Abstract
Background
Research activities aiming to investigate the genetic diversity are very crucial because they provide information for the breeding and germplasm conservation activities. Wheat is one of the most important cereal crops globally by feeding more than a third of the human population around the world.
Methods and results
During present investigation, a total of 74 Turkish bread wheat accessions (54 landraces and 20 cultivars) were used as plant material and iPBS-retrotransposons marker system was used for the molecular characterization. 13 polymorphic primers used for molecular characterization resulted a total of 152 bands. Range of calculated diversity indices like polymorphism information content (0.11–0.702), effective numbers of alleles (1.026–1.526), Shannon’s information index (0.101–0.247) and gene diversity (0.098–0.443) confirmed higher genetic variations in studied germplasm. Bread wheat landraces reflected higher genetic variations compared to commercial cultivars. Analysis of molecular variance resulted that higher (98%) genetic variations are present within populations. The model-based structure algorithm separated 74 bread wheat accessions in to two populations. Diversity indices based on structure evaluated population’s revealed population B as a more diverse population. The principal coordinate analysis and neighbor-joining analysis separated 74 bread wheat accessions according to their collection points. Genetic distance for 74 Turkish bread wheat accessions explored Bingol and Asure accessions as genetically diverse that can be used as parents for breeding activities.
Conclusions
The extensive diversity of bread wheat in Turkish germplasm might be used as genetic resource for the exhaustive wheat breeding program. For instance, accessions Bingol and Asure were found genetically diverse and can be used as parents for future breeding activities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The world is facing a big problem of food scarcity due to climate change and rapidly increasing world population. Khush et al. [1] stated that nearly 800 million people from the developing countries go to bed hungry. World population is increasing with rapid pace and estimated to reach 10 billion by 2050. Due to rapid increase in the world population and continuous changes in climatic conditions, there is a need to boost up world food production to meet food demands in 2050 [2, 3].
Wheat (Triticum aestivum L.) is hexaploid (2n = 6x = 42) is an important cereal crop belonging to Poaceae family and serving a source a food for millions of people [4, 5]. It is believed that bread wheat was originated through two polyploidization events between Triticum urartu (AA genome) and an Aegilops speltoides related species (BB genome) nearly 0.5 million years ago (hereafter Ma), resulting in the formation of Triticum turgidum ssp. diccocoides [6, 7]. Finally, hybridization between Triticum turgidum ssp. Durum (AABB genome) and Aegilops tauschii (DD genome) nearly 10,000 years ago in Fertile Crescent, in a region that nowadays comprises Northern Iran resulted in the formation of modern day hexaploid bread wheat (AABBDD) genome [6,7,8]. During 2019, wheat was cultivated globally on an area of 215,901,958 ha with a production of 765,769,635 tones [9]. Fertile Crescent which includes part of present-day Turkey is considered the origin and domestication center of wheat and its progenitors [10]. During 2019, wheat was cultivated globally on an area of 6,831,854 ha with a production of 19,000,000 tones [9]. As Turkey is a part of Fertile Crescent, therefore it is very important to explore and understand the genetic diversity in Turkish wheat and its wild relatives genetic resources for wheat improvement program [11]. Germplasm characterization is considered prerequisite for breeding activities as it facilitate novel genetic variations to the breeders that can be used for marker-assisted breeding [12]. Previous report confirmed that domestication, human selection, and breeding activities for improved traits resulted in genetic erosin and lowered the diversity wheat gene pool [13]. Therefore, studies regarding the assessment of genetic variations in wheat are very important for future breeding activities. Morphological and molecular markers are two widely used approaches for the characterization of germplasm. However, DNA based markers are more trustable and reproducible and not influenced by environmental factors compared to morphological markers [14].
Advancement in molecular markers techniques revolutionized the breeding activities [14]. Among these, retrotransposons are genetic elements having ability to copy their numbers, change their location and constitute major components of most eukaryotic genomes [15]. Long terminal repeat (LTR) and non-LTR retrotransposons are two major groups of retrotransposons. LTR- retrotransposons are in prevalence and more active in plants compared to non-LTR retrotransposons [16, 17]. However, limitations in both LTR and non-LTR retrotransposons leads the scientific community to develop inter primer binding site (iPBS) marker system [17]. Kalendar et al. [17] proposed iPBS-retrotransposons as a universal marker that can be used for the characterization of both animal and plant species. iPBS-retrotransposons markers has been used for the molecular characterization, phylogenetic and evolutionary study in various crop plants [18,19,20]. Previously, different molecular markers has been used for the molecular characterization of wheat germplasm [21,22,23]. However, there is scarcity of information about the characterization of bread wheat germplasm using iPBS-retrotransposons. Therefore, current study aimed to characterize Turkish bread wheat germplasm for the assessment of genetic diversity and to explore its population structure.
Materials and methods
Plant material and DNA isolation
74 bread wheat accessions including a total of 54 landraces and 20 commercial cultivars were used for the molecular characterization (Table 1). These landraces were collected from 14 provinces of Turkey (Fig. 1). To isolate the genomic DNA, all bread wheat accessions were sown in the greenhouse and their fresh and young leaves were harvested. The DNA extraction was achieved using CTAB protocol [24] and a specific protocol recommended by Diversity Arrays Technology (available at https://www.diversityarrays.com/orderinstructions/plant-dnaextraction-protocol-for-dart/). Quantification of isolated DNA samples was performed using 0.8% agarose gel and NanoDrop (DS11 FX, DeNovix Inc., Wilmington, DE, USA). 5 ng/μL was prepared as a final concentration for further polymerase chain reaction (PCR) analysis.
iPBS-retrotransposons PCR amplifications
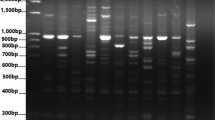
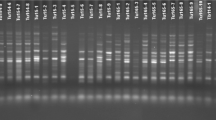
Initially, 75 iPBS primers were screened on randomly selected 10 bread wheat accessions. Among these 75 screened primers, 13 most polymorphic primers were evaluated for final PCR amplification of all 74 bread wheat accessions (Table 2). PCR amplification was executed according to the methodology of Kalendar et al. [17]. After PCR amplification, PCR product was run on 2% (w/v) agarose gel having TBE buffer (0.5×) at a stable voltage of 120 V for 220 min. Staining of gel was performed using ethidium bromide and graphics were taken through a UV Imager Gel Doc XR+system (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
Statistical analysis
As a dominant marker system, scoring was performed in binary fashion; 1 or 0 representing the presence and absence of a band respectively. Various diversity parameters like gene diversity (He) Shannon’s information index (I) and effective alleles number (Ne) were investigated through Popgene ver. 1.32 [25]. The Nei’s genetic distance among 74 Turkish bread wheat accessions was calculated using Popgene ver. 1.32 [25]. To explore genetic variations between landraces and cultivars, various diversity indices were also calculated using GenAlExV6.5 [26] software. The polymorphism information contents (PIC) was found using a formula i.e. PIC = 2fi (1 − fi), given by Roldán-Ruiz et al. [27]. Here fi represents the frequency of present loci of a molecular marker while (1 − fi) represents the frequency of absent loci. Principal coordinate analysis (PCoA) and analysis of molecular variance (AMOVA) were calculated through GenAlExV6.5 [26] software. To explore the genetic relationship among 74 Turkish bread wheat accessions, neighbor joining analysis was performed using R statistical software. Structure software was used to explore the population structure of Turkish bread wheat germplasm (Pritchard et al. [28]). The favorable numbers of clusters (K subpopulations) were estimated (1–10) by repeating analysis three time according to the report of Evanno et al. [29]. During each run, the burn-in and Markov Chain Monte Carlo (MCMC) were set to 50,000 each, and iterations were set to 10. Later, structure evaluated results were processed with STRUCTURE HARVESTER v.0.9.94 [30] to investigate most favorable K value. The pophelper an R package was used to visualize the most favorable ΔK [31].
Results
During this study, 13 iPBS-retrotransposons primers were used for the molecular characterization of Turkish bread wheat germplasm. These 13 primers resulted 152 bands and 11.69 bands were average bands/primer (Table 3). iPBS-2257 and iPBS-2257 produced maximum (16) bands, while minimum (4) bands were yielded with iPBS-2246. Among 152, 111 (73.2%) bands were found polymorphic, while 8.54 were average bands/primer. iPBS-2095 was found most polymorphic primer as it produced maximum numbers of polymorphic bands. The iPBS-2095 and iPBS-2381 primers showed maximum (100%) polymorphism. PIC value ranged 0.702 to 0.11 for iPBS-2074 and iPBS-2376 respectively, and mean PIC value was 0.42. The iPBS-2374 and iPBS-2376 produced maximum (1.526) and minimum (1.026) effective number of alleles respectively, while 1.312 was mean effective number of alleles during this study. Shannon’s information index ranged 0.101 for iPBS-2376 to 0.247 for iPBS-2374 and mean Shannon’s information index during this study was 0.165. Mean gene diversity was 0.256, while iPBS-2376 and iPBS-2374 resulted minimum (0.098) and maximum (0.443) gene diversity. Mean Nei’s genetic distance was 0.190, while maximum and minimum genetic distance was 0.427 (Bingol and Asure) and 0.04 (Van5 and Agri4). Various diversity indices were also calculated among bread wheat landraces and cultivars to explore the level of genetic variations (Table 4). Turkish bread wheat landraces reflected higher polymorphism (66.45%) and other calculated diversity indices compared to cultivars. Results of AMOVA reported the existence of higher (98%) genetic variation within population compared to among the populations (2%) (Table 5).
The genetic structure of Turkish bread wheat germplasm was separated into two groups as proposed by ΔK peak at K = 2 constructed in the structure harvester analysis (Fig. 2). The model-based structure algorithm separated 74 bread wheat accessions in to two populations on the basis of their collection points (Fig. 3). Population A clustered a total of 58 bread wheat accessions, while 16 accessions were present in population B. Various diversity indices and AMOVA was also calculated among structure evaluated populations (Table 6). Results showed that population B has more genetic variations compared to population A. The AMOVA also revealed the existence of higher genetic variations within population (92%) compared to among the populations (8%). The neighbor-joining analysis separated 74 bread wheat accessions into three population on the basis of their collection points (Fig. 4). The PCoA analysis strengthen the clustering of model-based structure algorithm by separating the 74 bread wheat accessions into two populations (Fig. 5).
Discussion
A reasonable studies have been documented for the characterization of bread wheat germplasm and its wild relatives using various types of molecular markers [21,22,23]. Regarding to retrotransposons based markers Demirel [32], used iPBS-retrotransposons marker for the molecular characterization of emmer and durum wheat. Queen et al. [33] used SSAP marker for linkage and genetic diversity analysis in bread wheat and its wild relatives. Similarly, Holasou et al. [21] used IRAP and REMAP markers for the molecular characterization of 49 Iranian bread wheat cultivars. However to best of knowledge, iPBS-retrotransposons markers are not used for the characterization of bread wheat germplasm.
During this study, 13 iPBS-retrotransposons primers yielded a total of 152 bands, among which 111 were found polymorphic (Table 3). Total and polymorphic bands reported in this study were higher than Nazarzadeh et al. [34] using RAPD and ISSR markers Kumar et al. [35], using ISSR marker Alshehri et al. [36], using SCoT and ISSR primers and Çifçi and Yağdi [37] using RAPD markers. The range and mean PIC value reported herein was found higher than earlier studies of Kumar et al. [35] using ISSR markers El-Sherbeny et al. [38], using ISSR markers and AL-Tamimi and AL-Janabi [39] using RAPD and ISSR markers. The resulted mean and range of effective number of alleles was higher than the Kumar et al. [35]. It was observed that iPBS-retrotransposons primers resulting the less number of alleles also resulted in low gene diversity. Similarly, higher gene diversity was observed for the primers producing higher alleles. This pattern was found similar with the Kumar et al. [35]. Mean gene diversity and Shannon’s information index observed in present report was higher than the Carvalho et al. [40]. Presence of higher values for various diversity indices in this study might be due to differences in germplasm and the nature of molecular marker. iPBS-retrotransposons marker system has been found highly reproducible and its universal nature has been already proven in various studies [18, 41]. Therefore, this marker system should be preferred for the molecular characterization of bread wheat germplasm compared to other dominant marker systems.
During this study, bread wheat landraces and cultivars were used as a plant material. Therefore, calculated diversity indices among landraces and cultivars showed the presence of higher genetic variations in landraces compared to cultivars. These results were in line with previous studies as they also reported the existence of higher genetic variations in wheat landraces compared to their cultivars [42, 43]. The AMOVA results revealed that maximum genetic variations in Turkish bread wheat germplasm are present within population. Results of AMOVA were also supported by previous studies as they also revealed higher genetic variations within populations [44, 45]. The Nei’s genetic distance revealed Bingol and Asure as genetically distinct accessions. Arystanbekkyzy et al. [41] stated that genetically distinct accessions are can be helpful to start breeding activities for favorable traits. Therefore, Bingol and Asure accessions can be used for future bread wheat breeding.
The model-based structure algorithm grouped 74 bread wheat accessions into two populations on the basis of their collection points (Fig. 3). Population A was found larger than population B by accounting 78.37% (58 accessions) accessions. Population B accounted a total of 16 accessions and 6 of these were commercial cultivars. The remaining 10 commercial cultivars were present in population A. It can be seen in structure results that accessions belonging to same province or their neighbor province were present showed similarity with each other. For example, accessions from Erzurum showed genetic similarity with Kars and similar was the case with the accessions from Bitlis and Van. It was also observed that accessions from east and north east provinces of Turkey were present in population B by making it a diverse population and showing their genetic similarity with each other’s. The neighbor-joining analysis grouped studied germplasm into three populations mainly on the basis of their collection points (Fig. 4). Population C was found larger than rest of the populations and clustered a total of 49 accessions. A total of 11 cultivars were grouped in population A, while population B and C accounted a total of 1 and 8 cultivars respectively. The neighbor-joining analysis showed admixture of accessions because accessions from various provinces were grouped under the same sub-groups. Therefore, preference was given to structure clustering because structure algorithm has been proven more trustable and much informative compared to other clustering algorithms [46, 47]. Different diversity parameters were also calculated for structure evaluated populations that revealed the existence of higher genetic variations in population B. The AMOVA analysis for structure evaluated population also confirmed the presence of higher genetic variations within populations compared to among the populations. Thus, it is stated that Turkish bread wheat germplasm has great level of genetic variations within the population that can be helpful for the breeding of this crop in future. The PCoA analysis supported the clustering of model-based structure algorithm and separated the Turkish bread wheat germplasm into two populations (Fig. 5).
Conclusion
This study provided a deep insight about genetic variations in Turkish bread wheat germplasm using iPBS-retrotransposons marker system. The Bingol and Asure were found genetically most diverse accessions and should be used for future breeding activities. Results of AMOVA explored higher genetic variations within populations compared to among the populations. Population A from structure clustering was found more diverse and accessions belonging to this population should be considered for future wheat breeding. The model-based structure algorithm and PCoA separated the studied germplasm into population mainly on the basis of their collection points. Present study also confirmed the applicability and universal nature of iPBS-retrotransposons markers that can be used for the investigation of genetic diversity of any crop.
Data availability
All data needed to conduct this study is provided within the manuscript.
References
Khush GS, Lee S, Cho JI, Jeon JS (2012) Biofortification of crops for reducing malnutrition. Plant Biotechnol Rep 6:195–202
Godfray HC, Beddington JR, Crute IR, Haddad L, Lawrence D, Muir JF, Pretty J, Robinson S, Thomas SM, Toulmin C et al (2010) Food security: the challenge of feeding 9 billion people. Science 327:812–818
Tilman D, Balzer C, Hill J, Befort BL (2011) Global food demand and the sustainable intensification of agriculture. PNAS 108:20260–20264
Bhatta M, Eskridge KM, Rose DJ, Santra DK, Baenziger PS, Regassa T et al (2017) Seeding rate, genotype, and topdressed nitrogen effects on yield and agronomic characteristics of winter wheat. Crop Sci 57:951–963
Marcussen T, Sandve SR, Heier L, Spannagl M, Pfeifer M, Jakobsen KS, Wulff BB, Steuernagel B, Mayer KF, Olsen OA (2014) International wheat genome sequencing consortium ancient hybridizations among the ancestral genomes of bread wheat. Science 345(6194):1250092. https://doi.org/10.1126/science.1250092
Feldman M, Lupton FG, Miller TE (1995) Wheat Triticum spp. (Gramineae-Triticinae). In: Smartt J, Simmonds NW (eds) Evolution of crop plants. Longman Scientific and Technical, New York, pp 184–192
Huang X, Wang L, Xu M, Röder M (2003) Microsatellite mapping of the powdery mildew resistance gene Pm5e in common wheat (Triticum aestivum L.). Theor Appl Genet 106(5):858–865
Dubcovsky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316(5833):1862–1866
Food and Agriculture Organization (2021) FAO statistical yearbook. Available at http://www.fao.org/docrep/017/i3138e/i3138e. Accessed 17 June 2021
Karagöz P, Özkan M (2014) Ethanol production from wheat straw by Saccharomyces cerevisiae and Scheffersomyces stipitis co-culture in batch and continuous system. Biores Technol 1(158):286–293
Morgounov AI, Keser M, Kan M, Kucukcongar M, Ozdemir F, Gummadov N, Muminjanov H, Zuev E, Qualset C (2016) Wheat landraces currently grown in Turkey: distribution, diversity, and use. Crop Science Society of America, Fitchburg
Nadeem MA, Gündoğdu M, Ercişli S, Karaköy T, Saracoğlu O, Habyarimana E, Lin X, Hatipoğlu R, Nawaz MA, Sameeullah M, Ahmad F, Baloch FS et al (2020) Uncovering phenotypic diversity and DArTseq marker loci associated with antioxidant activity in common bean. Genes. https://doi.org/10.3390/j.genes.2020.11010036
Haudry A, Cenci A, Ravel C, Bataillon T, Brunel D, Poncet C, Hochu I, Poirier S, Santoni S, Glémin S, David J et al (2007) Grinding up wheat: a massive loss of nucleotide diversity since domestication. Mol Biol Evol 24:1506–1517
Nadeem MA, Nawaz MA, Shahid MQ, Doğan Y, Comertpay G, Yıldız M, Hatipoğlu R, Ahmad F, Alsaleh A, Labhane N, Özkan H, Baloch FS et al (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32:261–285
Finnegan DJ (1989) Eukaryotic transposable elements and genome evolution. Trends Genet 5:103–107. https://doi.org/10.1016/0168-9525(89)90039-5
Alzohairy AM, Gyulai G, Ramadan MF, Edris S, Sabir JS, Jansen RK, Eissa HF, Bahieldin A et al (2014) Retrotransposon-based molecular markers for assessment of genomic diversity. Funct Plant Biol 41:781–789
Kalendar R, Antonius K, Smýkal P, Schulman AH (2010) iPBS: a universal method for DNA fingerprinting and retrotransposon isolation. Theor Appl Genet 121:1419–1430. https://doi.org/10.1007/s00122-010-1398-2
Karık Ü, Nadeem MA, Habyarimana E, Ercişli S, Yildiz M, Yılmaz A, Yang SH, Chung G, Baloch FS et al (2019) Exploring the genetic diversity and population structure of Turkish laurel germplasm by the iPBS-retrotransposon marker system. Agronomy. https://doi.org/10.3390/agronomy9100647
Baloch FS, Alsaleh A, de Miera LE, Hatipoğlu R, Çiftçi V, Karaköy T, Yıldız M, Özkan H et al (2015) DNA based iPBS-retrotransposon markers for investigating the population structure of pea (Pisum sativum) germplasm from Turkey. Biochem Syst Ecol 61:244–252
Nemli S, Kianoosh T, Tanyolac MB (2015) Genetic diversity and population structure of common bean (Phaseolus vulgaris L.) accessions through retrotransposon-based inter primer binding sites (iPBSs) markers. Turk J Agric For 39:940–948
Holasou HA, Rahmati F, Rahmani F, Imani M, Talebzadeh Z (2019) Elucidate genetic diversity and population structure of bread wheat (Triticum aestivum L.) cultivars using IRAP and REMAP markers. JCSB 22:139–151
Khadka K, Torkamaneh D, Kaviani M, Belzile F, Raizada MN, Navabi A et al (2020) Population structure of Nepali spring wheat (Triticum aestivum L.) germplasm. BMC Plant Biol. https://doi.org/10.1186/s12870-020-02722-8
Yadav MK, Chand P (2018) Assessment of genetic diversity among twenty Indian wheat (Triticum aestivum L.) cultivars using simple sequence repeat (SSR) markers. Int J Curr Microbiol App Sci 7:1708–1717
Doyle JJ, Doyle JL (1990) Isolation ofplant DNA from fresh tissue. Focus 12:39–40
Yeh FC, Yang R, Boyle TJ, Ye Z, Xiyan JM (2000) PopGene32, Microsoft Windows-based freeware for population genetic analysis, version 1.32. Molecular Biology and Biotechnology Centre, University of Alberta, Edmonton
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinforma
Roldán-Ruiz I, Dendauw J, Van Bockstaele E, Depicker A, De Loose M (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Mol Breed 6(2):125–134. https://doi.org/10.1023/A:1009680614564
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Earl DA (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Francis RM (2017) pophelper: an R package and web app to analyse and visualize population structure. Mol Ecol Resour 2017:27–32
Demirel F (2020) Genetic diversity of Emmer wheats using iPBS markers. Avrupa Bilim ve Teknoloji Dergisi. https://doi.org/10.31590/ejosat.814537
Queen RA, Gribbon BM, James C, Jack P, Flavell AJ et al (2004) Retrotransposon-based molecular markers for linkage and genetic diversity analysis in wheat. Mol Genet Genom 271(1):91–97
Nazarzadeh Z, Onsori H, Akrami S (2020) Genetic diversity of bread wheat (Triticum aestivum L.) genotypes using RAPD and ISSR molecular markers. J Genet Resour 6:69–76
Kumar P, Sharma V, Sanger RS, Kumar P, Yadav MK et al (2020) Analysis of molecular variation among diverse background wheat (Triticum aestivum L.) genotypes with the help of ISSR markers. IJCS 8:271–276
Alshehri MA, Alzahrani O, Aziza AT, Alasmari A, Ibrahim S, Bahattab O, Osman G, Alshamari AK, Alduaydi SA et al (2020) Correlation and genetic analyses of different characteristics in Saudi Arabian wheat reveal correlation networks and several trait-associated markers. J Anim Plant Sci 30(6):1486–1497
Çifçi EA, Yağdi K (2012) Study of genetic diversity in wheat (Triticum aestıvum) varities using random amplified polymorphic DNA (RAPD) analysis. Turk J Field Crop 17(1):91–95
El-Sherbeny GA, Omara MK, Farrage AA, Khaled AG (2020) Associations between ISSR markers and quantitative traits in bread wheat genotypes. Asian J Biol Sci 2(1):1–8
Al-Tamimi AJ, Al-Janabi AS (2019) Genetic diversity among bread wheat genotypes using RAPD and SSR markers. SABRAO J Breed Genet 51:325–339
Carvalho A, Lima-Brito J, Maçãs B, Guedes-Pinto H (2009) Genetic diversity and variation among botanical varieties of old Portuguese wheat cultivars revealed by ISSR assays. Biochem Genet 47:276–294
Arystanbekkyzy M, Nadeem MA, Aktas H, Yeken MZ, Zencirci N, Nawaz MA, Ali F, Haider MS, Tunc K, Chung G, Baloch FS et al (2019) Phylogenetic and taxonomic relationship of turkish wild and cultivated emmer (Triticum turgidum ssp. dicoccoides) revealed by iPBSretrotransposons markers. IJAB 21:155–163
Etminan A, Pour-Aboughadareh A, Mohammadi R, Ahmadi-Rad A, Noori A, Mahdavian Z, Moradi Z et al (2016) Applicability of start codon targeted (SCoT) and inter-simple sequence repeat (ISSR) markers for genetic diversity analysis in durum wheat genotypes. Biotechnol Biotechnol Equip 30:1075–1081
Shoaib A, Arabi MI (2006) Genetic diversity among Syrian cultivated and landraces wheat revealed by AFLP markers. Genet Resour Crop Evol 53:901–906
Mourad AM, Belamkar V, Baenziger PS (2020) Molecular genetic analysis of spring wheat core collection using genetic diversity, population structure, and linkage disequilibrium. BMC Genom. https://doi.org/10.1186/s12864-020-06835-0
Darvishzadeh R, Bernousi I (2012) Molecular similarity relationships among Iranian bread wheat cultivars and breeding lines using ISSR markers. Not Bot Horti Agrobot Cluj Napoca 40:254–260
Bouchet S, Pot D, Deu M, Rami JF, Billot C, Perrier X, Rivallan R, Gardes L, Xia L, Wenzl P, Kilian A et al (2012) Genetic structure, linkage disequilibrium and signature of selection in sorghum: lessons from physically anchored DArT markers. PLoS ONE. https://doi.org/10.1371/journal.pone.0033470
Newell MA, Cook D, Hofmann H, Jannink JL (2013) An algorithm for deciding the number of clusters and validation using simulated data with application to exploring crop population structure. Ann Appl Stat 7:1898–1916
Acknowledgements
The author is very grateful to Assoc. Prof. Faheem Shehzad Baloch from Faculty of Agricultural Sciences and Technologies, Sivas University of Science and Technology for providing plant material and supporting during whole experiment and reviewing the manuscript.
Funding
No funding was received to conduct this study.
Author information
Authors and Affiliations
Contributions
MAN conceptualized and established the methodology for this study, also performed molecular characterization of bread wheat germplasm, statistical analysis, and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author confirm that this article content has no conflict of interest.
Consent to publish
Author read the manuscript and showed his willingness to publish this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nadeem, M.A. Deciphering the genetic diversity and population structure of Turkish bread wheat germplasm using iPBS-retrotransposons markers. Mol Biol Rep 48, 6739–6748 (2021). https://doi.org/10.1007/s11033-021-06670-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06670-w