Abstract
An integral approach to decoding both culturable and uncultured microorganisms' metabolic activity involves the whole genome sequencing (WGS) of individual/complex microbial communities. WGS of culturable microbes, amplicon sequencing, metagenomics, and single-cell genome analysis are selective techniques integrating genetic information and biochemical mechanisms. These approaches transform microbial biotechnology into a quick and high-throughput culture-independent evaluation and exploit pollutant-degrading microbes. They are windows into enzyme regulatory bioremediation pathways (i.e., dehalogenase) and the complete bioremediation process of organohalide pollutants. While the genome sequencing technique is gaining the scientific community's interest, it is still in its infancy in the field of pollutant bioremediation. The techniques are becoming increasingly helpful in unraveling and predicting the enzyme structure and explore metabolic and biodegradation capabilities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ecological processes are associated with microbial activities, either indirectly or directly, where microbes perform vital functions in the overall biogeochemical cycling of nutrients and the degradation of persistent ecological pollutants [1, 2]. Microbes' use to remove natural or synthetic pollutants is the prevailing, low-cost green technology to treat different kinds of polluted environments [3, 4]. Microbes make fantastic pollutant degraders because they possess enzymes, for instance, dehalogenases. The enzymes use the environmental pollutants as nutrient or carbon sources, and their naturally small size enables facile contact with the pollutants [5]. However, the naturally occurring organohalide compounds are insignificant compared to anthropogenic ones [6]. Adding halogen atoms to organic molecules would significantly improve their properties, such as solubility and toxicity [6]. The change would result in valuable commercial product improvement but may impart serious effects on microbial metabolism [7]. At times, halogenation decreases the vulnerability of chemicals significantly to enzymatic attack and give rise to persistent compounds [8].
Consequently, bioremediation technologies to remove such chemicals from the environment are still growing. Several microbes have been identified as promising in the bioremediation of these pollutants [5, 9, 10]. However, the main questions remain, (i) what are the enzymes involved in pollutants degradation pathways? (ii) how do they carry out their function?, and (iii) how do they respond to various pollutants?. The questions are useful in forecasting the degradability and ability of the native microbial environment and provides valuable information about enzymes involved in degradation. However, the identification of critical regulatory genes, their structure, and the comprehension of their genetic responses remain challenging.
The arrival of molecular microbial tools has profoundly changed the microbial ecology area by providing a direct approach to environmental microbes' phylogeny and physiology, without depending on their cultivability [11]. This new microbial ecology era is defined by creating quick and high input techniques, which aids the culture-independent method of microbial communities dwelling in ecosystems. Hence, this incapacitates the limitations of cultivation-dependent methods. Molecular methods to evaluate microbial ecology facilitated the essential comprehension of microbial communities' structural and functional behaviors [12, 13], offering excellent prospects for novel bioremediation approaches. Molecular approaches permit immediate investigation of the transcripts, metabolites, genes, and protein of natural microbial communities in situ [12, 14]. It also gives insight into their relations that affect the attenuation of environmental pollutants, thereby restoring the polluted environments.
This review provides recent information on the bioremediation of environmental pollutants and the various genomic strategies involving the WGS, metagenomics, and single-cell genome analysis in pollutant degradation research. An overview of functional and structural arrangement and detailed application of the techniques to pollutant degradation research are also summarized. Moreover, information on gene arrangement, metabolic pathways, and the molecular mechanism of organohalide pollutants are lacking. The approaches may improve catabolic pathways and offer useful information on the genome, transcriptome, and proteome of in situ microbial communities without needing cultivation [12, 14].
Contaminants in natural environment
The extensive utilization of harmful organic compounds and the rise in pollutant concentrations in the environment are consequences of rapid industrialization, increased population, military activities, changed agricultural activities, and urbanization. Many pollutants and waste materials containing heavy metals, petroleum hydrocarbons, and halogenated organic compounds are released into the environment annually. The issue is exacerbated by their persistence and is amassed over a prolonged period. Their widespread distribution into air and groundwater has resulted in critical ecological and health challenges globally [15].
Categorically, common environmental pollutants are hydrocarbon, agricultural supplements (pesticide, herbicides, and synthetic insecticides), and heavy metals (Table 1). Petroleum hydrocarbons have multiple carbon bonds, which create strong, complex structures when bonded with other types of molecules. They exist in various forms, viz. short, medium, long aliphatic, aromatics, and polycyclic aromatic hydrocarbons (PAHs) of varying ratios [16]. Organohalides are also common environmental pollutants by spraying herbicides, fungicides, insecticides, hydraulic and heat fluids, plasticizers, and chemical intermediates. As a matter of fact, chlorinated phenolic compounds are one of the most abundant recalcitrant wastes discharged by the paper and pulp industry. Pesticides are globally applied in agriculture as well as in many public health sectors. Plastics and dyes are occasionally used, except for a few PAHs used in medicine [17]. However, the compounds are deemed as environmental pollutants in various terrestrial and aquatic ecosystems. Their detrimental impacts offset their usefulness, as they amass and are resistant to degradation. Toxicities of such compounds have been reported, including oxidative stress, changes in metabolic parameters, hepatic stress, and cellular necrosis [18, 19]. The biological consequences concede the survival, growth, reproduction, and the early-stage development of organisms [19,20,21].
The frequently used techniques to treat polluted environments include physical and chemical technologies such as burying, combusting, extracting soil vapor, soil washing, and dispersion [22]. The methods, however, are incapable of totally decomposing pollutants and are economically unfriendly. In some cases, more toxic chemicals are formed compared to before the treatment [22]. Due to the high cost and the lack of public acceptance of these methods, bioremediation seems to be an excellent alternative clean-up technology. Bioremediation is a method that biologically degrades organic wastes to a safe compound under regulated conditions or to levels lower than concentration limits established by regulating agencies [23]. The method is safe, cost-effective, and naturally eco-friendly, uses several enzymes from microbes to degrade toxic organic pollutants [10, 23, 24].
Classification of enzymes involve in bioremediation
Based on biodegradation ability, microbial enzymes are classified into oxidoreductases and hydrolases [24, 25]. Certain oxidoreductase-producing bacteria such as laccase, peroxidase, and oxygenase can degrade radioactive metals, chlorinated compounds, and petroleum containing hydrocarbons [21, 26, 27]. In contrast, hydrolases are common hydrolytic enzymes that bioremediate and detoxify agrochemical pollutants. For instance, oil spills, organohalides, organophosphates, and carbamate insecticides have been effectively degraded by dehalogenases, lipases, cellulases, carboxylesterases, and phosphodiesterases [10, 28].
The study of pollutant-degrading microbes, identifying their genetic and biochemistry, and developing techniques for their use have led to crucial human endeavors. Microbes stand preferable as agents to clean up the environment in terms of cost, chemical, and physical techniques. Nonetheless, microbial-driven clean up of polluted environments remains limited, following the lack of knowledge of the factors that control and regulate their metabolism, growth, and different microbial community dynamics. The recent use of innovative tools like proteomics, fluxomics, transcriptomics, genomics, and metabolomics has produced eco-friendlier pollutant treatment strategies [14]. Microbes can mineralize and degrade the contaminants into energy sources and then convert them into simpler intermediates. During this process, the microbes can transform toxic chemicals into nontoxic compounds or completely degrade them into carbon dioxide and water by converting them from one phase to another. The expression of the microbial enzymes’ genes relies on these toxic chemicals' presence, as the enzymes are substrate-specific and mobile due to their smaller size [24, 29, 30]. Gene expression of the enzyme depends on the substrate types present in the medium (Fig. 1).
An overview of gene expression in the biodegradation process in a bacteria system. The toxic chemicals of the halogenated compound must be able to enter the cells. Inside the cells, these toxic chemicals will trigger a specific gene in chromosomal DNA and allow for transcription and translation of a protein for an enzymatic reaction for biodegradation. The contaminants serve as carbon and/or energy source, and microbes can degrade/mineralize them into simpler intermediate
Bacterial dehalogenases
Degradation of organohalides by dehalogenases isolated from environmental bacteria and their basic mechanisms have been described in great detail [31]. Dehalogenases can be categorized into oxidative, reductive, hydrolytic, or thiolytic dehalogenases [10, 31]. Hydrolytic dehalogenases are further categorized into haloacid dehalogenases, haloalkane dehalogenases, fluoroacetate dehalogenases, and halohydrin dehalogenases that degrade aliphatic halogenated organic compounds. The haloacid dehalogenases are sub-divided into three kinds based on their substrate selectivity. The L-2-haloacid dehalogenases (L-DEX) and D-2-haloacid dehalogenases (D-DEX) catalyze L-2-haloalkanoate and D-2-haloalkanoate to produce L-2-hydroxyalkanoate and D-2-hydroxyalkanoate, respectively. D,L-2-haloacid dehalogenases (D,L-DEX) react with both D- and L-2-haloalkanoates, yielding 2hydroxyalkanoates, respectively [10, 31]. Haloacid dehalogenase (HAD) enzymes belong to a large phosphohydrolase superfamily, which include hydrolases, oxygenases and dehydrogenases that degrade polychlorinated biphenyls [10, 31]. Due to the rapid development of the WGS process, new microorganisms or enzymes (such as dehalogenase) can be easily identified and studied further by analyzing the full DNA sequence in genome databases.
Strategies for genome sequencing
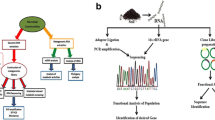
The application of genome sequencing methods is classified into amplicon sequencing, shotgun metagenomics, single-cell genomic sequencing, and whole-genome sequencing of cultured microbes, depending on the issue to be solved (Fig. 2). Progress in WGS has provided a better understanding of how microbes degrade pollutants and their environmental adaptation at genetic levels. Many enzymes, for instance, dehalogenases and their pathways responsible for pollutant degradation, have been described. Complete genome sequencing techniques have unveiled several enzymes that participated in pollutant degradation in different environments, as discussed in “Application of whole-genome sequencing techniques to pollutants degradation research” section. Although the various complete genome of distinct pollutant-degrading bacteria is known, the genetic basis of pollutant degradation (especially the regulatory function of haloacid dehalogenases) and environmental adaptation to the environments are yet to be clarified. Therefore, this article reviews the WGS approach in pollutant bioremediation by emphasizing the potential regulatory mechanism of haloacid dehalogenase. It is an excellent means for consolidating information on the complete bioremediation of organohalide pollutants. Identifying individual strains in the complex microbial communities can improve and, undoubtedly, expose more novel enzymes/genes capable of degrading pollutants from an individual cell.
Whole-genome sequencing (WGS) of cultured microbes
Whole Genome Sequencing (WGS) is one of the best methods for regaining the microbes' genetic and metabolic diversity. It becomes a problem when many microbes evade cultivation [32], making the complete genome less appreciated using traditional approaches. This technique uses sequencing platforms such as Illumina, PacBio, Nanopore, Qiagen, BGISEQ, IonTorrent, or other sequencers. Although this technique has flaws in genome sequencing, it can be rectified using a targeted enrichment method. The method enhances a particular microbe's isolation within a specified community, having the biochemical property of interest. Pertinently, the term ‘targeted enrichment’ refers to the isolation of a single microbe via a physically enhanced cell population based on patterns of phenotypic traits like size, density, shape, and the native spectral trait [33]. Subsequently, the isolated microbes are used for a complete genome assembly and sequencing.
However, this technique is not suitable for complex environmental samples because measured properties do not uniquely define them. This issue can be overcome by limited enrichment, enhancing the outcome of efforts on varying data sets of organisms. The procedure has successfully overcome the problem of low-yield in selected microbes [34]. Regardless of the availability of next-generation sequencing methods, improvement in bioinformatics, and the potential of sequencing data, the use of WGS of cultured microbes is still at a nascent stage. This method has allowed microbial ecologists to explore, compare, and characterize microbial communities since ∼ 98% of bacteria in an environmental sample are unculturable by traditional laboratory methods. The issue above can be overcome by metagenomics, and single-cell genomics, both of which allow scientists to access unculturable microbial genomes.
Metagenomic sequencing
Metagenomic sequencing reads DNA accurately from an environmental sample without cultivating individual colonies. The entire method involves the extraction of DNA from environmental samples, amplification, and high-throughput sequencing. DNA fragments produced by the sequencer are then suitably categorized (binned) and assembled into contigs by bioinformatics models. The qualified and accepted bins are then designated as metagenome-assembled genomes (MAGs) [35, 36]. According to the literature, metagenomic sequencing is grouped as amplicon metagenomic and shotgun metagenomic.
Amplicon sequencing
The partial 16S-based metagenomic technique, otherwise known as amplicon or targeted metagenomic, uses microbial marker genes like 16S rRNA, internal transcribed spacer, and other marker genes. The most general marker gene for amplicon or target sequencing is the 16S rRNA gene. It is used as a taxonomic marker that can resolve one significant query connected to microbial ecology as “who is there”, by conveying the sequence reads to a taxonomic ancestry based on identified 16S rRNA database, like Greengenes, SILVA or Ribosomal database project (RDP) [37]. However, the bacterial roles cannot be clearly defined when the 16S rRNA sequence reads fail to adequately resolve or detect microbes at the species/strain level. Also, the marker genes survey only emphasizes a few common genes, as they cannot precisely distinguish the microbes' functional or metabolic potentials [38,39,40].
Currently, projecting microbial functional capabilities from 16S rRNA gene is a prevalent substitute over the shotgun metagenomic technique. The cheaper 16S rRNA gene sequencing offers taxonomic structure but not functional interpretations. Several software packages can solve this challenge by identifying specific features to forecast the functional metagenomic abilities according to known 16S rRNA gene sequences linked to genomes. Piphillin, PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States), MelonnPan (Model-based Genomically Informed High-dimensional Predictor of Microbial Community Metabolic Profiles), and other similar tools are publicly accessible computational software for predicting microbial functional capabilities from detected 16S rRNA genes. The software uses direct nearest-neighbor matching or relies on reference phylogenetic trees of 16S rRNA gene amplicons to infer metagenomic function. The Kyoto Encyclopedia of Genes and Genomes (KEGG) and BioCyc are among the reference genome databases for predicting metagenomic functions [40, 41]. Shotgun metagenomic sequencing can validate the purported theory using phylogeny and functions [42].
Shotgun metagenomics
Shotgun metagenomics sequencing can reveal the potential of microbial communities and offers insights into their diversity, life cycle, and functions. The metagenomic reads encoding genes of interest offer functional annotation through gene fragment recruitment, de novo gene prediction, and protein family classification [43]. This technique provides a solution to two crucial questions associated with microbial ecology; “who is there” and “what are they doing.” Annotation is assigned to the reads to establish the functional gene, using databases like Non-Redundant (NR) and Kyoto Encyclopedia of Genes and Genomes (KEGG) eggNOG [44]. This approach offers a complete thoughtful of the community composition at a high resolution and possible metabolic lane related to the microbial community [45]. However, the assignment of functional annotation and metagenome assembly may prove challenging for shotgun metagenomic sequencing. The microbial classification structure is not available, unlike the 16S rRNA reports. To address this setback, RiboFR-Seq and epiPCR (emulsion, paired isolation, and concentration PCR) techniques can simultaneously capture both the 16S rRNA variable areas and their flanking protein-coding genes. In addition, they link functional genes and phylogenetic markers in uncultured single cells connected to metagenomic contigs and 16S rRNA profiles [46]. However, the methods only partly resolve the problem and are unable to connect all the functional genes of the microorganisms to their phylogeny.
There are currently two main methods in shotgun metagenomics, which focus on various parts of the microbial community within a defined environment. Firstly, the structural metagenomics' primary emphasis is to study uncultivated microbial composition and other properties, i.e., the complex metabolic network structure between community members [47]. Here, the microbial community composition is defined as the population structure, and its dynamics in a specific environment is related to factors such as pressures and spatiotemporal parameters. Greater insight is provided by observing the community composition in terms of interactions among the individual microbes within the community, vital for offering biochemical functions among their group members [47]. The studies of 16S rRNA and shotgun metagenomics are not equally exclusive. The methods establish a link between the 16S rRNA analysis to genes, which showed that the metabolic pathways are beneficial to determine the functional potential of a microbiome [48]. The approaches complement each other and permit a more in-depth investigation of pertinent biological queries in microbial ecosystems like “who are the community members?” and “what are their functional roles?”. The single-cell sequencing is recognized as an effective technique that provides sequencing information on target microbe at single-cell levels due to flaws in amplicon sequencing and shotgun metagenomic sequencing.
Single-cell genomics
This term signifies an individual cell's genomes, which might or might not include the complete genetic range in the microbiota. Single-cell sequencing of microbial cells is becoming an essential tool for the microbiologist. The technique complements other existing techniques, including traditional culture-based methods and metagenomic sequencing. Genome sequencing of individual cells is a novel culture-independent technique that provides an evolutionary record of the microbe and enables cell-to-cell variability studies in microbial populations. It links metabolic function to specific species and creates a high-quality genome for species with low richness [49].
The single-cell genome sequencing has steps that incorporate sample preparation and single-cell isolation by advanced isolation methods (such as micromanipulation, flow cytometry, microfluidics, and encapsulation in droplets). Other methods include DNA extraction, phylogenetic classification by 16S rRNA gene, WGA using Multiple Displacement Amplification (MDA), library preparation, sequencing, and data analysis [49]. In the case sequence analysis, the distinct steps comprise of quality assurance of raw reads, genome assembly (using a single-cell-specific assembler), automated and/or manual contaminant identification and removal, annotation, genome quality inspection, and categorization according to the Minimum Information on Single Amplified Genome (MISAG) standards [49] and database submission. The method improves complete genome sequence from a single bacterium, enabling novel discoveries of genes/pathways capable of degrading pollutants. This is made possible by studying how a specific gene is regulated by looking at the putative operon. Single-cell sequencing can unveil novel bacteria with an alternative genetic code, observe the gut microbial of cells that use host originated compounds, and quantify absolute taxon abundances in the gut microbiome [50].
Application of whole-genome sequencing techniques to pollutants degradation research
The use of WGS has led to novel discoveries on genes/pathways from microbes capable of degrading pollutants from the environment. Some of the most noticeable genes/pathways are summarized in Table 2. In 2009, Suenaga et al. found over 25 estradiol dioxygenase genes responsible for catalyzing the ring cleavage of dehydroxylated central intermediate compounds during aerobic degradation of aromatic hydrocarbon [51]. Monooxygenases, dioxygenases, hydroxylase, and ring-hydroxylating dioxygenases (RHD)-degrading aromatic polyaromatic hydrocarbons (benzoate, phenol, biphenyl, polychlorinated biphenyl, hexadecane, naphthalene, and phenanthrene) were also reported [52,53,54,55]. Some researchers detected several potential genes in dehalogenases, laccase, and cutinases. The genes were found to encode the degradation of halogenated hydrocarbons, chlorinated solvents, industrial dyes, and polyethylene terephthalate (PET) [9, 48, 56,57,58,59,60,61].
The complete genome sequence of pollutant-degrading bacteria has been extensively studied [62]. Bacterial isolates in oil/petroleum polluted sites, polluted marine water, wastewater, contaminated soils, and pharmaceutical-contaminated site (Table 1) have been sequenced, and the genetic sources of their pollutant mineralization potential had been revealed [62, 63]. In such cases, genome analysis of pollutant-degrading bacterial strains, pollutant degradation/uptake mechanism, and their genetic adaption for growing in the pollutant stressed environments might prove useful. Though pollutant degrading bacteria's complete pathway has been clarified, only a few reports have highlighted the degradation pathway [62]. Sequencing environmental/individual DNA via next-generation sequencing is rapid with minimum cost implications, and new innovative sequencing equipment is produced nearly every year. A large amount of data is available from next-generation sequencing platforms, but advances in bioinformatics analysis are lagging. For instance, reconstruction of a complete genome from most environments remains a challenge unless enrichment methods can reduce microbiome complexity in microcosms [64]. Till now, only genomes of dominants strain can be constructed from complex metagenomes [65]. The situation will only change if there is a computational improvement to restructure or complete genomes of rare taxa. Genomic techniques for addressing biochemical roles can help assign a functional and taxonomic unit to access the biodegradation activity [66]
Structural and functional characteristics of dehalogenase genes
The structural and regulatory function of dehalogenase genes can be understood using a cluster of genes called an operon. It is depicted as clusters of genes with related structures and functionally allows regulation of expression in microbes. Since the discovery of lac operon and various catabolic operons, microbes/enzymes' control strategies had been uncovered. Several microbial genomes show groups of genes within a single process that may be co-jointly transcribed and regulated in classical operons or with distinct promoters and regulators. However, the level of operon gene arrangement and gene clustering varies among species. In some bacteria, operons are reasonably unpreserved, and genes involved in one cellular process can be scattered in the genome [80]. Previously, gene cloning successfully revealed the gene structure of the haloacid dehalogenases in Rhizobium sp. RC1 (Fig. 3) is the only bacterium that produces three different haloacid dehalogenases, dehD, dehE, and dehL. The operon regulated a single regulatory gene (dehR), which controls all three structural genes of dehD, dehE, and dehL in Rhizobium sp. RC1. Rhizobium sp. RC1 dehalogenase is involved in the degradation of organohalide compounds and its expression is stimulated only in the presence of the pollutants in the environment [81].
A proposed genetic structure of haloacid dehalogenase genes of Rhizobium sp. RC1. The dehR represents a regulatory gene that controls all three dehalogenases. The dehP is the permease gene encoding the dehalogenase uptake protein. P1 and P2 represent promoter regions of the structural genes dehE and dehD/dehL, respectively. The ? represents an unknown gap between the two sets of genes. It can only be resolved when the full genome sequence of RC1 is obtained
Dehalogenase-coding genes are jointly clustered with regulatory genes, transport or uptake proteins, accessory genes, and other genes that partake in organohalides catabolism. Regulatory genes are usually located near dehalogenases and, often, in the opposite direction. They are a general feature of transcriptional factors in bacteria that enables intensive expression, production of transcriptional regulators nearby targeted genes, and favors efficient recognition of subsequent DNA-binding sites by averting spatial diffusion [82]. The entire genetic organization can only be studied by knowing the full genome sequence analysis.
Transcriptional factors involved in organohalide detection having the Helix-Turn-Helix (HTH) DNA binding domain are found in almost all known bacteria, transcriptional regulators. They make up the majority of regulators involved in bioremediation [83]. The binding of small ligands to this site induces conformational changes in transcriptional factors, thereby affecting their DNA-binding properties [84]. Some transcriptional factors have been associated with specific organohalide compounds, for instance, transcriptional regulators of the lysR family (like CatR, ClcR, LinR, PcpR). They typically detect aromatic or aliphatic organohalide pollutants such as chlorobenzoate, chlorophenol, chloroethane, and chloroethene [62]. The NtrC family of transcriptional regulators (DmpR, MopR, XylR) can identify both aromatic compounds substituted with or without halogens, although with low affinity [84]. The MarR family transcriptional factors, on the other hand, regulate aliphatic organohalide solvents [85]. However, the complete genome of Norcadia soli strain Y48 was found to contain the above said transcriptional factors, in addition to TetR, GntR, and Ic1R-family transcriptional regulators [86].
Dehalogenase expression may also be affected by transport/uptake proteins, which could be part of its operon. Genes encoding for transporters or uptake proteins in the uptake of organohalides are often located near dehalogenase genes (Fig. 3) [87]. Generally, many enzymes/genes involved in pollutants degradation and transcriptional factors that respond to organohalide pollutants are not fully characterized [88]. The survival of bacteria and their ability to catalyze under harsh conditions hinge on organohalides and nutrient uptake systems. With regards to organohalides biodegradation, genomics and metagenomics approaches have successfully identified genes in several microbes that code for enzymes related to their biodegradation. Information on the whole-genome analysis of genes of pollutant-degrading bacteria structure and function is described in “Structural and functional characteristics of dehalogenase genes” section.
Table 3 presents the complete genomic survey of bacteria capable of degrading organohalide pollutants. Many aspects of regulatory pathways remain unexplored compared to extensive information on genes and enzymes responsible for organohalides degradation. Hence, it is unsurprising that some aspects of the regulatory pathways are responsible for limiting organohalides' practical degradation in unpolluted or polluted environments.
Genomics is a powerful computer-based technology used to understand the structure and feature of all genes in an organism [101]. In this review, the structure and regulation of haloacid dehalogenases are illustrated using the complete genome of organohalide-degrading Burkholderia caribensis MBA4, Pseudomonas aeruginosa N002 and Sphingobium chlorophenolicum strain L-1. Burkholderia caribensis MBA4 specifically degrades monochloroacetic acid and D,L-2-bromopropionic (D,L-2-BP) acid, but acts weakly on D, L-2- chloropropionic acid (D,L2-DCP) [56, 58]. The haloacid utilizing operon comprising of dehalogenase deh4a and permease deh4p genes was discovered in replicon CP012747. In contrast, the deh4a eight other genes were annotated as haloacid dehalogenase or haloacid dehalogenase-like proteins for the whole genome (Fig. 4a) [58]. The role of permease deh4p is to transport monochloroacetic acid into the cell. Figure 4b illustrates the hydrolysis of glycolate by a glycolate oxidase (an enzyme that has 3 subunits; viz GlcD, GlcE and GlcF.) where the genes clustered as an operon. In the case of Burkholderia caribensis MBA4, three glycolate oxidase operons were identified [58], of which one gene is located downstream of the deh4a in replicon CP012747 (Fig. 4b). This operon has a downstream malate synthase gene (glcB) and an upstream regulatory gene (glcC) in the opposite strand. Another glcDEF containing an upstream glcC was discovered in replicon CP012746 with neither adjacent glcC nor glcB (Fig. 4b). Hence, the findings conveyed that glycolate oxidase could utilize glycolate in three ways [58].
In the second example, the complete genome of Pseudomonas aeruginosa N002 contained several genes involved in the degradation of alkane, alkene, aromatic hydrocarbon, other crude oil products and organohalides (including chloroalkane, chloroalkene, chlorocyclohexane and chlorobenzene) [62]. The 2-haloacid dehalogenase (had-2) at position A222_04245 was clustered with other proteins such as, alcohol dehydrogenases (frmA, Fe-adh, adhP), transcriptional regulator (lysR), and other related proteins (Fig. 5). Other pollutants-degrading enzymes were also detected.
Genome organisations of 2-haloacid dehalogenase (had-2) of Pseudomonas aeruginosa N002 in cluster with other genes. NAD-dependent aldehyde dehydrogenases (exaC), transcriptional regulator (lysR), class III alcohol dehydrogenase (frmA), predicted hydrolases (dehH), NAD-dependent aldehyde dehydrogenases (aldedh), cytochrome C (cytC), alcohol dehydrogenase, class IV (Fe-adh) and Zn-dependent alcohol dehydrogenases (adhP)
Thirdly, whole-genome analysis of Sphingobium chlorophenolicum strain L-1 by Copley et al. (2012) [99] showed that pentachlorophenol (PCP) was metabolized by three different enzymes. The dehalogenase (PcpC) is clustered with structural proteins (PcpBD, PcpA and PcpE), transcriptional regulators (PcpR and PcpM), transporter systems, and other proteins that partake in PCP hydrolysis (Fig. 6).
Conclusion
Halogenated organic compounds present a critical global environmental problem due to their toxicity and persistence. Anthropogenic activities release an enormous quantity of pollutants, and it is expected that a significant amount will remain in the environment. The problem can be overcome by using natural microbes to restore a polluted environment. Many newly discovered pollutant-degrading enzymes represent new tools for environmental biotechnology. This broad knowledge acquired has allowed us to discover numerous exclusively dehalogenase-producing bacteria, profoundly improving our understanding of different microbes' capabilities to degrade pollutants in a wide range of environments. Scientists can better comprehend the complete degradative potential, interactions, and functions of unculturable microbes. However, before using whole-genome approaches for uncovering microbial usability in bioremediation, certain areas must first be addressed, for instance, the specific gene’s structure, function, and regulation. Therefore, combining these genomic techniques with the data delivered by high-throughput technologies is now possible, capable of accelerating the discovery of enzyme regulatory bioremediation pathways (i.e., dehalogenase). Consequently, integrating these techniques with mechanistic information of bioremediation processes, the elucidation of structure–function relationship and knowledge on the microbes' regulatory pathways will provide the basis for successful biodegradation processes and leading to improved intervention strategies for bioremediation. The technology will allow a better understanding of the complete bioremediation process of organohalide pollutants.
References
Singh L, Singh VP (2015) Textile dyes degradation: a microbial approach for biodegradation of pollutants. Microbial degradation of synthetic dyes in wastewaters. Springer, Cham, pp 187–204
Alka S, Shahir S, Ibrahim N, Chai TT, Bahari ZM, Abd Manan F (2020) The role of plant growth promoting bacteria on arsenic removal: a review of existing perspectives. Environ Technol Innov 17:100602
Kumar V, Shahi SK, Singh S (2018) Bioremediation: an eco-sustainable approach for restoration of contaminated sites. Microbial bioprospecting for sustainable development. Springer, Singapore, pp 115–136
Davoodi SM, Miri S, Taheran M, Brar SK, Galvez-Cloutier R, Martel R (2020) Bioremediation of unconventional oil contaminated ecosystems under natural and assisted conditions: a review. Environ Sci Technol 54(4):2054–2067
Saldarriaga-Hernandez S, Hernandez-Vargas G, Iqbal HM, Barceló D, Parra-Saldívar R (2020) Bioremediation potential of Sargassum sp. biomass to tackle pollution in coastal ecosystems: circular economy approach. Sci Total Environ 715:136978
Jing R, Kjellerup BV (2020) Predicting the potential for organohalide respiration in wastewater: comparison of intestinal and wastewater microbiomes. Sci Total Environ 705:135833
Ghersa F, Pessah S, Vera ACD, Ferraro DO (2020) Theory and practice for environmental risk assessment understanding the trade off between the benefits and risks behind herbicide use as tool for designing sustainable weed management systems. Decision support systems for weed management. Springer, Cham, pp 161–189
Bagiu IC, Sarac I, Radu F, Bostan C, Butnariu M, Bagiu RV (2020) Ecotechnologies for persistent pollutants. Bioremediation and biotechnology. Springer, Cham, pp 105–138
Oyewusi HA, Wahab RA, Kaya Y, Edbeib MF, Huyop F (2020) Alternative bioremediation agents against haloacids, haloacetates and chlorpyrifos using novel halogen-degrading bacterial isolates from the hypersaline lake tuz. Catalysts 10(6):651
Oyewusi HA, Wahab RA, Huyop F (2020) Dehalogenase-producing halophiles and their potential role in bioremediation. Mar Pollut Bull 160:111603
Pérez-Losada M, Arenas M, Galán JC, Bracho MA, Hillung J, García-González N et al (2020) High-throughput sequencing (HTS) for the analysis of viral populations. Infect Genet Evol 80:104208
Shekhar SK, Godheja J, Modi DR (2020) Molecular technologies for assessment of bioremediation and characterization of microbial communities at pollutant-contaminated sites. Bioremediation of industrial waste for environmental safety. Springer, Singapore, pp 437–474
Oyewusi HA, Wahab RA, Huyop F (2020) Molecular docking and molecular dynamics simulation of Bacillus thuringiensis dehalogenase against haloacids haloacetates and chlorpyrifos. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1835727
Malla MA, Dubey A, Yadav S, Kumar A, Hashem A, AbdAllah EF (2018) Understanding and designing the strategies for the microbe-mediated remediation of environmental contaminants using omics approaches. Front Microbiol 9:1132
Carpenter A (2019) Oil pollution in the North Sea: the impact of governance measures on oil pollution over several decades. Hydrobiologia 845(1):109–127
Steliga T, Jakubowicz P, Kapusta P (2012) Changes in toxicity during in situ bioremediation of weathered drill wastes contaminated with petroleum hydrocarbons. Bioresour Technol 125:1–10
White E (1998) Realizing remediation: a summary of contaminated sediment remediation activities in the Great Lakes Basin: US Environmental Protection Agency, Great Lakes National Program Office
Alomar C, Sureda A, Capó X, Guijarro B, Tejada S, Deudero S (2017) Microplastic ingestion by Mullus surmuletus Linnaeus, 1758 fish and its potential for causing oxidative stress. Environ Res 159:135–142
Pannetier P, Morin B, Le Bihanic F, Dubreil L, Clérandeau C, Chouvellon F et al (2020) Environmental samples of microplastics induce significant toxic effects in fish larvae. Environ Int 134:105047
Liberti A, Bertocci I, Pollet A, Musco L, Locascio A, Ristoratore F et al (2020) An indoor study of the combined effect of industrial pollution and turbulence events on the gut environment in a marine invertebrate. Mar Environ Res 158:104950
Sharma B, Shukla P (2020) Lead bioaccumulation mediated by Bacillus cereus BPS-9 from an industrial waste contaminated site encoding heavy metal resistant genes and their transporters. J Hazard Mater 401:123285
Ferronato N, Torretta V (2019) Waste mismanagement in developing countries: a review of global issues. Int J Environ Res Public Health 16(6):1060
Gupta C, Prakash D (2020) Novel bioremediation methods in waste management: novel bioremediation methods. Waste management: concepts, methodologies, tools, and applications. IGI Global, Philadelphia, pp 1627–1643
Sharma B, Dangi AK, Shukla P (2018) Contemporary enzyme based technologies for bioremediation: a review. J Environ Manag 210:10–22
Bilal M, Iqbal HM, Barceló D (2019) Persistence of pesticides-based contaminants in the environment and their effective degradation using laccase-assisted biocatalytic systems. Sci Total Environ 695:133896
Okino-Delgado CH, Zanutto-Elgui MR, do Prado DZ, Pereira MS, Fleuri LF (2019) Enzymatic bioremediation: current status, challenges of obtaining process, and applications. Microbial metabolism of xenobiotic compounds. Springer, Singapore, pp 79–101
Mishra B, Varjani S, Agrawal DC, Mandal SK, Ngo HH, Taherzadeh MJ et al (2020) Engineering biocatalytic material for the remediation of pollutants: a comprehensive review. Environ Technol Innov 101063
Kumar SS, Ghosh P, Malyan SK, Sharma J, Kumar V (2019) A comprehensive review on enzymatic degradation of the organophosphate pesticide malathion in the environment. J Environ Sci Health Part C 37(4):288–329
Wright RJ, Gabriel E, Vinko Z, Mira L, Joseph AC (2020) Marine plastic debris: a new surface for microbial colonization. Environ Sci Technol 54(19):11657–11672
Arora PK (2020) Bacilli-mediated degradation of xenobiotic compounds and heavy metals. Front Bioeng Biotechnol 8:1100
Adamu A, Wahab RA, Aliyu F, Aminu AH, Hamza MM, Huyop F (2020) Haloacid dehalogenases of Rhizobium sp. and related enzymes: catalytic properties and mechanistic analysis. Process Biochem 92:437–446
Rappé MS, Giovannoni SJ (2003) The uncultured microbial majority. Annu Rev Microbiol 57(1):369–394
Sekar R, Fuchs BM, Amann R, Pernthaler J (2004) Flow sorting of marine bacterioplankton after fluorescence in situ hybridization. Appl Environ Microbiol 70(10):6210–6219
Podar M, Abulencia CB, Walcher M, Hutchison D, Zengler K, Garcia JA et al (2007) Targeted access to the genomes of low-abundance organisms in complex microbial communities. Appl Environ Microbiol 73(10):3205–3214
Alneberg J, Karlsson CM, Divne AM, Bergin C, Homa F, Lindh MV et al (2018) Genomes from uncultivated prokaryotes: a comparison of metagenome-assembled and single-amplified genomes. Microbiome 6(1):173
Parks DH, Rinke C, Chuvochina M, Chaumeil P-A, Woodcroft BJ, Evans PN et al (2017) Recovery of nearly 8000 metagenome-assembled genomes substantially expands the tree of life. Nat Microbiol 2(11):1533–1542
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A et al (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6(3):610
Donia MS, Fischbach MA (2015) Small molecules from the human microbiota. Science 349(6246):1254766
Mallick H, Franzosa EA, Mclver LJ, Banerjee S, Sirota-Madi A, Kostic AD et al (2019) Predictive metabolomic profiling of microbial communities using amplicon or metagenomic sequences. Nat Commun 10(1):1–11
Wixon J, Kell D (2000) The kyoto encyclopedia of genes and genomes–KEGG. Yeast (Chichester, England) 17(1):48–55
Caspi R, Billington R, Ferrer L, Foerster H, Fulcher CA, Keseler IM et al (2016) The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res 44(D1):D471–D480
Imchen M, Kumavath R, Barh D, Azevedo V, Ghosh P, Viana M et al (2017) Searching for signatures across microbial communities: metagenomic analysis of soil samples from mangrove and other ecosystems. Sci Rep 7(1):1–13
Sharpton TJ (2014) An introduction to the analysis of shotgun metagenomic data. Front Plant Sci 5:209
Huerta-Cepas J, Szklarczyk D, Forslund K, Cook H, Heller D, Walter MC et al (2016) eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic prokaryotic and viral sequences. Nucleic Acids Res 44(D1):D286–D293
Liu J, Wang H, Yang H, Zhang Y, Wang J, Zhao F et al (2013) Composition-based classification of short metagenomic sequences elucidates the landscapes of taxonomic and functional enrichment of microorganisms. Nucleic Acids Res 41(1):e3–e3
Zhang Y, Ji P, Wang J, Zhao F (2016) RiboFR-Seq: a novel approach to linking 16S rRNA amplicon profiles to metagenomes. Nucleic Acids Res 44(10):e99–e99
Alves LDF, Westmann CA, Lovate GL, de Siqueira GMV, Borelli TC, Guazzaroni ME (2018) Metagenomic approaches for understanding new concepts in microbial science. Int J Genomics. https://doi.org/10.1155/2018/2312987
Mukherjee A, Chettri B, Langpoklakpam JS, Basak P, Prasad A, Mukherjee AK et al (2017) Bioinformatic approaches including predictive metagenomic profiling reveal characteristics of bacterial response to petroleum hydrocarbon contamination in diverse environments. Sci Rep 7(1):1–22
Labonté JM, Swan BK, Poulos B, Luo H, Koren S, Hallam SJ et al (2015) Single-cell genomics-based analysis of virus–host interactions in marine surface bacterioplankton. ISME J 9(11):2386–2399
Props R, Kerckhof F-M, Rubbens P, De Vrieze J, Sanabria EH, Waegeman W et al (2017) Absolute quantification of microbial taxon abundances. ISME J 11(2):584–587
Suenaga H, Koyama Y, Miyakoshi M, Miyazaki R, Yano H, Sota M et al (2009) Novel organization of aromatic degradation pathway genes in a microbial community as revealed by metagenomic analysis. ISME J 3(12):1335–1348
Imperato V, Portillo-Estrada M, McAmmond BM, Douwen Y, Van Hamme JD, Gawronski SW et al (2019) Genomic diversity of two hydrocarbon-degrading and plant growth-promoting pseudomonas species isolated from the oil field of bóbrka (Poland). Genes 10(6):443
Chemerys A, Pelletier E, Cruaud C, Martin F, Violet F, Jouanneau Y (2014) Characterization of novel polycyclic aromatic hydrocarbon dioxygenases from the bacterial metagenomic DNA of a contaminated soil. Appl Environ Microbiol 80(21):6591–6600
Silva CC, Hayden H, Sawbridge T, Mele P, De Paula SO, Silva LC et al (2013) Identification of genes and pathways related to phenol degradation in metagenomic libraries from petroleum refinery wastewater. PLoS ONE 8(4):e61811
Iwai S, Chai B, Sul WJ, Cole JR, Hashsham SA, Tiedje JM (2010) Gene-targeted-metagenomics reveals extensive diversity of aromatic dioxygenase genes in the environment. ISME J 4(2):279–285
Oyewusi HA, Abdul Wahab R, Edbeib MF, Mohamad MAN, Abdul Hamid AA, Kaya Y et al (2021) Functional profiling of bacterial communities in Lake Tuz using 16S rRNA gene sequences. Biotechnol Biotechnol Equip 35(1):1–10
Weigold P, El-Hadidi M, Ruecker A, Huson DH, Scholten T, Jochmann M et al (2016) Metagenomic-based survey of microbial (de) halogenation potential in a German forest soil. Sci Rep 6(1):1–13
Pan Y, Kong KF, Tsang JS (2015) Complete genome sequence and characterization of the haloacid–degrading Burkholderia caribensis MBA4. Stand Genomic Sci 10(1):114
Hug LA, Edwards EA (2013) Diversity of reductive dehalogenase genes from environmental samples and enrichment cultures identified with degenerate primer PCR screens. Front Microbiol 4:341
Sulaiman S, Yamato S, Kanaya E, Kim JJ, Koga Y, Takano K et al (2012) Isolation of a novel cutinase homolog with polyethylene terephthalate-degrading activity from leaf-branch compost by using a metagenomic approach. Appl Environ Microbiol 78(5):1556–1562
Ferrer M, Golyshina OV, Chernikova TN, Khachane AN, Reyes-Duarte D, Santos VAMD et al (2005) Novel hydrolase diversity retrieved from a metagenome library of bovine rumen microflora. Environ Microbiol 7(12):1996–2010
Das D, Baruah R, Roy AS, Singh AK, Boruah HPD, Kalita J et al (2015) Complete genome sequence analysis of Pseudomonas aeruginosa N002 reveals its genetic adaptation for crude oil degradation. Genomics 105(3):182–190
Elufisan TO, Lozano L, Bustos P, Rodríguez-Luna IC, Sánchez-Varela A, Oyedara OO et al (2019) Complete genome sequence of stenotrophomonas maltophilia strain SVIA2, isolated from crude oil-contaminated soil in Tabasco, Mexico. Microbiol Resour Announc 8(30):e00529-e619
Delmont TO, Eren AM, Maccario L, Prestat E, Esen ÖC, Pelletier E et al (2015) Reconstructing rare soil microbial genomes using in situ enrichments and metagenomics. Front Microbiol 6:358
Kantor RS, Wrighton KC, Handley KM, Sharon I, Hug LA, Castelle CJ et al (2013) Small genomes and sparse metabolisms of sediment-associated bacteria from four candidate phyla. MBio 4(5):e00708-e713
Nkongolo KK, Narendrula-Kotha R (2020) Advances in monitoring soil microbial community dynamic and function. J Appl Genet 61(2):249–263
Rath CM, Janto B, Earl J, Ahmed A, Hu FZ, Hiller L et al (2011) Meta-omic characterization of the marine invertebrate microbial consortium that produces the chemotherapeutic natural product ET-743. ACS Chem Biol 6(11):1244–1256
Bargiela R, Mapelli F, Rojo D, Chouaia B, Tornés J, Borin S et al (2015) Bacterial population and biodegradation potential in chronically crude oil-contaminated marine sediments are strongly linked to temperature. Sci Rep 5:11651
Korlević M, Zucko J, Dragić MN, Blažina M, Pustijanac E, Zeljko TV et al (2015) Bacterial diversity of polluted surface sediments in the northern Adriatic Sea. Syst Appl Microbiol 38(3):189–197
Oh S, Tandukar M, Pavlostathis SG, Chain PS, Konstantinidis KT (2013) Microbial community adaptation to quaternary ammonium biocides as revealed by metagenomics. Environ Microbiol 15(10):2850–2864
Fonti V, Beolchini F, Rocchetti L, Dell’Anno A (2015) Bioremediation of contaminated marine sediments can enhance metal mobility due to changes of bacterial diversity. Water Res 68:637–650
Si D, Xiong Y, Li X, Ma L, Deng X, Wang Y et al (2017) Isolation and genome sequence analysis of a bacterium degrading dexamethasone. Biomed Res 28(11):976–1683
Chauhan A, Pathak A, Ewida AY, Griffiths Z, Stothard P (2016) Whole genome sequence analysis of an alachlor and endosulfan degrading pseudomonas strain W15Feb9B isolated from ochlockonee River, Florida. Genom Data 8:134–138
Tabata M, Ohhata S, Kawasumi T, Nikawadori Y, Kishida K, Sato T et al (2016) Complete genome sequence of a γ-hexachlorocyclohexane degrader, Sphingobium sp. strain TKS, isolated from a γ-hexachlorocyclohexane-degrading microbial community. Genome Announc. https://doi.org/10.1128/genomeA.00247-16
Tabata M, Ohhata S, Nikawadori Y, Kishida K, Sato T, Kawasumi T et al (2016) Comparison of the complete genome sequences of four γ-hexachlorocyclohexane-degrading bacterial strains: insights into the evolution of bacteria able to degrade a recalcitrant man-made pesticide. DNA Res 23(6):581–599
Zhang C, Hao Q, Zhang Z, Zhang X, Pan H, Zhang J et al (2019) Whole genome sequencing and analysis of chlorimuron-ethyl degrading bacteria klebsiella pneumoniae 2N3. Int J Mol Sci 20(12):3053
Lee Y, Lee Y, Jeon CO (2019) Biodegradation of naphthalene, BTEX, and aliphatic hydrocarbons by Paraburkholderia aromaticivorans BN5 isolated from petroleum-contaminated soil. Sci Rep 9(1):1–13
Qiu Q, Wang P, Kang H, Wang Y, Tian K, Huo H (2019) Genomic analysis of a new estrogen-degrading bacterial strain Acinetobacter sp. DSSKY-A-001. Int J Genom 2019:1–13
Yao Y, Tang H, Su F, Xu P (2015) Comparative genome analysis reveals the molecular basis of nicotine degradation and survival capacities of arthrobacter. Sci Rep 5:8642
Binnewies TT, Motro Y, Hallin PF, Lund O, Dunn D, La T et al (2006) Ten years of bacterial genome sequencing: comparative-genomics-based discoveries. Funct Integr Genomics 6(3):165–185
Adamu A, Wahab RA, Huyop F (2016) l-2-Haloacid dehalogenase (DehL) from Rhizobium sp. RC1. SpringerPlus 5(1):695
Weng X, Xiao J (2014) Spatial organization of transcription in bacterial cells. Trends Genet 30(7):287–297
Ray S, Gunzburg MJ, Wilce M, Panjikar S, Anand R (2016) Structural basis of selective aromatic pollutant sensing by the effector binding domain of MopR, an NtrC family transcriptional regulator. ACS Chem Biol 11(8):2357–2365
Rivera-Gómez N, Segovia L, Pérez-Rueda E (2011) Diversity and distribution of transcription factors: their partner domains play an important role in regulatory plasticity in bacteria. Microbiology 157(8):2308–2318
Pop SM, Kolarik RJ, Ragsdale SW (2004) Regulation of anaerobic dehalorespiration by the transcriptional activator CprK. J Biol Chem 279(48):49910–49918
Yang R, Liu G, Chen T, Li S, An L, Zhang G et al (2019) Characterization of the genome of a nocardia strain isolated from soils in the Qinghai-Tibetan plateau that specifically degrades crude oil and of this biodegradation. Genomics 111(3):356–366
Su X, Tsang JS (2013) Existence of a robust haloacid transport system in a Burkholderia species bacterium. Biochim Biophys Acta (BBA) 1828(2):187–192
Bringel F, Besaury L, Amato P, Kröber E, Kolb S, Keppler F et al (2019) Methylotrophs and methylotroph populations for chloromethane degradation. Curr Issues Mol Biol 33:149–172
Zang H, Wang H, Miao L, Cheng Y, Zhang Y, Liu Y et al (2020) Carboxylesterase, a de-esterification enzyme, catalyzes the degradation of chlorimuron-ethyl in Rhodococcus erythropolis D310-1. J Hazard Mater. https://doi.org/10.1016/j.jhazmat.2019.121684
Pathak A, Chauhan A, Ewida AY, Stothard P (2016) Whole genome sequence analysis of an Alachlor and Endosulfan degrading Micrococcus sp. strain 2385 isolated from Ochlockonee River, Florida. J Genomics 4:42–47
Pan X, Xu T, Xu H, Fang H, Yu Y (2017) Characterization and genome functional analysis of the DDT-degrading bacterium Ochrobactrum sp. DDT-2. Sci Total Environ 592:593–599
Pan X, Lin D, Zheng Y, Zhang Q, Yin Y, Cai L et al (2016) Biodegradation of DDT by Stenotrophomonas sp. DDT-1: characterization and genome functional analysis. Sci Rep 6(1):21332
Ohtsubo Y, Sato T, Kishida K, Tabata M, Ogura Y, Hayashi T et al (2014) Complete genome sequence of Pseudomonas aeruginosa MTB-1, isolated from a microbial community enriched by the technical formulation of hexachlorocyclohexane. Genome Announc 2(1):e01130-e1213
Nagata Y, Ohtsubo Y, Endo R, Ichikawa N, Ankai A, Oguchi A et al (2010) Complete genome sequence of the representative γ-hexachlorocyclohexane-degrading bacterium Sphingobium japonicum UT26. J Bacteriol 192(21):5852–5853
Kaur J, Verma H, Tripathi C, Khurana J, Lal R (2013) Draft genome sequence of a hexachlorocyclohexane-degrading bacterium, Sphingobium baderi strain LL03T. Genome Announc 1(5):e00751-e813
Saxena A, Nayyar N, Sangwan N, Kumari R, Khurana J, Lal R (2013) Genome sequence of Novosphingobium lindaniclasticum LE124T, isolated from a hexachlorocyclohexane dumpsite. Genome Announc 1(5):e00715-e813
Negi V, Lata P, Sangwan N, Gupta SK, Das S, Rao D et al (2014) Draft genome sequence of hexachlorohexane (HCH)-degrading Sphingobium lucknowense strain F2T, isolated from an HCH dumpsite. Genome Announc 2(4):e00788-e814
Chaussonnerie S, Saaidi P-L, Ugarte E, Barbance A, Fossey A, Barbe V et al (2016) Microbial degradation of a recalcitrant pesticide: chlordecone. Front Microbiol 7:2025
Copley SD, Rokicki J, Turner P, Daligault H, Nolan M, Land M (2012) The whole genome sequence of Sphingobium chlorophenolicum L-1: insights into the evolution of the pentachlorophenol degradation pathway. Genome Biol Evol 4(2):184–198
Azam S, Parthasarathy S, Singh C, Kumar S, Siddavattam D (2019) Genome organization and adaptive potential of archetypal organophosphate degrading Sphingobium fuliginis ATCC 27551. Genome Biol Evol 11(9):2557–2562
Giani AM, Gallo GR, Gianfranceschi L, Formenti G (2020) Long walk to genomics: history and current approaches to genome sequencing and assembly. Comput Struct Biotechnol 18:9–19
Acknowledgements
The authors would like to express their gratitude to the Department of Biosciences, Faculty of Science, Universiti Teknologi Malaysia, for their facilities. This work was funded by Fundamental Research Grant Scheme (FRGS)-Ministry of Education RJ130000.7854.5F189 (FRGS/1/2019/STG05/UTM/01/1) for financial assistance and the Ministry of Education through Tertiary Education Trust Fund (TETFund) Nigeria for the studentship.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Oyewusi, H.A., Wahab, R.A. & Huyop, F. Whole genome strategies and bioremediation insight into dehalogenase-producing bacteria. Mol Biol Rep 48, 2687–2701 (2021). https://doi.org/10.1007/s11033-021-06239-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06239-7