development of durable bacterial blight resistant lines of Samba Mahsuri possessing Xa33, Xa21, Xa13&Xa5. Progressive Res. 9:1224–1227
Babu R, Nair SK, Prasanna B, Gupta H (2004) Integrating marker-assisted selection in crop breeding-prospects and challenges. Curr Sci. 87:607–619
Hsu YC, Chiu CH, Yap R, Tseng YC, Wu YP (2020) Pyramiding bacterial blight resistance genes in Tainung82 for broad-spectrum resistance using marker-assisted selection. Int J Mol Sci. 21(4):1281. https://doi.org/10.3390/ijms21041281. PMID: 32074964; PMCID: PMC7072918
Pradhan SK, Nayak DK, Pandit E, Behera L, Anandan A, Lenka S, Barik DP (2016) Incorporation of bacterial leaf blight resistance genes into low-land rice cultivars through marker-assisted breeding. Phytopathol. 106:710–718
Wang GL, Song WY, Ruan DL, Sideris S, Ronald PC (1996) The cloned gene, Xa21, confers resistance to multiple Xanthomonas oryzae pv. Oryzae isolates in transgenic plants. Mol Plant-Microbe Interact. 9:850–855
Iyer AS, McCouch SR (2004) The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol Plant Microbe Interact. 17:1348–1354
Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL (2006) Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev. 20:1250–1255
Sanchez A, Brar D, Huang N, Li Z, Khush G (2000) Sequence tagged site marker-assisted selection for three bacterial blight resistance genes in rice. Crop Sci. 40:792–797
Gopalakrishnan S, Sharma RK, Rajkumar KA, Joseph M, Singh VP, Singh AK, Bhat KV, Singh NK, Mohapatra T (2008) Integrating marker-assisted background analysis with foreground selection for identification of superior bacterial blight resistant recombinants in Basmati rice. Plant Breed. 127:131–139
Pradhan SK, Pandit E, Pawar S, Baksh SY, Mukherjee AK, Mohanty SP (2019) Development of flash-flood tolerant and durable bacterial blight resistant versions of mega rice variety ‘swarna’ through marker-assisted backcross breeding. Sci Rep. 9:12810
Baliyan N, Malik R, Rani R, Mehta K, Vashisth U, Dhillon S, Boora KS (2018) Integrating marker-assisted background analysis with foreground selection for pyramiding bacterial blight resistance genes into Basmati rice. C R Biologies. 341:1–8
Ellur RK, Khanna A, Bhowmick PK, Vinod KK, Nagarajan M, Mondal KK, Singh NK, Singh K, Prabhu KV, Singh AK (2016) Marker-aided incorporation of Xa38, a Novel Bacterial Blight Resistance Gene, in PB1121 and comparison of its resistance spectrum with xa13 + Xa21. Sci Rep. 6:29188
Chen JM, Fu ZY, Quan BQ, Tian DG, Li G, Wang F (2009) Breeding hybrid rice restoring line with double resistance to rice blast and bacterial blight by marker-assisted selection. Mol Plant Breed. 7:465–470
Funding
The authors acknowledge the support of IRRI-OUAT project (Pyramiding of resistance genes for BLB and submergence tolerance into Pratikshya: A popular rice variety of Odisha).
Author information
Authors and Affiliations
Contributions
DB, KCS supervised the work. MP, DB developed the genetic materials. MP, DB, KCS, MD, and JPS performed and supported the molecular analyses. MP, DB performed the field evaluations. MP, DB, and JPS performed and supported the analyses of data. MP, DB, KCS, MD, and JPS contributed in writing and critically revised the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pradhan, M., Bastia, D., Samal, K.C. et al. Pyramiding resistance genes for bacterial leaf blight (Xanthomonas oryzae pv. Oryzae) into the popular rice variety, Pratikshya through marker assisted backcrossing. Mol Biol Rep 50, 9047–9060 (2023). https://doi.org/10.1007/s11033-023-08805-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08805-7