Abstract
Background
Shark species are overfished at a global scale, as they are poached for the finning industry or are caught as bycatch. Efficient conservation measures require fine-scale spatial and temporal studies to characterize shark habitat use, infer migratory habits, analyze relatedness, and detect population genetic differentiation. Gathering these types of data is costly and time-consuming, especially when it requires collection of shark tissue samples.
Methods and results
Genetic tools, such as microsatellite markers, are the most economical sampling method for collecting genetic data, as they enable the estimation of genetic diversity, population structure and parentage relationships and are thus an efficient way to inform conservation strategies. Here, a set of 45 microsatellite loci was tested on three blacktip reef shark (Carcharhinus melanopterus) populations from three Polynesian islands: Moorea, Morane and Tenararo. The set was composed of 10 previously published microsatellite markers and 35 microsatellite markers that were developed specifically for C. melanopterus as part of the present study. The 35 novel and 10 existing loci were cross-amplified on eight additional shark species (Carcharhinus amblyrhynchos, C. longimanus, C. sorrah, Galeocerdo cuvier, Negaprion acutidens, Prionacea glauca, Rhincodon typus and Sphyrna lewini). These species had an average of 69% of successful amplification, considered if at least 50% of the individual samples being successfully amplified per species and per locus.
Conclusions
This novel microsatellite marker set will help address numerous knowledge gaps that remain, concerning genetic stock identification, shark behavior and reproduction via parentage analysis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over a third (37.5%) of all Chondrichthyan species are threatened with extinction [8, 15, 16, 50]. All shark species are currently threatened by overfishing and bycatch [1, 23]. Sharks are mesopredators in marine ecosystems, as they have a key role in maintaining predation pressure on lower trophic levels [35, 44]. The loss of sharks in marine ecosystems has ramifications, such as inducing trophic cascades [23].
Blacktip reef sharks (Carcharhinus melanopterus) live in coral reef ecosystems and have a tropical and subtropical Indo-Pacific distribution from eastern Africa to the Red Sea and in French Polynesia where they are poached for their fins used in traditional Asian meals [10]. French Polynesia is the world’s largest shark sanctuary, spanning an area of 4.7 million km2 [56]. Blacktip reef shark populations are very abundant around all islands and atolls of French Polynesia, where they were found to be genetically isolated among the different archipelagos, isolated islands, and atolls [55]. The limited gene flow combined with existing anthropogenic threats on sharks suggests the threat of depletion of isolated local populations [7, 11, 55]. In this context, understanding how to optimize blacktip reef shark conservation strategies by targeting protection of certain types of habitats such as shark nurseries or adult individuals that have a high reproductive output is essential. Genetic analysis tools have the potential to reveal habitat use or identify lineages to improve measures aimed at increasing juvenile shark survival, as demonstrated in other vertebrates [36, 46]. Genetic markers enable the estimation of the genetic effective sizes of populations, demonstrate inter-island gene-flow and allow parentage analyses to be conducted [9, 13].
Microsatellites are short, simple DNA sequences composed of a specific motif that is repeated between 7 to 50 times [4] and dispersed throughout the eukaryotic nuclear genome [12, 28]. While such markers were developed for particular species, because of the time and resources that are required for their development, testing the cross-species transferability of these markers is essential to facilitate their use in further studies [5, 33, 40]. For C. melanopterus, several previous studies used 14 microsatellites originally developed for other shark species, such as C. limbatus [25], C. plumbeus [41] and Negaprion brevirostris [17], to explore population structure and to conduct parentage analysis. To date, no microsatellite markers were developed specifically for C. melanopterus [13]. While recent studies urge the use of single nucleotide polymorphism (SNP) markers for population genetic structure analyses, microsatellite markers remain the most cost-efficient in terms of library preparation and scoring [52, 19]. Moreover, microsatellites are especially successful when focusing on individual segregation, fingerprinting and more generally in parentage analysis as they are abundant in the genome and highly polymorphic [20, 18, 26, 14, 29].
The aim of this study was to develop a microsatellite library specifically for C. melanopterus that would provide enough microsatellite markers to study genetic structure and parentage links in blacktip reef shark populations in French Polynesia. Further, we conducted cross-amplification tests of the newly developed markers for C. melanopterus on eight other shark species.
Material and methods
Sampling
A total of 120 individuals sampled around three Pacific islands were used in this study: Moorea (an island located in the Society archipelago; 17° 29′ 31″ S, 149° 50′ 08″ O), Morane (an atoll located in the south of the Tuamotu and Gambier archipelago; 23° 10′ 0″ S, 137° 8′ 0″ W) and Teneraro (an atoll located in the Acteon Group, in the south of the Tuamotu; 21° 18′ 0″ S, 136° 45′ 0″ W). Moorea is separated from Morane and Tenararo by 1460 km and 1435 km, respectively. Morane and Tenararo are separated by 208 km. Samples from each of the respective islands were collected to ensure the representativeness of the genetic diversity of blacktip reef sharks across French Polynesia (Table S2).
For cross-species amplification tests, eight species of sharks were selected to test the transferability of the microsatellite markers, including: 15 C. amblyrhynchos, 5 C. longimanus, 15 C. sorrah, 15 Galocerdo cuvier, 8 Negaprion acutidens, 8 Prionace glauca, 15 Rhincodon typus and 15 Shyrna lewini. Samples were selected from the shark fin-clip collection at the Center for Island Research and Environmental Observatory (CRIOBE), in Moorea. Members of the CRIOBE collected C. melanopterus and N. acutidens fin clips using a gillnet with a 5 cm mesh size that was set perpendicular to shore around Moorea Island. Samples of adult sharks were collected using a fishing rod with a barbless hook or with a biopsy probe [34]. Other samples used for cross-species amplification were collected by international partner institutions (see Acknowledgments).
DNA extraction and design of C. melanopterus-specific microsatellites
Total DNA was extracted from eight C. melanopterus individuals with the QIAcube HT DNA extraction robot (QIAGEN, Hilden, Germany) and sent to GenoScreen (Lille, France) for microsatellite library preparation and sequencing. A total of 1 μg from an equimolar DNA pool (41.69 ± 0.035 ng/μL) of the eight specimens was used for the development of Illumina MiSeq Nano library, sequenced on the MiSeq platform Illumina (San Diego, CA). The obtained sequences were merged with the software PrinSeq [47], and the final analysis and primer design were completed with the software QDD v.3 [31]. Among 1,650,409 raw sequences, 867,286 merged sequences were obtained, 9355 primer sets were designed, and 1675 primer sets were selected. Based on this dataset, 50 primer pairs were selected, according to their repeat number (≥ 9), motif, and PCR product size (≥ 100 bp), and then tested. Among the 50 primer pairs, 35 loci were successfully amplified on C. melanopterus DNA through PCR amplification. Moreover, ten previously published primer pairs initially developed for Carcharhinus limbatus (Cli111, Cli103, Cli102, Cli12, Cli107), Negaprion acutidens (LS20, LS75, LS32, LS54) and C. plumbeus (Cpl128) were acquired from previously published studies (Table 1) and tested for PCR amplification on C. melanopterus.
Molecular analysis
DNA was extracted from C. melanopterus samples to characterize and test the newly developed microsatellites, using the QIAcube HT DNA extraction robot (Qiagen, Hilden, Germany) from small (< 0.5 cm3) shark fin clips. PCR amplifications were performed using theType-it Microsatellite PCR kit (Qiagen, Hilden, Germany) in 12 μL total volume reactions containing 4 μL Type-it Multiplex PCR Master Mix 2X (contains HotStarTaq® Plus DNA polymerase, Type-it Microsatellite PCR buffer with 6 mM MgCl2 and dNTPs), 6μL RNase-free water, 1 μL of primers (2 μM forward and reverse primers diluted in TE pH 8 buffer) and 1 μL of DNA template at 40 ng/μL. Forward primers were labelled with a fluorescent dye (YAKYE, 6FAM, ATTO550, or ATTO565, Applied Biosystems). Amplifications were carried out as follows; 5 min at 95 °C; followed by 45 cycles of 30 s at 95 °C, 1 min 30 s at optimal annealing temperature (53–60 °C, depending on locus, Table 1), and 30 s at 72 °C; and a final extension step of 30 min at 60 °C. The 45 loci were combined into eight multiplexes according to their size range and primer annealing temperature to perform PCR (Table 1). Each PCR product was run through a 1.5% agarose gel and visualized using ethidium bromide to verify amplifications. PCR products were sent to GenoScreen (Lille, France) and allele sizes were assessed using an Applied Biosystems 3730 Sequencer. For accurate sizing, an internal size ladder (GeneScan 500 LIZ, Applied Biosystems) was used. The same protocol was used to amplify each of the 45 markers on the eight other shark species. An additional low temperature was tested per multiplex (multiplexes 1 and 5 were amplified at 60 °C; multiplexes 2, 3, 7 and 8 were amplified at 53 °C, multiplexes 4 and 6 were amplified 55 °C).
Data analysis
The resulting electropherograms were scored using the program Geneious® v2022.1 (Geneious Prime, San Diego, CA). For the 120 blacktip reef shark samples, the software MICRO-CHECKER v.2.2.3 [53] was used to test for the presence of null alleles, scoring errors and large-allele dropouts, as the number of samples was sufficient. The R package LEA v.3.9.0 [21] was used to analyze population structure among the three islands by estimating the least-square estimates of ancestry proportions. The K ancestral populations were determined through an entropy criterion that evaluates the quality of fit of the statistical model with the data and helps select the number of ancestral populations that best explain the genotypic data [21]. The software GENODIVE v.3.0 [32] was used to validate the K number of ancestral populations found with the package LEA v.3.9.0. The software GENODIVE defines the optimal clustering by using the sum of squares to calculate the Bayesian Information Criterion (BIC), the lowest BIC value (within the clustering analysis from K = 2 to K = 10) indicates the optimal K number of ancestral populations [32]. The software GENODIVE v.3.0 allowed for the calculation of the Fst index (population structure indicator, p value ≤ 0.001, [32]) using a pairwise differentiation analysis. Allele frequencies, the total number of alleles (Na) and the number of private alleles (NPA) were estimated through the software GenAlEx v.6.503 [38]. The estimations of the observed (Ho) and expected (He) heterozygosities and the inbreeding coefficient (Fis) were performed in GENETIX v4.05.2 [6]. GENETIX also was used to compute the linkage disequilibrium (LD), after a sequential Bonferroni correction. The R package strataG v.2.4.905 [2] was used to test for the effects of bottlenecks in the populations in which significant linkage disequilibrium was found. For population where significant LD was detected, the m-ratio [22] was calculated to test for recent bottlenecks [49]. For the other shark species, selection of the highest number of individuals that were successfully amplified between the two amplification temperatures was used to compute the number of alleles, number of private alleles and optimal annealing temperature for each locus (Table S1).
Results
A total of 44 loci showed clear amplification profiles in C. melanopterus (Table S4 and Table S5), while Cli107 presented ambiguous or no peak profiles for all individuals. According to MICRO-CHECKER analysis, loci Cm7137 and Cli12 were likely to include null alleles along with low heterozygosity levels and above-average fixation indexes for Moorea and Tenararo samples (Table 2). Morane samples showed the same results for Cm7137, but Cli12 did not seem to present null alleles. Shark individuals from Morane showed null alleles for locus LS75, presenting a high fixation index and a large difference between the observed and expected heterozygosities. No evidence was found for null alleles, scoring errors or large allele drop-out at any other loci. Slight linkage disequilibrium was detected among all loci in C. melanopterus individuals from Moorea, as only 3.8% of pairwise locus combinations had a significant disequilibrium after a sequential Bonferroni correction (Table S3). For loci Cm7137 and Cli12, which presented null alleles, a significant linkage disequilibrium was recorded in 14% of the pairwise loci combinations after sequential Bonferroni correction for Moorea (Table S3). For the blacktip reef shark samples from Moorea, 35 loci were polymorphic and presented 2 to 20 alleles, while 9 loci (Cm14931, Cm14813, Cm12415, Cm20205, Cm18780, Cm11610, Cm20294, Cm14866 and Cm16167; Table 2) appeared monomorphic. The shark samples from Tenararo and Morane atolls were also monomorphic at these 9 loci and presented an additional monomorphic locus (Cm5929) (Table S5). For C. melanopterus individuals from Moorea, microsatellite markers presented a total of 195 alleles with an average of 4.4 ± 3.5 alleles per locus. Dinucleotide markers had a total of 162 alleles with an average of 5.5 ± 3.7 alleles. Trinucleotide repeats represented a total of 18 alleles with an average of 2.2 ± 1.6 alleles. Tetranucleotide repeats represented a total of 15 alleles and an average of 2.1 ± 1.3 alleles per locus.
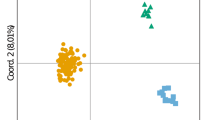
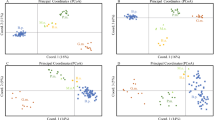
Three distinct populations were identified between the islands Moorea, Morane and Tenararo (BIC = 846, for K = 3, Fig. 1). The Fst values were high and significant (p-value ≤ 0.001): 0.153 between Moorea and Morane, 0.127 between Moorea and Tenararo, and 0.112 between Morane and Tenararo.
Cross-species amplification
All 45 microsatellite markers were tested for cross-species amplification in eight additional shark species (Table 3). Each microsatellite marker was amplified at two different annealing temperatures (the temperature for which it was successfully amplified in C. melanopterus and one additional temperature), and the temperature that supported the successful amplification of the highest number of individuals was used to compute the results (Table 3). The percentage of successfully amplified samples per locus was considered notable above 50%. Carcharhinus sorrah had the highest percentage of samples that amplified at least at 50% of individuals per locus with a total of 93%, and Sphyrna lewini had the lowest percentage of samples amplified above 50% per locus with 36%. The average total number of alleles per species was 150.6 ± 37.5, with C. sorrah accounting for the highest total number of alleles (210 alleles) and C. longimanus the lowest (95 alleles) (Table S3). The average total number of private alleles was 38.8 ± 16.2, with C. sorrah accounting for the highest number of private alleles (53) and C. longimanus the lowest (17) (Table S3).
Discussion
Microsatellite marker amplification
Overall, a total of 35 new microsatellite markers were successfully amplified for blacktip reef sharks (C. melanopterus) together with 9 published markers initially designed for other shark species. Among these 35 new loci, 57% were dinucleotides, 23% were trinucleotides and 20% were tetranucleotides, which is consistent with the findings of Richards et al. [43]. The great white shark (Carcharodon carcharias) had similar di, tri and tetranucleotide microsatellite proportions (di = 7467 (88.9%), tri = 864 (10.3%), tetra = 73 (0.9%), [43]) as blacktip reef sharks, in contrast with other fish species (Nothobranchius furzeri exhibited a di- > tetra- > trinucleotide pattern, [27]) or chimpanzees (the distribution pattern in Pan troglodytes was di- > tetra- > trinucleotide, [58]).
While it has been shown that trinucleotides are easier to genotype than dinucleotides [43], developing microsatellite markers has proven time-consuming due to their scarcity in shark genomes [48]. Dinucleotides were found to have a higher level of polymorphism (5.58 ± 3.78 alleles) than trinucleotides (2.25 ± 1.67 alleles) and tetranucleotides (2.14 ± 1.35 alleles) in blacktip reef sharks surrounding the islands of Moorea, Tenararo and Morane. Polymorphism levels for di-, tri- and tetranucleotides seem to be variable across shark species, as Galeocerdo cuvier had a higher polymorphism for tetranucleotide microsatellite marker (Na = 8, N = 101, [39]) and Carcharhinus plumbeus for dinucleotide microsatellite markers (Na = 4–39, N = 47, [40]). Moreover, all microsatellite markers that were monomorphic in blacktip reef sharks (i.e., 9 loci) were polymorphic in at least one of the eight other tested species, emphasizing the value of testing monomorphic loci on closely related species [26].
Genetic diversity
The average observed heterozygosities were 0.447 for Moorea, 0.337 for Morane and 0.351 for Tenararo population and the average expected heterozygosities were 0.442, 0.337 and 0.382, respectively. Vignaud et al. [55] analyzed the genetic structure of blacktip reef sharks across French Polynesia and found a higher expected heterozygosity for Moorea (He = 0.6, N = 38) and Tenararo (He = 0.51, N = 51), like Mourier and Planes [37] who also found a slightly higher average expected heterozygosity (Moorea, He = 0.55, N = 247). The number of microsatellite markers used (33 polymorphic loci in the present study vs. 14 loci in Vignaud et al. [55] and 17 in Mourier and Planes [37]) might be the reason for these differences. For the island of Moorea, the sample size of N = 247 used by Mourier and Planes [37], might be the cause of the slightly higher expected heterozygosity. Moreover, Moorea is a larger island (132 km2) than Morane (2 km2) and Tenararo (2.72 km2), with a more extensive reef area, and may thus have a larger effective population size. Further, Moorea’s proximity to Tahiti and Tetiaroa might provide higher genetic exchange. In contrast, the islands of Morane and Tenararo were found to be isolated atolls (Fst = 0.112) in the south of the Tuamotu Archipelago, limiting gene exchange with other blacktip reef shark populations.
Two microsatellite markers, Cm7137 and Cli12, repeatedly presented null alleles, leading to heterozygote deficiency and a significant departure from Hardy–Weinberg Equilibrium. Moreover, these loci had an above average level of LD, with 14% of their pairwise loci combinations exhibiting significant disequilibrium (the average LD of the sample of Moorea was of 3.8% which could originate from a bottleneck in the population as the M-ratio for Moorea was 0.75 ± 0.21); thus, they should be removed from further analyses. To date, no linkage disequilibrium in the pairwise locus combination was reported in other shark species (Carcharhinus limbatus, [25]; Rhincodon typus, [42]; Carcharhinus leucas, [39]; Ginglymostoma cirratum, [24]). The LD measured in blacktip reef sharks in Moorea might be caused by the structure of the population, with related subgroups within the population due to non-random mating among individuals [3, 45].
Cross-species amplification
Cross-species amplification of the 44 microsatellites had an average success of 69% (with a threshold set at 50% of the samples to define successful amplification per locus) among the eight shark species. Indeed, cross-species amplification in chondrichthyan species were often found successful due to highly conserved microsatellite flanking sequences [51]. The species that had the highest percentage of markers that were amplified in at least 50% of the samples per species were also from the genus Carcharhinus (between 82 and 93%, Table 3). These results are consistent with the phylogeny of sharks as these three shark species are closely related to C. melanopterus [30, 54]. The existing shark phylogeny includes three major clades: (i) Carcharhinus melanopterus, C. sorrah, C. longimanus, Prionace glauca and Negaprion acutidens are in the cluster Galeomorphii, in the Carcharhiniformes group, while (ii) Rhincodon typus belongs to the Orectolobiformes within the Galeomorphii group, and (iii) Galeocerdo cuvier and Sphyrna lewini belong to a separate cluster in the Carcharhiniformes group [54]. Phylogenetic inference using microsatellites might be limited for the species having frequent mutations and recombinations which can decrease the frequency of microsatellite loci [30].
Study of the eight shark species targeted for cross-species amplification, as well as blacktip reef sharks, will benefit from the use of this new set of 44 microsatellite markers. These markers will support critical conservation-based research, such as estimating population sizes, deciphering parentage links between individuals, and analyzing structure and demographic history of shark populations.
References
Alves LMF, Nunes M, Marchand P, le Bizec B, Mendes S, Correia JPS, Lemos MFL, Novais SC (2016) Blue sharks (Prionace glauca) as bioindicators of pollution and health in the Atlantic Ocean: contamination levels and biochemical stress responses. Sci Total Environ 563–564:282–292. https://doi.org/10.1016/j.scitotenv.2016.04.085
Archer FI, Adams PE, Schneiders BB (2017) StrataG: An R package for manipulating, summarizing and analyzing population genetic data. Mol Ecol Resour 17(1):5–11. https://doi.org/10.1111/1755-0998.12559
Baird SJE (2015) Exploring linkage disequilibrium. Mol Ecol Resour 15(5):1017–1019. https://doi.org/10.1111/1755-0998.12424
Balaresque P (2007) Microsatellites: life cycle concept and neutrality issues. Medicine/Sciences 23(8–9):729–734. https://doi.org/10.1051/medsci/20072389729
Barbará T, Palma-Silva C, Paggi GM, Bered F, Fay MF, Lexer C (2007) Cross-species transfer of nuclear microsatellite markers: potential and limitations. Mol Ecol 16(18):3759–3767. https://doi.org/10.1111/j.1365-294X.2007.03439.x
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (1996) GENETIX 4.05, logiciel sous WindowsTM pour la génétique des populations. Laboratoire Génome, Populations, Interactions, CNRS UMR 5000, Université de Montpellier II, Montpellier (France)
Boussarie G, Momigliano P, Robbins WD, Cornu LB, Fauvelot J-F, Kiszka C, Manel S, Mouillot D, Vigliola L (2021) Identifying barriers to gene flow and hierarchical conservation units from seascape genomics: a modelling framework applied to a marine predator. Ecography. https://doi.org/10.1101/2021.10.25.465682
Cardeñosa D, Shea SK, Zhang H, Fischer GA, Simpfendorfer CA, Chapman DD (2022) Two-thirds of species in a global shark fin trade hub are threatened with extinction: conservation potential of international trade regulations for coastal sharks. Conserv Lett. https://doi.org/10.1111/conl.12910
Chaix G, Gerber S, Razafimaharo V, Vigneron P, Verhaegen D, Hamon S (2003) Gene flow estimation with microsatellites in a Malagasy seed orchard of Eucalyptus grandis. Theor Appl Genet 107(4):705–712. https://doi.org/10.1007/s00122-003-1294-0
Clarke SC, Harley SJ, Hoyle SD, Rice JS (2013) Population trends in Pacific oceanic sharks and the utility of regulations on shark finning. Conserv Biol 27(1):197–209. https://doi.org/10.1111/j.1523-1739.2012.01943.x
Dapp DR, Huveneers C, Walker TI, Reina RD (2017) Physiological response and immediate mortality of gill-net-caught blacktip reef sharks (Carcharhinus melanopterus). Mar Freshw Res 68(9):1734. https://doi.org/10.1071/MF16132
DeWoody JA, Avise JC (2000) Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J Fish Biol 56(3):461–473. https://doi.org/10.1111/j.1095-8649.2000.tb00748.x
Domingues RR, Hilsdorf AWS, Gadig OBF (2018) The importance of considering genetic diversity in shark and ray conservation policies. Conserv Genet 19(3):501–525. https://doi.org/10.1007/s10592-017-1038-3
Dudgeon CL, Blower DC, Broderick D, Giles JL, Holmes BJ, Kashiwagi T, Krück NC, Morgan JAT, Tillett BJ, Ovenden JR (2012) A review of the application of molecular genetics for fisheries management and conservation of sharks and rays. J Fish Biol 80(5):1789–1843. https://doi.org/10.1111/j.1095-8649.2012.03265.x
Dulvy NK, Pacoureau N, Rigby CL, Pollom RA, Jabado RW, Ebert DA, Finucci B, Pollock CM, Cheok J, Derrick DH, Herman KB, Sherman CS, VanderWright WJ, Lawson JM, Walls RHL, Carlson JK, Charvet P, Bineesh KK, Fernando D et al (2021) Overfishing drives over one-third of all sharks and rays toward a global extinction crisis. Curr Biol 31(21):4773-4787.e8. https://doi.org/10.1016/j.cub.2021.08.062
Dwyer RG, Krueck NC, Udyawer V, Heupel MR, Chapman D, Pratt HL, Garla R, Simpfendorfer CA (2020) Individual and population benefits of marine reserves for reef sharks. Curr Biol. https://doi.org/10.1016/j.cub.2019.12.005
Feldheim KA, Gruber SH, Ashley MV (2002) Population genetic structure of the lemon shark (Negaprion brevirostris) in the western Atlantic: DNA microsatellite variation. Mol Ecol 10(2):295–303. https://doi.org/10.1046/j.1365-294X.2001.01182.x
Feldheim KA, Gruber SH, Ashley MV (2004) Reconstruction of parental microsatellite genotypes reveals female polyandry and philopatry in the lemon shark, Negaprion brevirostris. Evolution 58(10):2332–2342. https://doi.org/10.1111/j.0014-3820.2004.tb01607.x
Feldheim KA, Jabado RW, Chapman DD, Cardeñosa D, Maddox JD (2020) Microsatellite primer development in elasmobranchs using next generation sequencing of enriched libraries. Mol Biol Rep 47(4):2669–2675. https://doi.org/10.1007/s11033-020-05357-y
Flanagan SP, Jones AG (2019) The future of parentage analysis: from microsatellites to SNPs and beyond. Mol Ecol 28(3):544–567. https://doi.org/10.1111/mec.14988
Frichot E, François O (2015) LEA: an R package for landscape and ecological association studies. Methods Ecol Evol 6(8):925–929. https://doi.org/10.1111/2041-210X.12382
Garza JC, Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10(2):305–318
Graham NAJ, Spalding MD, Sheppard CRC (2010) Reef shark declines in remote atolls highlight the need for multi-faceted conservation action. Aquat Conserv Mar Freshwat Ecosyst 20(5):543–548. https://doi.org/10.1002/aqc.1116
Karl SA, Castro ALF, Garla RC (2012) Population genetics of the nurse shark (Ginglymostoma cirratum) in the western Atlantic. Mar Biol 159(3):489–498. https://doi.org/10.1007/s00227-011-1828-y
Keeney DB, Heist EJ (2003) Characterization of microsatellite loci isolated from the blacktip shark and their utility in requiem and hammerhead sharks. Mol Ecol Notes 3(4):501–504. https://doi.org/10.1046/j.1471-8286.2003.00492.x
Larson SE, Daly-Engel TS, Phillips NM (2017) Review of current conservation genetic analyses of Northeast Pacific sharks. Adv Mar Biol 77:79–110. https://doi.org/10.1016/bs.amb.2017.06.005
Lei Y, Zhou Y, Price M, Song Z (2021) Genome-wide characterization of microsatellite DNA in fishes: survey and analysis of their abundance and frequency in genome-specific regions. BMC Genomics 22(1):421. https://doi.org/10.1186/s12864-021-07752-6
Li Y-C, Korol AB, Fahima T, Beiles A, Nevo E (2002) Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Mol Ecol 11(12):2453–2465. https://doi.org/10.1046/j.1365-294X.2002.01643.x
Maduna SN, Rossouw C, Roodt-Wilding R, van der Merwe AEB (2014) Microsatellite cross-species amplification and utility in southern African elasmobranchs: a valuable resource for fisheries management and conservation. BMC Res Notes. https://doi.org/10.1186/1756-0500-7-352
Martin AP, Pardini AT, Noble LR, Jones CS (2002) Conservation of a dinucleotide simple sequence repeat locus in sharks. Mol Phylogenet Evol 23(2):205–213. https://doi.org/10.1016/S1055-7903(02)00001-5
Meglécz E, Pech N, Gilles A, Dubut V, Hingamp P, Trilles A, Grenier R, Martin J-F (2014) QDD version 3.1: a user-friendly computer program for microsatellite selection and primer design revisited: experimental validation of variables determining genotyping success rate. Mol Ecol Resour 14(6):1302–1313. https://doi.org/10.1111/1755-0998.12271
Meirmans PG (2020) GENODIVE version 3.0: Easy to use software for the analysis of genetic data of diploids and polyploids. Mol Ecol Resour 20(4):1126–1131. https://doi.org/10.1111/1755-0998.13145
Mendes NJ, Cruz VP, Ashikaga FY, Camargo SM, Oliveira C, Piercy AN, Burgess GH, Coelho R, Santos MN, Mendonça FF, Foresti F (2016) Microsatellite loci in the tiger shark and cross-species amplification using pyrosequencing technology. PeerJ 4:e2205. https://doi.org/10.7717/peerj.2205
Meyer L, Fox A, Huveneers C (2018) Simple biopsy modification to collect muscle samples from free-swimming sharks. Biol Conserv 228:142–147. https://doi.org/10.1016/j.biocon.2018.10.024
Momigliano P, Harcourt R, Robbins WD, Stow A (2015) Connectivity in grey reef sharks (Carcharhinus amblyrhynchos) determined using empirical and simulated genetic data. Sci Rep 5(1):13229. https://doi.org/10.1038/srep13229
Moore JA, Draheim HM, Etter D, Winterstein S, Scribner KT (2014) Application of large-scale parentage analysis for investigating natal dispersal in highly vagile vertebrates: a case study of American black bears (Ursus americanus). PLoS ONE 9(3):e91168. https://doi.org/10.1371/journal.pone.0091168
Mourier J, Planes S (2013) Direct genetic evidence for reproductive philopatry and associated fine-scale migrations in female blacktip reef sharks (Carcharhinus melanopterus) in French Polynesia. Mol Ecol 22(1):201–214. https://doi.org/10.1111/mec.12103
Peakall R, Smouse PE (2006) GenAlex 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6(1):288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Pirog A, Jaquemet S, Blaison A, Soria M, Magalon H (2016) Isolation and characterization of eight microsatellite loci from Galeocerdo cuvier (tiger shark) and cross-amplification in Carcharhinus leucas, Carcharhinus brevipinna, Carcharhinus plumbeus and Sphyrna lewini. PeerJ 4:e2041. https://doi.org/10.7717/peerj.2041
Portnoy DS, Heist EJ (2012) Molecular markers: progress and prospects for understanding reproductive ecology in elasmobranchs. J Fish Biol 80(5):1120–1140. https://doi.org/10.1111/j.1095-8649.2011.03206.x
Portnoy DS, McDowell JR, Thompson K, Musick JA, Graves JE (2006) Isolation and characterization of five dinucleotide microsatellite loci in the sandbar shark, Carcharhinus plumbeus. Mol Ecol Notes 6(2):431–433. https://doi.org/10.1111/j.1471-8286.2006.01261.x
Ramírez-Macías D, Shaw K, Ward R, Galván-Magaña F, Vázquez-Juárez R (2009) Isolation and characterization of microsatellite loci in the whale shark (Rhincodon typus). Mol Ecol Resour 9(3):798–800. https://doi.org/10.1111/j.1755-0998.2008.02197.x
Richards VP, Suzuki H, Stanhope MJ, Shivji MS (2013) Characterization of the heart transcriptome of the white shark (Carcharodon carcharias). BMC Genomics 14(1):697. https://doi.org/10.1186/1471-2164-14-697
Roff G, Doropoulos C, Rogers A, Bozec Y-M, Krueck NC, Aurellado E, Priest M, Birrell C, Mumby PJ (2016) The ecological role of sharks on coral reefs. Trends Ecol Evol 31(5):395–407. https://doi.org/10.1016/j.tree.2016.02.014
Roncallo PF, Beaufort V, Larsen AO, Dreisigacker S, Echenique V (2019) Genetic diversity and linkage disequilibrium using SNP (KASP) and AFLP markers in a worldwide durum wheat (Triticum turgidum L. var. durum) collection. PLoS ONE 14(6):0218562. https://doi.org/10.1371/journal.pone.0218562
Salles OC, Pujol B, Maynard JA, Almany GR, Berumen ML, Jones GP, Saenz-Agudelo P, Srinivasan M, Thorrold SR, Planes S (2016) First genealogy for a wild marine fish population reveals multigenerational philopatry. Proc Natl Acad Sci USA 113(46):13245–13250. https://doi.org/10.1073/pnas.1611797113
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011(27):863–864. https://doi.org/10.1093/bioinformatics/btr026
Scotti I, Magni F, Paglia G, Morgante M (2002) Trinucleotide microsatellites in Norway spruce (Picea abies): their features and the development of molecular markers. Theor Appl Genet 106(1):40–50. https://doi.org/10.1007/s00122-002-0986-1
Sharma SP, Ghazi MG, Katdare S, Dasgupta N, Mondol S, Gupta SK, Hussain SA (2021) Microsatellite analysis reveals low genetic diversity in managed populations of the critically endangered gharial (Gavialis gangeticus) in India. Sci Rep. https://doi.org/10.1038/s41598-021-85201-w
Simpfendorfer C, Yuneni RR, Tanay D, Seyha L, Haque AB, Fahmi, Bin Ali AD, Bineesh KK, Gautama DA, Maung A, Sianipar A, Utzurrum JAT, Voye VQ (2020) Carcharhinus melanopterus. The IUCN Red List of Threatened Species. https://doi.org/10.2305/IUCN.UK.2020-3.RLTS.T39375A58303674.en
Staden M, Gledhill KS, Gennari E, McCord ME, Parkinson M, Watson RGA, Rhode C, Bester-van der Merwe AE (2020) Microsatellite development and detection of admixture among three sympatric Haploblepharus species (Carcharhiniformes: Scyliorhinidae). Aquat Conserv Mar Freshwat Ecosyst 30(12):2336–2350. https://doi.org/10.1002/aqc.3406
Sturm AB, Eckert RJ, Méndez JG, González-Díaz P, Voss JD (2020) Population genetic structure of the great star coral, Montastraea cavernosa, across the Cuban archipelago with comparisons between microsatellite and SNP markers. Sci Rep 10(1):15432. https://doi.org/10.1038/s41598-020-72112-5
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4(3):535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Vélez-Zuazo X, Agnarsson I (2011) Shark tales: a molecular species-level phylogeny of sharks (Selachimorpha, Chondrichthyes). Mol Phylogenet Evol 58(2):207–217. https://doi.org/10.1016/j.ympev.2010.11.018
Vignaud TM, Mourier J, Maynard JA, Leblois R, Spaet JLY, Clua E, Neglia V, Planes S (2014) Blacktip reef sharks, Carcharhinus melanopterus, have high genetic structure and varying demographic histories in their Indo-Pacific range. Mol Ecol 23(21):5193–5207. https://doi.org/10.1111/mec.12936
Ward-Paige CA, Worm B (2017) Global evaluation of shark sanctuaries. Glob Environ Chang. https://doi.org/10.1016/j.gloenvcha.2017.09.005
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38(6):1358. https://doi.org/10.2307/2408641
Xu Y, Li W, Hu Z, Zeng T, Shen Y, Liu S, Zhang X, Li J, Yue B (2018) Genome-wide mining of perfect microsatellites and tetranucleotide orthologous microsatellites estimates in six primate species. Gene 643:124–132. https://doi.org/10.1016/j.gene.2017.12.00
Acknowledgements
We thank the French NGO Ailerons for providing fin clips of adult Prionacea glauca collected in the Mediterranean Sea, B. Holmes (University of Queensland) for providing us with Galeocerdo cuvier fin clips collected on the Great Barrier reef off the Australian coast, M. Meekan (Australian Institute of Marine Science) for providing Rhincodon typus samples collected on the Ningaloo reef off the west coast of Australia, and J. Ovenden (Southern Fisheries Center, Australia) for providing Carcharhinus sorrah samples collected on the Great Barrier reef. The Sphyrna lewini samples were collected by J. B. Galves in Madagascar near Nosy Be Island.
Funding
Funding was provided by École Pratique des Hautes Etudes (EPHE).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no conflict of interest.
Ethical approval
Sampling of blacktip reef sharks was conducted by the Center for Island Research and Environmental Observatory (CRIOBE) under the permit N°9524 issued by the Ministère de la Promotion des Langues, de la Culture, de la Communication et de l’Environnement of the French Polynesian government in October 2015.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Eustache, K.B., Boissin, É., Tardy, C. et al. Characterization of 35 new microsatellite markers for the blacktip reef shark (Carcharhinus melanopterus) and cross-species amplification in eight other shark species. Mol Biol Rep 50, 3205–3215 (2023). https://doi.org/10.1007/s11033-022-08209-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-08209-z