Abstract
Calcium signals play critical functions in regulating diverse arrays of plant growth and development and mediating a variety of biotic and abiotic stress responses as a second messenger. Calcineurin B-like (CBL) proteins were involved with plant-specific Ca2+signaling as calcium sensors. In this work, we retrieved 152 CBL gene members from 15 different grass species, surveyed their phylogenetic relationships and sequence features and also performed expression patterns and functional analyses of rice CBLs. Phylogenetic analysis indicated that grass CBLs fall into four different groups (Group A–D). Sequence analysis showed that CBL proteins harboring four conserved calcium-binding EF-hand have key amino acid residues Asp and Glu which had relatively high proportion in the average abundance. Molecular evolutionary analyses revealed that group A, B and C CBLs in their evolution process suffered the purifying selection, while group D CBLs were subjected to positive selection. Moreover, expression analyses showed significant divergent expression patterns of OsCBLs in various organs and under different hormones and abiotic stresses. Furthermore, tolerance analysis revealed that OsCBL3 and OsCBL8 overexpression transgenic rice seedlings improved salt tolerance and OsCBL5, OsCBL6 and OsCBL7 positively regulated drought stress. In general, the domain and base sequence of the CBL gene family is highly conserved in grasses. OsCBL genes had specific gene expression profiles and function in different stresses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Plants must sense and respond to various perturbations or stresses by properly altering their cellular and physiological status to maintain normal growth and development in such adverse environments. As is well known, this adaptation process is associated with two critical cellular components, calcium ion (Ca2+) and Ca2+-binding proteins. This external stimulus is transduced into a ubiquitous and distinct Ca2+ signature through regulating to specific temporal and spatial changes in cytosolic calcium concentration [(Ca2+)cyt] in plant cells (Evans et al. 2001; Sanders et al. 2002). Although the Ca2+ signature plays a vital role in stimulus–response coupling, Ca2+-binding proteins decode and relay the information encoded by the Ca2+ signature, which serves as a Ca2+ sensor, and perceives the specific Ca2+ signature and transmits these signals to downstream pathways to response-specific stress (Luan et al. 2002; Weinl and Kudla 2009).
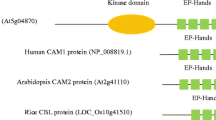
The Ca2+ sensor harboring the canonical elongation factor hand (EF-hand) Ca2+-binding motifs can be classified into four families: calmodulin (CaM), CaM-like proteins (CMLs), calcineurin B-like (CBL) proteins and Ca2+-dependent protein kinases (CDPKs)(Luan 2009; McCormack et al. 2005; Schulz et al. 2013). Among these, CDPKs possess a kinase domain modulated by an EF-hand domain, thereby forming ‘sensor-response’ modules (Harper and Harmon 2005; Sanders et al. 2002). By contrast, the rest of the family and their interacting kinases are divided into two modules: Ca2+-binding function and kinase activity (Burstenbinder et al. 2017). Therefore, CBL proteins that were activated by binding the calcium signature can physically interact with the NAF (Asn-Ala-Phe) domain of the CBL-interacting protein kinases (CIPKs) (Chaves-Sanjuan et al. 2014). As a calcium sensor protein for Ca2+ binding, CBL in plants are similar to neuronal calcium sensors and calcineurin B (CNB) from animals (Kudla et al. 1999), harboring four EF-hand domains that are characterized by a conserved Asp (D) and Glu (E) residue with a completely constant spacing (Batistic and Kudla 2009; Kolukisaoglu et al. 2004). In addition, the short N-terminal of CBL proteins (CBL1, − 4, − 5, − 8 and − 9) contain conserved MGCXXS/T motifs that contributed to the CBLs to anchor in the membrane to transduce Ca2+ signal after lipid modification of myristoylation (Batistic et al. 2008; Weinl and Kudla 2009). While the extended N-terminus of CBL proteins (vacuolar-targeted CBL2 and -6) had another recognizable lipid modification motif (S-acylation) (Batistic et al. 2012; Zhang et al. 2017). Moreover, the C-terminal of CBL proteins harbor a conserved FPSF domain (because of the presence of conserved P, M, L, F, P and F residues) that are the target of phosphorylation by CIPK, which might generate additional specificity to the CBL–CIPK interaction (Mohanta et al. 2015; Sanyal et al. 2015).
To date, the CBL protein was first identified in Arabidopsis thaliana (Kolukisaoglu et al. 2004), which was followed by identification in Populus (Zhang et al. 2008), rice (Gu et al. 2008), cotton (Lu et al. 2017), grapevine (Xi et al. 2017), canola (Zhang et al. 2014), eggplant (Li et al. 2016), Physcomitrella patens (Weinl and Kudla 2009) and Selaginella moellendorffii (Weinl and Kudla 2009). The ten CBLs encoded in the Arabidopsis and rice genome whose four introns are absolutely conserved in phase and position (Kolukisaoglu et al. 2004). CBL1 mediates cold and low K+ stresses response via CIPK-coupled signaling in Arabidopsis (Huang et al. 2011; Li et al. 2006). Moreover, CBL1 also contributes to rescue the salt and osmotic stress in Sedirea japonica (Cho et al. 2018). CBL1 and CBL9 play an important role in regulating K+ homeostasis in root and stomata through CIPK23 (Cheong et al. 2007) and also affects pollen development (Mahs et al. 2013). It is noteworthy that CBL10 modulates K+ homeostasis by directly interacting with Arabidopsis K+ transporter 1 (AKT1) (Ren et al. 2013). In addition, OsCBL10 confers flooding tolerance during seed germination with enhanced Ca2+ flow (Ye et al. 2018). CBL1 and CBL4 specifically mediate salt stress signaling and membrane H+ transport (Liu and Zhu 1998). CBL2 and CBL3 not only controls ion homeostasis (Tang et al. 2012), but also modulates plant seed size and embryonic development (Eckert et al. 2014). CBL5 can increase plant’s drought stress tolerance (Cheong et al., 2010) and CBL7 can affect plant responses to low nitrate in Arabidopsis (Ma et al. 2015).
The grass family consists of large and nearly ubiquitous monocotyledonous plants that can be used as forage, building materials, fuel, and food. A great deal of effort has been made to investigate the role of the CBL gene family in some plant species, but a more detailed evolution and phylogeny of CBL protein in monocot, including grasses, has not been described yet. Here, we investigated the phylogenetic relationships and adaptive evolution of the 152 CBL genes from 15 grass species at the genome-wide level. Additionally, the expression patterns and the function in response to salt and drought stresses of these genes are examined in rice.
Materials and Methods
Identification of CBL Gene Family Members in Grasses
The ten CBL gene information from the model plant Oryza sativa were searched from the TIGR rice Genome Annotation Resources database (https://rice.plantbiology.msu.edu/) (Ouyang et al. 2007). To identify CBL genes in other grass species, BLASTP searches were performed against orthologous protein sequences using rice CBLs as independent probes (Altschul et al. 1997) in phytozome (https://www.phytozome.net/) (Goodstein et al. 2012) and public PLAZA (https://bioinformatics.psb.ugent.be/plaza/) (Van Bel et al. 2018). The CBL gene was only considered as a candidate when it harbored four EF-hand domains, which was subjected to domain analyses by scanning in InterPro software (de Castro et al. 2006). Moreover, all information was examined for redundancy and no alternative splice variants were selected. All retrieved non-redundant coding and genomic sequences were collected in 15 grass species.
Sequence Alignment, Protein Motif, Phylogenetic Tree and Molecular Evolution Analyses
Multiple sequence alignments of the protein or DNA sequences of all CBL genes were performed by the Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/) with default settings. The alignment logos of the protein conserved domain were viewed by WebLogo (https://weblogo.threeplusone.com/). The domain analyses were executed using InterProScan against the protein database (https://www.ebi.ac.uk/interpro/). The theoretical pI (isoelectric point) and Mw (molecular weight) of CBLs were measured using Compute pI/Mw tool online (https://web.expasy.org/compute_pi/). Phylogenetic analyses were performed by the maximum-likelihood (ML) method with a Jones–Taylor–Thornton (JTT) model and the 2000 iterations bootstrap test in the MEGA 6.0 software (Tamura et al. 2013). To insure the topology of the ML tree with more divergent domains, all positions with 95% site coverage were eliminated. The aligned genomic sequences were applied to assess the value of Ka and Ks as well as their ratios by the DNASP v5.10 software (Librado and Rozas 2009). To study the genetic divergence between each group, we counted the genetic distances based on the amino acid sequences with the JTT model in the MEGA 6.0 (Tamura et al. 2013). Their overall mean distances were also measured according to the related MEGA file of phylogenetic tree of grass CBLs.
Plant Growth, Treatments and Expression Analysis
Gene expression analyses of rice cultivar Zhonghua 11 (ZH11) were investigated in different tissues and response to abiotic stresses and hormone treatments. It is important that high-frequency genetic transformation is established for the generation of transgenic lines and phenotype analyses in our laboratory. Pre-germinated seeds were placed in water for 2 days that were then transferred into a soil mix. All rice plants have a growth condition with a cycle of 26 °C and 14 h light (> 3000 lx)/22 °C and 10 h dark. For real-time quantitative PCR (RT-qPCR) analyses, 2-week-old rice plants were used for harvesting root, shoot, leaf blade, leaf sheath, shoot apical meristem (SAM) at vegetative or reproductive stage and nine different flower stages. For phytohormone analysis, 2-week-old seedlings were treated in MS liquid medium containing 50 μM TIBA, 100 µM SA, 50 µM IAA, 100 µM ABA and 50 µM GA for 0.5 h, 1 h, 3 h, 6 h, 12 h and 24 h, respectively. For abiotic stress treatment, 2-week-old seedlings were treated in MS liquid medium containing 50 µM CuSO4, 25 µM K2Cr2O7, 20% PEG 6000 and 300 mM NaCl for 0.5 h, 1 h, 3 h, 6 h, 12 h and 24 h, respectively. In addition, seedlings were transferred to a growth chamber at 4 °C for cold treatment. Untreated plants were used as control samples. The seedlings were sampled at different time points after treatment and stored at − 80 °C until use. Total RNA was extracted using TRIzol reagent and reverse-transcribed into cDNA using PrimeScript RT Master Mix Perfect Real Time (TaKaRa) followed the manufacturer’s instructions. The total RNA was quantified using Nanodrop1000 and its integrity was assessed by electrophoresis in 1.5% (w/v) agarose gel. The RT-qPCR was performed in a 10 µl reaction with 5–50 ng of first-stand cDNA products (4 µl), 5 pmol of each primer (0.4 µl), 5 μl SYBR Green Master Mix (2X) and 0.2 µl ROX as a passive reference standard to normalize the SYBR fluorescent signal. The thermal profile for RT-qPCR was: initial activation at 95 °C for 10 min, followed by 40 cycles of 95 °C for 30 s, 60 °C for 30 s, and 72 °C for 30 s. The rice actin gene was used as an internal reference for all RT-qPCR analyses. Each treatment was conducted in triplicate independently using three biological replicates. The relative expression of OsCBL genes was calculated using the 2−ΔΔCt method. The primer sets are listed in Table S1.
Binary Vector Construction, Rice Transformation and Characterization of the Transgenic Lines
To make OsCBLs constitutive overexpression, the modified pCAMBIA1301 vector P1301-UbiNos under the control of the maize ubiquitin promoter was constructed by inserting the coding sequence of OsCBLs which was amplified with primers of OsCBLsOE-F and OsCBLsOE-R (Table S2). Screening of transgenic plants was attained by planting on 1/2 Murashige and Skoog (MS) medium containing 25 µg/L hygromycin. The presence of the transgenic insertion was confirmed through PCR in the transgenic plants. Moreover, seeds were collected and sown for phenotyping from T1 transgenic rice plants. More than 20 independent plant lines from each transformation were picked for morphologic observation, and ZH11 was used as control.
Phenotype Analyses for Drought and Salt Tolerance
For drought stress assay, 2-week-old OsCBLs overexpression (OsCBLs-OE) transgenic plants and wild-type (WT) plants located in same growth condition were subjected to drought stress by 20% PEG 6000 for 10 days. The length of of the shoot and root of at least 20 seedlings of each line were measured at 10 days after treatment, respectively. Likewise, 2-week-old transgenic and non-transgenic plants grown in the same environment were transferred into 1/2 MS containing 300 mM NaCl. At 10 days after treatment, the length of the root and shoot were measured, respectively.
Results
Identification of CBL Protein Sequences in 15 Grass Species Based on the Conserved EF-Hand Domain
To survey the evolution of CBL protein architecture across grass plants, 15 publicly available genomes of Poales were selected for CBL gene family identification analyses (Table 1). We found that the numbers of CBLs varied from species to species across the grass plants. Finally, 152 non-redundant CBL sequences containing the conserved EF-hand domain were retrieved in total (Table 1, Table S3). Individual CBLs were assigned names using the ortholog based on the evolutionary relationship with ten Arabidopsis CBLs. The hexaploid wheat genome contained the largest number CBL genes, whereas Zoysia japonica only contained six, but most of them were around ten (Table 1, Table S3).
Phylogenetic Relationships of Grass CBL Genes
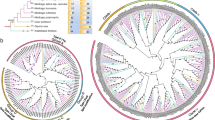
To investigate the evolution history among grass CBL gene families, the ML phylogenetic tree was reconstructed based on the full-length amino acid of the 152 CBL protein sequences (Fig. 1). In the phylogenetic tree, the grass CBLs can be divided into four main clades and designated as group A, B, C and D, respectively (Fig. 1, Table S3), which is in accordance with the Arabidopsis and rice reported CBLs (Albrecht et al. 2001; Kolukisaoglu et al. 2004). The groups A, B, C and D contain CBL9/10, CBL2/3/6, CBL1 and CBL4/5/7/8, respectively (Fig. 1). In addition, to assess the difference between whole sequences and functional motifs, we also reconstructed the phylogenetic tree using the EF-hand motif sequences (Fig. S1). As expected, the phylogenetic tree using EF-hand motifs of CBL protein displayed similar phylogenetic relationship compared with whole sequences, indicating that the evolutionary relationship of the CBL proteins was mainly determined by functional motifs, especially their characteristic EF-hand motif.
Common Conserved Domain Compositions and Genomic Analysis of CBL Proteins in Grass
To better obtain an overview of the characteristics among grass CBLs, their common conserved domains were further analyzed. As expected, three main common motifs of CBL proteins were subjected to the domain analyses using InterProScan database, suggestive of functional similarities within gene family (Fig. 2). As calcium sensors, CBL proteins harbor four typical EF-hands (IPR002048) as a structural basis for calcium binding (Albrecht et al. 2001) (Fig. 2a). Generally, each EF-hand is composed of 12 amino acid residues, in which 6 amino acids at positions 1(X), 3(Y), 5(Z), 7(− X), 9(− Y) and 12(-Z) are highly conserved and involved in binding of calcium ion (Fig. 2b). It is notable that the EF1 loop contains 14 amino acid residues caused by an insertion between position 1 and 3, and AtCBL6 lacked four amino acids and AtCBL7 had an insertion of five additional amino acids (Kolukisaoglu et al. 2004). Surprisingly, CBL4 can bind up to four Ca2+ ions; however, only two EF-hands bind Ca2+ in CBL4-CIPK24 complex (Fig. 2a) (Sanchez-Barrena et al. 2013). Interestingly, some N-terminus of CBL proteins possessed a highly conserved N-myristoylation motif that favored the attachment of CBL to the plasma membrane and transmitted the calcium signal, while the second glycine residue of the consensus sequence MGCXXSK/T is involved in protein myristoylation (Fig. 2a, c). Moreover, all CBLs similarly harbor a highly conserved C-FPSF domain (Fig. 2d).
Analyses of motif structures and compositions of all grass CBLs identified in this study. a Schematic depiction of the typical domain structure of grass CBLs. The overall structure of CBLs consists of four EF-hands (boxes with numbers), which the first and fourth bind two Ca2+ ions. In all CBL proteins, the first EF-hand has an unconventional structure, encompassing 14 amino acids instead of the 12 in a canonical EF hand (black box) flanked by two alpha helices and the amino acids at position 1(X), 3(Y), 5(Z), 7(− X), 9(− Y) and 12(− Z) of the loop co-ordinate and bind the Ca2+ ion. The position of lipid modification sites (N-myristoylation and S-acylation) and the C-terminal FPSF domain was shown by jagged lines and red box, respectively. b Sequence features shown in the form of web logos representing the EF hands of all CBL sequences. Spacing of EF hands in all CBLs is invariable. c Detailed comparisons of N-myristoylation motif sequences of grass CBLs. d Detailed comparisons of C-terminal FPSF domain sequences of grass CBLs
In addition, we also investigated the exon/intron organizations of different CBL genes according to the genomic information received. The majority of CBL genes contain seven introns occupying the main points (55%), which mean that the exon/intron organizations of grass CBL genes have a conserved model in their evolutionary process, while only one CBL gene contains a maximum of 20 introns (PeCBL1-4) or a minimum of being intronless (HvCBL6-2) (Table S3). Group A CBLs possess introns ranging from 5 (HvCBL10) to 11 (AcCBL9) and dominated by 8 (64%), while the numbers of introns in group B CBLs ranged from 0 (HvCBL6-2) to 9 (AcCBL6) and 7 occupied the main points (61%; Table S3). Remarkably, most groups C and D CBLs contain seven introns (Table S3). Afterward, we further determined the Mw and pI of different CBL proteins using the online version of Compute pI/Mw tool. The Mw of CBL proteins ranged from 12.3 (SiCBL7) to 80.115 (PeCBL1-4) kDa and the pI varied from 4.26 (SiCBL7) to 10.13 (SiCBL1) (Table S3). It is noteworthy that the pI of other CBLs removed SiCBL1were in the slightly acidic ranges. The average amino acid composition of CBL proteins ranged from 0.53 (tryptophan) to 10.83 (leucine) (Table S4). Remarkably, the average abundance of the most important amino acids aspartic and glutamic (EF-hand) which were responsible for the Ca2+-binding function of CBLs were 8.31 and 8.87, respectively. Moreover, the average abundance of glycine (N-myristoylation motif) that is required for protein myristoylation was 4.08 (Table S4). Furthermore, the average abundance of the hydrophobic amino acids in CBLs was relatively higher than ones of other amino acids such as alanine (6.63), isoleucine (5.68), leucine (10.83) and valine (6.27) (Table S4).
Previous research have reported that several consensus sequences play vital roles in their structure or function, including the EF-hand, N-myristoylation motif and the FPSF domain in C-terminal region. In particular, the aspartate and glutamate in EF-hand is the structural basis of function for the calcium sensor. To better illustrate consensus sequences that might be characteristic of the phylogenetic group, we then executed sequence logos motif analyses using WebLogo 3 online tool for each group. For example, Fig. 2b displayed the consensus sequence of four EF-hand from all group CBLs. The residues marked stars were required for the calcium binding function (Fig. S2). The CBLs contained group-specific conserved first non-canonical EF-hand motif, S–C–S-I-I-D/N-D-G-L-I-H–K-E-E (group A), S–S-A-V-I/V-D-D-G-L-I-N-K-E-E (group B), S-G-S-V-I-D-D-G-L-I-N-K-E-E (group C) and S-X-S-I-X-K-D-G-L-I-H–K-E-E (group D), respectively (Fig. S2). In addition, grass CBLs also harbored a group-specific conserved FPSF domain in the C-terminal,, F-P-S-F-V/I-F-N-T-X-V-E-D (group A), F-P-S-F-V-F–H/N-S-Q–V-D/E-D (group B), F-P-S-F-V-F–H/N-S-E-V-D-D (group C) and F-P-S-F-V/I-X-X-S-E/G-X-X-D/E (group D), respectively (Fig.S3). The presence of group-specific EF-hand and/or FPSF domain in CBLs indicated that group-specific CBL proteins may show distinct affinities to calcium ion or substrates (CIPKs).
Molecular Evolutionary Analyses
The genetic distance between groups A and D CBLs (0.279) was the lowest compared with other counterparts, suggesting that their sequence similarity was higher between the group A and D CBLs (Table S5), which was in accordance with their evolutionary relationship (Fig. 1). On the contrary, the highest genetic distance was between group B and C CBLs, indicating that the sequence of two group CBLs showed greater difference than other group CBLs. In addition, if there is no significant difference in genetic distance between each group, it implies that there is inconspicuous sequence divergence between each group CBLs. Furthermore, the average overall mean distance of CBL was 0.662 (standard error 0.343).
To elucidate the evolutionary basis of each group CBLs, we assessed the nonsynonymous-to-synonymous rate ratio using full-length DNA sequences (ώ = dN/dS) (Table 2) under different codon substitution-based evolutionary models. The mean ώ values were less than 1 for group A (ώ = 0.403), group B (ώ = 0.710) and group C (ώ = 0.674) CBLs, suggesting strong purifying selection during the CBLs evolution (Fig. 1). However, the average ώ values of group D CBLs was 1.293 that is greater than 1, indicating that these genes show positive selection during their evolution process (Table 2). Moreover, to compare the difference between whole sequences and functional motifs, we evaluated the ώ values employing the motif sequences under the same evolutionary models (Table S6). As expected, there were equally low ώ values in group A, B and C CBLs when the sequences of EF1 were performed, suggesting that EF1 are well conserved during the evolution process, which was also in accordance with the result for four EF-hand motifs together, as supported by a previous study (Table S6). Interestingly, there were diverse results in the analyses of the FPSF domain. The mean ώ values were 2.028 in group B CBLs, while it were 0.306 in group D (Table S6), indicating that the FPSF domain has more variations compared to EF-hand, which may be associate with their roles in the calcium signal transduction process.
Expression Patterns of OsCBLs
Expression patterns would provide evidence of their functional divergence among all members of a gene family (Whittle and Krochko 2009). In this analysis, the expression patterns of OsCBL genes in various stages of flower development were investigated using RT-qPCR. OsCBL1, OsCBL2, OsCBL3 and OsCBL6 revealed higher expression in all the tissues tested. OsCBL4 showed a very high expression only in leaf blade, but low expression in other organs. OsCBL9 is moderately expressed in root, shoot, leaf blade and leaf sheath, while it shows high expression in other tissues (Fig. 3). In addition, OsCBL5, OsCBL7, OsCBL8 and OsCBL10 showed a relatively lower expression in all organs other than roots, but it was a little different. OsCBL5 and OsCBL8 displayed relatively low expression in all organs detected and OsCBL10 moderate expression in whole flower developmental stages, while OsCBL7 showed moderate expression in the leaf sheath (Fig. 3). These results indicated that these genes may have distinct functions in the corresponding tissues.
We also had special concerns about the rice group B CBL members on account of them lacking myristoylation motifs which are involved in protein anchoring to the membrane to perceive and transduce the calcium signal, especially OsCBL3 and OsCBL6. To explore their roles in response to hormones, the expression profiles in rice seedlings were examined by RT-qPCR under TIBA, SA, IAA, GA and ABA (Fig. 4a). OsCBL3 and OsCBL6 exhibited increases after TIBA and SA treatments and peaked at 1 h, then remained at relatively high levels under TIBA, and recovered to near normal levels at 3 h after SA treatment. Moreover, OsCBL3 and OsCBL6 were induced by ABA within 1 h and then suppressed at 3 h after treatment. Furthermore, OsCBL3 and OsCBL6 were not significantly affected after IAA and GA treatments. We also analyzed their expression profiles under PEG 6000, salt, cold, CuSO4 and K2Cr2O7 applications (Fig. 4b). OsCBL3 was up-regulated at 1 h after K2Cr2O7 treatment, and then recovered to near normal levels, while OsCBL6 showed no obvious change. Likewise, OsCBL3 and OsCBL6 peaked at 3 h after CuSO4 application and then OsCBL3 maintained relatively high expression level, while OsCBL6 recovered to normal condition. Cold treatment had no apparent influence on OsCBL3 expression, but strongly induced OsCBL6 expression up to 12 h. Salt treatment slightly suppressed OsCBL3 expression, while OsCBL6 had mild up-regulated expression. Drought treatment suppressed OsCBL3 and OsCBL6 expression from 3 h, staying up to 12 h and then had a recovery at 24 h up to normal levels.
Time course quantitative gene expression profiles of OsCBL3 and OsCBL6 under hormone treatments and abiotic stresses. a Clustering of OsCBL3 and OsCBL6 according to their expression profiles in the 2-week-old seedling of ZH11 plants after different phytohormone treatments. b Clustering of OsCBL3 and OsCBL6 according to their expression profiles in the 2-week-old seedling of ZH11 plants under abiotic stresses
Tolerance Analyses of OsCBLs-OE Plants Under Salt and Drought Condition
To elucidate the putative roles of OsCBLs in plant, their overexpression transgenic rice plants were generated. To confirm whether the CBL genes were integrated into the genomic DNA, PCR amplification with their specific primers to produce whole coding sequence was performed (Fig. S4). In addition, GUS staining assays in the root of transgenic rice plants were also executed (Fig. S5). Moreover, we also detected their expression levels in transgenic rice plants using RT-qPCR methods (Fig. S6). These results showed that the mRNA of OsCBL genes was increased more than in WT plants (Fig. S6). We further tested the tolerance of OsCBL-OE plants under salt and drought stresses. For salt tolerance analyses, the growth status of WT and OsCBL-OE plants was compared on pre- and post-treatment (Fig. 5a). The root lengths of OsCBL3-OE and OsCBL8-OE plants were significantly higher than those of WT seedlings, while other OsCBLs-OE plants had no apparent change (Fig. 5b). Moreover, the shoot growth of OsCBL4-OE and OsCBL6-OE plants was obviously suppressed by salt stress, while other OsCBLs-OE plants had no effect (Fig. 5b). The result indicated that OsCBL3 and OsCBL8 might improve the tolerance to high-salt stress in rice, while OsCBL4 and OsCBL6 were likely hypersensitive to it.
Characterization of the OsCBLs-overexpressing transgenic rice lines under salt stresses. a Growth performance of the OsCBLs-OE and WT seedlings grown on 1/2MS medium supplemented with 300 mM NaCl. Scale bar = 1 cm. b Root and shoot length of the OsCBLs-OE and WT seedlings grown on 1/2MS medium supplemented with or without 300 mM NaCl. Values are expressed as mean ± SD (n = 3 experiments). Asterisk indicate significant difference at p < 0.05 level between WT and OsCBLs-OE plants
We next examined the drought tolerance of OsCBLs-OE and WT plants with 20% PEG 6000 1/2 MS medium as mimetic drought environment. The morphologic observation of WT and OsCBLs-OE plants was executed on pre- and post-treatment (Fig. 6a). The root lengths of OsCBL5-OE, OsCBL6-OE and OsCBL7-OE plants were significantly higher than those of WT after treatment, while those of OsCBL8-OE, OsCBL9-OE and OsCBL10-OE plants were apparently lower (Fig. 6b). However, the shoot length of all OsCBLs-OE plants had no evident influence except for OsCBL8-OE, OsCBL9-OE and OsCBL10-OE plants which were suppressed by drought stress (Fig. 6b). The data suggested that OsCBL5, OsCBL6 and OsCBL7 might confer resistance to drought, while OsCBL8, OsCBL9 and OsCBL10 had the opposite effect.
Characterization of the OsCBLs-overexpressing transgenic rice lines under drought stresses. a Growth performance of the OsCBLs-OE and WT seedlings grown on 1/2MS medium supplemented with 20℅ PEG6000. Scale bar = 1 cm. b Root and shoot length of the OsCBLs-OE and WT seedlings grown on 1/2MS medium supplemented with or without 20℅ PEG6000. Values are expressed as mean ± SD (n = 3 experiments). Asterisk indicate significant difference at p < 0.05 level between WT and OsCBLs-OE plants
Discussion
Phylogeny and Evolution of CBL Gene Family in Grass
In this study, the evolution of CBL gene family in grass was addressed. For this purpose, we first identified 152 CBLs from 15 Poales and then reconstructed their ML phylogenetic tree (Fig. 1). Strikingly, all CBLs except Z. japonica can be divided into four groups, which might be involved in the incomplete annotation of the genome sequence (Table S1). In addition, the size of the CBL gene family in different grass species had no apparent difference and was not associated with genome size and mostly had near ten CBL genes (Table 1). For instance, O. sativa, Setaria italica and Hordeum vulgare all had 10 CBL genes, while their genome size is 372 Mb, 441.7 Mb and 5100 Mb, respectively. Nevertheless, the gene expansion might be involved with polyploidy or ancient polyploidization events (Jiao et al. 2011). For example, 21 CBL genes, rather than 10, had been identified in hexaploid bread wheat genome which arose in two polyploidization events (Mayer et al. 2014). Therefore, we can speculate that whole genome duplication contributed to gene expansion of CBLs, which is similar to the patterns of MKK and rhomboid gene families (Jiang and Chu 2018; Li et al. 2015). Moreover, except group D CBLs, the mean ώ values were all less than 1 and showed purifying selection during their evolution process and suggested that duplicated or lost genes were easily affected by positive selection (Table 2). Furthermore, domain analyses revealed that CBL genes were highly conserved in the study investigated including EF-hand motif, N-myristoylation motif and the FPSF domain (Fig. 2). As the basis of calcium binding, the average abundance of amino acid residues D and E contained a relatively high proportion (Table S4). Compared to canonical sequences in CaMs, EF-hands of each group CBLs have variable degrees of conservation (Fig.S2), implying that specific signal transduction is likely to depend on different Ca2+ affinities that is not uniquely governed by the EF-hand sequence (Nagae et al. 2003; Sanyal et al. 2015). Above all, these results suggested that the CBL gene family is highly conserved in grasses.
Differential Expression of OsCBLs Genes
So far, numerous CBL genes have been characterized from the higher plants. However, tissue-specific distribution patterns may allow CBL proteins to have corresponding functions in different pathways. For instance, AtCBL4 has specific expression in roots (Guo et al. 2001), while the expression of AtCBL10 is limited to shoot tissues (Kim et al. 2007). These expression patterns enabled CBL4-CIPK24 to effectively mediate Na+ extrusion from roots, and CBL10-CIPK24 in shoot tissues modulated sequestration of Na+ into vacuoles (Weinl and Kudla 2009). Therefore, we investigated the expression profiles in different rice organs. OsCBL4 was highly expressed in leaf, while OsCBL5, OsCBL7 and OsCBL8 were mainly expressed in roots (Fig. 3), implying that rice might have a different signaling pathway from Arabidopsis CBL4-CIPK24 that contributed to response to salt stress. Moreover, OsCBL3, OsCBL6 and OsCBL9 had a high expression in various flower stages (Fig. 3). Hence, we have reason to speculate that it exist a distinct Ca2+ signaling pathway in flower developmental stages.
Function of Rice CBLs in Salt and Drought Stress Responses
In previous studies, expression of AtCBL1 was induced by various stimuli such as salt, drought, cold, wounding and ABA exposure (Fotster et al. 2019; Kudla et al. 1999). AtCBL9 negatively modulates low-temperature stress (Gao and Zhang 2019) and renders plants hypersensitive to ABA in Arabidopsis (Pandey et al. 2004). In addition, overexpression of SpCBL6 improved cold tolerance and reduced drought tolerance (Zhou et al. 2016), while turnip BrrCBL9.2 enhanced salt tolerance in transgenic Arabidopsis plants (Yin et al. 2017). Interestingly, the expression of OsCBL3 was slightly down-regulated under salt condition (Fig. 4), while their overexpression plants enhanced the tolerance of salt stress (Fig. 5). The result indicated that OsCBL3 positively mediated salt tolerance. Likewise, OsCBL6 expression was suppressed (Fig. 4) and their overexpression in plants conferred tolerance to drought stress (Fig. 6). In addition, OsCBL6 was induced by salt and was hypersensitive to it (Fig. 5). These results suggested that OsCBL6 positively improved drought tolerance and negatively regulated salt stress in rice. Unlike CBL4, both OsCBL3 and OsCBL6 lack N-myristoylation motif which favors carrying out the function in membrane targeting of the CBL–CIPK complex (Ishitani et al. 2000). Their structural difference may be the reasons for distinct function in response to salt stress, CBL4 positively improves salt tolerance, while OsCBL6 may have opposite functions.
Conclusion
This study showed that a total of 152 CBL genes were identified among 15 grass plant species based on the presence of a conserved EF-hand domain. Based on the phylogenetic and domain analyses, the CBL gene family can be divided into four groups. It was noteworthy that only Group D CBLs had positive selection during their evolution process. Expression analysis showed that OsCBL3, OsCBL4, OsCBL6 and OsCBL9 had apparent modulated during various tissues in O. sativa. Tolerance analyses revealed that OsCBL3 and OsCBL8 enhanced salt tolerance in rice, while OsCBL4 and OsCBL6 negatively regulated to it. In addition, OsCBL5, OsCBL6 and OsCBL7 might positively modulate drought stress, while OsCBL8, OsCBL9 and OsCBL10 were hypersensitive to it.
Data availability
Not applicable.
Abbreviations
- CBL:
-
Calcineurin B-like
- [Ca2+]cyt:
-
Cytosolic calcium concentration
- EF-hand:
-
Elongation factor hand
- CaM:
-
Calmodulin
- CMLs:
-
CaM-like proteins
- CDPKs:
-
Ca2+-dependent protein kinases
- CIPKs:
-
CBL-interacting protein kinases
- CNB:
-
Calcineurin B
- AKT1:
-
Arabidopsis K+ transporter 1
- pI:
-
Isoelectric point
- Mw:
-
Molecular weight
- ML:
-
Maximum-likelihood
- JTT:
-
Jones-Taylor-Thornton
- ZH11:
-
Zhonghua 11
- RT-qPCR:
-
Real-time quantitative PCR
- SAM:
-
Shoot apical meristem
- MS:
-
Murashige and Skoog
- WT:
-
Wild type
- ώ = dN/dS :
-
Nonsynonymous-to-synonymous rates ratio
References
Albrecht V, Ritz O, Linder S, Harter K, Kudla J (2001) The NAF domain defines a novel protein-protein interaction module conserved in Ca2+-regulated kinases. Embo J 20:1051–1063
Altschul SF, Madden TL, Schaffer AA, Zhang JH, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Batistic O, Kudla J (2009) Plant calcineurin B-like proteins and their interacting protein kinases. Bba-Mol Cell Res 1793:985–992
Batistic O, Sorek N, Schultke S, Yalovsky S, Kudla J (2008) Dual fatty acyl modification determines the localization and plasma membrane targeting of CBL/CIPK Ca2+ signaling complexes in Arabidopsis. Plant Cell 20:1346–1362
Batistic O, Rehers M, Akerman A, Schlucking K, Steinhorst L, Yalovsky S, Kudla J (2012) S-acylation-dependent association of the calcium sensor CBL2 with the vacuolar membrane is essential for proper abscisic acid responses. Cell Res 22:1155–1168
Van Bel M, Diels T, Vancaester E, Kreft L, Botzki A, Van de Peer Y, Coppens F, Vandepoele K (2018) PLAZA 4.0: an integrative resource for functional, evolutionary and comparative plant genomics. Nucleic Acids Res 46:D1190–D1196
Burstenbinder K, Moller B, Plotner R, Stamm G, Hause G, Mitra D, Abel S (2017) The IQD family of calmodulin-binding proteins links calcium signaling to microtubules, membrane subdomains, and the nucleus. Plant Physiol 173:1692–1708
de Castro E, Sigrist CJA, Gattiker A, Bulliard V, Langendijk-Genevaux PS, Gasteiger E, Bairoch A, Hulo N (2006) ScanProsite: detection of PROSITE signature matches and ProRule-associated functional and structural residues in proteins. Nucleic Acids Res 34:W362–W365
Chaves-Sanjuan A, Sanchez-Barrena MJ, Gonzalez-Rubio JM, Moreno M, Ragel P, Jimenez M, Pardo JM, Martinez-Ripoll M, Quintero FJ, Albert A (2014) Structural basis of the regulatory mechanism of the plant CIPK family of protein kinases controlling ion homeostasis and abiotic stress. P Natl Acad Sci USA 111:E4532–E4541
Cheong YH, Pandey GK, Grant JJ, Batistic O, Li L, Kim BG, Lee SC, Kudla J, Luan S (2007) Two calcineurin B-like calcium sensors, interacting with protein kinase CIPK23, regulate leaf transpiration and root potassium uptake in Arabidopsis. Plant J 52:223–239
Cheong YH, Sung SJ, Kim BG, Pandey GK, Cho JS, Kim KN, Luan S (2010) Constitutive overexpression of the calcium sensor CBL5 confers osmotic or drought stress tolerance in Arabidopsis. Mol Cells 29:159–165
Cho JH, Choi MN, Yoon KH, Kim KN (2018) Ectopic expression of SjCBL1, Calcineurin B-Like 1 gene from Sedirea japonica, rescues the salt and osmotic stress hypersensitivity in Arabidopsis cbl1 mutant. Front Plant Sci 9:1188
Eckert C, Offenborn JN, Heinz T, Armarego-Marriott T, Schultke S, Zhang CX, Hillmer S, Heilmann M, Schumacher K, Bock R, Heilmann I, Kudla J (2014) The vacuolar calcium sensors CBL2 and CBL3 affect seed size and embryonic development in Arabidopsis thaliana. Plant J 78:146–156
Evans NH, McAinsh MR, Hetherington AM (2001) Calcium oscillations in higher plants. Curr Opin Plant Biol 4:415–420
Fotster S, Schmidt LK, Kopic E, Anschutz U, Huang SG, Schlucking K, Koster P, Waadt R, Larrieu A, Batistic O, Rodriguez PL, Grill E, Kudla J, Becker D (2019) Wounding-induced stomatal closure requires jasmonate-mediated activation of GORK K+ channels by a Ca2+ sensor-kinase CBL1-CIPK5 complex. Dev Cell 48:87–89
Gao YL, Zhang GZ (2019) A calcium sensor calcineurin B-like 9 negatively regulates cold tolerance via calcium signaling in Arabidopsis thaliana. Plant Signal Behav 14:e1573099
Goodstein DM, Shu SQ, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186
Gu ZM, Ma BJ, Jiang Y, Chen ZW, Su X, Zhang HS (2008) Expression analysis of the calcineurin B-like gene family in rice (Oryza sativa L.) under environmental stresses. Gene 415:1–12
Guo Y, Halfter U, Ishitani M, Zhu JK (2001) Molecular characterization of functional domains in the protein kinase SOS2 that is required for plant salt tolerance. Plant Cell 13:1383–1399
Harper JF, Harmon A (2005) Plants, symbiosis and parasites: a calcium signalling connection. Nat Rev Mol Cell Bio 6:555–566
Huang CL, Ding S, Zhang H, Du H, An LZ (2011) CIPK7 is involved in cold response by interacting with CBL1 in Arabidopsis thaliana. Plant Sci 181:57–64
Ishitani M, Liu JP, Halfter U, Kim CS, Shi WM, Zhu JK (2000) SOS3 function in plant salt tolerance requires N-myristoylation and calcium binding. Plant Cell 12:1667–1677
Jiang M, Chu ZQ (2018) Comparative analysis of plant MKK gene family reveals novel expansion mechanism of the members and sheds new light on functional conservation. BMC Genomics 19:407
Jiao YN, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang HY, Soltis PS, Soltis DE, Clifton SW, Schlarbaum SE, Schuster SC, Ma H, Leebens-Mack J, dePamphilis CW (2011) Ancestral polyploidy in seed plants and angiosperms. Nature 473:97–U113
Kim BG, Waadt R, Cheong YH, Pandey GK, Dominguez-Solis JR, Schultke S, Lee SC, Kudla J, Luan S (2007) The calcium sensor CBL10 mediates salt tolerance by regulating ion homeostasis in Arabidopsis. Plant J 52:473–484
Kolukisaoglu U, Weinl S, Blazevic D, Batistic O, Kudla J (2004) Calcium sensors and their interacting protein kinases: Genomics of the Arabidopsis and rice CBL–CIPK signaling networks. Plant Physiol 134:43–58
Kudla J, Xu Q, Harter K, Gruissem W, Luan S (1999) Genes for calcineurin B-like proteins in Arabidopsis are differentially regulated by stress signals. Proc Natl Acad Sci USA 96:4718–4723
Li LG, Kim BG, Cheong YH, Pandey GK, Luan S (2006) A Ca2+ signaling pathway regulates a K+ channel for low-K response in Arabidopsis. Proc Natl Acad Sci USA 103:12625–12630
Li Q, Zhang N, Zhang LS, Ma H (2015) Differential evolution of members of the rhomboid gene family with conservative and divergent patterns. New Phytol 206:368–380
Li J, Jiang MM, Ren L, Liu Y, Chen HY (2016) Identification and characterization of CBL and CIPK gene families in eggplant (Solanum melongena L.). Mol Genet Genomics 291:1769–1781
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Liu JP, Zhu JK (1998) A calcium sensor homolog required for plant salt tolerance. Science 280:1943–1945
Lu TT, Zhang GF, Sun LR, Wang J, Hao FS (2017) Genome-wide identification of CBL family and expression analysis of CBLs in response to potassium deficiency in cotton. Peer J 5:e3653
Luan S (2009) The CBL–CIPK network in plant calcium signaling. Trends Plant Sci 14:37–42
Luan S, Kudla J, Rodriguez-Concepcion M, Yalovsky S, Gruissem W (2002) Calmodulins and calcineurin B-like proteins: calcium sensors for specific signal response coupling in plants. Plant Cell 14:S389–S400
Ma Q, Tang RJ, Zheng XJ, Wang SM, Luan S (2015) The calcium sensor CBL7 modulates plant responses to low nitrate in Arabidopsis. Biochem Bioph Res Commun 468:59–65
Mahs A, Steinhorst L, Han JP, Shen LK, Wang Y, Kudla J (2013) The calcineurin B-like Ca2+ Sensors CBL1 and CBL9 function in pollen germination and pollen tube growth in Arabidopsis. Mol Plant 6:1149–1162
Mayer KFX, Rogers J, Dolezel J, Pozniak C, Eversole K, Feuillet C, Gill B, Friebe B, Lukaszewski AJ, Sourdille P, Endo TR, Dolezel J, Kubalakova M, Cihalikova J, Dubska Z, Vrana J, Sperkova R, Simkova H, Rogers J, Febrer M, Clissold L, McLay K, Singh K, Chhuneja P, Singh NK, Khurana J, Akhunov E, Choulet F, Sourdille P, Feuillet C, Alberti A, Barbe V, Wincker P, Kanamori H, Kobayashi F, Itoh T, Matsumoto T, Sakai H, Tanaka T, Wu JZ, Ogihara Y, Handa H, Pozniak C, Maclachlan PR, Sharpe A, Klassen D, Edwards D, Batley J, Olsen OA, Sandve SR, Lien S, Steuernagel B, Wulff B, Caccamo M, Ayling S, Ramirez-Gonzalez RH, Clavijo BJ, Steuernagel B, Wright J, Pfeifer M, Spannagl M, Mayer KFX, Martis MM, Akhunov E, Choulet F, Mayer KFX, Mascher M, Chapman J, Poland JA, Scholz U, Barry K, Waugh R, Rokhsar DS, Muehlbauer GJ, Stein N, Gundlach H, Zytnicki M, Jamilloux V, Quesneville H, Wicker T, Mayer KFX, Faccioli P, Colaiacovo M, Pfeifer M, Stanca AM, Budak H, Cattivelli L, Glover N, Martis MM, Choulet F, Feuillet C, Mayer KFX, Pfeifer M, Pingault L, Mayer KFX, Paux E, Spannagl M, Sharma S, Mayer KFX, Pozniak C, Appels R, Bellgard M, Chapman B, Pfeifer M, Pfeifer M, Sandve SR, Nussbaumer T, Bader KC, Choulet F, Feuillet C, Mayer KFX, Akhunov E, Paux E, Rimbert H, Wang SC, Poland JA, Knox R, Kilian A, Pozniak C, Alaux M, Alfama F, Couderc L, Jamilloux V, Guilhot N, Viseux C, Loaec M, Quesneville H, Rogers J, Dolezel J, Eversole K, Feuillet C, Keller B, Mayer KFX, Olsen OA, Praud S (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
McCormack E, Tsai YC, Braam J (2005) Handling calcium signaling: Arabidopsis CaMs and CMLs. Trends Plant Sci 10:383–389
Mohanta TK, Mohanta N, Mohanta YK, Parida P, Bae HH (2015) Genome-wide identification of Calcineurin B-Like (CBL) gene family of plants reveals novel conserved motifs and evolutionary aspects in calcium signaling events. BMC Plant Biol 15:189
Nagae M, Nozawa A, Koizumi N, Sano H, Hashimoto H, Sato M, Shimizu T (2003) The crystal structure of the novel calcium-binding protein AtCBL2 from Arabidopsis thaliana. J Biol Chem 278:42240–42246
Ouyang S, Zhu W, Hamilton J, Lin H, Campbell M, Childs K, Thibaud-Nissen F, Malek RL, Lee Y, Zheng L, Orvis J, Haas B, Wortman J, Buell CR (2007) The TIGR Rice genome annotation resource: improvements and new features. Nucleic Acids Res 35:D883–D887
Pandey GK, Cheong YH, Kim KN, Grant JJ, Li LG, Hung W, D'Angelo C, Weinl S, Kudla J, Luan S (2004) The calcium sensor calcineurin B-Like 9 modulates abscisic acid sensitivity and biosynthesis in Arabidopsis. Plant Cell 16:1912–1924
Ren XL, Qi GN, Feng HQ, Zhao S, Zhao SS, Wang Y, Wu WH (2013) Calcineurin B-like protein CBL10 directly interacts with AKT1 and modulates K+ homeostasis in Arabidopsis. Plant J 74:258–266
Sanchez-Barrena MJ, Martinez-Ripoll M, Albert A (2013) Structural biology of a major signaling network that regulates plant abiotic stress: The CBL–CIPK mediated pathway. Int J Mol Sci 14:5734–5749
Sanders D, Pelloux J, Brownlee C, Harper JF (2002) Calcium at the crossroads of signaling. Plant Cell 14:S401–S417
Sanyal SK, Pandey A, Pandey GK (2015) The CBL–CIPK signaling module in plants: a mechanistic perspective. Physiol Plantarum 155:89–108
Schulz P, Herde M, Romeis T (2013) Calcium-dependent protein kinases: hubs in plant stress signaling and development. Plant Physiol 163:523–530
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tang RJ, Liu H, Yang Y, Yang L, Gao XS, Garcia VJ, Luan S, Zhang HX (2012) Tonoplast calcium sensors CBL2 and CBL3 control plant growth and ion homeostasis through regulating V-ATPase activity in Arabidopsis. Cell Res 22:1650–1665
Weinl S, Kudla J (2009) The CBL–CIPK Ca2+-decoding signaling network: function and perspectives. New Phytol 184:517–528
Whittle CA, Krochko JE (2009) Transcript profiling provides evidence of functional divergence and expression networks among ribosomal protein gene Paralogs in Brassica napus. Plant Cell 21:2203–2219
Xi Y, Liu JY, Dong C, Cheng ZM (2017) The CBL and CIPK gene family in grapevine (Vitis vinifera): genome-wide analysis and expression profiles in response to various abiotic stresses. Front Plant Sci 8:978
Ye NH, Wang FZ, Shi L, Chen MX, Cao YY, Zhu FY, Wu YZ, Xie LJ, Liu TY, Su ZZ, Xiao S, Zhang H, Yang JC, Gu HY, Hou XX, Hu QJ, Yi HJ, Zhu CX, Zhang JH, Liu YG (2018) Natural variation in the promoter of rice calcineurin B-like protein10 (OsCBL10) affects flooding tolerance during seed germination among rice subspecies. Plant J 94:612–625
Yin X, Wang QL, Chen Q, Xiang N, Yang YQ, Yang YP (2017) Genome-wide identification and functional analysis of the calcineurin B-like protein and calcineurin b-like protein-interacting protein kinase gene families in turnip (Brassica rapa var. rapa). Front Plant Sci 8:1191
Zhang HC, Yin WL, Xia XL (2008) Calcineurin B-Like family in Populus: comparative genome analysis and expression pattern under cold, drought and salt stress treatment. Plant Growth Regul 56:129–140
Zhang HF, Yang B, Liu WZ, Li HW, Wang L, Wang BY, Deng M, Liang WW, Deyholos MK, Jiang YQ (2014) Identification and characterization of CBL and CIPK gene families in canola (Brassica napus L.). BMC Plant Biol 14:8
Zhang CX, Beckmann L, Kudla J, Batistic O (2017) N-terminal S-acylation facilitates tonoplast targeting of the calcium sensor CBL6. Febs Lett 591:3745–3756
Zhou YL, Cheng Y, Yang YQ, Li X, Supriyo B, Sun XD, Yang YP (2016) Overexpression of SpCBL6, a calcineurin B-like protein of Stipa purpurea, enhanced cold tolerance and reduced drought tolerance in transgenic Arabidopsis. Mol Biol Rep 43:957–966
Funding
This research was funded by the National Natural Science Foundation of China (81360611) and Shanghai Sailing Program (19YF1414800). The funding bodies had no role in study design, analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
MJ and GW conceived and designed the project. MJ executed the bioinformatics analysis, conducted the experiments and wrote the manuscript. CZ, MZ, YL and GW retrieved gene sequence data and performed the bioinformatics analysis. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Ethical approval
Not applicable.
Informed consent
Not applicable.
Electronic supplementary material
Below is the link to the electronic supplementary material.
12374_2020_9240_MOESM7_ESM.tif
Fig. S1. Maximum Likelihood phylogenetic trees were reconstructed using EF-hand motifs of CBL proteins in 15 grasses. Phylogenetic analysis was carried with EF-hand motifs of protein sequences for 152 CBL proteins from 15 grass species identified in this study Supplementary file7 (TIF 8180 kb)
12374_2020_9240_MOESM8_ESM.tif
Fig. S2. Sequence features shown in the form of web logos representing the conserved EF-hand motif of each group. The red stars indicate residues of functional or structural importance based on phylogenetic conservations. Logos were generated using the Weblogo3 application (http://weblogo.threeplusone.com/) Supplementary file8 (TIF 4225 kb)
12374_2020_9240_MOESM9_ESM.tif
Fig. S3. Sequence features shown in the form of web logos representing the conserved FPSF domain in C-terminal region of each group Supplementary file9 (TIF 1744 kb)
12374_2020_9240_MOESM10_ESM.tif
Fig. S4. Molecular analysis of OsCBLs transgenic rice plants in the level of DNA. 1: OsCBL1,2: OsCBL2,3: OsCBL3,4: OsCBL4,5: OsCBL5,6: OsCBL6,7: OsCBL7,8: OsCBL8,9: OsCBL9,10: OsCBL10, Marker:100 bp plus DNA ladder Supplementary file10 (TIF 2635 kb)
12374_2020_9240_MOESM12_ESM.tif
Fig. S6. Real-time PCR analysis of OsCBLs T3 transgenic rice lines. A: OsCBL1-OE, B:OsCBL2-OE, C:OsCBL3-OE, D: OsCBL4-OE, E: OsCBL5-OE, F: OsCBL6-OE, G: OsCBL7-OE, H:OsCBL8-OE, I: OsCBL9-OE, J:OsCBL10-OE Supplementary file12 (TIF 7048 kb)
Rights and permissions
About this article
Cite this article
Jiang, M., Zhao, C., Zhao, M. et al. Phylogeny and Evolution of Calcineurin B-Like (CBL) Gene Family in Grass and Functional Analyses of Rice CBLs. J. Plant Biol. 63, 117–130 (2020). https://doi.org/10.1007/s12374-020-09240-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-020-09240-y