Abstract
Background
Vega Island is located off the eastern tip of the Antarctic Peninsula (Maritime Antarctica), in the Weddell Sea. In this study, we used metabarcoding to investigate green algal DNA sequence diversity present in sediments from three lakes on Vega Island (Esmeralda, Copépodo, and Pan Negro Lakes).
Methods and results
Total DNA was extracted and the internal transcribed spacer 2 region of the nuclear ribosomal DNA was used as a DNA barcode for molecular identification. Green algae were represented by sequences representing 78 taxa belonging to Phylum Chlorophyta, of which 32% have not previously been recorded from Antarctica. Sediment from Pan Negro Lake generated the highest number of DNA reads (11,205), followed by Esmeralda (9085) and Copépodo (1595) Lakes. Esmeralda Lake was the richest in terms of number of taxa (59), with Copépodo and Pan Negro Lakes having 30 taxa each. Bray–Curtis dissimilarity among lakes was high (~ 0.80). The Order Chlamydomonadales (Chlorophyceae) gave the highest contribution in terms of numbers of taxa and DNA reads in all lakes. The most abundant taxon was Chlorococcum microstigmatum.
Conclusions
The study confirms the utility of DNA metabarcoding in assessing potential green algal diversity in Antarctic lakes, generating new Antarctic records.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Antarctic Peninsula (Maritime Antarctica) consists of a north–south oriented chain of heavily glaciated mountains which drive strong climatic differences between its warmer and moister western coastal regions and colder eastern coast in the Weddell Sea [1]. Various lakes are present in this region, varying greatly in terms of trophic status and salinity [2, 3]. Vega Island is located off the eastern tip of the Antarctic Peninsula, in the Weddell Sea. Although there is a broad general literature on the algal communities of lakes and ponds of the Antarctic Peninsula [4,5,6,7,8], information on lakes from Vega Island is largely restricted to abiotic aspects [1, 9], excepting two recent studies exploring diatom paleo-assemblages [10] and sediment fungal communities [11].
Green algae (Viridiplantae) are noted in all studies of Antarctic lake algal communities, represented mainly by the Phylum Chlorophyta [12], with zygnematophycean green algae and other taxa phylogenetically close to higher plants (Streptophyta) being less common. De Wever et al. [8] highlighted the wide phylogenetic diversity of apparently endemic Antarctic lineages of microscopic green algae, consistent with hypotheses of strong regionalization and long-term evolutionary isolation within Antarctica, even of microbial diversity [13,14,15]. According to De Wever et al. [8], their findings, supported by molecular analyses, contrasted with most previous morphological studies, which generally concluded that Antarctic green algae were mostly represented by cosmopolitan species. While these algae are a ubiquitous group present in ecosystems globally [12], many green algae are also known to tolerate extreme conditions such as high salinity and low temperatures, particularly within the order Chlamydomonadales [16, 17]. Several new green algal species have recently been described from Antarctica [18], highlighting the presence of many psychrotrophic or psychrophilic taxa [19, 20]. Green algae are also the primary constituent of the vividly coloured snow algal communities that are well represented in polar regions [16, 21]. Snow algae can decrease the albedo of snow, accelerating the rate of snowmelt [22]. Snow algal species (e.g., Chlamydomonas nivalis) have been also reported in other Antarctic habitats such as lake sediments and soils, in both morphological [5,6,7] and molecular [23] studies.
Considerable advances in the assessment of microbial diversity in environmental samples have been made possible thanks to recent developments in molecular biology [24]. In the Antarctic continent, although a number of recent studies of green algal diversity on soil and rock substrates using molecular tools have been published [23, 25,26,27], the use of such approaches on freshwater communities currently remains limited [8]. As with other microbial groups, some algae cannot be grown in culture, while reliance on morphology alone can also be misleading as high levels of morphological conservation typify algae present in the harsh environmental conditions of Antarctica; this is particularly true with regard to green algae, whose relatively high level of cryptic species and phenotypic plasticity may hamper conclusive diagnosis based solely on morphological features [28]. Furthermore, resting stages and spores cannot usually be identified using traditional methods and, consequently, morphology-based techniques fail to adequately represent the range of microalgal diversity in the natural environment [29]. DNA metabarcoding using high throughput sequencing (HTS) represents a powerful method for the detection of rare species [25, 29]. In this study, we used metabarcoding to investigate green algal (Viridiplantae) DNA sequence diversity present in sediments obtained from three lakes on Vega Island, generating new information about the biodiversity of these poorly known lakes.
Methods
Study area
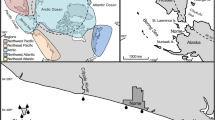
Vega Island is located in the James Ross Archipelago, of the north-east tip of the Antarctic Peninsula. It has an area of 253 km2, more than 80% covered by permanent ice. The climate of the island is cold and semi-arid, with precipitation (water equivalent) of ca. 300–500 mm year−1 [30] and a mean annual temperature of ca. − 7 °C [31]. Sediment samples were obtained from three freshwater lakes on Vega Island: Esmeralda (63° 52′ 21.4″ S; 57° 36′ 24.1″ W), Copépodo (63° 52′ 41.9″ S; 57° 35′ 43.8″ W) and Pan Negro (63° 52′ 04.6″ S; 57° 37′ 12.6″ W). The three lakes are located at Cape Lamb, at the south-west tip of Vega Island (Supplementary File 1). The cape has the largest ice-free area on the island, with a maximum elevation of 482 m a.s.l [10]. All three lakes are surrounded by moraine deposits and located in endorheic basins, which retain water from snow melt in their catchments and have no outflow. A summary of lake characteristics is given in Table 1.
Sampling
Sampling was conducted during the austral summer of 2016/17, from the littoral zone of each lake. One sediment core of 30 cm length was collected manually from each lake using PVC pipes (60 mm diameter × 50 cm length) previously disinfected to avoid contamination following the protocol described by [11]. The cores were kept at − 20 °C in sterilized plastic bags after collection and during transportation to the laboratory at the Federal University of Minas Gerais, Brazil. There, the cores were thawed and cut into 5 cm sections; only the top, middle and base sections of the core were used for analysis. One subsample of each core section (top, middle and base) was processed together into the same DNA extraction in order to increase DNA yield. Water physical and chemical parameters (temperature, pH, electrical conductivity and total dissolved solids) were measured in situ simultaneously with sediment sampling in each lake, using a Hanna multi-parameter probe HI 9828 (Hanna Instruments, USA).
DNA extraction, Illumina library construction and sequencing
Total DNA was extracted from ca. 0.5 g of soil using the QIAGEN Power Soil Kit (QIAGEN, Carlsbad, USA), following the manufacturer’s instructions. DNA quality was analyzed by agarose gel electrophoresis (1% agarose in 1 × Trisborate-EDTA) and then quantified using the Quanti-iT™ Pico Green dsDNA Assay (Invitrogen). The internal transcribed spacer 2 region (ITS2) of the nuclear ribosomal DNA was used as a DNA barcode for molecular species identification [32] using the universal primers ITS3 and ITS4 [33]. Library construction and DNA amplification were performed using the Herculase II Fusion DNA Polymerase Nextera XT Index Kit V2, following the Illumina 16S Metagenomic Sequencing Library Preparation Part #15044223 Rev. B protocol. Paired-end sequencing (2 × 300 bp) was performed on a MiSeq System (Illumina) by Macrogen Inc. (South Korea).
Data analysis and taxa assignment
The PLANiTS2 database [34] was used for identification of the Viridiplantae sequences obtained. Raw fastq files were filtered using BBDuk version 38.34 (BBMap – Bushnell B. – sourceforge.net/projects/bbmap/) to remove Illumina adapters, known Illumina artefacts and the PhiX Control v3 Library. Quality read filtering was carried out using Sickle version 1.33 -q 30 -l 50 [35] and to trim 3ʹ or 5ʹ ends with low Phred quality score. Sequences shorter than 50 bp were also discarded. The remaining sequences were imported to QIIME2 version 2019.10 (https://qiime2.org/) for bioinformatics analyses [36].
For assessing the sequence data obtained against the PLANiTS2 database, the pipeline was executed for merged paired-ended sequences with the following plug-ins: vsearch join-pairs [37], vsearch dereplicate-sequences, quality-filter q-score-joined [38], vsearch cluster-features-de-novo 97% identity limit and vsearch uchime-denovo. Taxonomic assignments were determined for operational taxonomic units (OTUs) using the feature-classifier [39] classify-sklearn against the PLANiTS2 database [34] trained with Naïve Bayes classifier. For comparative purposes we used the number of DNA reads as a proxy for relative abundance [40], and each OTU was considered as a different taxon. It is important to recognize that the assignment of a detected DNA sequence does not confirm the presence of the organism in a viable form, while assignment is also limited by the level of diversity coverage of the available databases [23, 41]. Taxa assignment into higher taxonomical ranks followed Leliaert et al. [12] and Algaebase [42].
Diversity analyses
Diversity analyses were performed using R software version 4.0.2 [43]. Rarefaction curves were prepared with the iNEXT (iNterpolation and EXTrapolation) package [44]. Alpha diversity was assessed using Shannon (H’) and Pielou (J’ Equitability) indices and natural logarithms. For assessment of dissimilarity between locations, the Bray–Curtis Index was used. A Venn diagram was also prepared to compare OTU occurrences across the three lakes [45].
Results
The rarefaction curves reached asymptote, indicating that the sampling effort was sufficient to represent the sequence diversity present within the DNA reads at each sampling location (Supplementary File 2). A total of 1,194,202 paired-end ITS2 DNA reads were generated in the sequencing run, of which 243,373 remained after quality filtering. The large majority of these reads represented other taxonomic groups not considered in the present study (e.g., fungi). Pan Negro Lake generated the highest number of green algal DNA reads (11,205), followed by Esmeralda (9085) and Copépodo (1595) Lakes. Esmeralda Lake contained the highest number of green algal taxa assigned (59), with Copépodo and Pan Negro Lakes each containing 30 taxa. Altogether 78 taxa were assigned in the three lakes (Table 2).
The highest diversity of green algae in Esmeralda Lake was corroborated by its Shannon Index (2.92), followed by Copépodo Lake (2.63). The lowest Shannon Index (1.50) and Equitability Index (0.44) were obtained in Pan Negro Lake. In this lake, the single Chlamydomonadales species Chlorococcum microstigmatum contributed 60% of the total number of DNA reads (Fig. 1), decreasing the evenness; in the other lakes, the distribution of DNA reads among the different taxa was more uniform, resulting in an Equitability Index above 0.70 (0.72 at Esmeralda Lake and 0.77 at Copépodo Lake) (Fig. 1).
The highest contributing green algae in sediments from three lakes on Vega Island, Antarctica, in terms of DNA reads (%) (the six taxa shown for each lake represent at least 60% of the total local abundance). Color bar: green = Class Chlorophyceae; gray = Class Trebouxiophyceae; blue = Class Ulvophyceae
All green algal taxa assigned in this study belonged to Phylum Chlorophyta, representing three classes (Chlorophyceae, Trebouxiophyceae and Ulvophyceae) and nine orders (Fig. 2). Of the 78 taxa assigned, 25 (32%) are not listed in the available literature and are likely first records from Antarctica. These included the genera Chlamydocapsa, Paulschulzia, Auxenochlorella, Didymogenes, Jaagichlorella, Desmococcus, Lobosphaera, Ploeotila and Desmochloris (Table 2).
The Order Chlamydomonadales (Chlorophyceae) represented the highest contribution in terms of numbers of both taxa and DNA reads in all lakes (Fig. 2). Its relative abundance was above 80% in Copépodo and Pan Negro Lakes. In Esmeralda Lake, the relative contribution of the different orders was more uniform compared to Copépodo and Pan Negro Lakes, both in terms of number of taxa and number of reads (Fig. 2).
Eleven taxa (14%) were present in all three lakes (Fig. 3). Pan Negro Lake was the most dissimilar, presenting Bray–Curtis Index values of 0.88 and 0.83 with Copépodo and Esmeralda Lakes, respectively; between these two lakes, the Bray–Curtis Index was 0.79.
Discussion
This study revealed a relatively high diversity of green algal sequences in sediments of Antarctic lakes (78 taxa), while recognizing that the assignment of a DNA sequence does not confirm the presence of the organism in a viable form [26, 44]. In general, green algal richness reported in previous studies of Antarctic algae from different habitats has ranged from 12 to 48 taxa, in studies based on morphological data [4,5,6,7] or from isolated cultures [8]. Consistent with this previous literature, our study confirmed the dominance of Chlorophyta and its “UTC” core group (Ulvophyceae, Trebouxiophyceae and Chlorophyceae) over green algae representing the Streptophyta [12].
A majority of the green algal taxa assigned here have previously been reported in Antarctic studies either using traditional morphological approaches or in metabarcoding studies [23, 42]. Amongst the potentially new Antarctic records, the majority of taxa originate in the Northern Hemisphere [42]. Incorrect taxonomic assignments (e.g., synonyms) may be a factor here. For example, Hoham & Mullet [46] consider Scotiella cryophila, assigned here as a first Antarctic record, a synonym of Chloromonas nivalis and S. antarctica, known Antarctic species [4].
The green algal diversity assigned in the sediments examined here included both typically planktonic genera (e.g., Chlamydomonas, Desmodesmus, Coenochloris, Micractinium, Koliella) [4, 7], genera typically growing on substrates or found in lichenized forms (e.g., Stigeoclonium, Stichococcus, Bracteacoccus, Prasiola, Elliptochloris, Chlorococcum, Coccomyxa, Lobosphaera, Myrmecia, Trebouxia, Desmochloris, Ulothrix, Pseudendochlonium) [5, 6, 47] and snow algal genera (e.g., Scotiella, Chloromonas, Chlamydomonas) [17, 48]. Some marine taxa (e.g., Pseudendoclonium) were also detected, which is not uncommon in Antarctic studies of this type [23] considering the common occurrence of high winds and local bird populations that can link terrestrial/freshwater and marine environments.
The three maritime Antarctic lakes studied here shared abiotic features more similar to some continental Antarctic lakes [49], especially in terms of high values of electrical conductivity (680 to 1250 µS cm−1 in our study). As they have no outflow, over time they will concentrate salts washed into them in melt water, leading gradually to increased salinity. On King George Island, located north-west of the Antarctic Peninsula, most lakes have electrical conductivity around 130 µS cm−1 [3]. As our lakes are not directly exposed to sea spray, these differences when compared to other maritime lakes from the western side of the Peninsula may also be contributed to by the low precipitation that is typical of the Vega Islands, where they are located. Copépodo Lake, in particular, showed the most extreme conditions, with the highest electrical conductivity (1250 µS cm−1) and an alkaline pH (8.1). Copépodo Lake generated the lowest number of DNA reads in our study; it is possible that these more extreme conditions may limit the diversity of green algal communities in this lake.
The use of DNA metabarcoding as applied here was effective in the detection of rare species, enhancing knowledge of potential green algal diversity in Antarctic environments. Câmara et al. [23] detected 65 distinct green algal OTUs in soil samples from two protected and non-protected sites on Deception Island (South Shetland Islands) using the same approach, of which 71% (45 taxa) were common to both sites. In contrast, in the current study, only 14% of the taxa detected were shared by all three lakes, highlighting the dissimilarity (high beta diversity) among the lakes studied. The geological and hydrological differences among lakes might have contributed to this relatively high dissimilarity. However, we recognize that the limited replication possible in the current study does not permit detailed examination of the diversity differences between the lakes, and that future studies to understand the nature of dissimilarity between such lakes will require greater replication.
Pan Negro Lake, which had the same taxon richness as Copépodo Lake, generated the highest number of DNA reads, and was the most distinct lake. The number of taxa in this lake is likely to be underestimated, as 17% of the DNA reads were identified within the Order Chlamydomonadales, and not to species level. This Order originally comprised flagellated green algae (e.g., Chlamydomonas, Chloromonas); later, molecular studies expanded it to include some coccoid taxa, such as the genus Chlorococcum [50]. This genus was particularly abundant in the lakes studied here, especially Pan Negro Lake. Chlamydomonas spp. also dominate in the phytoplankton of Antarctic hypertrophic lakes [7]; the presence of the Chlamydomonas group at high relative abundance in these lake sediments is consistent with them acting as a repository of genetic information on taxa present in the overlying water column. Members of Chlamydomonas are frequently reported in Antarctic phycological studies, with many species well adapted to low-temperature environments [20]. In the current study, besides C. nivalis, a key component of snow algal communities [16, 21], the species C. raudensis was among the most abundant in all three lakes. This psychrophilic species (also referred to in the literature as Chlamydomonas sp. UWO231) was originally described from samples collected in the permanently ice-covered Lake Bonney in the Victoria Land Dry Valleys [19]. It is very plausible that other as yet unknown species of green algae are present in the lakes of Vega Island, and not represented in current sequence databases.
Conclusions
Diverse assemblages of green algal DNA sequences were detected in sediments obtained from three lakes on Vega Island, including many assignments to taxa not previously recorded from the region. The study gives information about this part of the Maritime Antarctica and confirms the utility of DNA metabarcoding as a tool for assessing potential green algal diversity in Antarctic lakes, including the recognition of taxa not previously recorded from the continent. There is a clear need for increased effort to collect and sequence green algal voucher specimens in this region in order to reconcile molecular and morphological diversity studies, and confirm the presence of currently unrecorded or undescribed species as strongly implied by our data.
Availability of data and material
All data generated or analysed during this study are included in this published article.
Code availability
Not applicable.
References
Píšková A, Roman M, Bulínová M et al (2019) Late-Holocene palaeoenvironmental changes at Lake Esmeralda (Vega Island, Antarctic Peninsula) based on a multi-proxy analysis of laminated lake sediment. Holocene 29(7):1155–1175. https://doi.org/10.1177/0959683619838033
Vincent WF, Vincent CL (1982) Response to nutrient enrichment by the plankton of Antarctic coastal lakes and the inshore Ross Sea. Polar Biol 1:159–165
Vinocur A, Unrein F (2000) Typology of lentic water bodies at Potter Peninsula (King George Island, Antarctica) based on physical-chemical characteristics and phytoplankton communities. Polar Biol 23:858–870. https://doi.org/10.1007/s003000000165
Vinocur A, Izaguirre I (1994) Freshwater algae (excluding Cyanophyceae) from nine lakes and pools of Hope Bay, Antarctica Peninsula. Antarct Sci 6(4):483–489. https://doi.org/10.1017/S0954102094000738
Vinocur A, Pizarro H (1995) Periphyton flora of some lotic and lentic environments of Hope Bay (Antarctic Peninsula). Polar Biol 15:401–414. https://doi.org/10.1007/BF00239716
Vinocur A, Pizarro H (2000) Microbial mats of twenty-six lakes from Potter Peninsula, King George Island, Antarctica. Hydrobiologia 437:171–185. https://doi.org/10.1023/A:1026511125146
Izaguirre I, Allende L, Schiaffino MR (2020) Phytoplankton in Antarctic lakes: biodiversity and main ecological features. Hydrobiologia 847:1–31
De Wever A, Leliaert F, Verleyen E, Vanormelingen P, Van der Gucht K, Hodgson DA, Sabbe K, Vyverman W (2009) Hidden levels of phylodiversity in Antarctic green algae: further evidence for the existence of glacial refugia. Proc R Soc B 276:3591–3599. https://doi.org/10.1098/rspb.2009.0994
Chaparro MAE, Chaparro MAE, Córdoba FE, Lecomte KL, Gargiulo JD, Barrios AM, Urán JM, Czalbowski NTM, Lavat A, Böhnel HN (2017) Sedimentary analysis and magnetic properties of Lake Anónima, Vega Island. Antarct Sci 29:429–444. https://doi.org/10.1017/S0954102017000116
Bulínová M, Kohler TJ, Kavan J, Van de Vijver B, Nývlt D, Nedbalová L, Coria SH, Lirio JM, Kopalová K (2020) Comparison of diatom paleo-assemblages with adjacent limno-terrestrial communities on Vega Island, Antarctic Peninsula. Water 12:1340. https://doi.org/10.3390/w12051340
Ogaki MB, Câmara PEAS, Pinto OHB, Lirio JM, Coria SH, Vieira E, Carvalho-Silva M, Convey P, Rosa CA, Rosa LH (2021) Diversity of fungal DNA in lake sediments on Vega Island, north-east Antarctic Peninsula assessed using DNA metabarcoding. Extremophiles 25:257–265. https://doi.org/10.1007/s00792-021-01226-z
Leliaert F, Smith DR, Moreau H, Herron MD, Verbruggen H, Delwiche CF, De Clerck O (2012) Phylogeny and molecular evolution of the green algae. Crit Rev Plant Sci 31:1–46. https://doi.org/10.1080/07352689.2011.615705
Laybourn-Parry J, Pearce DA (2007) The biodiversity and ecology of Antarctic lakes: models for evolution. Philos T R Soc B 362:2273–2289. https://doi.org/10.1098/rstb.2006.1945
Convey P, Biersma EM, Casanova-Katny A, Maturana CS (2020) Refuges of Antarctic diversity. In: Oliva J, Ruiz-Fernández J (eds) Past Antarctica. Academic Press, Burlington, pp 181–200. https://doi.org/10.1016/B978-0-12-817925-3.00010-0
Verleyen E, Van de Vijver B, Tytgat B et al (2021) Diatoms define a novel freshwater biogeography of the Antarctic. Ecography 44:548–560. https://doi.org/10.1111/ecog.05374
Davey MP, Norman L, Sterk P et al (2019) Snow algae communities in Antarctica: metabolic and taxonomic composition. New Phytol 222:1242–1255. https://doi.org/10.1111/nph.15701
Peng Z, Liu G, Huang K (2021) Cold adaptation mechanisms of a snow alga Chlamydomonas nivalis during temperature fluctuations. Front Microbiol 11:611080. https://doi.org/10.3389/fmicb.2020.611080
Chae H, Lim S, Kim HS, Choi H-G, Kim JH (2019) Morphology and phylogenetic relationships of Micractinium (Chlorellaceae, Trebouxiophyceae) taxa, including three new species from Antarctica. Algae 34(4):267–275. https://doi.org/10.4490/algae.2019.34.10.15
Pocock T, Lachance M-A, Pröschold T, Priscu JC, Kim SS, Huner NPA (2004) Identification of a psychrophilic green alga from Lake Bonney Antarctica: Chlamydomonas raudensis Ettl. (UWO241) Chlorophyceae. J Phycol 40:1138–1148. https://doi.org/10.1111/j.1529-8817.2004.04060.x
Zhang X, Cvetkovska M, Morgan-Kiss R, Hüner NPA, Smith DR (2021) Draft genome sequence of the Antarctic green alga Chlamydomonas sp. UWO241. IScience 24(2):102084. https://doi.org/10.1016/j.isci.2021.102084
Gray A, Fretwell P, Smith AG, Convey P, Peck LS, Krolikowski M, Mendelova M, Davey MP (2020) Remote sensing reveals Antarctic green snow algae as important terrestrial carbon sink. Nat Commun. https://doi.org/10.1038/s41467-020-16018-w
Lutz S, Anesio AM, Raiswell R, Edward A, Newton RJ, Gill F, Benning LG (2016) The biogeography of red snow microbiomes and their role in melting Arctic glaciers. Nat Commun 7:11968. https://doi.org/10.1038/ncomms11968
Câmara PEAS, Carvalho-Silva M, Pinto OHB, Amorim ET, Henriques DK, Silva TH, Pellizzari F, Convey P, Rosa LH (2021) Diversity and ecology of Chlorophyta (Viridiplantae) assemblages in protected and non-protected sites in Deception Island (Antarctica, South Shetland Islands) assessed using an NGS approach. Microb Ecol 81:323–334. https://doi.org/10.1007/s00248-020-01584-9
Taş N, Jong AEE, Li Y, Trubl G, Xue Y, Dove NC (2021) Metagenomic tools in microbial ecology research. Curr Opin Biotechnol 67:184–191
Rippin M, Borchhardt N, Williams L, Colesie C, Jung P, Büdel B, Karsten U, Becker B (2018) Genus richness of microalgae and cyanobacteria in biological soil crusts from Svalbard and Livingston Island: morphological versus molecular approaches. Polar Biol 41:909–923. https://doi.org/10.1007/s00300-018-2252-2
Fraser CI, Connell L, Lee CK, Cary SC (2018) Evidence of plant and animal communities at exposed and subglacial (cave) geothermal sites in Antarctica. Polar Biol 41:417–421. https://doi.org/10.1007/s00300-017-2198-9
Garrido-Benavent I, Pérez-Ortega S, Durán J, Ascaso C, Pointing SB, Rodríguez-Cielos R, Navarro F, de los Ríos A (2020) Differential colonization and succession of microbial communities in rock and soil substrates on a maritime Antarctic glacier forefield. Front Microbiol. https://doi.org/10.3389/fmicb.2020.00126
Huss V, Frank C, Hartmann EC, Hirmer M (1999) Biochemical taxonomy and molecular phylogeny of the genus Chlorella sensu lato (Chlorophyta). J Phycol 35(3):587–598. https://doi.org/10.1046/j.1529-8817.1999.3530587.x
Ruppert K, Kline RJ, Rahman MS (2019) Past, present, and future perspectives of environmental DNA (eDNA) metabarcoding: a systematic review in methods, monitoring, and applications of global eDNA. Glob Ecol Conserv 17:1–29. https://doi.org/10.1016/j.gecco.2019.e00547
Van Lipzig NPM, King JC, Lachlan-Cope TA (2004) Precipitation, sublimation, and snow drift in the Antarctic Peninsula region from a regional atmospheric model. J Geophys Res 109:D24106
Hrbáček F, Nývlt D, Láska K (2017) Active layer thermal dynamics at two lithologically different sites on James Ross Island, Eastern Antarctic Peninsula. CATENA 149:592–602
Hadi SIIA, Santana H, Brunale PPM, Gomes TG, Oliveira MD, Matthiensen A, Oliveira MEC, Silva FCP, Brasil BSAF (2016) DNA barcoding green microalgae isolated from Neotropical inland waters. PLoS ONE 11(2):e0149284. https://doi.org/10.1371/journal.pone.0149284
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, London, pp 515–322
Banchi E, Ametrano CG, Greco S, Stanković D, Muggia L, Pallavicini A (2020) PLANiTS: a curated sequence reference dataset for plant ITS DNA metabarcoding. Database. https://doi.org/10.1093/database/baz155
Joshi NA, Fass JN (2011) Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33) [Software]. https://github.com/najoshi/sickle. https://doi.org/10.1080/0028825X.1968.10428810.
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open-source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10(1):57–59. https://doi.org/10.1038/nmeth.2276
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Boylern E, Knight R, Huttley GA, Caporaso JG (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6:90–107. https://doi.org/10.1186/s40168-018-0470-z
Giner CR, Forn I, Romac S, Logares RC, Massana R (2016) Environmental sequencing provides reasonable estimates of the relative abundance of specific picoeukaryotes. Appl Environ Microbiol 82:4757–4766. https://doi.org/10.1128/AEM.00560-16
Darling JA, Mahon AR (2011) From molecules to management: adopting DNA-based methods for monitoring biological invasions in aquatic environments. Environ Res 111:978–988. https://doi.org/10.1016/j.envres.2011.02.001
Guiry MD, Guiry GM (2021) AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. http://www.algaebase.org. Accessed 01 Jan 2021
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Hsieh TC, Ma KH, Chao A (2016) iNEXT: an R package for interpolation and extrapolation of species diversity (Hill numbers). Methods Ecol Evol 7:1451–1456. https://doi.org/10.1111/2041-210X.12613
Bardou P, Mariette J, Escudié F, Djemiel C, Klopp C (2014) jvenn: an interactive Venn diagram viewer. BMC Bioinform 15:293. https://doi.org/10.1186/1471-2105-15-293
Hoham RW, Mullet JE (1978) Chloromonas nivalis (Chod.) Hoh. & Mull. comb. nov., and additional comments on the snow alga, Scotiella. Phycologia 17(1):106–107
Kováčik L, Pereira AB (2001) Green alga Prasiola crispa and its lichenized form Mastodia tesselata in Antarctic environment: general aspects. Nova Hedwigia 123:465–478
Remias D, Procházková L, Holzinger A, Nedbalová L (2018) Ecology, cytology and phylogeny of the snow alga Scotiella cryophila K-1 (Chlamydomonadales, Chlorophyta) from the Austrian Alps. Phycologia 57(5):581–592. https://doi.org/10.2216/18-45.1
Sabbe K, Hodgson DA, Verleyen E, Taton A, Wilmotte A, Vanhoutte K, Vyverman W (2004) Salinity, depth and the structure and composition of microbial mats in continental Antarctic lakes. Freshw Biol 49:296–319. https://doi.org/10.1111/j.1365-2427.2004.01186.x
Buchheim MA, Lemieux C, Otis C, Gutell RR, Chapman RL, Turmel M (1996) Phylogeny of the Chlamydomonadales (Chlorophyceae): a comparison of ribosomal RNA gene sequences. Mol Phylogenet Evol 5(2):391–402
Acknowledgements
This study received financial support from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Programa Antártico Brasileiro (PROANTAR), Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Institutos Nacionais de Ciência e Tecnologia (INCT) Criosfera 2. P. Convey is supported by Natural Environment Research Council (NERC) core funding to the British Antarctic Survey (BAS) ‘Biodiversity, Evolution and Adaptation’ Team. Thanks also to congresswoman Jô Moraes, to Instituto de Ciências Biológicas da Universidade de Brasília and to the Brazilian Navy and Air Force. We thank the Instituto Antártico Argentino for the logistical and financial support of the Antarctic campaign to Vega Island and to the Lagos Field Group.
Funding
National Council for Scientific and Technological Development (CNPq), Brazilian Antarctic Program (PROANTAR), Research Foundation of the State of Minas Gerais (FAPEMIG), Coordination for the Improvement of Higher Education Personnel (CAPES), National Institutes of Science and Technology (INCT) Criosfera 2, Natural Environment Research Council (NERC) core funding to the British Antarctic Survey (BAS) ‘Biodiversity, Evolution and Adaptation’ Team.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Paulo Eduardo Aguiar Saraiva Câmara, Mayara Baptistucci Ogaki, Otávio Henrique Bezerra Pinto, Juan Manuel Lirio, Silvia H. Coria, Rosemary Vieira, Micheline Carvalho-Silva, Eduardo Toledo Amorim, Peter Convey, Luiz Henrique Rosa and Bárbara Medeiros Fonseca. The first draft of the manuscript was written by Bárbara Medeiros Fonseca and Paulo Eduardo Aguiar Saraiva Câmara and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
All authors gave their consent to participate.
Consent for publication
All authors gave their consent for publication.
Research involved humans and/or animals
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fonseca, B.M., Câmara, P.E.A.S., Ogaki, M.B. et al. Green algae (Viridiplantae) in sediments from three lakes on Vega Island, Antarctica, assessed using DNA metabarcoding. Mol Biol Rep 49, 179–188 (2022). https://doi.org/10.1007/s11033-021-06857-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06857-1