Abstract
Background
WRKY transcription factor is involved in regulation of plant growth and development, response to biotic and abiotic stresses, including homologous WRKY3 and WRKY4 genes which play a vital role in regulating plants defense response to pathogen and drought stress.
Methods and results
To investigate the function of AtWRKY3 and AtWRKY4 genes in regulating salt and Me-JA stresses, the loss-of-function mutations were generated by clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated 9 (Cas9) system in Arabidopsis thaliana. Several independent transgenic lines with single or double mutations were obtained via Agrobacterium-mediated transformation. The knockout lines of AtWRKY3 and AtWRKY4 genes were successfully achieved and confirmed by qRT-PCR technology. Expression analysis showed that AtWRKY3 and AtWRKY4 genes had significantly up-regulated under salt and Me-JA stresses. The growth of double mutant plants under salt or Me-JA stresses were significantly inhibited compared with corresponding wild type (WT) plants, especially their root lengths. Moreover, the double mutant plants displayed salt and Me-JA sensitivity phenotypic characteristics, such as the increased relative electrolyte leakage (REL) and a substantial reduce in the activities of antioxidant enzymes, including superoxide dismutase (SOD), peroxidase (POD) and catalase (CAT) activities.
Conclusion
Taken together, these data suggested that the simultaneous modification of homologous gene copies of WRKY are established using CRISPR/Cas9 system in A. thaliana and the loss of AtWRKY3 and AtWRKY4 has an effect on ROS scavenging pathways to reduce stress tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants have evolved complicated system that can counteract the influence of environmental fluctuations including biotic and abiotic stresses through sophisticated network of transcriptional or hormonal components. The transcriptional elements may include different transcription factors (TFs), such as WRKY, AP2 (APETALA2)/ERF (ethylene-responsive factor) and NAC [no apical meristem (NAM), Arabidopsis thaliana transcription activation factor [ATAF1/2] and cup-shaped cotyledon (CUC2)], are plant-specific and they particularly play critical and unique roles in the control of plant life activities, such as growth and development, and response to diverse environmental stresses [1]. WRKY TFs comprise one of the largest TF families in plants and are characterized by a long DBD (DNA binding domain) with a highly conserved WRKYGQK heptapeptide sequence and a C2H2 or C2HC Zinc finger motif [2]. These conserved sequences are known as the WRKY domains which are indispensable for WRKY specific binding to W-box (C/T)TGAC(T/C) in the promoter regions [2, 3]. The first identified WRKY TF–SWEET POTATO FACTOR1 (SPF1) negatively regulate the sporamin and β-amylase expression in sweet potato (Ipomoea batatas)[4]. So far, 109, 74 and 86 WRKY superfamily members in Oryza sativa, A. thaliana and Brachypodium distachyon have been reported, respectively [5, 6]. WRKY TFs play essential roles in various abiotic stresses, including wounding, drought, soil salinity, heat, cold, and heavy metal contamination [7,8,9]. Moreover, WRKY TFs have been demonstrated that is induced by a variety of signaling substances including salicylic acid (SA), jasmonic acid (JA) and abscisic acid (ABA) [10,11,12]. To date, WRKY TFs have been implicated in various physiological processes including plant growth, root development, metabolism, embryo morphogenesis, senescence, and germination [13, 14]. Importantly, the physiological functions of WRKY3 and WRKY4 genes have been demonstrated in response to pathogen infection and abiotic stresses. For instance, the NaWRKY3 in tobacco is involved in defense response to wounding [15]. MuWRKY3 enhances drought stress tolerance in groundnut by regulating the expression of stress-responsive genes and the activity of reactive oxygen species (ROS) scavenging enzymes [16]. Arabidopsis WRKY3 and WRKY4 genes enhance plant resistance to the necrotrophic pathogens [17]. However, the role of the two WRKY transcription factors in response to salt stress and JA has not been directly analyzed with their loss-of-function mutants.
More recently, the clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein 9 (CRISPR/Cas9) system was used for gene editing to produce mutants has been rapidly developed [18, 19]. Compared to RNAi technology, CRISPR/Cas9 technology has prominent advantages including complete, high efficiency and stable activation of gene expression, which can provide strong technical support for plant gene functional research and crop molecular breeding. However, the biological functional redundancy usually occurs into two or more homologous genes. If only one gene is knockout, the expected phenotype may be not generated. The different single guide RNA (sgRNA) mix with same Cas9 protein to cut specific corresponding sites for generating double-strand breaks (DSB) have been developed [20]. The method can attain simultaneously editing of homologous genes and solve these issues mentioned above. For example, seven gene heritable mutations have produced using this method via single transformation events in rice [21]. More importantly, a sgRNA designed with their conserved sequence of homologous genes can direct Cas9 to specific corresponding sites to simultaneously generate multiple mutations.
In recent years, with the shortage of freshwater resource and the aggravation of soil salinization, salt-tolerance breeding has become a hot spot in the world. Plants response to salt stress signaling and Na+ tolerance through a calcium-dependent protein kinase pathway namely the Salt-Overly-Sensitive (SOS) pathway [22]. In this pathway, the EF-hand Ca2+-binding protein SOS3 (also called calcineurin B-like, CBL) that are activated by binding the cytosolic calcium signal elicited by salt stress can physically interacts with and activates SOS2 (also namely CBL-interacting protein kinases, CIPK) [23, 24]. Subsequently, the activated SOS2 phosphorylates and activates the SOS1 that is a Na+/H+ antiporter located at the plasma membrane [25]. In addition, JA is an important signaling molecule in plant signal pathway that plays a significant role in regulating plant defense response and abiotic stress [26, 27]. For instance, Arabidopsis WRKY50 and WRKY51 proteins mediate low oleic acid-derived repression of JA signaling pathway for defense responses [28]. There are other physiological processes involved with the JA, such as senescence [29], elicitors of plant secondary metabolism [30], floral nectar synthesis and secretion [31], and so on.
Although studies of WRKY3 and WRKY4 in A. thaliana reveal their functions to varying degrees, they have not been functionally characterized in response to salt and Me-JA stresses. In this study, we characterized their roles of the AtWRKY3 and AtWRKY4 in response to salt and Me-JA stresses based on the mutants generated by CRISPR/Cas9 system. Our results revealed that the loss of AtWRKY3 and AtWRKY4 functions resulted in an attenuated tolerance of salt and Me-JA in A. thaliana, especially in the double mutants, demonstrating that AtWRKY3 and AtWRKY4 play a positive role and functional redundancy in salt and Me-JA stress responses.
Materials and methods
Plant materials, growth conditions and treatments
The A. thaliana Columbia-0 (Clo-0) ecotype was used in this study. Seedlings were grown in a soil mix (clay is mixed with soil at 3:1 ratio) and placed in a growth room with 22 °C and 70% room humidity under a 16-h-light (> 3000 lx)/8-h-dark photoperiod. For quantitative real-time PCR (qRT-PCR) analyses, nine-day-old seedlings were treated with 100 mM NaCl and 300 μM Me-JA for 4 h and 24 h, respectively. The whole plants were collected on the corresponding time point and frozen in liquid nitrogen for further RNA extraction.
For salt stress treatment, nine-day-old transgenic seedlings were placed with 50 or 100 mM NaCl solution, then each length of the main root of the WT and mutant plants were measured and analyzed at two days after treatments. Me-JA treatment was achieved by exposing nine-day-old transgenic plants to 100 or 300 μM Me-JA solution for two days, and the growing status were photographed and the root length were measured. For all the treatments and experiments above, Clo-0 and mutants were grown in 1/2 MS medium under identical controlled conditions. The seedling samples were collected after treatments and stored at − 80 °C until use.
CRISPR/Cas9 vector construction with specific sgRNA and A. thaliana transformation
A detailed description of pYAO:hSpCas9-target-sgRNA fusion plasmid used in the present study was reported previously [32]. The WRKY3 (At2g03340) and WRKY4 (At1g13960) gene information were downloaded from the Arabidopsis information resources database (TAIR: http://www.arabidopsis.org/) [33]. Multiple sequence alignment of the coding sequence (CDS) of AtWRKY3 and AtWRKY4 genes for searching appropriate conserved sequence were performed by the Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo/). Finally, the sequence upstream adjacent to protospacer adjacent motif (PAM) which contained a specific restriction enzyme site (NdeI) was selected as the target to facilitate subsequent identification of positive plants. Subsequently, ‘TGATT’ and ‘AAAC’ oligonucleotide sequences which were the cohesive terminals of BsaI site was added at the 5′ end as sgRNA upper/lower primer, respectively (Table S1). The DNA oligonucleotides designed was synthesized by Sangon Biotechnologies, Inc. (shanghai, CN) and annealed to generate dimers. The pYAO:hSpCas9-target-sgRNA vector was digested by BsaI and then inserted into annealed DNA to generate pYAO:hSpCas9-WRKY3/4-sgRNA, and the sequence integrity was confirmed by Sanger sequencing. Then the positive vectors were introduced into Agrobacterium tumefacien strain EHA105 using electric shock and used for transformation by the floral-dip to generate the single or double mutant of AtWRKY3 and AtWRKY4 in transgenic A. thaliana.
Identification of AtWRKY3/WRKY4 knockout transgenic plants using PCR and sequencing analyses
The transgenic T0 seeds mentioned above were sterilized with 75% ethanol for 1 min and then 10% bleach for 10 min, washed at least three times with sterile water. The sterile seeds were sown on the 1/2 MS medium containing 25 mg/L hygromycin at 4 °C for three days in dark and transferred to the greenhouse at 22 °C under 16-h-light/8-h-dark photoperiod. After two to three weeks, Hyg-resistant seedlings were potted in soil until they grown to 4–6 leaf period. Genomic DNA was extracted from T1 transgenic plant leaves using CTAB method for PCR using a pair of specific primers (Table S1). Likewise, to eliminate the effect on the multi-allelic patterns, the T2 whole transgenic seedlings were also used for mutation identification of AtWRKY3 and AtWRKY4 genes. The PCR product was amplified from WT and transgenic genomic DNA with specific primers mentioned above and followed by NdeI full digestion and then separated by 1% agarose gel electrophoresis. No digested products selected were purified from the gel and ligated to pGEM®-T Easy Vector Systems (Promega, Madison, WI, USA) for sequencing. The sequences of transgenic and WT plants were aligned using program Vector NTI Advanced v9.0 to characterize the detailed mutations induced by CRISPR/Cas9 system.
Gene expression analysis
To evaluate the expression patterns of AtWRKY genes under salt and Me-JA stresses, we also executed the experiments by qRT-PCR. Twelve AtWRKY genes including AtWRKY2, -3, -4, -8, -16, -28, -31, -34, -42, -52, -64, and -66 were selected and detected. For the abiotic stress treatment, nine-day-old seedlings were dipped in 1/2 MS liquid medium containing 100 mM NaCl and 300 μM Me-JA, and then the whole plants were harvested after treatment for 4 h and 24 h, respectively. To examine whether AtWRKY3 and AtWRKY4 have been broken in mutants mentioned above, their expression levels using qRT-PCR method was performed. The reaction mixture for PCR is as follows: 5 μL SYBR Premix Ex Tap (2×), 0.4 μL PCR Forward Primer (1 μmol /mL), 0.4 μL PCR Reverse Primer (1 μmol /mL), 4 μL cDNA template. The program is: 95 °C 30 s; 95 °C 5 s, 60 °C 34 s, Repeat steps for 40 cycles; 95 °C 15 s, 60 °C 1 min; 95 °C 15 s, ROX Reference Dye (50×) 0.2 μL. The Actin (At3g18780) gene of A. thaliana was used as the reference gene for all qRT-PCR analyses. Each treatment was conducted with three independent biological replicates. The relative expression levels were calculated by the 2−ddCT comparative method. The primer sets were listed in Table S1.
Physiological and biochemical measurements
For physiological and biochemical analysis, the whole mutants and WT plants were collected from each the salt and Me-JA stress assays. Briefly, after growth for nine days in plate, the seedlings were transferred to 1/2 MS medium with 50 mM or 100 mM NaCl; 100 or 300 μM Me-JA for two days and harvested for next analysis. Superoxide dismutase (SOD), peroxidase (POD), catalase (CAT) activities and relative electrolyte leakage (REL) were determined spectrophotometrically following previously described protocols [34,35,36]. At least 15 seedlings were contained in each sample, and each treatment was implemented with three biological replicates. Data analyses were performed using DPS 7.05 Data Processing System Software. For all analyses, the significance of differences was calculated at p < 0.05. Sample variability is given as the standard deviation (SD) of the mean.
Results
Selection of target sites and vector construction
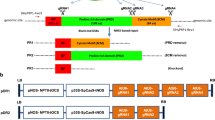
The CRISPR-Cas9-sgRNA vector was used to express the target sequences of guide RNA driven by U6-26 promoter and Cas9 enzyme driven by YAO promoter (Fig. 1A). In order to obtain the simultaneous knockout mutations in the AtWRKY3 and AtWRKY4 transcription factors, the sequence of target sites (TS) that is high conserved used for guide RNA was selected. Moreover, the sgRNA contained a NdeI restriction enzyme site for subsequent identification of positive plants (Fig. 1B). Therefore, the 19 bp sequence “CTGTTGTAACAACATATGA” was selected for the generation of sgRNA (Fig. 1B). The TS of both sequences were almost the same only with a difference of a single nucleotide that is threonine (T) in WRKY3 changed into alanine (A) in WRKY4 (Fig. 1B). The sequence closer to PAM has higher homology and lower off-target effects. Therefore, the selected sgRNA theoretically meet the requirements of follow-up study. The sequences of TS reside in the C-terminal WRKY domain correspond to amino residues 460–467 in WRKY3 and 452–459 in WRKY4, respectively (Fig. 1C).
Construction of AtWRKY3/4-pYAO-Cas9 binary vector. A Structure of the CRISPR/Cas9 binary vector for A. thaliana transformation by floral-dip method. The hSpCas9 cassette is driven by the YAO promoter, while sgRNA is controlled by the AtU6-26 promoter. NLS, nuclear localization sequence; LB, left border; RB, right border. B Schematic representation of sgRNA primer of AtWRKY3/4. PAM, protospacer adjacent motif; NdeI, NdeI enzyme site. C Location of target sites (TS) in protein. Blue rectangle and black line indicated exons and introns, respectively. (Color figure online)
Identification of transgenic positive plants
The sgRNA designed was synthesized and cloned into the BsaI restriction site of the hSpCas9 vector (pYAO:hSpCas9-WRKY3/4-sgRNA), which was introduced into the A. thaliana genome by Agrobacterium-mediated transformation with floral-dip method. Next, the T1 seeds were generated to detect the situations of mutation in genome editing at TS. To determine the detailed information of insertion–deletion (InDel) modifications, the T1 plants were screened by hygromycin and then genomic DNA of anti-hygromycin plant leaf was extracted. Subsequently, the specific fragment contained TS was amplified and digested by NdeI enzyme. PCR products should have a single band because the genomes of transgenic plants had mutations at the predicted Cas9 cleavage site (3 nt upstream of the PAM sequence) in the AtWRKY3 or AtWRKY4 gene sequence, which resulted in the damage of NdeI enzymatic site (Fig. S1). The result showed that 12 lines had a single band at 692 bp in the AtWRKY3 sequence including lines 2#, 3#, 4#, 8#, 9#, 11#, 12#, 13#, 16#, 19#, 20#, and 22# (Fig. S1). The rest was partially cut off by the enzyme, indicating that these plants were chimeric. What's more, the similar results also had a single band at 455 bp in the AtWRKY4 sequence including lines 8# and 19# (Fig. S1), indicating that the two plants likely was homozygous. The sequencing results exhibited that there were various mutations in the AtWRKY3 and AtWRKY4 genomic sequence (Fig. S1). In addition, we also obtained the single mutants of AtWRKY3 or AtWRKY4 gene, such as, lines 24#, 25#, and 28# had only mutations of AtWRKY3, while lines 23#, 32#, and 35# in AtWRKY4 (Fig. S2).
To eliminate the multi-allelic patterns and further validate the mutation situation, the whole seedlings of T2 transgenic plants were also analyzed with same methods mentioned above. Electrophoretic results displayed that seven lines had a single band at 692 bp for AtWRKY3, and only 8#, 13#, and 16# were single band at 455 bp for AtWRKY4, suggesting that these plants were probable homozygous (Fig. 2A and B). Moreover, the sequencing results showed that mutations had different profiles in the AtWRKY3 and AtWRKY4 genomic sequence. For example, there was a deletion of 15 bp in AtWRKY3 genomic sequence in 2# plant, while 16# and 19# plants had a single thymine (T) or adenine (A) insertion at 3 nt upstream of the PAM sequence, respectively (Fig. 2C and E). 20# plants deleted 4 bp at 3 nt upstream of the PAM sequence, while 8# and 13# plants showed a deletion of 2 bp in the sgRNA TS of AtWRKY4 (Fig. 2D and F). Moreover, four “CATA” deletions for AtWRKY3 at same position in 8# plants were also observed, leading to a frameshift mutation (Fig. 2E). 19# plants had similar situation with a single “A” insertion for AtWRKY4 at same position (Fig. 2F). In general, these results indicated that targeting AtWRKY3 and AtWRKY4 by CRISPR/Cas9 system could efficiently bring to loss-of-function modifications.
CRISPR/Cas9-mediated the mutation circumstances of AtWRKY3 and AtWRKY4 genes in T2 generation of transgenic plants. A, B Identification of PCR products digested with restriction enzyme NdeI by gel electrophoresis. NdeI, PCR products digested with NdeI. The length of PCR products of NdeI digested are marked by arrow heads. The number on the lanes represents the different transgenic plant lines. Negative sign represents the no enzyme digestion, while plus sign enzyme digestion. M, DNA marker. C, D Comparisons and analyses of the PCR sequencing results of different transgenic plants. The gRNA sequence is labeled with light yellow and the PAM sequence is marked with a box. E, F Detailed mutation information of the AtWRKY3 and AtWRKY4 transgenic line (T2), respectively. DNA fragments around the target sequences were amplified by PCR and then subjected to sequencing analysis. The PAM and the sgRNA target sequences are indicated by blue and red lines, respectively. Green arrows indicated the insertion of an alanine (A) nucleotide. The arrow indicated the position of a deleted A or threonine (T) nucleotide. (Color figure online)
Expression analysis of AtWRKY genes under salt and Me-JA stresses
Expression patterns maybe offer clues to their functional difference among all members of a gene family [37]. To illustrate the role in responses to environmental stress, the expression patterns in A. thaliana seedlings were examined by qRT-PCR under salt and Me-JA conditions. AtWRKY3, -4, and -64 all showed significant increases after salt and Me-JA treatments and peaked at 24 h, while AtWRKY2, -8, -16, and -52 had no obvious changes (Fig. 3). Moreover, other AtWRKY genes exhibited up-regulated expression patterns only in a certain treatment. For example, AtWRKY28, -31 and -42 had high expression under Me-JA condition and no apparent changes in high salt treatment, while AtWRKY66 was just opposite with it (Fig. 3). Remarkably, AtWRKY3 was more rapidly induced by NaCl compared with Me-JA, while AtWRKY4 induced strongly by NaCl and Me-JA than AtWRKY3 (Fig. 3). These results indicated that AtWRKY3 and AtWRKY4 were involved in response to salt or Me-JA stresses, and might have distinct functions through different pathways.
The loss of AtWRKY3/WRKY4 decreases salt and Me-JA stress tolerance to transgenic plants
To further confirm the gene knockout to the effect on plant growth and development in transgenic plants, the expression of AtWRKY3 and AtWRKY4 were detected by qRT-PCR method. The result showed a very lower expression of AtWRKY3 in wrky3wrky4 double mutants than those of WT, while the expression of AtWRKY4 in mutants have different expression levels compared with WT plants (Fig. S3). The expression of AtWRKY3 in lines 2#, 11#, 19# showed no obvious change than WT, while lines 8#, 13# and 16# were significant decreased compared with WT (Fig. S3). From the comprehensive evaluation of the result mentioned above, we may speculate that the CRISPR-Cas9 technology can not only edit AtWRKY3 and AtWRKY4 genes at DNA level, but also the change of transcript levels was resulted from the action CRISPR-Cas9 system.
Their mutation situations, expression levels and stress response patterns were comprehensively taken into accounts, the lines 8#, 13#, and 16#, were ultimately selected for further phenotypic analyses under salt and Me-JA stresses. To analyze the function of AtWRKY3 and AtWRKY4 in salt stress, nine-day-old T2 transgenic plants and WT plants were transplanted to 1/2 MS medium supplemented with 50 mM NaCl and 100 mM NaCl, respectively. After two days, photographs were taken and recorded. The growth of WT and double mutants showed significant difference, especially their root lengths (Fig. 4A and B). As expected, the root lengths of mutants in 50 mM NaCl were longer than those in 100 mM NaCl (Fig. 4A and B). However, the difference with double mutants, the single mutants had no obvious change under similar condition (Fig. S4A and B). Moreover, WT, 8#, 13# and 16# plants all displayed similar POD and CAT activities before salt stress treatment. However, the activity of SOD in Atwrky3wrky4 plants was lower than that in WT, indicating that WRKY3 and WRKY4 as TFs can positively regulate transcription and enzyme activity of SOD (Fig. 4C–E). Furthermore, the decreases of SOD, POD and CAT activity in Atwrky3wrky4 plants were evident at two days after salt stress treatment as compared to the corresponding WT (Fig. 4C–E). It was noted that the activity of POD in Atwrky3wrky4 8# plants had no obvious difference as compared to that in WT, while 13# plant had significant decreases under salt stresses including 50 and 100 mM NaCl treatment (Fig. 4D). In addition, there was no significant difference in POD and CAT activities under normal conditions between WT and Atwrky3wrky4 plants, while their activities were significantly changed in 50 mM and 100 mM NaCl treatments (Fig. 4D and E). As expected, the Atwrky3wrky4 plants showed higher REL contents than WT plants after salt treatments (Fig. 4F). However, there were no obvious difference in single mutants of AtWRKY3 or AtWRKY4 under salt stress, including SOD, POD, CAT activity, and REL contents (Fig. S4C–F). In general, these data suggested that the loss of AtWRKY3 and AtWRKY4 might be attenuate the salt stress tolerance via cellular oxidative damage.
Decreased salt tolerance in AtWRKY3 and AtWRKY4 knockout plants. A Phenotypic characteristics of wrky3/wrky4 and WT seedlings at different concentration during the salt stress experiments. Growth of the A. thaliana plant on 1/2 MS medium (control) and 1/2 MS medium supplied with NaCl. B The root lengths were measured at eleven days after germination (i.e. after treatment with NaCl two days). C–F SOD, POD, CAT activity, and REL of the mutant and WT seedlings under salt stress treatment, respectively. Control represents a normal growth condition. The error bars indicate the standard deviation from triplicate experiments. *indicate significant difference (p ≤ 0.05) compared with WT plants with no treatment. WT, wild type; 8#, wrky3/wrky4-8; 13#, wrky3/wrky4-13; 16#, wrky3/wrky4-16; SOD, superoxide dismutase; POD, peroxidase; CAT, catalase; REL, relative electrolyte leakage
In addition, we next analyzed the function of AtWRKY3 and AtWRKY4 using nine-day-old transgenic plants and WT plants with 100 or 300 μM Me-JA stresses, respectively. Then photographs were taken and root lengths were recorded after two days treatments. Compared with control plants, the growth and development of mutants and WT plants displayed striking difference, especially the root lengths (Fig. 5A and B). However, compared with double mutants, the single mutants had no obvious difference under 100 μM or 300 μM Me-JA stress (Fig. S5A and B). Moreover, we also investigated the activities of SOD, POD and CAT after Me-JA stress treatments. As similar with salt stress treatments, 8#, 13# and 16# plants all displayed the evident decrease of SOD, POD and CAT activity in Atwrky3wrky4 plants at two days after Me-JA stress treatment as compared to those in corresponding WT (Fig. 5C–E). Furthermore, the activity of POD and CAT have no obvious difference in normal condition between WT and Atwrky3wrky4 plants, while significant change in both 100 μM and 300 μM Me-JA treatments (Fig. 5D and E), suggesting that the loss of Atwrky3wrky4 led to the decrease of antioxidant enzymes activities in plants in response to Me-JA stress. Likewise, the double plants exhibited higher REL contents than the corresponding WT plants, indicating that the loss of AtWRKY3/4 decreased the Me-JA tolerance (Fig. 5F). However, the SOD, POD, CAT activity, and REL contents had no significant change in single mutants of AtWRKY3 or AtWRKY4 under Me-JA stress (Fig. S5C–F). Thus, there results suggested that silencing of WRKY3 and WRKY4 decreased the Arabidopsis to Me-JA stress tolerance.
Decreased of Me-JA tolerance in AtWRKY3 and AtWRKY4 knockout plants. A Phenotypic characteristics of wrky3/wrky4 and WT seedlings at different concentration during the Me-JA stress experiments. Growth of the A. thaliana plant on 1/2 MS medium (control) and 1/2 MS medium supplied with Me-JA. B The root lengths were calculated from 25 seedlings studied at eleven days after germination (i.e. after treatment with Me-JA two days). C–F Analyses of SOD, POD, CAT activities, and REL contents, respectively. Control represents a normal growth condition. All experiments were repeated three times and an asterisk above the columns indicated significant differences at p ≤ 0.05 levels with corresponding WT with no Me-JA treatment. WT, wild type; 8#, wrky3/wrky4-8; 13#, wrky3/wrky4-13; 16#, wrky3/wrky4-16; SOD, superoxide dismutase; POD, peroxidase; CAT, catalase; REL, relative electrolyte leakage
Furthermore, we next examined the expression of SOD, POD and CAT genes in WT and mutant plants. A total of nine genes were selected for analysis including three SOD genes (At3g10920, At5g18100 and At2g28190), three POD genes (At2g18140, At5g66390 and At5g58400) and three CAT genes (At1g20630, At4g35090 and At1g20620), which have been confirmed previously to be associated with SOD, POD or CAT activity. Under salt or Me-JA stress treatment, the transcripts of all SODs and PODs showed a decreased significantly levels in mutant plants, while the expression levels of CATs had no apparent changes in mutants even moderately increase (Fig. 6). These results suggested that these SODs and PODs are induced by AtWRKY3 and/or AtWRKY4 to improve SOD and POD activity in plants, while CATs may have other pathway to response to salt or Me-JA stress treatment. Taken together, these results indicated that the loss of AtWRKY3 and AtWRKY4 all attenuated the activities of SOD, POD and CAT in plants exposed to salt or Me-JA stress treatment. The CRISPR/Cas9 system can efficiently knockout AtWRKY3 and AtWRKY4 genes, these mutants obtained by this method help us preliminarily analyze the role of AtWRKY3 and AtWRKY4 under the salt and Me-JA stress.
Expression profiles of SODs, PODs and CATs. The plants were treated with different concentration of NaCl or Me-JA for 48 h, and the plants grown under normal conditions were harvested at the corresponding time points as controls. Data are means ± SD from three independent experiments. The asterisk indicates the fold changes of expression levels is greater 2 or less 0.5 than WT plants
Discussion
Acquisition of multiplex mutants by CRISPR/Cas9 system with conserved sequence of homologous genes as sgRNA
The identification of gene function based on mutants acquisition that mainly contains natural mutagenesis, physical or chemical mutation or random insertion of T-DNA [38]. These methods have obvious disadvantages, such as low efficiency, uncontrollable result, time-consuming and laborious identify. Therefore, the researcher developed a genome editing technique which introduces DNA mutations with insertions and/or deletions (indels) or base substitutions in target sequences. Until now, the CRISPR/Cas9 system has been the best choice for genome editing with high efficiency, accuracy, and ease of use compared with other technologies such as Zinc finger nucleases (ZFNs) system, transcription activator-like effector nucleases (TALENs) system, etc. [39, 40]. More importantly, we sometimes need to edit simultaneously multiple gene loci, such as mutation of multiple members of gene families, functionally redundant genes or related genes in the same biochemical pathway [21]. Therefore, we designed a single sgRNA with conserved sequences of homologous genes AtWRKY3 and AtWRKY4 as targets to direct Cas9 to specific corresponding sites (Fig. 1) and results in achieving simultaneously the gene editing AtWRKY3 and AtWRKY4 (Fig. 2). This situation is similar with from that reported for other genes. For example, three homoalleles that encode MILDEW-RESISTANCE LOCUS (MLO) proteins were yielded simultaneously mutations using CRISPR-Cas9 technology [41]. In addition, the expression of AtWRKY3 and AtWRKY4 genes was significant down-regulated (Fig. S3). These results indicated that AtWRKY3 and AtWRKY4 genes have been successfully knockout by CRISPR/Cas9 gene editing technology.
AtWRKY3 and AtWRKY4 genes response to abiotic stresses
The WRKY TFs constitute a quite large family with 74 members in A. thaliana and play important regulator roles in various plant developmental and physiological processes [2, 6]. For example, A. thaliana WRKY45 regulates plant tolerance to abiotic stress in soil through mediating the expression of a phosphated transporter [42]. Arabidopsis WRKY57 interacting with jasmonate zimdomain and auxin/indole-3-acetic acid proteins manages the balance between jasmonate and auxin signaling in leaf senescence [43]. A. thaliana WRKY12, -23, -33, and -44 characterize in regulating specialized metabolite production [44,45,46,47]. Similarly, AtWRKY25 plays an important role in the host defense to the bacterial pathogen Pseudomonas syringae [48]. AtWRKY75 involving in a tripartite amplification loop induced by SA and ROS accelerates leaf senescence [49]. However, AtWRKY54 and AtWRKY70 synergistically negatively regulate leaf senescence [50]. AtWRKY6 play vital roles in ABA signaling during seed germination and early seedling development by directly down-regulated RAV1 expression [51]. However, the biological function of most WRKY genes had not been functionally characterized so far. In the present study, we demonstrated that AtWRKY3 and AtWRKY4 play important roles in salt and Me-JA stress tolerance in A. thaliana. Our results displayed that the expression of AtWRKY3 and AtWRKY4 had significant increase under salt and Me-JA stress (Fig. 3). In addition, both WT and mutant plants all grew normally and had a normal phenotype, suggesting that knockout of AtWRKY3 and AtWRKY4 had no significant effect on plant growth or phenotype (Figs. 4 and 5). However, the wrky3/wrky4 double mutants all grew slowly and badly under salt and Me-JA stresses, especially the root lengths (Figs. 4A, B and 5A, B). This characteristic is similar to that reported by WRKY3 and WRKY4 in other plant species. For instance, ThWRKY4 can improve A. thaliana tolerance to salt and its development and growth, protect from the loss of chlorophyll, modulate ROS [52]. Hence, AtWRKY3 and AtWRKY4 play positive roles in regulating stress tolerance. Moreover, previous studies shown that over-expression of WRKY significantly enhanced SOD and POD activity and accumulated much more ROS scavenging enzyme [52, 53]. As expected, knockout of AtWRKY3 and AtWRKY4 plants displayed that the activities of SOD, POD and CAT were both lower than corresponding WT plants under stress treatments (Figs. 4C–E and 5C–E). The transcripts of the SOD and POD genes were positively correlated with the transcript levels of AtWRKY3 and/or AtWRKY4 (Fig. 6). Moreover, the double plants displayed higher REL contents than corresponding WT plants (Figs. 4F and 5F). Taken together, these results indicated that knockout of AtWRKY3 and AtWRKY4 attenuated SOD, POD and CAT activities, and increased the REL contents, resulting in obviously decreased stress tolerance functions.
Conclusion
Based on our results, we can speculate that AtWRKY3 and AtWRKY4 have no obvious influences to the plant development and growth and act as a positive regulator in response to salt and Me-JA stresses, according to the negative effect on SOD, POD and CAT activities, and increased REL contents, of their mutant plants under abiotic stresses compared with corresponding WT plants. Ultimately, the double mutants of Atwrky3wrky4 generated by CRISPR/Cas9 system showed decrease tolerance to salt and Me-JA stresses.
Abbreviations
- CRISPR:
-
Clustered regularly interspaced short palindromic repeat
- Cas9:
-
CRISPR-associated 9
- WT:
-
Wide type
- SOD:
-
Superoxide dismutase
- POD:
-
Peroxidase
- CAT:
-
Catalas
- REL:
-
Relative electrolyte leakage
- TFs:
-
Transcription factors
- AP2:
-
APETALA2
- ERF:
-
Ethylene-responsive factor
- NAM:
-
No apical meristem
- ATAF1/2:
-
Arabidopsis thaliana transcription activation factor
- CUC2:
-
Cup-shaped cotyledon
- DBD:
-
DNA binding domains
- SPF1:
-
SWEET POTATO FACTOR1
- SA:
-
Salicylic acid
- JA:
-
Jasmonic acid
- ABA:
-
Abscisic acid
- ROS:
-
Reactive oxygen species
- sgRNA:
-
Single guide RNA
- PAM:
-
Protospacer adjacent motif
- DSB:
-
Double-strand breaks
- Clo-0:
-
Columbia-0
- qRT-PCR:
-
Quantitative real-time PCR
- MS:
-
Murashige and Skoog
- PAM:
-
Protospacer adjacent motif
- ZFNs:
-
Zinc finger nucleases
- TALENs:
-
Transcription activator-like effector nucleases
- MLO:
-
MILDEW-RESISTANCE LOCUS
References
Jiang Y, Zeng B, Zhao H, Zhang M, Xie S, Lai J (2012) Genome-wide transcription factor gene prediction and their expressional tissue-specificities in maize. J Integr Plant Biol 54(9):616–630
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5(5):199–206
Hussain A, Li X, Weng Y, Liu Z, Ashraf MF, Noman A, Yang S, Ifnan M, Qiu S, Yang Y, Guan D, He S (2018) CaWRKY22 acts as a positive regulator in pepper response to Ralstonia solanacearum by constituting networks with CaWRKY6, CaWRKY27, CaWRKY40, and CaWRKY58. Int J Mol Sci 19(5):1426
Ishiguro S, Nakamura K (1994) Characterization of a cDNA encoding a novel DNA-binding protein, SPF1, that recognizes SP8 sequences in the 5’ upstream regions of genes coding for sporamin and beta-amylase from sweet potato. Mol Gen Genet 244(6):563–571
Jiang J, Ma S, Ye N, Jiang M, Cao J, Zhang J (2017) WRKY transcription factors in plant responses to stresses. J Integr Plant Biol 59(2):86–101
Wen F, Zhu H, Li P, Jiang M, Mao W, Ong C, Chu Z (2014) Genome-wide evolutionary characterization and expression analyses of WRKY family genes in Brachypodium distachyon. DNA Res 21(3):327–339
Chen L, Zhang L, Li D, Wang F, Yu D (2013) WRKY8 transcription factor functions in the TMV-cg defense response by mediating both abscisic acid and ethylene signaling in Arabidopsis. Proc Natl Acad Sci USA 110(21):E1963–E1971
Tripathi P, Rabara RC, Rushton PJ (2014) A systems biology perspective on the role of WRKY transcription factors in drought responses in plants. Planta 239(2):255–266
Niu C, Jiang M, Li N, Cao JG, Hou MF, Ni DA, Chu ZQ (2019) Integrated bioinformatics analysis of As, Au, Cd, Pb and Cu heavy metal responsive marker genes through Arabidopsis thaliana GEO datasets. Peer J 7(1):e6495
Qiu D, Xiao J, Ding X, Xiong M, Cai M, Cao Y, Li X, Xu C, Wang S (2007) OsWRKY13 mediates rice disease resistance by regulating defense-related genes in salicylate- and jasmonate-dependent signaling. Mol Plant Microbe Interact 20(5):492–499
Ullah A, Sun H, Hakim YX, Zhang X (2018) A novel cotton WRKY gene, GhWRKY6-like, improves salt tolerance by activating the ABA signaling pathway and scavenging of reactive oxygen species. Physiol Plant 162(4):439–454
Tang Y, Kuang JF, Wang FY, Chen L, Hong KQ, Xiao YY, Xie H, Lu WJ, Chen JY (2013) Molecular characterization of PR and WRKY genes during SA- and MeJA-induced resistance against Colletotrichum musae in banana fruit. Postharvest Biol Technol 79:62–68
Li GZ, Wang ZQ, Yokosho K, Ding B, Fan W, Gong QQ, Li GX, Wu YR, Yang JL, Ma JF, Zheng SJ (2018) Transcription factor WRKY22 promotes aluminum tolerance via activation of OsFRDL4 expression and enhancement of citrate secretion in rice (Oryza sativa). New Phytol 219(1):149–162
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15(5):247–258
Skibbe M, Qu N, Galis I, Baldwin IT (2008) Induced plant defenses in the natural environment: Nicotiana attenuata WRKY3 and WRKY6 coordinate responses to herbivory. Plant Cell 20(7):1984–2000
Kiranmai K, Rao GL, Pandurangaiah M, Nareshkumar A, Reddy VA, Lokesh U, Venkatesh B, Johnson AMA, Sudhakar C (2018) A novel WRKY transcription factor, MuWRKY3 (Macrotyloma uniflorum Lam. Verdc.) enhances drought stress tolerance in transgenic groundnut (Arachis hypogaea L.) plants. Front Plant Sci 9(1):346
Lai Z, Vinod K, Zheng Z, Fan B, Chen Z (2008) Roles of Arabidopsis WRKY3 and WRKY4 transcription factors in plant responses to pathogens. BMC Plant Biol 8:68
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157(6):1262–1278
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823
Ma XL, Zhang QY, Zhu QL, Liu W, Chen Y, Qiu R, Wang B, Yang ZF, Li HY, Lin YR, Xie YY, Shen RX, Chen SF, Wang Z, Chen YL, Guo JX, Chen LT, Zhao XC, Dong ZC, Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8(8):1274–1284
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167(2):313–324
Chaves-Sanjuan A, Sanchez-Barrena MJ, Gonzalez-Rubio JM, Moreno M, Ragel P, Jimenez M, Pardo JM, Martinez-Ripoll M, Quintero FJ, Albert A (2014) Structural basis of the regulatory mechanism of the plant CIPK family of protein kinases controlling ion homeostasis and abiotic stress. Proc Natl Acad Sci USA 111(42):E4532–E4541
Jiang M, Zhao CL, Zhao MF, Li YZ, Wen GS (2020) Phylogeny and evolution of calcineurin B-like (CBL) gene family in grass and functional analyses of rice CBLs. J Plant Biol 63(2):117–130
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Adie BAT, Perez-Perez J, Perez-Perez MM, Godoy M, Sanchez-Serrano JJ, Schmelz EA, Solano R (2007) ABA is an essential signal for plant resistance to pathogens affecting JA biosynthesis and the activation of defenses in Arabidopsis. Plant Cell 19(5):1665–1681
Pieterse CMJ, Leon-Reyes A, Van der Ent S, Van Wees SCM (2009) Networking by small-molecule hormones in plant immunity. Nat Chem Biol 5(5):308–316
Gao QM, Venugopal S, Navarre D, Kachroo A (2011) Low oleic acid-derived repression of jasmonic acid-inducible defense responses requires the WRKY50 and WRKY51 proteins. Plant Physiol 155(1):464–476
Seltmann MA, Stingl NE, Lautenschlaeger JK, Krischke M, Mueller MJ, Berger S (2010) Differential impact of lipoxygenase 2 and jasmonates on natural and stress-induced senescence in Arabidopsis. Plant Physiol 152(4):1940–1950
De Geyter N, Gholami A, Goormachtig S, Goossens A (2012) Transcriptional machineries in jasmonate-elicited plant secondary metabolism. Trends Plant Sci 17(6):349–359
Radhika V, Kost C, Boland W, Heil M (2010) Towards elucidating the differential regulation of floral and extrafloral nectar secretion. Plant Signal Behav 5(7):924–926
Yan L, Wei S, Wu Y, Hu R, Li H, Yang W, Xie Q (2015) High-efficiency genome editing in Arabidopsis using YAO promoter-driven CRISPR/Cas9 system. Mol Plant 8(12):1820–1823
Lamesch P, Berardini TZ, Li DH, Swarbreck D, Wilks C, Sasidharan R, Muller R, Dreher K, Alexander DL, Garcia-Hernandez M, Karthikeyan AS, Lee CH, Nelson WD, Ploetz L, Singh S, Wensel A, Huala E (2012) The Arabidopsis information resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40(D1):D1202–D1210
Wang Y, Gao C, Liang Y, Wang C, Yang C, Liu G (2010) A novel bZIP gene from Tamarix hispida mediates physiological responses to salt stress in tobacco plants. J Plant Physiol 167(3):222–230
Mittova V, Volokita M, Guy M, Tal M (2000) Activities of SOD and the ascorbate-glutathione cycle enzymes in subcellular compartments in leaves and roots of the cultivated tomato and its wild salt-tolerant relative Lycopersicon pennellii. Physiol Plant 110(1):42–51
Li JB, Luan YS, Liu Z (2015) SpWRKY1 mediates resistance to Phytophthora infestans and tolerance to salt and drought stress by modulating reactive oxygen species homeostasis and expression of defense-related genes in tomato. Plant Cell Tissue Organ Cult 123(1):67–81
Whittle CA, Krochko JE (2009) Transcript profiling provides evidence of functional divergence and expression networks among ribosomal protein gene paralogs in Brassica napus. Plant Cell 21(8):2203–2219
Zhang Y, Ma X, Xie X, Liu YG (2017) CRISPR/Cas9-based genome editing in plants. Prog Mol Biol Transl Sci 149:133–150
Mao YF, Botella JR, Liu YG, Zhu JK (2019) Gene editing in plants: progress and challenges. Natl Sci Rev 6(3):421–437
Gaj T, Gersbach CA, Barbas CF (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31(7):397–405
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32(9):947–951
Wang H, Xu Q, Kong YH, Chen Y, Duan JY, Wu WH, Chen YF (2014) Arabidopsis WRKY45 transcription factor activates PHOSPHATE TRANSPORTER1;1 expression in response to phosphate starvation. Plant Physiol 164(4):2020–2029
Jiang Y, Liang G, Yang S, Yu D (2014) Arabidopsis WRKY57 functions as a node of convergence for jasmonic acid- and auxin-mediated signaling in jasmonic acid-induced leaf senescence. Plant Cell 26(1):230–245
Grunewald W, De Smet I, Lewis DR, Lofke C, Jansen L, Goeminne G, Vanden Bossche R, Karimi M, De Rybel B, Vanholme B, Teichmann T, Boerjan W, Van Montagu MC, Gheysen G, Muday GK, Friml J, Beeckman T (2012) Transcription factor WRKY23 assists auxin distribution patterns during Arabidopsis root development through local control on flavonol biosynthesis. Proc Natl Acad Sci USA 109(5):1554–1559
Mao G, Meng X, Liu Y, Zheng Z, Chen Z, Zhang S (2011) Phosphorylation of a WRKY transcription factor by two pathogen-responsive MAPKs drives phytoalexin biosynthesis in Arabidopsis. Plant Cell 23(4):1639–1653
Wang H, Avci U, Nakashima J, Hahn MG, Chen F, Dixon RA (2010) Mutation of WRKY transcription factors initiates pith secondary wall formation and increases stem biomass in dicotyledonous plants. Proc Natl Acad Sci USA 107(51):22338–22343
Johnson CS, Kolevski B, Smyth DR (2002) TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 14(6):1359–1375
Zheng Z, Mosher SL, Fan B, Klessig DF, Chen Z (2007) Functional analysis of Arabidopsis WRKY25 transcription factor in plant defense against Pseudomonas syringae. BMC Plant Biol 7:2
Guo P, Li Z, Huang P, Li B, Fang S, Chu J, Guo H (2017) A tripartite amplification loop involving the transcription factor WRKY75, salicylic acid, and reactive oxygen species accelerates leaf senescence. Plant Cell 29(11):2854–2870
Besseau S, Li J, Palva ET (2012) WRKY54 and WRKY70 co-operate as negative regulators of leaf senescence in Arabidopsis thaliana. J Exp Bot 63(7):2667–2679
Huang Y, Feng CZ, Ye Q, Wu WH, Chen YF (2016) Arabidopsis WRKY6 transcription factor acts as a positive regulator of abscisic acid signaling during seed germination and early seedling development. Plos Genet 12(2):e1005833
Zheng L, Liu G, Meng X, Liu Y, Ji X, Li Y, Nie X, Wang Y (2013) A WRKY gene from Tamarix hispida, ThWRKY4, mediates abiotic stress responses by modulating reactive oxygen species and expression of stress-responsive genes. Plant Mol Biol 82(4–5):303–320
Wang Z, Feng R, Zhang X, Su Z, Wei J, Liu J (2019) Characterization of the Hippophae rhamnoides WRKY gene family and functional analysis of the role of the HrWRKY21 gene in resistance to abiotic stresses. Genome 62(10):689–703
Acknowledgements
We are grateful to Dr. Qi Xie (Institute of Genetics and Developmental Biology, VAS) for providing us with the vector pYAO:hSpCas9-target-sgRNA.
Funding
This research was supported by the Shanghai Sailing Program (19YF1414800). The funding body had no role in study design, analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
MJ conceived, designed and funded the research. MJ, PL and XL performed the experiments, analyzed the data and wrote the manuscript. MJ revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human and/or animal participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
All authors consent to publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11033_2021_6541_MOESM2_ESM.tif
Supplementary file2 CRISPR/Cas9-mediated the mutations of AtWRKY3 and AtWRKY4 genes in T1 generation of different transgenic plant lines. (A-B) Identification of PCR products digested with restriction enzyme NdeI by gel electrophoresis. NdeI, PCR products digested with NdeI. The length of PCR products of NdeI digested are marked by red arrow heads. The number on the lanes represents the different transgenic plant lines. e Sign “N” represents the no enzyme digestion, while other lanes represents enzyme digestion. M, DNA marker. (C-D) Comparisons and analyses of the sequencing results. The gRNA sequence is labeled with light yellow and the PAM sequence is marked with a box. (E-F) Detailed mutation information of the AtWRKY3 and AtWRKY4 transgenic line (T1), respectively. DNA fragments around the target sequences were amplified by PCR and then subjected to sequencing analysis. The PAM and the sgRNA target sequences are indicated by blue and red lines, respectively. Green arrows indicated the insertion of an alanine (A) nucleotide. The blue arrow indicated the position of a deleted A or threonine (T) nucleotide (TIF 1808 kb)
11033_2021_6541_MOESM3_ESM.jpg
Supplementary file3 CRISPR/Cas9-mediated the single mutations of AtWRKY3 (A) or AtWRKY4 (B) gene in T2 generation of different transgenic plant lines. The number plus pound sign represents the different transgenic plant lines (JPG 130 kb)
11033_2021_6541_MOESM4_ESM.tif
Supplementary file4 Relative expression levels of AtWRKY3 and AtWRKY4 in mutants by qRT-PCR. The asterisk indicates the fold changes of expression levels is less 0.5 than WT plants (TIF 3041 kb)
11033_2021_6541_MOESM5_ESM.jpg
Supplementary file5 Physiological indies measurement under salt stress. (A) Phenotypes of wrky3 plants under salt stress. (B) Root lengths. (C) SOD activity. (D) POD activity. (E) CAT activity. (F) REL contents. Control represents a normal growth condition. The number plus pound sign represents the different transgenic plant lines. WT, wild type; 24#, wrky3-24; 25#, wrky3-25; 28#, wrky3-28; 23#, wrky4-23; 32#, wrky4-32; 35#, wrky4-35; SOD, superoxide dismutase; POD, peroxidase; CAT, catalase; REL, relative electrolyte leakage. Mean and S.D. values were obtained from three independent experiments (JPG 127 kb)
11033_2021_6541_MOESM6_ESM.jpg
Supplementary file6 Physiological indies measurement under Me-JA stress. (A) Phenotypes of wrky4 plants under Me-JA stress. (B) Root lengths. (C) SOD activity. (D) POD activity. (E) CAT activity. (F) REL contents. Control represents a normal growth condition. The number plus pound sign represents the different transgenic plant lines. WT, wild type; 24#, wrky3-24; 25#, wrky3-25; 28#, wrky3-28; 23#, wrky4-23; 32#, wrky4-32; 35#, wrky4-35; SOD, superoxide dismutase; POD, peroxidase; CAT, catalase; REL, relative electrolyte leakage. Mean and S.D. values were obtained from three independent experiments (JPG 129 kb)
Rights and permissions
About this article
Cite this article
Li, P., Li, X. & Jiang, M. CRISPR/Cas9-mediated mutagenesis of WRKY3 and WRKY4 function decreases salt and Me-JA stress tolerance in Arabidopsis thaliana. Mol Biol Rep 48, 5821–5832 (2021). https://doi.org/10.1007/s11033-021-06541-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06541-4