Abstract
Background
Bone marrow mesenchymal stem cells (BMSCs) can differentiate into osteoblasts and thus present a tremendous therapeutic potential in osteoporosis. Here, we elucidated the involvement of long non-coding RNAs (lncRNAs) HOXA transcript antisense RNA, myeloid-specific 1 (HOTAIRM1) in the osteogenic differentiation of BMSCs.
Methods and results
The expression levels of HOTAIRM1, miR-152-3p, ETS proto-oncogene 1 (ETS1), runt-related transcription factor 2 (RUNX2), Osterix, and osteocalcin (OCN) were determined by a quantitative real-time polymerase chain reaction (qRT-PCR) or western blot method. Targeted relationship between miR-152-3p and HOTAIRM1 or ETS1 was confirmed by dual-luciferase reporter and RNA pull-down assays. The activity of alkaline phosphatase (ALP) was measured by the ALP Activity Assay Kit. The extent of the calcium deposition was assessed by Alizarin Red Staining. Our data showed that HOTAIRM1 and ETS1 levels were up-regulated and miR-152-3p expression was down-regulated during osteogenic differentiation of human BMSCs (HBMSCs). HOTAIRM1 overexpression enhanced osteogenic differentiation of HBMSCs, and decreased level of HOTAIRM1 suppressed osteogenic differentiation of HBMSCs. HOTAIRM1 directly targeted miR-152-3p. ETS1 was identified as a direct and functional target of miR-152-3p. Furthermore, HOTAIRM1 functioned as a post-transcriptional regulator of ETS1 expression by miR-152-3p.
Conclusion
The findings in this paper identify HOTAIRM1 as a novel regulator of osteogenic differentiation of BMSCs by the regulation of miR-152-3p/ETS1 axis, uncovering HOTAIRM1 as a promising therapeutic strategy for osteoporosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteoporosis is a common skeletal disorder characterized by low bone mass and increased bone fragility that is one of the leading causes for bone fractures in the population older than 50 years, especially postmenopausal women [1]. The incidence of osteoporosis has been increasing with the rapidly aging population [2]. Bone marrow mesenchymal stem cells (BMSCs) are a heterogeneous class of cells with the capacity of self-renewal and differentiation and have been unraveled the differentiation potential into osteoblasts [3]. A better understanding for the mechanisms underlying osteogenic differentiation of BMSCs will provide a new opportunity for osteoporosis management.
Long non-coding RNAs (lncRNAs) play critical roles in a wide variety of biological processes [4]. Moreover, lncRNAs can influence gene expression by functioning as sponges of certain microRNAs (miRNAs) and thus regulate osteogenic differentiation of BMSCs [5, 6]. As an example, Shang et al. identified TCONS_00041960 as a strong promoter for osteogenesis of rat BMSCs via miR-125a-3p and miR-204-5p [7]. Wang et al. illuminated that growth arrest specific 5 (GAS5) contributed to osteogenic differentiation of BMSCs in osteoporotic mice through the regulation of miR-135a-5p [8]. Conversely, maternally expressed 3 (MEG3) was validated as a negative regulator of BMSC osteogenic differentiation by reducing miR-133a-3p activity [9]. Recent work demonstrates that HOXA transcript antisense RNA, myeloid-specific 1 (HOTAIRM1) can promote osteogenesis of MSCs isolated from menstrual blood and umbilical cord [10]. However, whether HOTAIRM1 is implicated in BMSC osteogenic differentiation remains unclear.miRNAs function as post-transcriptional modulators of gene expression and have been implicated in the differentiation of BMSCs into osteoblasts [11, 12]. Previous work substantiated that miR-152 was underexpressed during BMSC osteogenic differentiation and acted as a negative modulator of osteogenic differentiation of BMSCs [13, 14]. Moreover, Han et al. demonstrated the involvement of miR-152 in osteogenesis of BMSCs by operating as a molecular mediator of hsa_circ_0076690 [14]. Nonetheless, it is still underexplored whether miR-152-3p is involved in the modulation of HOTAIRM1 in BMSC osteogenic differentiation. For these reasons, we sought to identify the precise parts of HOTAIRM1 in osteogenic differentiation of BMSCs.
Materials and methods
Cell culture and differentiation
Human BMSCs (HBMSCs) were purchased from Procell (Wuhan, China) and propagated at 37 °C with 5% CO2 using complete growth medium (Procell). 293 T cells (American Type Culture Collection, ATCC, Rockville, MD, USA) were cultured using standard protocols provided by ATCC. For osteogenic induction, cultured HBMSCs at 60% confluence were maintained in differentiation medium consisted of complete growth medium, 0.05 mM L-ascorbic acid (Chemegen, Shanghai, China), 100 mM dexamethasone (Sigma-Aldrich, St Louis, MO, USA) and 10 mM β-glycerophosphate (Sigma-Aldrich) for 7–14 days. Fresh medium was replaced every other day.
Transient transfection of cells
Chemically synthetic HOTAIRM1-shRNA vectors (sh-HOTAIRM1#1, sh-HOTAIRM1#2 and sh-HOTAIRM1#3), ETS proto-oncogene 1 (ETS1)-shRNA vector (sh-ETS1) and a nontarget-shRNA vector (sh-NC) were obtained from Ribobio (Shanghai, China) and their sequences were: sh-HOTAIRM1#1: 5′-GGAGACUGGUAGCUUAUUATT-3′, sh-HOTAIRM1#2: 5′-GAUUAAUCAACCACACUGATT-3′, sh-HOTAIRM1#3: 5′-AGAAACUCCGUGUUACUCATT-3′, sh-ETS1: 5′-UUCUUGUUUGAUAGCAAAGUA-3′ and sh-NC: 5′-AAGACAUUGUGUGUCCGCCTT-3′. The mature miR-152-3p sequence (5′-UCAGUGCAUGACAGAACUUGG-3′, Ribobio) was used as the corresponding mimic, and the sequence of miR-152-3p inhibitor was the exact antisense of the mature miR-152-3p sequence (5′-CCAAGUUCUGUCAUGCACUGA-3′, Ribobio). Chemically modified scrambled oligonucleotides (mimic NC, 5′-CGAUCGCAUCAGCAUCGAUUGC-3′ and inhibitor NC, 5′-CUAACGCAUGCACAGUCGUACG-3′) were designed as matched negative controls. The plasmids expressing miRNA mimics and inhibitors were also obtained from Ribobio. The expression plasmids (oe-HOTAIRM1 and oe-ETS1) were generated by cloning the coding sequence of human HOTAIRM1 (Accession: NR_038367.1) and ETS1 (Accession: NM_001330451.2) into the pcDNA3.1 vector (Invitrogen, Carlsbad, CA, USA), respectively. The nontarget pcDNA3.1 plasmid (vector) was used as the negative control.
HBMSCs of ~ 50% confluence in 24-well plates were transiently transfected with 200 ng of plasmids using Lipofectamine 3000 reagent as per the manufacturing instructions (Invitrogen). After 24 h transfection, the cells were cultured in differentiation medium for 7 days and then harvested for quantitative real-time polymerase chain reaction (qRT-PCR) and western blot analyses.
Cell osteogenic differentiation assay
Transfected HBMSCs were maintained in differentiation medium for 7–14 days to induce their osteogenic differentiation. On day 7 of incubation, the ALP activity of cultured HBMSCs was analyzed using a colorimetric ALP Activity Assay Kit (Elabscience, Houston, TX, USA). The absorption at 405 nm was measured, and the results were expressed relative to the absorption of total protein. On day 14 of incubation, the extent of calcium deposition of cultured HBMSCs was evaluated with Alizarin Red Staining (ARS) using 0.1% ARS staining solution (Wkbio, Tianjin, China). Quantification of cellular staining was performed using the AxioVision LE64 software (Carl Zeiss, Oberkochen, Germany).
RNA preparation and qRT-PCR
RNA was prepared from transfected or untransfected HBMSCs with Trizol (Invitrogen) as recommended by the manufacturer. To determine gene expression levels, RNA extracts (100 ng) were reverse-transcribed into cDNA with a Bio-Rad iSCRIPT Kit (Bio-Rad, Hercules, CA, USA) with random primers or miScript Reverse Transcription Kit with stem-loop RT PCR primers (Qiagen, Hilden, Germany) based on the protocols of manufacturers. cDNA was subjected to qRT-PCR using iQSYBR Green (Bio-Rad) or miScript SYBR Green PCR Kit (Qiagen) on a Rotor-Gene Q instrument (Qiagen). Results were analyzed using the 2−ΔΔCt method [15] and normalized to the expression of glyceraldehyde 3-phosphate dehydrogenase (GAPDH) or U6. The sequences of primers were presented in Supplementary Table S1.
Western blot
Total protein was prepared from transfected or untransfected HBMSCs using RIPA lysis buffer (Solarbio, Beijing, China) supplemented with protease inhibitor cocktails (Roche, Sussex, UK). Total cell extracts (30 µg) were resolved on 10% SDS polyacrylamide gels and then blotted onto polyvinylidene fluoride membranes (GE Healthcare, Toyko, Japan). Immunoblotting was done using anti-RUNX2 (ab236639), anti-OCN (ab93876), anti-Osterix (ab209484), anti-ETS1 (ab225868) and anti-GAPDH (ab9485) primary antibodies (shown in Supplementary Table S2). After incubation with horseradish peroxidase-labeled anti-rabbit IgG secondary antibody (ab97051), the immunoreactive complexes were detected with an enhanced chemiluminescence substrate (ECL, Bio-Rad). Quantification of the blots was carried out using Multi-Gauge software (Fujifilm, Toyko, Japan). All antibodies were purchased from Abcam (Cambridge, UK) and used as per the accompanying guidance.
Bioinformatics
The targeted miRNAs of HOTAIRM1 and the molecular targets of miR-152-3p were predicted using the online software starBase v.3 available at http://starbase.sysu.edu.cn/.
Dual-luciferase reporter assay
Firefly luciferase reporter constructs were generated by cloning the fragments of HOTAIRM1 and ETS1 3′UTR encompassing the miR-152-3p-binding sequence or miss-matched target sequence into the pmirGLO vector (Promega, Madison, WI, USA) downstream from the firefly luciferase coding region, respectively. 293 T cells of 60% confluence were cotransfected using Lipofectamine 3000 with each reporter construct (100 ng), the pRL-SV40 Renilla control plasmid (1 ng, Promega), and miR-152-3p mimic (30 nM) or mimic NC. The luciferase activity was gauged after 48 h transfection using the Dual-luciferase Reporter Assay (Promega). Firefly luciferase activity was normalized to that of Renilla luciferase.
RNA pull-down assays
HBMSCs were induced in differentiation medium for 7 days and then lysed with RIPA lysis buffer. Cell lysates were incubated with biotin-labeled miR-152-3p mimic (Bio-miR-152-3p, Ribobio) or negative control Bio-NC at 4 °C for 4 h before adding the streptavidin beads (Sigma-Aldrich) for 4 h. Beads were harvested and bound RNA was isolated to examine the enrichment levels of HOTAIRM1 and EST1 mRNA by qRT-PCR.
Statistical analysis
In this study, Prism version 7.0 software (GraphPad, La Jolla, CA, USA) was used for all statistical analyses. Differences were compared using a two-tailed Student’s t-test or analysis of variance (ANOVA) with Tukey’s post hoc test. Data were shown as mean ± standard deviation (SD) from at least three independent experiments, and error bars depicted SD. The p values < 0.05 were considered significant.
Results
HOTAIRM1 is up-regulated during osteogenic differentiation of HBMSCs
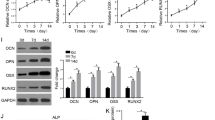
To validate the involvement of HOTAIRM1 in osteogenic differentiation of HBMSCs, we firstly detected its expression level by qRT-PCR during the process. Remarkably, HOTAIRM1 expression was increased in a time-dependent manner during osteogenic differentiation of HBMSCs (Fig. 1a). We also assessed the levels of osteoblast-associated factors (OCN, RUNX2 and Osterix) during osteogenic differentiation. As expected, the mRNA (Fig. 1b–d) and protein (Fig. 1e–g) levels of RUNX2, OCN and Osterix were significantly augmented during osteogenic differentiation of HBMSCs.
HOTAIRM1 is overexpressed during osteogenic differentiation of HBMSCs. A–G HBMSCs were cultured in differentiation medium for 21 days. A At the indicated time points (0, 3, 7, 14 and 21 days), HOTAIRM1 level was detected by qRT-PCR. B–D The mRNA levels of RUNX2, OCN and Osterix were gauged by qRT-PCR at the indicated time points. E–G The protein levels of RUNX2, OCN and Osterix were determined by western blot at the indicated time points. *P < 0.05
HOTAIRM1 positively regulates osteogenic differentiation of HBMSCs
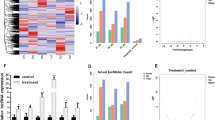
To elucidate the biological role of HOTAIRM1 in osteogenic differentiation of HBMSCs, we manipulated its expression in HBMSCs using a HOTAIRM1-shRNA (sh-HOTAIRM1) vector or HOTAIRM1 overexpression plasmid (oe-HOTAIRM1). Transfection of sh-HOTAIRM1#1, sh-HOTAIRM1#2 or sh-HOTAIRM1#3, but not sh-NC, significantly decreased HOTAIRM1 expression (Fig. 2a), and oe-HOTAIRM1 introduction led to a clear elevation of HOTAIRM1 expression in HBMSCs (Fig. 2b). Since sh-HOTAIRM1#1 caused the most significant down-regulation in HOTAIRM1 expression, we selected it for the further experiments. Interestingly, silencing of HOTAIRM1 resulted in reduced expression of RUNX2, OCN and Osterix at both mRNA and protein, while overexpression of HOTAIRM1 significantly elevated the levels of RUNX2, OCN and Osterix mRNA and protein (Fig. 2c–f). Moreover, HOTAIRM1 silencing suppressed the activity of osteogenic marker ALP, while enforced expression of HOTAIRM1 enhanced its activity (Fig. 2g, h). Using Alizarin Red Staining, we found that HOTAIRM1-silenced HBMSCs showed lower calcium deposition than controls, whereas HBMSCs stably expressing HOTAIRM1 exhibited increased calcium deposition (Fig. 2i, j). All these data indicate that HOTAIRM1 regulates osteogenic differentiation of HBMSCs.
HOTAIRM1 positively modulates the differentiation of HBMSCs into osteoblast. A, B HBMSCs transfected with sh-NC, sh-HOTAIRM1#1, sh-HOTAIRM1#2 or sh-HOTAIRM1#3 (A), and vector or oe-HOTAIRM1 (B) were cultured in differentiation medium for 7 days, and then HOTAIRM1 expression was assessed by qRT-PCR. C–J HBMSCs were transfected with sh-NC, sh-HOTAIRM1#1, vector or oe-HOTAIRM1 and then cultured in differentiation medium for 7 or 14 days, followed by the measurement of RUNX2, OCN and Osterix mRNA levels by qRT-PCR on day 7 (C, D), RUNX2, OCN and Osterix protein levels by western blot on day 7 (E, F), ALP activity using the ALP Activity Kit on day 7 (G, H), the extent of calcium deposition with Alizarin Red Staining on day 14 (I, J). *P < 0.05
HOTAIRM1 directly targets miR-152-3p
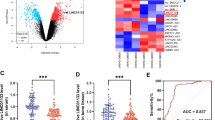
The data of qRT-PCR showed that miR-152-3p expression was significantly reduced during osteogenic differentiation of HBMSCs (Fig. 3a). Using starBase v.3 software, a putative target sequence for miR-152-3p was predicted within HOTAIRM1 (Fig. 3b). To confirm the validity of the target sequence for interaction, we cloned HOTAIRM1 fragment encompassing the miR-152-3p target sequence or mutated seed region into a luciferase plasmid (Fig. 3b). Transfection of the wild-type reporter construct (HOTAIRM1 wt) induced about a 70% reduction of relative luciferase activity in the presence of miR-152-3p mimic vector, and this effect was abrogated by the mutant-type reporter (HOTAIRM1 mut) (Fig. 3c). RNA pull-down assays showed that incubation of cell lysates with Bio-miR-152-3p led to a significant augmentation of HOTAIRM1 enrichment level in HBMSCs (Fig. 3d). These findings suggest the direct relationship between HOTAIRM1 and miR-152-3p.
HOTAIRM1 targets miR-152-3p by directly binding to miR-152-3p. A HBMSCs were cultured in differentiation medium for 21 days, and miR-152-3p expression was detected by qRT-PCR at the indicated time points. B Schematic of the putative target sequence for miR-152-3p within HOTAIRM1 predicted by starBase v.3 software and mutated the miR-152-3p-binding region. C Dual-luciferase reporter assays were carried out in 293 T cells. D RNA pull-down assays were performed using Bio-NC or Bio-miR-152-3p in HBMSCs cultured in differentiation medium for 7 days. E HBMSCs were transfected with inhibitor NC vector, miR-152-3p inhibitor vector, mimic NC vector or miR-152-3p mimic vector and then cultured in differentiation medium for 7 days, followed by the assessment of relative miR-152-3p expression by qRT-PCR. *P < 0.05
HOTAIRM1 regulates osteogenic differentiation of HBMSCs through miR-152-3p
We next examined whether miR-152-3p is involved in HOTAIRM1-mediated regulation in osteogenic differentiation of HBMSCs. The transfection efficiencies of plasmids expressing miR-152-3p inhibitor and miR-152-3p mimic in HBMSCs cultured in differentiation medium were determined by qRT-PCR. The results showed that miR-152-3p expression was suppressed by miR-152-3p inhibitor vector and elevated by miR-152-3p mimic vector (Fig. 3e). By contrast, miR-152-3p expression was significantly elevated in HOTAIRM1-silenced HBMSCs, and miR-152-3p inhibitor vector strongly abolished the expression augmentation induced by HOTAIRM1 silencing (Fig. 4a). Moreover, the expression of miR-152-3p was markedly reduced in HBMSCs as a result of HOTAIRM1 overexpression, and this effect was reversed by miR-152-3p mimic vector (Fig. 4b). Strikingly, miR-152-3p down-regulation reversed HOTAIRM1 silencing-mediated repression on RUNX2, OCN and Osterix expression, and re-expression of miR-152-3p abrogated oe-HOTAIRM1-driven augmentation of RUNX2, OCN and Osterix levels of HBMSCs (Fig. 4c–f). Furthermore, miR-152-3p down-regulation abated HOTAIRM1 silencing-mediated suppression on ALP activity and calcium deposition, while miR-152-3p re-expression reversed oe-HOTAIRM1-driven enhancement of ALP activity and calcium deposition of HBMSCs (Fig. 4g–j). Together, these results demonstrate that miR-152-3p is a molecular mediator of HOTAIRM1 in regulating osteogenic differentiation of HBMSCs.
HOTAIRM1 modulates osteogenic differentiation of HBMSCs by miR-152-3p. A–J HBMSCs were transfected with sh-NC, sh-HOTAIRM1#1, sh-HOTAIRM1#1 + inhibitor NC, sh-HOTAIRM1#1 + miR-152-3p inhibitor, vector, oe-HOTAIRM1, oe-HOTAIRM1 + mimic NC or oe-HOTAIRM1 + miR-152-3p mimic and then cultured in differentiation medium for 7 or 14 days, followed by the detection of miR-152-3p level by qRT-PCR on day 7 (A, B), RUNX2, OCN and Osterix mRNA levels by qRT-PCR on day 7 (C, D), RUNX2, OCN and Osterix protein levels by western blot on day 7 (E, F), ALP activity using the ALP Activity Kit on day 7 (G, H), the extent of calcium deposition with Alizarin Red Staining on day 14 (I and J). *P < 0.05
miR-152-3p directly interacts with the 3′UTR of ETS1
The data of qRT-PCR and western blot also revealed that ETS1 expression was significantly up-regulated at both mRNA and protein during osteogenic differentiation of HBMSCs (Fig. 5a, b). The software starBase v.3 showed that ETS1 3′UTR harbored a putative region that was partially complementary to miR-152-3p (Fig. 5c). Luciferase assays revealed that transfection of miR-152-3p mimic vector significantly reduced the luciferase activity of the wild-type ETS1 3′UTR reporter (ETS1 3′UTR wt), and site-directed mutation (ETS1 3′UTR mut) remarkably abolished the repression of miR-152-3p (Fig. 5d). Moreover, incubation of Bio-miR-152-3p caused a striking elevation of ETS1 mRNA enrichment level compared with the Bio-NC control (Fig. 5e). Hence, ETS1 is a direct target of miR-152-3p.
miR-152-3p directly targets ETS1 through ETS1 3′UTR. A, B HBMSCs were cultured in differentiation medium for 21 days, and ETS1 mRNA and protein levels were detected by qRT-PCR and western blot at the indicated time points. C Schematic of the miR-152-3p-binding region within ETS1 3′UTR and the mutant of the seed region. D Dual-luciferase assays were done in 293 T cells cotransfected with ETS1 3′UTR wt or ETS1 3′UTR mut and miR-152-3p mimic vector or mimic NC. E RNA pull-down assays were conducted using Bio-miR-152-3p or Bio-NC in HBMSCs cultured in differentiation medium for 7 days. *P < 0.05
ETS1 is a functional target of miR-152-3p in regulating osteogenic differentiation of HBMSCs
The transfection efficiencies of sh-ETS1 vector and oe-ETS1 in HBMSCs cultured in differentiation medium were gauged by qRT-PCR and western blot. The data revealed that the mRNA and protein levels of ETS1 were reduced by sh-ETS1 vector and conversely enhanced by oe-ETS1 (Fig. 6a, b). In cultured HBMSCs, enforced expression of miR-152-3p significantly decreased the mRNA and protein levels of ETS1, whereas miR-152-3p depletion led to a marked increase in ETS1 expression (Fig. 6c–f), reinforcing that ETS1 was a direct target of miR-152-3p. Moreover, enforced expression of miR-152-3p strongly attenuated the expression of RUNX2, OCN and Osterix, while miR-152-3p depletion enhanced their expression (Fig. 6g–j). Additionally, the ALP activity and calcium deposition of HBMSCs were reduced by miR-152-3p overexpression, while they were conversely enhanced by miR-152-3p depletion (Fig. 6k–n).
miR-152-3p negatively modulates osteogenic differentiation of HBMSCs by ETS1. A, B HBMSCs were transfected with sh-NC, sh-ETS1, vector or oe-ETS1 and then cultured in differentiation medium for 7 days, followed by the detection of ETS1 mRNA expression by qRT-PCR and ETS1 protein level by western blot. C–N HBMSCs were transfected with mimic NC vector, miR-152-3p mimic vector, miR-152-3p mimic + vector, miR-152-3p mimic + oe-ETS1, inhibitor NC vector, miR-152-3p inhibitor vector, miR-152-3p inhibitor + sh-NC or miR-152-3p inhibitor + sh-ETS1 and then cultured in differentiation medium for 7 or 14 days, followed by the determination of ETS1 mRNA level by qRT-PCR (C, D), ETS1 protein expression by western blot (E and F), RUNX2, OCN and Osterix mRNA levels by qRT-PCR (G, H), RUNX2, OCN and Osterix protein levels by western blot (I, J), ALP activity using the ALP Activity Kit on day 7 (K, L), the extent of calcium deposition with Alizarin Red Staining on day 14 (M, N). *P < 0.05
To elucidate whether ETS1 represents an importantly functional target of miR-152-3p in regulating osteogenic differentiation of HBMSCs, we increased ETS1 expression with oe-ETS1 in miR-152-3p-overexpressing cells and down-regulated ETS1 expression in miR-152-3p-silenced cells using sh-ETS1 vector (Fig. 6c–f). Notably, re-expression of ETS1 abolished the reduction of miR-152-3p overexpression on RUNX2, OCN and Osterix levels, while decreased ETS1 level reversed miR-152-3p depletion-mediated elevation of RUNX2, OCN and Osterix expression of HBMSCs (Fig. 6g–j). Consistently, ETS1 re-expression reversed miR-152-3p overexpression-imposed suppression on ALP activity and calcium deposition, whereas ETS1 down-regulation counteracted miR-152-3p depletion-mediated enhancement of ALP activity and calcium deposition of HBMSCs (Fig. 6k–n). These results together establish that ETS1 is a functional downstream effector of miR-152-3p.
HOTAIRM1 regulates ETS1 expression via miR-152-3p
Based on the above findings, we wanted to investigate whether HOTAIRM1 was a post-transcriptional modulator of ETS1 expression in HBMSCs cultured in differentiation medium. As expected, the mRNA and protein levels of ETS1 were significantly down-regulated by HOTAIRM1 silencing (Fig. 7a, b), while they were conversely elevated as a result of HOTAIRM1 overexpression (Fig. 7c, d). Furthermore, transfection of miR-152-3p inhibitor vector reversed the down-regulation of HOTAIRM1 silencing on ETS1 expression (Fig. 7a, b), and elevated expression of miR-152-3p abolished oe-HOTAIRM1-mediated ETS1 expression augmentation in HBMSCs (Fig. 7c, d). All these data show that HOTAIRM1 regulates ETS1 expression through miR-152-3p.
HOTAIRM1 controls the expression of ETS1 by targeting miR-152-3p. A, B HBMSCs were transfected with sh-NC, sh-HOTAIRM1#1, sh-HOTAIRM1#1 + inhibitor NC or sh-HOTAIRM1#1 + miR-152-3p inhibitor before the culture in differentiation medium for 7 days, and then ETS1 mRNA and protein levels were evaluated by qRT-PCR and western blot, respectively. C, D HBMSCs transfected with vector, oe-HOTAIRM1, oe-HOTAIRM1 + mimic NC vector or oe-HOTAIRM1 + miR-152-3p mimic vector were cultured in differentiation medium for 7 days, followed by the assessment of ETS1 mRNA level by qRT-PCR and ETS1 protein expression by western blot. *P < 0.05
Discussion
BMSCs can differentiate into osteoblasts and thus present a tremendous therapeutic potential for osteoporosis [3]. Nevertheless, the mechanisms underlying osteogenic differentiation of BMSCs are still largely unclear. Recently, many lncRNAs have been shown to be implicated in the differentiation of BMSCs into osteoblasts [16]. In this paper, we sought to elucidate the biological role of HOTAIRM1 in osteogenic differentiation of BMSCs and identify a novel lncRNA/miRNA/mRNA regulatory network governing it.
Several previous studies have uncovered the conflicting roles of HOTAIRM1 in tumor development [17,18,19,20]. These contradictory findings might partially due to the different tumor models in these studies, where HOTAIRM1 enhances the tumorigenesis of glioblastoma [17, 18] and represses the progression of hepatocellular carcinoma [19] and colorectal cancer [20]. Moreover, HOTAIRM1 is able to regulate differentiation of various types of cells, such as neuron, monocyte and myeloid cells [21,22,23]. Chen et al. demonstrated that HOTAIRM1 contributed to osteogenesis of dental follicle stem cells [24]. Moreover, abnormal expression of HOTAIRM1 has been discovered to influence osteogenic differentiation of MSCs isolated from menstrual blood and umbilical cord [10]. Our results showed the overexpression of HOTAIRM1 during osteogenic differentiation of HBMSCs. Furthermore, gain- and loss-of-function phenotypes of HOTAIRM1 uncovered that HOTAIRM1 is a positive regulator of osteogenic differentiation of HBMSCs.
Emerging evidence has demonstrated the crucial involvement of miR-152-3p in human cancers, such as chronic myeloid leukemia and prostate cancer [25, 26]. miR-152-3p is also involved in the pathogenesis of ischemic stroke, diabetic nephropathy and acute kidney injury [27,28,29]. Here, we first confirmed that HOTAIRM1 directly targets miR-152-3p. Similar to previous studies [13, 14], we validated that miR-152-3p acts as a negative regulator in HBMSC osteogenic differentiation. Furthermore, our study tied in the regulation of HOTAIRM1 in osteogenic differentiation of HBMSCs through miR-152-3p.
ETS1 is a member of the ETS domain family of transcription factors and serves as a key factor in cancer progression [30]. Increasing evidence has shown that ETS1 is involved in the process of osteogenesis [31, 32]. For instance, Qi et al. illuminated the important regulation of ETS1 in osteogenic differentiation of MSCs [33]. Fan et al. reported that ETS1 mediated the repressive effect of miR-532-3p on osteogenic differentiation of MC3T3-E1 cells [34]. Moreover, Hua et al. demonstrated that the MALAT1/miR-155-5p axis controlled osteogenic differentiation of human periodontal ligament stem cells by targeting ETS1 [35]. In this paper, we were first to identify that ETS1 is a functionally important target of miR-152-3p in regulating osteogenic differentiation of HBMSCs. Furthermore, our current study identified that HOTAIRM1 acts as a post-transcriptional regulator of ETS1 expression through miR-152-3p, suggesting the regulation of the HOTAIRM1/miR-152-3p/ETS1 cascade in osteogenic differentiation of BMSCs. Similarly, Ren and colleagues uncovered the involvement of the LINC00963/miR-760/ETS1 network in osteogenic differentiation of BMSCs [36]. These findings suggest that the HOTAIRM1/miR-152-3p/ETS1 and LINC00963/miR-760/ETS1 axes might be two interactional or paralleled regulatory networks in osteogenic differentiation of BMSCs. ETS1 can regulate high glucose-induced osteogenic differentiation of MC3T3-E1 cells through the up-regulation of Runx2 [37]. Additionally, ETS1 inhibits miR-128 transcription and thus enhances osteogenic differentiation of MC3T3-E1 cells via HOXA13/β-Catenin cascade [38]. Therefore, more researches will be performed to determine whether Runx2 or miR-128/HOXA13/β-Catenin cascade is involved in the regulation of the HOTAIRM1/miR-152-3p/ETS1 axis in osteogenic differentiation of BMSCs.
The efficacy of BMSCs in bone tissue engineering and regenerative medicine-based applications is under intensive exploration at present [39, 40]. Crucial regulators of osteogenic differentiation of BMSCs, including lncRNAs and miRNAs, have been proposed as potential ancillary factors for the use of BMSCs [6, 11]. Our findings suggest that targeting HOTAIRM1/miR-152-3p/ETS1 axis might be a promising strategy for the efficacy of BMSCs in bone tissue engineering and clinical treatment.
In summary, our study identifies that HOTAIRM1 functions as a novel regulator of osteogenic differentiation of BMSCs by targeting the miR-152-3p/ETS1 axis, as illustrated in Fig. 8. Our findings provide a novel theoretical and experimental rationale for the use of BMSCs in clinical treatment for osteoporosis.
Schematic model of the HOTAIRM1/miR-152-3p/ETS1 axis in regulating osteogenic differentiation. During osteogenic differentiation, HOTAIRM1 was up-regulated and miR-152-3p was down-regulated, thereby resulting in increased ETS1 expression. Overexpression of ETS1 enhanced osteogenic differentiation of HBMSCs
Data availability
The datasets generated during the present study are available from the corresponding author on reasonable request.
References
Ensrud KE, Crandall CJ (2017) Osteoporosis. Ann Intern Med 167(3):Itc17–Itc32. https://doi.org/10.7326/aitc201708010
Alejandro P, Constantinescu F (2018) A review of osteoporosis in the older adult: an update. Rheum Dis Clin North Am 44(3):437–451. https://doi.org/10.1016/j.rdc.2018.03.004
Wang C, Meng H, Wang X, Zhao C, Peng J, Wang Y (2016) Differentiation of bone marrow mesenchymal stem cells in osteoblasts and adipocytes and its role in treatment of osteoporosis. Med Sci Monit 22:226–233. https://doi.org/10.12659/msm.897044
Quinn JJ, Chang HY (2016) Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet 17(1):47–62. https://doi.org/10.1038/nrg.2015.10
Yang Q, Jia L, Li X, Guo R, Huang Y, Zheng Y et al (2018) Long noncoding RNAs: new players in the osteogenic differentiation of bone marrow- and adipose-derived mesenchymal stem cells. Stem Cell Rev Rep 14(3):297–308. https://doi.org/10.1007/s12015-018-9801-5
Wang J, Liu S, Shi J, Liu H, Li J, Zhao S et al (2020) The role of lncRNAs in osteogenic differentiation of bone marrow mesenchymal stem cells. Curr Stem Cell Res Ther 15(3):243–249. https://doi.org/10.2174/1574888x15666191227113742
Shang G, Wang Y, Xu Y, Zhang S, Sun X, Guan H et al (2018) Long non-coding RNA TCONS_00041960 enhances osteogenesis and inhibits adipogenesis of rat bone marrow mesenchymal stem cell by targeting miR-204-5p and miR-125a-3p. J Cell Physiol 233(8):6041–6051. https://doi.org/10.1002/jcp.26424
Wang X, Zhao D, Zhu Y, Dong Y, Liu Y (2019) Long non-coding RNA GAS5 promotes osteogenic differentiation of bone marrow mesenchymal stem cells by regulating the miR-135a-5p/FOXO1 pathway. Mol Cell Endocrinol 496:110534. https://doi.org/10.1016/j.mce.2019.110534
Wang Q, Li Y, Zhang Y, Ma L, Lin L, Meng J et al (2017) LncRNA MEG3 inhibited osteogenic differentiation of bone marrow mesenchymal stem cells from postmenopausal osteoporosis by targeting miR-133a-3p. Biomed Pharmacother 89:1178–1186. https://doi.org/10.1016/j.biopha.2017.02.090
Fu L, Peng S, Wu W, Ouyang Y, Tan D, Fu X (2019) LncRNA HOTAIRM1 promotes osteogenesis by controlling JNK/AP-1 signalling-mediated RUNX2 expression. J Cell Mol Med 23(11):7517–7524. https://doi.org/10.1111/jcmm.14620
Wang J, Liu S, Li J, Zhao S, Yi Z (2019) Roles for miRNAs in osteogenic differentiation of bone marrow mesenchymal stem cells. Stem Cell Res Ther 10(1):197. https://doi.org/10.1186/s13287-019-1309-7
Li B (2018) MicroRNA regulation in osteogenic and adipogenic differentiation of bone mesenchymal stem cells and its application in bone regeneration. Curr Stem Cell Res Ther 13(1):26–30. https://doi.org/10.2174/1574888x12666170605112727
Li Q, Xing W, Gong X, Wang Y, Sun H (2019) Astragalus polysaccharide promotes proliferation and osteogenic differentiation of bone mesenchymal stem cells by down-regulation of microRNA-152. Biomed Pharmacother. https://doi.org/10.1016/j.biopha.2019.108927
Han S, Kuang M, Sun C, Wang H, Wang D, Liu Q (2020) Circular RNA hsa_circ_0076690 acts as a prognostic biomarker in osteoporosis and regulates osteogenic differentiation of hBMSCs via sponging miR-152. Aging (Albany NY) 12(14):15011–15020. https://doi.org/10.18632/aging.103560
di Val R, Cervo P, Lena AM, Nicoloso M, Rossi S, Mancini M, Zhou H et al (2012) p63-microRNA feedback in keratinocyte senescence. Proc Natl Acad Sci USA 109(4):1133–1138. https://doi.org/10.1073/pnas.1112257109
Del Real A, López-Delgado L, Sañudo C, García-Ibarbia C, Laguna E, Perez-Campo FM et al (2020) Long noncoding RNAs as bone marrow stem cell regulators in osteoporosis. DNA Cell Biol. https://doi.org/10.1089/dna.2020.5672
Li Q, Dong C, Cui J, Wang Y, Hong X (2018) Over-expressed lncRNA HOTAIRM1 promotes tumor growth and invasion through up-regulating HOXA1 and sequestering G9a/EZH2/Dnmts away from the HOXA1 gene in glioblastoma multiforme. J Exp Clin Cancer Res 37(1):265. https://doi.org/10.1186/s13046-018-0941-x
Liang Q, Li X, Guan G, Xu X, Chen C, Cheng P et al (2019) Long non-coding RNA, HOTAIRM1, promotes glioma malignancy by forming a ceRNA network. Aging (Albany NY) 11(17):6805–6838. https://doi.org/10.18632/aging.102205
Zhang Y, Mi L, Xuan Y, Gao C, Wang YH, Ming HX et al (2018) LncRNA HOTAIRM1 inhibits the progression of hepatocellular carcinoma by inhibiting the Wnt signaling pathway. Eur Rev Med Pharmacol Sci 22(15):4861–4868. https://doi.org/10.26355/eurrev_201808_15622
Ren T, Hou J, Liu C, Shan F, Xiong X, Qin A et al (2019) The long non-coding RNA HOTAIRM1 suppresses cell progression via sponging endogenous miR-17-5p/B-cell translocation gene 3 (BTG3) axis in 5-fluorouracil resistant colorectal cancer cells. Biomed Pharmacother 117:109171. https://doi.org/10.1016/j.biopha.2019.109171
Rea J, Menci V, Tollis P, Santini T, Armaos A, Garone MG et al (2020) HOTAIRM1 regulates neuronal differentiation by modulating NEUROGENIN 2 and the downstream neurogenic cascade. Cell Death Dis 11(7):527. https://doi.org/10.1038/s41419-020-02738-w
Xin J, Li J, Feng Y, Wang L, Zhang Y, Yang R (2017) Downregulation of long noncoding RNA HOTAIRM1 promotes monocyte/dendritic cell differentiation through competitively binding to endogenous miR-3960. Onco Targets Ther 10:1307–1315. https://doi.org/10.2147/ott.s124201
Chen ZH, Wang WT, Huang W, Fang K, Sun YM, Liu SR et al (2017) The lncRNA HOTAIRM1 regulates the degradation of PML-RARA oncoprotein and myeloid cell differentiation by enhancing the autophagy pathway. Cell Death Differ 24(2):212–224. https://doi.org/10.1038/cdd.2016.111
Chen Z, Zheng J, Hong H, Chen D, Deng L, Zhang X et al (2020) lncRNA HOTAIRM1 promotes osteogenesis of hDFSCs by epigenetically regulating HOXA2 via DNMT1 in vitro. J Cell Physiol. https://doi.org/10.1002/jcp.29695
Wang L, Wang Y, Lin J (2018) MiR-152-3p promotes the development of chronic myeloid leukemia by inhibiting p27. Eur Rev Med Pharmacol Sci 22(24):8789–8796. https://doi.org/10.26355/eurrev_201812_16646
Feng F, Liu H, Chen A, Xia Q, Zhao Y, Jin X et al (2019) miR-148-3p and miR-152-3p synergistically regulate prostate cancer progression via repressing KLF4. J Cell Biochem 120(10):17228–17239. https://doi.org/10.1002/jcb.28984
Zhang A, Qian Y, Qian J (2019) MicroRNA-152-3p protects neurons from oxygen-glucose-deprivation/reoxygenation-induced injury through upregulation of Nrf2/ARE antioxidant signaling by targeting PSD-93. Biochem Biophys Res Commun 517(1):69–76. https://doi.org/10.1016/j.bbrc.2019.07.012
Roux M, Perret C, Feigerlova E, Mohand Oumoussa B, Saulnier PJ, Proust C et al (2018) Plasma levels of hsa-miR-152-3p are associated with diabetic nephropathy in patients with type 2 diabetes. Nephrol Dial Transpl 33(12):2201–2207. https://doi.org/10.1093/ndt/gfx367
Ma P, Zhang C, Huo P, Li Y, Yang H (2020) A novel role of the miR-152-3p/ERRFI1/STAT3 pathway modulates the apoptosis and inflammatory response after acute kidney injury. J Biochem Mol Toxicol. https://doi.org/10.1002/jbt.22540
Dittmer J (2015) The role of the transcription factor Ets1 in carcinoma. Semin Cancer Biol. https://doi.org/10.1016/j.semcancer.2015.09.010
Gao Y, Ganss BW, Wang H, Kitching RE, Seth A (2005) The RING finger protein RNF11 is expressed in bone cells during osteogenesis and is regulated by Ets1. Exp Cell Res 304(1):127–135. https://doi.org/10.1016/j.yexcr.2004.10.031
Koyama T, Kamemura K (2015) Global increase in O-linked N-acetylglucosamine modification promotes osteoblast differentiation. Exp Cell Res 338(2):194–202. https://doi.org/10.1016/j.yexcr.2015.08.009
Qi MC, Hu J, Zou SJ, Chen HQ, Zhou HX, Han LC (2008) Mechanical strain induces osteogenic differentiation: Cbfa1 and Ets-1 expression in stretched rat mesenchymal stem cells. Int J Oral Maxillofac Surg 37(5):453–458. https://doi.org/10.1016/j.ijom.2007.12.008
Fan Q, Li Y, Sun Q, Jia Y, He C, Sun T (2020) miR-532-3p inhibits osteogenic differentiation in MC3T3-E1 cells by downregulating ETS1. Biochem Biophys Res Commun 525(2):498–504. https://doi.org/10.1016/j.bbrc.2020.02.126
Hua L, Zhang X (2021) MALAT1 regulates osteogenic differentiation of human periodontal ligament stem cells through mediating miR-155-5p/ETS1 axis. Tissue Cell 73:101619. https://doi.org/10.1016/j.tice.2021.101619
Ren L, Guo L, Kou N, Lv J, Wang Z, Yang K (2021) LncRNA LINC00963 promotes osteogenic differentiation of hBMSCs and alleviates osteoporosis progression by targeting miRNA-760/ETS1 axis. Autoimmunity 54(6):313–325. https://doi.org/10.1080/08916934.2021.1922890
Xia W, Han X, Wang L (2021) E26 transformation-specific 1 is implicated in the inhibition of osteogenic differentiation induced by chronic high glucose by directly regulating Runx2 expression. J Biomed Res 36(1):39–47. https://doi.org/10.7555/jbr.35.20210123
Li R, Dong Y, Li F (2021) ETS proto-oncogene 1 suppresses MicroRNA-128 transcription to promote osteogenic differentiation through the HOXA13/β-catenin axis. Front Physiol 12:626248. https://doi.org/10.3389/fphys.2021.626248
Arthur A, Gronthos S (2020) Clinical application of bone marrow mesenchymal stem/stromal cells to repair skeletal tissue. Int J Mol Sci. https://doi.org/10.3390/ijms21249759
Jiang Y, Zhang P, Zhang X, Lv L, Zhou Y (2021) Advances in mesenchymal stem cell transplantation for the treatment of osteoporosis. Cell Prolif 54(1):e12956. https://doi.org/10.1111/cpr.12956
Funding
None.
Author information
Authors and Affiliations
Contributions
All authors made substantial contribution to conception and design, acquisition of the data, or analysis and interpretation of the data; take part in drafting the article or revising it critically for important intellectual content; gave final approval of the revision to be published; and agree to be accountable for all aspect of the work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no financial conflict of interest.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Written informed consents were obtained from all participants and this study was permitted by the Ethics Committee of Tianjin Medical University General Hospital.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Liu, Y. & Lei, P. LncRNA HOTAIRM1 promotes osteogenic differentiation of human bone marrow-derived mesenchymal stem cells by targeting miR-152-3p/ETS1 axis. Mol Biol Rep 50, 5597–5608 (2023). https://doi.org/10.1007/s11033-023-08466-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08466-6