Abstract
Background
Breast cancer (BC) is the most prevalent cancer among females worldwide. Numerous studies suggest that specific RNAs play a crucial role in carcinogenesis. The primate-specific microRNA gene cluster located on the 19q27.3 region of chromosome 19 (C19MC) could potentially regulate tumor cell proliferation, migration, and invasion.
Objective
The objective of this study was to compare the expression of miRNAs from the C19MC cluster in breast cancer tumor and non-tumor samples, as well as in the serum of individuals affected by BC and healthy individuals.
Methods
Peripheral blood was collected from 100 BC patients and 100 healthy individuals, and breast cancer samples including tumor and margin tissues were obtained. After RNA extraction, Real-time PCR was employed to investigate the expression of C19MC, specifically mir-515–1, mir-515–2, mir-516-A1, mir-516-A2, mir-516-B1, mir-516-B2, mir-517-A, mir-517-B, mir-517-C, and mir-518-A1, in the serum and tissue of BC patients and tumor margins. Statistical analyses and ROC curves were generated using GraphPad Prism software (v8.04), with a significance level set at p < 0.05.
Results
Our findings demonstrate a strong correlation between high expression of all C19MC miRNAs mentioned, except for mir-517-B, mir-517-C and mir- 518 in BC. These miRNAs show potential as notable non-invasive tumor markers.
Conclusion
The data obtained from our study support the overall impact of C19MC miRNAs in BC detection and emphasize the potential role of several C19MC members in this process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer remains one of the most prevalent cancers among females worldwide, with over 2 million new cases diagnosed in 2021 [1]. Several risk factors contribute to breast cancer, including alcohol consumption, radiation exposure, hormonal imbalance, a sedentary lifestyle, and hereditary genetic factors [1, 2]. Extensive research has linked the expression of specific miRNAs to cancer, indicating their potential as cancer biomarkers [3]. miRNAs are non-coding RNAs (17 to 25 bp) that can suppress mRNA expression by binding to the 3’ or 5’ UTR, regulating various biological activities such as cell cycle control and apoptosis, and playing crucial roles in cancer initiation and progression. These short non-coding single-stranded RNAs act post-transcriptionally to regulate gene expression. During miRNA biogenesis, the pre-miRNA precursor arms can generate either 5p or 3p miRNA species [4,5,6]. The selective maturation or co-existence of the 5p and 3p species is biologically significant as they target different sets of genes [7, 8]. miRNAs, particularly those within the same family, are often clustered in specific chromosomal locations to provide regulatory advantages [9]. One such human miRNA cluster, the chromosome 19 miRNA cluster (C19MC), is mapped on chromosome 19 and comprises 46 highly homologous miRNA genes [9, 10]. C19MC is a primate-specific miRNA cluster that emerged late in primate evolution, with bioinformatics analyses suggesting its crucial roles in primate reproduction, development, and differentiation compared to lower vertebrates [11, 12]. While C19MC is predominantly expressed in reproductive tissues and certain cancers like colorectal cancer, breast cancer, and primitive neuroectodermal brain tumors [13,14,15], its role in EMT gene regulation remains unknown [16,17,18]. Certain miRNAs, such as mir-224 and mir-373, have been shown to correlate with breast cancer [19, 20].
Extracellular RNAs (exRNAs), including miRNAs, present in biofluids are proposed as non-invasive biomarkers for monitoring organ function and can be quantitatively analyzed. In plasma, miRNAs exist as soluble ribonucleoproteins enclosed in exRNA carriers and transported through the vasculature. However, comprehensive studies involving healthy individuals are necessary to gain insights into the abundance and compositional variability of plasma miRNAs [21,22,23].
The aim of this study is to compare the expression of miRNAs from the C19MC cluster in breast cancer tumor and non-tumor samples, as well as in the serum of individuals affected by breast cancer and healthy individuals. The study aims to investigate the potential role of C19MC miRNAs as non-invasive tumor markers and their overall impact on breast cancer detection.
Methods and materials
Samples
In this case–control study, we enrolled 100 patients diagnosed with breast cancer as the case group, and peripheral venous blood samples were collected from healthy individuals as the control group. Tumor tissue samples and adjacent non-tumor tissues were collected from the 100 breast cancer patients, 20 of whom had a family history of cancer. Due to limitations in sample collection and the necessity to minimize test errors and unwanted variables, we collected both tissue and serum samples from the same breast cancer-affected patients.The patients’ ages ranged from 25 to 81, with an average age of 48 years. The study was approved by the Ethics Committee of the Department of Biology at the University of Tabriz (14,000,907-198-7). It followed ethical guidelines outlined in the 1975 Declaration of Helsinki, and full written consent was obtained from all participants.
RNA extraction and Real-time assay (qPCR)
RNA extraction was carried out using a commercially available kit from ZiAViZ company, which utilizes magnetic nanoparticles. Samples were analyzed using the one-step real-time RT-PCR method, which involves four primers: RP1, RP2, P1, and P2. RP1 contains P1 and an 11-base sequence that is complementary to the 3’ end of the target sncRNA, while RP2 contains P2 and an 11-base sequence that is identical to the 5’ end of the target sncRNA. The detection process includes three stages: in Stage 1, reverse transcription occurs, where RP1 hybridizes with the target sncRNA, which is then extended in the presence of Reverse Transcriptase M-MLV (RNase H-) and dNTPs at 37 °C. In Stage 2, RP2 hybridizes with the cDNA of the miRNA at 37 °C, and both sequences are extended in the presence of hot-start Taq polymerase (HS Taq) and dNTPs at 60 °C after denaturation at 95 °C. Stage 3 involves conventional PCR with the primers P1 and P2, and the amplification of cDNA is continuously monitored in real-time using SYBR green I (Table 1).
Statistical analyses
Statistical analyses were performed using GraphPad Prism software (v8.04) with a p-value of 0.05 considered significant. Receiver operating characteristic (ROC) curves were constructed to assess the diagnostic accuracy of identified miRNAs as potential biomarkers for breast cancer. The area under the curve (AUC) was calculated to assess diagnostic performance, with a higher AUC value indicating better diagnostic accuracy.
Results
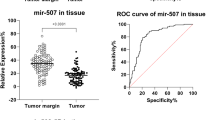
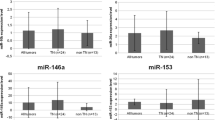
The findings showed that certain C19MC miRNAs, including mir-515–1, mir-515–2, mir-516-A1, mir-516-A2, mir-516-B1, mir-516-B2, mir-517-A were highly expressed in breast cancer tissue and serum samples (Table 2). Additionally, the study revealed that the expression of several of these miRNAs was significantly different between tumor margins and breast cancer tumors, indicating the potential role of these miRNAs in driving the breast cancer phenotype.
Importantly, ROC curve analysis showed that mir-515–1, mir-515–2, mir-516-A1, mir-516-A2, mir-516-B1, mir-516-B2, and mir-517-A had high sensitivity and specificity and could serve as potential tumor markers for breast cancer detection. The expression of mir-515–2, mir-516-A2, and mir-516-B2 in patient serum was also significantly different from that of the control group, indicating their potential as non-invasive biomarkers for breast cancer detection. In the context of biomarkers, a high AUC (Area Under the Curve) value of these microRNA in the ROC curve indicates the predictive performance of a biomarker in distinguishing between two groups or conditions (Figs. 1 and 2).
Discussion
Breast cancer is a prevalent cancer among females, and various risk factors, including non-coding RNAs, have been identified as essential in tumor initiation and progression [24,25,26]. C19MC is a micro-non-coding RNA cluster expressed predominantly in various tissues of the human genome [27]. This study aimed to analyze the expression of C19MC miRNAs in breast cancer tissue and tumor margins, as well as measure their concentration in serum samples of both cases and controls.
The findings revealed significant differences in the expression levels of several C19MC miRNAs, including miR-515–2, miR-516-A2, miR-516-B2, miR-516-B1, and miR-517-A, between breast cancer tumors and tumor margins. Notably, the expression levels of these miRNAs were higher in both breast cancer tissue and serum samples compared to tumor margins. Additionally, when comparing miRNA expression levels with their genomic positions, two different trends emerged: in breast cancer, miRNA expression slightly increased towards the 5ʹ end of the cluster, while in margin tissues, expression levels showed a decreasing tendency [26,27,28,29].
Furthermore, analysis of the data revealed differential expression between the two tissue sample sets. Extracellular RNAs (exRNAs) in biofluids are proposed as non-invasive biomarkers for monitoring organ function, and cell-lineage-specific microRNAs are present in plasma as soluble ribonucleoproteins enclosed in exRNA carriers and transported through the vasculature. However, more extensive studies involving healthy individuals are needed to gain insights into plasma miRNA abundance and composition variability.
The study also highlighted the functional roles of individual C19MC miRNAs in diverse systems. For example, mir-520-h has been found to inhibit migration and invasion of pancreatic cells [28], mir-520-b suppresses the migration of breast cancer cells by targeting hepatitis B X-interacting protein and IL-8 [29, 30], mir-520-c-3p inhibits hepatocellular carcinoma cell invasion by targeting glypican-3 [31] and abrogates breast cancer cell metastasis by suppressing TGFBR2 [32]. However, mir-520-c-3p may promote invasion and metastasis in breast tumor cells by suppressing CD44 [33].
Various studies suggest that miRNAs such as mir-512-3p, mir-512-5p, mir-516a-5p, mir-516b-5p, and mir-498-5p could play pivotal roles in the development of cancer therapies [34,35,36,37]. Future studies should focus on elucidating the molecular processes and pathways regulated by these miRNAs. Although C19MC is the largest human miRNA cluster, its functions are still relatively unknown [38,39,40]. With further understanding, it is reasonable to predict that C19MC could provide a variety of diagnostic and therapeutic targets in the field of cancer [41].
In conclusion, this study demonstrates significant differences in the expression of several C19MC miRNAs between tumor margins and breast cancer tumors, highlighting their potential role in breast cancer detection. C19MC miRNAs have the potential to be utilized as important and non-invasive biomarkers in breast cancer screening. However, many aspects of C19MC still need to be unraveled, posing a challenge for future research in this area. Further studies are necessary to elucidate the molecular processes and pathways involved.
Data availability
All data analyzed during this study are included in this published article. Further raw data from the current study are available from the corresponding author upon request.
References
Gopinath A, Cheema AH, Chaludiya K, Khalid M, Nwosu M, Agyeman WY et al (2022) The impact of dietary fat on breast cancer incidence and survival: a systematic review. Cureus 14(10):e30003. https://doi.org/10.7759/cureus.30003. ((PubMed: 36381753))
Kolyvas EA, Caldas C, Kelly K, Ahmad SS (2022) Androgen receptor function and targeted therapeutics across breast cancer subtypes. Breast Cancer Res 24(1):79. https://doi.org/10.1186/s13058-022-01574-4. ((PubMed: 36376977))
Chakrabortty A, Patton DJ, Smith BF, Agarwal P (2023) miRNAs: potential as biomarkers and therapeutic targets for cancer. Genes 14(7):1375. https://doi.org/10.3390/genes14071375
Fridrichova I, Zmetakova I (2019) MicroRNAs contribute to breast cancer invasiveness. Cells 8(11):1361. https://doi.org/10.3390/cells8111361. ((PubMed: 31683635))
Choo KB, Soon YL, Nguyen PNN, Hiew MSY, Huang CJ (2014) MicroRNA-5p and -3p co-expression and cross-targeting in colon cancer cells. J Biomed Sci 21:95
Pan G, Liu Y, Shang L, Zhou F, Yang S (2021) EMT-associated microRNAs and their roles in cancer stemness and drug resistance. Cancer Commun (Lond) 41(3):199–217. https://doi.org/10.1002/cac2.12138. ((PubMed: 33506604))
Huang CJ, Nguyen PNN, Choo KB, Sugii S, Wee K, Cheong SK et al (2014) Frequent co-expression of miRNA-5p and -3p species and cross-targeting in induced pluripotent stem cells. Int J Med Sci 11:82
Razak SRA, Ueno K, Takayama N, Nariai N, Nagasaki M, Saito R et al (2013) Profiling of microRNA in human and mouse ES and iPS cells reveals overlapping but distinct microRNA expression patterns. PLoS ONE 8:e73532
Anokye-Danso F, Trivedi CM, Juhr D, Gupta M, Cui Z, Tian Y et al (2011) Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency. Cell Stem Cell 8:376–388
Zhang S, Shan C, Kong G, Du Y, Ye L, Zhang X (2012) MicroRNA-520e suppresses the growth of hepatoma cells by targeting the NF-κB-inducing kinase (NIK). Oncogene 31:3607–3620
Keklikoglou I, Koerner C, Schmidt C, Zhang JD, Heckmann D, Shavinskaya A et al (2012) MicroRNA-520/373 family functions as a tumor suppressor in estrogen receptor-negative breast cancer by targeting NF-κB and TGF-β signaling pathways. Oncogene 31:4150–4163
Kleinman CL, Gerges N, Papillon-Cavanagh S, Sin-Chan P, Pramatarova A, Quang D-AK et al (2014) Fusion of TTYH1 with the C19MC microRNA cluster drives expression of a brain-specific DNMT3B isoform in the embryonal brain tumor ETMR. Nat Genet 46:39–44
Cui W, Zhang Y, Hu N, Shan C, Zhang S, Zhang W et al (2010) miRNA-520b and mir-520e sensitize breast cancer cells to complement attack via directly targeting 3’UTR of CD46. Cancer Biol Ther 10:232–241
Min D, Lv X, Wang X, Zhang B, Meng W, Yu F et al (2013) Downregulation of mir-302c and mir-520c by 1,25(OH)2D3 treatment enhances the susceptibility of tumor cells to natural killer cell-mediated cytotoxicity. Br J Cancer 109:723–730
Su C-M, Wang M-Y, Hong C-C, Chen H-A, Su Y-H, Wu C-H et al (2016) mir-520h is crucial for DAPK2 regulation and breast cancer progression. Oncogene 35(9):1134–1142
Zhang J, Liu L, Sun Y, Xiang J, Zhou D, Wang L et al (2016) MicroRNA-520g promotes epithelial ovarian cancer progression and chemoresistance via DAPK2 repression. Oncotarget 7:26516–26534
Wang J, Haubrock M, Cao K-M, Hua X, Zhang C-Y, Wingender E et al (2011) Regulatory coordination of clustered microRNAs based on microRNA-transcription factor regulatory network. BMC Syst Biol 5:199
Lin S, Cheung WKC, Chen S, Lu G, Wang Z, Xie D et al (2010) Computational identification and characterization of primate-specific microRNAs in the human genome. Comput Biol Chem 34:232–241
Zhou J-Y, Zheng S-R, Liu J, Shi R, Yu H-L, Wei M (2016) Mir-519d facilitates the progression and metastasis of cervical cancer through direct targeting Smad7. Cancer Cell Int 16:21
Kan H, Guo W, Huang Y, Liu D (2015) MicroRNA-520g induces epithelial-mesenchymal transition and promotes metastasis of hepatocellular carcinoma by targeting SMAD7. FEBS Lett 589:102–109
Noguer-Dance M, Abu-Amero S, Al-Khtib M, Lefe A, Coullin P, Moore GE (2010) The primate-specific microRNA gene cluster (C19MC) is imprinted in the placenta. Hum Mol Genet 19:3566–3582
Sugii S, Kida Y, Berggren WT, Evans RM (2011) Feeder-dependent and feeder-independent iPS cell derivation from human and mouse adipose stem cells. Nat Protoc 6:346–358
Choo KB, Tai L, Hymavathee KS, Wong CY, Nguyen PNN, Huang CJ et al (2014) Oxidative stress-induced premature senescence in Wharton’s jelly-derived mesenchymal stem cells. Int J Med Sci 11:1201–1207
Ma W, Yu Q, Jiang J, Du X, Huang L, Zhao L et al (2016) mir-517a is an independent prognostic marker and contributes to cell migration and invasion in human colorectal cancer. Oncol Lett 11:2583–2589
Ren G, Wang L (2016) Study on the relationship between mir-520g and the development of breast cancer. Eur Rev Med Pharmacol Sci 20:657–663
Tsai C-H, Tsai H-C, Huang H-N, Hung C-H, Hsu C-J, Fong Y-C et al (2014) Resistin promotes tumor metastasis by down-regulation of mir-519d through the AMPK/p38 signaling pathway in human chondrosarcoma cells. Oncotarget 6:258–270
Brownlie RJ, Zamoyska R (2013) T cell receptor signaling networks: branched, diversified and bounded. Nat Rev Immunol 13:257–269
Danielsen SA, Eide PW, Nesbakken A, Guren T, Leithe E, Lothe RA (2015) Portrait of the PI3K/AKT pathway in colorectal cancer. Biochim Biophys Acta 1855:104–121
Guan R, Cai S, Sun M, Xu M (2017) Upregulation of miR-520b promotes ovarian cancer growth. Oncol Lett 14:3155–3161. https://doi.org/10.3892/ol.2017.6552
Hu N, Zhang J, Cui W, Kong G, Zhang S, Yue L, Bai X, Zhang Z, Zhang W, Zhang X, Ye L (2011) miR-520b regulates migration of breast cancer cells by targeting hepatitis B X-interacting protein and interleukin-8. J Biol Chem 286(15):13714–13722
Menon A, Abd-Aziz N, Khalid K, Poh CL, Naidu R (2022) miRNA: a promising therapeutic target in cancer. Int J Mol Sci 23(19):11502. https://doi.org/10.3390/ijms231911502
Li M, Lee KF, Lu Y, Clarke I, Shih D, Eberhart C et al (2009) Frequent amplification of a chr19q13.41 microRNA polycistron in aggressive primitive neuroectodermal brain tumors. Cancer Cell 16:533–546
Flor I, Neumann A, Freter C, Helmke BM, Langenbuch M, Rippe V et al (2012) Abundant expression and hemimethylation of C19MC in cell cultures from placenta-derived stromal cells. Biochem Biophys Res Commun 422:411–416
Pinho FG, Frampton AE, Nunes J (2013) Downregulation of microRNA-515-5p by the estrogen receptor modulates sphingosine kinase 1 and breast cancer cell proliferation. Can Res 73:5936–5948
Jagadeeswaran G, Zheng Y, Sumathipala N, Jiang H, Arrese EL, Soulages JL et al (2010) Deep sequencing of small RNA libraries reveals dynamic regulation of conserved and novel microRNAs and microRNA-stars during silkworm development. BMC Genomics 11:1–18
Li S-C, Liao Y-L, Ho M-R, Tsai K-W, Lai C-H, Lin W (2012) miRNA arm selection and isomiR distribution in gastric cancer. BMC Genomics 13(Suppl 1):S13
Carroll AP, Goodall GJ, Liu B (2014) Understanding principles of miRNA target recognition and function through integrated biological and bioinformatics approaches. Wiley Interdisc Rev 5:361–379
Akbari Moqadam F, Pieters R, den Boer ML (2013) The hunting of targets: challenge in miRNA research. Leukemia 27:16–23
Jalvy-Delvaille S, Maurel M, Majo V, Pierre N, Chabas S, Combe C et al (2012) Molecular basis of differential target regulation by mir-96 and mir-182: the glypican-3 as a model. Nucleic Acids Res 40:1356–1365
Bar M, Wyman SK, Fritz BR, Qi J, Garg KS, Parkin RK et al (2008) MicroRNA discovery and profiling in human embryonic stem cells by deep sequencing of small RNA libraries. Stem Cells 26:2496–2505
Portt L, Norman G, Clapp C, Greenwood M, Greenwood MT (2011) Anti-apoptosis and cell survival: a review. Biochem Biophys Acta 1813:238–259
Acknowledgements
We thank all patients and individuals for participating in this project.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
SARA Experimented and wrote the manuscript with support from MH and MAHF. MH supervised the work and contributed to the design and implementation of the research, the analysis of the results, and the writing of the manuscript. MAHF Helped supervise the project. All authors discussed the results and contributed and reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors confirm that there are no known conflicts of interest associated with this publication, and there has been no significant financial support for this work that could have influencedits outcome.
Ethical approval
This study was approved by the Department of Biology committee at the University of Tabriz (14000907–198-7).
Consent to participate
All stages of sample collection have been done with the consent of patients and healthy people, and written consent forms have been obtained.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Altalebi, S.A.R., Haghi, M. & Hosseinpour Feizi, M.A. Expression study of microRNA cluster on chromosome 19 (C19MC) in tumor tissue and serum of breast cancer patient. Mol Biol Rep 50, 9825–9831 (2023). https://doi.org/10.1007/s11033-023-08801-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08801-x