Abstract
Cyanobacteria are an ancient group of photoautotrophic prokaryotes, and play an essential role in the global carbon cycle. They are also model organisms for studying photosynthesis and circadian regulation, and metabolic engineering and synthetic biology strategies grants light-driven biotechnological applications to cyanobacteria, especially for engineering cyanobacteria cells to achieve an efficient light-driven system for synthesizing any product of interest from renewable feedstocks. However, lower yield limits the potential of industrial application of cyanobacterial synthetic biology, and some key limitations must be overcome to realize the full biotechnological potential of these versatile microorganisms. Although genetic engineering toolkits for cyanobacteria have made some progress, the tools available still lag behind conventional heterotrophic microorganism. Consequently, this study describes the current situations and limitations of genetic engineering in cyanobacteria, and further improvements are proposed to improve the output of targeted products. We believe that cyanobacteria-mediated light-driven platforms towards efficient synthesis of green chemicals could unlock a bright future by developing the tools for strain manipulation and novel chassis organisms with excellent performance for biotechnological applications, which could also accelerate the advancement of bio-manufacturing industries.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With the deepening of social industrialization, a large number of chemical products, polymers and other products that improve our life quality are manufactured by petroleum-derived feedstocks, which will be accompanied by serious problems [1, 2]. Scientific researchers are committed to developing sustainable production processes towards efficient synthesis of green chemicals, and significant progress has been achieved on traditional heterotrophic microorganisms, which use organic carbon sources as the energy source to power the synthesis of NADPH and ATP, leading to increased chemical synthesis costs. Recently, the increasing concentration of carbon dioxide in the atmosphere has led to global warming and ocean acidification [3], and sequestration by photosynthetic microorganisms that converts atmospheric CO2 into valuable chemical products has attracted extensive attention due to the fact that CO2 is a cheap, abundant and sustainable carbon source and photosynthesis of microorganisms will help to reduce the accumulation of CO2 in the environment. Among the various photosynthetic microorganisms, microalgae have become a research focus due to their strong environmental adaptability and abundant metabolic diversity and have been applied in many fields, including renewable energy, pharmaceutical industry, environmental monitoring, food industry, biotechnology, animal feed and environmental purification (Fig. 1). Adhering to the advantage of microalgae’s natural carbon fixation capacity may help achieve the goal of “carbon neutrality and carbon emission peak”.
Microalgae have attracted much attention due to their strong environmental adaptability and abundant metabolic diversity, and have been applied in many fields, including renewable energy, pharmaceutical industry, environmental monitoring, food industry, biotechnology, animal feed and environmental purification
As an important component of microalgae, cyanobacteria are an ancient group of photoautotrophic prokaryotes, and play an essential role in the global carbon cycle [4]. Compared with traditional higher plants and eukaryotic algae, cyanobacteria have the following advantages: (I) High energy conversion efficiency. Cyanobacteria can convert up to 9.0% of solar energy into biomass, while the solar energy conversion efficiency of higher plants is only 0.5-3.0%. (II) High carbon sequestration ability. Cyanobacteria can use bicarbonate to efficiently capture CO2 in the environment through its unique carbon concentration mechanism, further improving the utilization rate of CO2 by cyanobacteria. (III) Simple nutritional demands and rapid growth rate. The growth of cyanobacteria only requires sunlight, water, carbon dioxide and trace element compounds, and the simple nutritional demands and photosynthetic structure enable them to grow at a faster rate. In addition, cyanobacteria could be discovered in different niches, providing algae strains with unique stress resistance. (IV) Independency of fertile land. Cyanobacteria can be cultured in flasks at the laboratory scale or grown in bioreactors or runway ponds at outdoor scales, which could avoid competition with food crops for arable land. (V) Tractable genetics. Simple genomic information and convenient genetic manipulation of cyanobacterial cells pave the way for targeted regulation of intracellular metabolic flow through metabolic engineering. Consequently, cyanobacteria have served as an important model organism for studying photosynthesis and are of considerable interest for applications in light-driven biotechnological applications.
Although cyanobacteria own the above advantages, especially high carbon fixation ability, they lack the original materials for the efficient synthesis of valuable chemical compounds. To date, the current studies focus on photosynthesis and circadian regulation in non-modified cyanobacteria [5,6,7], and there are still several challenges in producing non-natural chemicals whose biosynthetic routes have not been identified in wild-type cyanobacterial strains. Metabolic engineering and synthetic biology have attracted extensive attention to redesign and reconstruct the metabolic pathway of model algae, by which the designated DNA fragments can be assembled for the production of commodity chemicals widely used in medical, health care and cosmetics. To date, many de novo synthesis metabolic pathways have been explored to synthesize different kinds of valuable chemical products in model freshwater cyanobacteria with the drawback of lower yield [8, 9]. There are still only a few examples of having produced highly effective photosynthetic chemicals with titers of more than 1 g/L. For instance, the highest production titer of ethanol was reported in Synechocystis sp. PCC6803 (PCC6803) with a titer of 5.5 g/L [10], and photosynthetic 2,3-butanediol and isobutyraldehyde production have achieved titers of 2.4 g/L and 1.1 g/L, respectively [3, 11]. Consequently, genetic engineering should be used to improve the photosynthetic titers of chemicals [8, 12], and some key limitations must be overcome to realize the full biotechnological potential of these versatile microorganisms.
Genetic manipulation tools for precise gene expression control is lacking
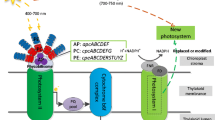
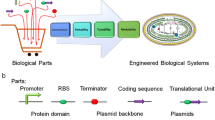
Cyanobacteria lack genetic manipulation tools for precise gene expression control, making it difficult to reach the theoretical yield and limiting the industrialization process of algae. Efficient expressions of heterologous genes play a key role in synthesizing biological products in cyanobacteria (Fig. 2A). A previous study proved that the expression ability of heterogenic genes is far less than that of endogenous genes, and its highest expression level only accounts for about 3% of the total soluble protein in cells [13], which significantly limits the application potential of cyanobacteria in the production of chemical products. Thus, there is an urgency to develop a series of genetic toolboxes that could precisely regulate gene expression in cyanobacteria to overcome the challenges mentioned above.
Genetic engineering tools of cyanobacteria towards efficient synthesis of green chemicals. (A) A library of synthetic parts such as promoter and RBS could be designed and characterized for tuneability of heterologous gene in cyanobacteria. (B) Schematic drawing of genome editing in cyanobacteria via the traditional homologous recombination mediated cloning vector or self-replicating plasmids
The expression of heterologous genes was mainly regulated by transcription and translation, while the former played an important role in prokaryotes [14]. In general, the transcription rates were controlled by changing the strength and accessibility of promoters [15]. Previous studies have shown that mutagenesis-based constitutive promoter library and inducible promoter system have been successfully established in Escherichia coli and Saccharomyces cerevisiae [16,17,18]. However, after testing the promoter elements from E. coli and S. cerevisiae in cyanobacteria cells, it was found that they did not obtain the desired results, and even the opposite results would appear. For example, the Ptrc1O promoter derived from E. coli displayed high efficiency in PCC6803, which was 3-fold higher than the endogenous PrbcL promoter. Nevertheless, the other promoters derived from E. coli, such as Plac, Ptet and λPR, showed low efficiency in PCC6803 [19], and the reason for this phenomenon was that the composition of RNA polymerase in cyanobacteria was significantly different from that of non-photosynthetic bacteria [20].
At present, a series of natural promoters have been excavated from essential genes and highly expressed genes in cyanobacteria, which are mainly involved in the fixation of carbon dioxide (PrbcL, Pcmp and Psbt), photosystem I (PpsaA and PpsaD), photosystem II (PpsbA1 and PpsbA2), and phycocyanin synthesis (Pcpc) [21, 22]. Besides, a natural nickel-induced promoter PnrsB was also successfully used to express the phage cleavage gene in PCC6803 [23]. Meanwhile, several groups have made great strides in developing a range of inducible promoter systems in PCC6803 and Synechococcus sp. PCC 7942 (PCC7942), including photo-responsive promoters, IPTG-responsive promoters, nutrients-responsive promoters, and metal-responsive promoters [24,25,26,27]. Despite some progresses achieved, many problems remain. For example, the strength of the same promoter showed different performances between different cyanobacterial strains [28]. Consequently, it is urgent to design and excavate more promoter elements to promote the application potential of the cyanobacteria expression system.
In addition, the regulation of gene expression at the translational level remained comparatively essential. The ribosome binding site (RBS) is a small nucleotide sequence, and the Shine-Dalgarno (SD) sequence in RBS could interact with 16 S rRNA via complementary pairing. Different RBS sequences have different affinities with ribosomes, and the speed of RNA translation could be controlled by changing the adaptability of the RBS sequence. Presently, the screening of different RBS sequences has shown great insights in PCC6803 and Synechococcus sp. PCC7002 (PCC7002) [29, 30], however, the same RBS sequence expressed in different strains had different translation initiation rates [20]. In addition to the SD sequences, an appropriately sized spacer region between the SD sequences and start codon also significantly affected the activity of RBS [31], and thermodynamic models were also used to predict the translation efficiency of the given RBS sequence in different organisms [32]. Until now, only few RBS sequences have been characterized in cyanobacteria, and an RBS library should be constructed to improve the translation efficiency of heterologous genes in cyanobacteria.
The genome editing of cyanobacteria is simple, the cycle is relatively slow
Although the genome editing of cyanobacteria is simple, the cycle is relatively slow (Fig. 2B). An optimum chassis for synthetic biology applications should satisfy the following requirements. On the one hand, the complete genome annotation of the chassis should be obtained. On the other hand, the chassis should allow for genome editing with simplicity and high efficiency. To date, more than 40 cyanobacteria genomes have been sequenced, and the inherent advantages of cyanobacteria should be combined to further develop synthetic biology tools, which could be applied to the genome editing of cyanobacteria with important theoretical value and potential applications [22]. Furthermore, the neutral sites on chromosomes and the endogenous plasmids are often used as target sites to express the heterologous genes in cyanobacteria [33, 34]. The common method to edit the cyanobacteria genome by natural transformation and homologous recombination in cyanobacteria genetic engineering is time-consuming (multiple antibiotic screening), an inefficient method (complete segregation in multi-copy plasmid), and may sometimes cause the loss of foreign genes.
The integrative expression on chromosomes could increase the genetic stability of heterologous genes compared to free plasmids, which is conducive to the long-term subculture of transgenic lines. It was reported that gene copy numbers positively influenced the overall expression of a heterologous protein in several endogenous plasmids, which is crucial for optimum chassis with high production performance. For example, Nozzi et al. (2017) explored integrating the heterologous pathway of 2,3-butanediol into the chromosome and endogenous plasmid pAQ1 of PCC7002, and the results showed that the titer of 2,3-butanediol in the transgenic strain with an integrative expression on the endogenous plasmid was higher, demonstrating the superiority of heterologous gene expression via endogenous plasmid [35]. However, the long growth cycle and multi-copy genome provides a longer period to obtain transgenic strain with the complete segregation for each copy, which slows down the efficiency of genetic manipulation in cyanobacteria. To overcome this challenge, CRISPR-based technology, a promising genome editing tool, has been used. CRISPR-based genome editing technology has been widely applied in various cyanobacteria strains [36, 37]. However, the major challenge for CRISPR system application was off-target effects, which could influence the genome editing efficiency [38]. In perspective, CRISPR-based genome editing is a promising strategy in the field of cyanobacterial genetics due to its potential to solve existing difficulties.
Furthermore, introducing heterologous genes into cyanobacteria cells with the help of self-replicating plasmids overcame the limitations in homologous recombination, such as low segregation efficiency and difficult integration of large inserted fragments into the genome. Meanwhile, the expression intensity of heterologous genes derived from self-replicating plasmids was usually higher than that derived from chromosomes [39]. Currently, self-replicating plasmids can be classified into two types, broad host-range plasmid [40] and cryptic plasmid [41], and have been designed to conduct a rapid functional study of novel genes or metabolic pathways [42]. Meanwhile, broad host-range plasmids were widely used to efficiently express heterologous genes in cyanobacteria. For instance, Miao et al. (2017) constructed a series of self-replicating plasmids derived from the broad host-range vector pEEK2 to express the synthesis route of isobutanol in PCC6803 [43]. The hydrocarbon biosynthetic pathway was also constructed via the broad-host-range vector pVZ321 in PCC7002 [44]. However, the plasmid stability remained to be considered when using self-replicating plasmids. Up to now, the self-replicating plasmids used in cyanobacteria were located on the RSF1010-derived plasmids, and the copy number of these plasmids was slightly higher than that of the chromosome in Synechocystis [45]. However, RSF1010-derived plasmids tended to be slowly eliminated from cells, and antibiotic-based selection pressure should be used to maintain the functions of these plasmids. Huang et al. (2010) constructed the broad host-range vector pPMQAK1 derived from the RSF1010 plasmid to express the GFPmut3B gene in cyanobacteria, which could be stored in cells for three months under appropriate antibiotic selection pressure [19]. Further, a series of shuttle expression vectors have been constructed in model cyanobacteria in recent years [39, 46]. Although the number of shuttle expression vectors increased with the rapid development of synthetic biology and metabolic engineering in cyanobacteria, developing stable shuttle expression vectors in non-model cyanobacteria still faced some challenges, among which screening a suitable replication initiation site for specific cyanobacteria strain remains particularly important.
Conclusions and perspectives
Several cyanobacterial chassis strains offer great promise to produce green chemicals with the drawback of lower yield, and further improvements are required. Firstly, cyanobacteria are difficult to reach the theoretical yield due to the deficiency of genetic manipulation tools, and novel promoters or ribosome binding sites for precise gene expression control should be further excavated. Secondly, the traditional homologous recombination mediated approach should integrate heterologous expression module into chromosomes or endogenous plasmids with the challenge of time-consuming, and the use of broad host-range vector faces the problem of compatibility. Further, developing stable shuttle expression vectors in cyanobacteria will provide us with better research capabilities in cyanobacteria-mediated biotechnological applications. Furthermore, optimization of CRISPR-based technologies should be used to improve the efficiency of genome modification. Thirdly, positive selection and maintenance of engineered strains often requires the use of antibiotics, and only a limited number of antibiotic markers are available. Due to the toxicity of antibiotics to algae cells and their environmentally unfriendly properties, strategies based on alternative nutrient for selection pave the way for the practical resource for positive selection of engineered strains [47, 48]. Meanwhile, two-stage selection/counter-selection and site-specific recombinase systems have been used in cyanobacteria [49, 50], and more marker-less editing strategies should be further developed. Fourth, a large number of heterologous metabolic pathways have been attempted in model freshwater cyanobacteria, however, large-scale cultivation of these model organisms using finite fresh water resources is not the best option from the perspective of economic and environmental protection. Several marine microalgae species should be excavated to overcome the above shortcomings. Finally, the omics technology should be further explored, and strategies based on dynamic metabolic regulation could also be applied to redirect the carbon flux to low-flux pathways to improve the output of targeted products. By developing the tools for strain manipulation and novel chassis organisms with excellent performance for biotechnological applications, cyanobacteria-mediated light-driven platforms towards efficient synthesis of green chemicals could unlock a bright future, which could also accelerate the advancement of bio-manufacturing industries.
Data Availability
No data available.
References
Hernández N, Williams RC, Cochran EW (2014) The battle for the “green” polymer. Different approaches for biopolymer synthesis: bioadvantaged vs. bioreplacement. Org Biomol Chem 12(18):2834–2849. https://doi.org/10.1039/C3OB42339E
Kircher M (2015) Sustainability of biofuels and renewable chemicals production from biomass. Curr Opin Chem Biol 29:26–31. https://doi.org/10.1016/j.cbpa.2015.07.010
Oliver JWK, Machado IMP, Yoneda H, Atsumi S (2014) Combinatorial optimization of cyanobacterial 2,3-butanediol production. Metab Eng 22:76–82. https://doi.org/10.1016/j.ymben.2014.01.001
Knoot CJ, Ungerer J, Wangikar PP, Pakrasi HB (2018) Cyanobacteria: promising biocatalysts for sustainable chemical production. J Biol Chem 293(14):5044–5052. https://doi.org/10.1074/jbc.R117.815886
Bell-Pedersen D, Cassone VM, Earnest DJ, Golden SS, Hardin PE, Thomas TL, Zoran MJ (2005) Circadian rhythms from multiple oscillators: lessons from diverse organisms. Nat Rev Genet 6(7):544–556. https://doi.org/10.1038/nrg1633
Paddock ML, Boyd JS, Adin DM, Golden SS (2013) Active output state of the Synechococcus Kai circadian oscillator. Proc Natl Acad Sci USA 110(40):E3849–3857. https://doi.org/10.1073/pnas.1315170110
Vinyard DJ, Mayfield SP, Gimpel J, Ananyev GM, Cornejo MA, Golden SS, Dismukes GC (2013) Natural variants of photosystem II subunit D1 tune photochemical fitness to solar intensity. J Biol Chem 288(8):5451–5462. https://doi.org/10.1074/jbc.M112.394668
Hirokawa Y, Kubo T, Soma Y, Saruta F, Hanai T (2020) Enhancement of acetyl-CoA flux for photosynthetic chemical production by pyruvate dehydrogenase complex overexpression in Synechococcus elongatus PCC7942. Metab Eng 57:23–30. https://doi.org/10.1016/j.ymben.2019.07.012
Savakis P, Hellingwerf KJ (2015) Engineering cyanobacteria for direct biofuel production from CO2. Curr Opin Biotechnol 33:8–14. https://doi.org/10.1016/j.copbio.2014.09.007
Gao Z, Zhao H, Li Z et al (2012) Photosynthetic production of ethanol from carbon dioxide in genetically engineered cyanobacteria[J]. Energ Environ Sci 5(12):9857–9865. https://doi.org/10.1039/C2EE22675H
Atsumi S, Higashide W, Liao JC (2009) Direct photosynthetic recycling of carbon dioxide to isobutyraldehyde. Nat Biotechnol 27(12):1177–1180. https://doi.org/10.1038/nbt.1586
Choi SY, Lee HJ, Choi J et al (2016) Photosynthetic conversion of CO2 to farnesyl diphosphate-derived phytochemicals (amorpha-4, 11-diene and squalene) by engineered cyanobacteria[J]. Biotechnol biofuels 9(1):1–12. https://doi.org/10.1186/s13068-016-0617-8
Shestakov SV, Khyen NT (1970) Evidence for genetic transformation in blue-green alga Anacystis nidulans. Molec Gen Genetics 107(4):372–375. https://doi.org/10.1007/BF00441199
Makrides SC (1996) Strategies for achieving high-level expression of genes in Escherichia coli. Microbiol Mol Biol Rev 60(3):512–538. https://doi.org/10.1006/mpat.1996.0056
Balzer S, Kucharova V, Megerle J, Lale R, Brautaset T, Valla S (2013) A comparative analysis of the properties of regulated promoter systems commonly used for recombinant gene expression in Escherichia coli. Microb Cell Fact 12(1):26–39. https://doi.org/10.1186/1475-2859-12-26
Alper H, Fischer C, Nevoigt E, Stephanopoulos G (2005) Tuning genetic control through promoter engineering. Proc Natl Acad Sci U S A 102(36):12678–12683. https://doi.org/10.1073/pnas.0504604102
Curran KA, Crook NC, Karim AS, Gupta A, Wagman AM, Alper HS (2014) Design of synthetic yeast promoters via tuning of nucleosome architecture. Nat Commun 5:4002–4009. https://doi.org/10.1038/ncomms5002
Brosius J, Erfle M, Store J (1985) Spacing of the – 10 and – 35 regions in the tac promoter. Effect on its in vivo activity. J Biol Chem 260(6):3539–3541. https://doi.org/10.1016/0165-022X(85)90070-3
Huang HH, Camsund D, Lindblad P, Heidorn T (2010) Design and characterization of molecular tools for a synthetic biology approach towards developing cyanobacterial biotechnology. Nucleic Acids Res 38(8):2577–2593. https://doi.org/10.1093/nar/gkq164
Heidorn T, Camsund D, Huang H-H, Lindberg P, Oliveira P, Stensjo K, Lindblad P (2011) Synthetic biology in cyanobacteria engineering and analyzing novel functions. Methods Enzymol 497:539–579. https://doi.org/10.1016/B978-0-12-385075-1.00024-X
Takahama K, Matsuoka M, Nagahama K, Ogawa T (2003) Construction and analysis of a recombinant cyanobacterium expressing a chromosomally inserted gene for an ethylene-forming enzyme at the psbAI locus. J Biosci Bioeng 95:302–305. https://doi.org/10.1016/S1389-1723(03)80034-8
Wang B, Wang J, Zhang W, Meldrum DR (2012) Application of synthetic biology in cyanobacteria and algae. Front Microbiol 3:344–358. https://doi.org/10.3389/fmicb.2012.00344
Liu X III RC (2009) Nickel-inducible lysis system in Synechocystis sp. PCC6803. Proc Natl Acad Sci USA 27(10):21550–21554. https://doi.org/10.1073/pnas.0911953106
Berto P, D’Adamo S, Bergantino E, Vallese F, Giacometti GM, Costantini P (2011) The cyanobacterium Synechocystis sp. PCC 6803 is able to express an active [FeFe]-hydrogenase without additional maturation proteins. Biochem Biophys Res Commun 405(4):678–683. https://doi.org/10.1016/j.bbrc.2011.01.095
Camsund D, Heidorn T, Lindblad P (2014) Design and analysis of LacI-repressed promoters and DNA-looping in a cyanobacterium. J Biol Eng 8(1):4–26. https://doi.org/10.1186/1754-1611-8-4
Qi Q, Hao M, Ng W-o, Slater SC, Baszis SR, Weiss JD, Valentin HE (2005) Application of the Synechococcus nirA promoter to establish an inducible expression system for engineering the Synechocystis tocopherol pathway. Appl Environ Microbiol 71(10):5678–5684. https://doi.org/10.1128/AEM.71.10.5678-5684.2005
Peca L, Kos PB, Vass I (2007) Characterization of the activity of heavy metal-responsive promoters in the cyanobacterium Synechocystis PCC6803. Acta Biol Hung 58:11–22. https://doi.org/10.1556/ABiol.58.2007.Suppl.2
Berla BM, Saha R, Immethun CM, Maranas CD, Moon TS, Pakrasi HB (2013) Synthetic biology of cyanobacteria: unique challenges and opportunities. Front Microbiol 4:246–259. https://doi.org/10.3389/fmicb.2013.00246
Ma J, Campbell A, Karlin S (2002) Correlations between Shine-Dalgarno sequences and gene features such as predicted expression levels and operon structures. J Bacteriol 184(20):5733–5745. https://doi.org/10.1128/JB.184.20.5733-5745.2002
Markley AL, Begemann MB, Clarke RE, Gordon GC, Pfleger BF (2015) Synthetic biology toolbox for controlling gene expression in the cyanobacterium Synechococcus sp. strain PCC7002. ACS Synth Biol 4(5):595–603. https://doi.org/10.1021/sb500260k
Chen H, Bjerknes M, Kumar R, Jay E (1994) Determination of the optimal aligned spacing between the Shine–Dalgarno sequence and the translation initiation codon of Escherichia coli mRNAs. Nucleic Acids Res 22:4953–4957. https://doi.org/10.1093/nar/22.23.4953
Salis HM, Mirsky EA, Voigt CA (2009) Automated design of synthetic ribosome binding sites to control protein expression. Nat Biotechnol 27(10):946–950. https://doi.org/10.1038/nbt.1568
Ng AH, Berla BM, Pakrasi HB (2015) Fine-tuning of photoautotrophic protein production by combining promoters and neutral sites in the cyanobacterium Synechocystis sp. Strain PCC6803. Appl Environ Microbiol 81(19):6857–6863. https://doi.org/10.1128/AEM.01349-15
Xu Y, Alvey RM, Byrne PO, Graham JE, Shen G, Bryant DA (2011) Expression of genes in cyanobacteria: adaptation of endogenous plasmids as platforms for high-level gene expression in Synechococcus sp. PCC7002. Methods Mol Biol 684:273–293. https://doi.org/10.1007/978-1-60761-925-3_21
Nozzi NE, Case AE, Carroll AL, Atsumi S (2017) Systematic approaches to efficiently produce 2,3-butanediol in a marine cyanobacterium. ACS Synth Biol 6(11):2136–2144. https://doi.org/10.1021/acssynbio.7b00157
Behler J, Vijay D, Hess WR, Akhtar MK (2018) CRISPR-based technologies for metabolic engineering in cyanobacteria. Trends Biotechnol 36(10):996–1010. https://doi.org/10.1016/j.tibtech.2018.05.011
Niu TC, Lin GM, Xie LR, Wang ZQ, Xing WY, Zhang JY, Zhang CC (2019) Expanding the potential of CRISPR-Cpf1-based genome editing technology in the cyanobacterium Anabaena PCC7120. ACS Synth Biol 8(1):170–180. https://doi.org/10.1021/acssynbio.8b00437
Li B, Zhao W, Luo X, Zhang X, Li C, Zeng C, Dong Y (2017) Engineering CRISPR-Cpf1 crRNAs and mRNAs to maximize genome editing efficiency. Nat Biomed Eng 1(5):1–10. https://doi.org/10.1038/s41551-017-0066
Chen Y, Taton A, Go M, London RE, Pieper LM, Golden SS, Golden JW (2016) Self-replicating shuttle vectors based on pANS, a small endogenous plasmid of the unicellular cyanobacterium Synechococcus elongatus PCC7942. Microbiology 162(12):2029–2041. https://doi.org/10.1099/mic.0.000377
Lin PC, Saha R, Zhang F, Pakrasi HB (2017) Metabolic engineering of the pentose phosphate pathway for enhanced limonene production in the cyanobacterium Synechocystis sp. PCC6803. Sci rep 7(1):1–10. https://doi.org/10.1038/s41598-017-17831-y
Deng M, Coleman JR (1999) Ethanol synthesis by genetic engineering in cyanobacteria. Appl Environ Microbiol 65(2):523–528. https://doi.org/10.1128/AEM.65.2.523-528.1999
Liu D, Liberton M, Yu J, Pakrasi HB, Bhattacharyya-Pakrasi M (2018) Engineering nitrogen fixation activity in an oxygenic phototroph. mBio 9(3):18–29. https://doi.org/10.1101/316547
Miao R, Liu X, Englund E, Lindberg P, Lindblad P (2017) Isobutanol production in Synechocystis PCC6803 using heterologous and endogenous alcohol dehydrogenases. Metab Eng Commun 5:45–53. https://doi.org/10.1016/j.meteno.2017.07.003
Knoot CJ, Pakrasi HB (2019) Diverse hydrocarbon biosynthetic enzymes can substitute for olefin synthase in the cyanobacterium Synechococcus sp. PCC7002. Sci Rep 9(1):1360–1371. https://doi.org/10.1038/s41598-018-38124-y
Eaton-Rye JJ (2004) The construction of gene knockouts in the cyanobacterium Synechocystis sp. PCC 6803. Photosynthesis Res Protocols Humana Press 309–324. https://doi.org/10.1002/bit.25713
Liu D, Pakrasi HB (2018) Exploring native genetic elements as plug-in tools for synthetic biology in the cyanobacterium Synechocystis sp. PCC6803. Microb Cell Fact 17(1):48–55. https://doi.org/10.1186/s12934-018-0897-8
Polyviou D, Hitchcock A, Baylay AJ, Moore CM, Bibby TS (2015) Phosphite utilization by the globally important marine diazotroph Trichodesmium. Environ Microbiol Rep 7:824–830. https://doi.org/10.1111/1758-2229.12308
Selao TT, Włodarczyk A, Nixon PJ, Norling B (2019) Growth and selection of the cyanobacterium Synechococcus sp. PCC7002 using alternative nitrogen and phosphorus sources. Metab Eng 54:255–263. https://doi.org/10.1016/j.ymben.2019.04.013
Lea-Smith DJ, Vasudevan R, Howe CJ (2016) Generation of marked and markerless mutants in model cyanobacterial species. J Vis Exp 111:54001. https://doi.org/10.3791/54001
Tan X, Liang F, Cai K, Lu X (2013) Application of the FLP/FRT recombination system in cyanobacteria for construction of markerless mutants. Appl Microbiol Biotechnol 97:6373–6382. https://doi.org/10.1007/s00253-013-4837-6
Acknowledgements
The authors thank the timely help given by Dr. Wei Xu, Chaobo Zhang, Jiashun Li, and Jiachao Xin during the revision process of this manuscript. Our deepest gratitude goes to the editors and anonymous reviewers for their careful work and thoughtful suggestions that have helped improve this paper substantially. Jie Cheng wants to thank, in particular, for the patience, care, and support received from Xiongyan Du over the years.
Funding
This work was financially supported by the Natural Science Foundation of Hainan Province (grant #422QN265) and the National Key Research and Development Program of China (2019YFA0904600; 2022YFC3401803).
Author information
Authors and Affiliations
Contributions
Jie Cheng and Kaidian Zhang conceived and designed the project. Yuyong Hou performed constructive discussions and interpreted the significance of this study. Jie Cheng, Kaidian Zhang and Yuyong Hou prepared the draft and finalized the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, J., Zhang, K. & Hou, Y. The current situations and limitations of genetic engineering in cyanobacteria: a mini review. Mol Biol Rep 50, 5481–5487 (2023). https://doi.org/10.1007/s11033-023-08456-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08456-8