Abstract
Ribosomal protein S3 (RPS3) is a component of the 40S ribosomal subunit. It is known to function in ribosome biogenesis and as an endonuclease. RPS3 has been shown to be over expressed in colon adenocarcinoma but its role in colon cancer is still unknown. In this study, we aim at determining the expression levels of RPS3 in a colon cancer cell line Caco-2 compared to a normal colon mucosa cell line NCM-460 and study the effects of targeting this protein by siRNA on cellular behavior. RPS3 was found to be expressed in both cell lines. However, siRNA treatment showed a more protruding effect on Caco-2 cells compared to NCM-460 cells. RPS3 knockdown led to a significant decrease in the proliferation, survival, migration and invasion and an increase in the apoptosis of Caco-2 cells. Western blot analysis demonstrated that these effects correlated with an increase in the level of the tumor suppressor p53 and a decrease in the level and activity of lactate dehydrogenase (LDH), an enzyme involved in the metabolism of cancer cells. No significant effect was shown in normal colon NCM-460 cells. Targeting p53 by siRNA did not affect RPS3 levels indicating that p53 may be a downstream target of RPS3. However, the concurrent knockdown of RPS3 and p53 showed no change in LDH level in Caco-2 cells suggesting an interesting interplay among the three proteins. These findings might present RPS3 as a selective molecular marker in colon cancer and an attractive target for colon cancer therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ribosomal protein S3 (RPS3) is a component of the small ribosomal subunit 40S. It is involved in the 40S ribosomal maturation and initiation of translation through the interaction with various initiation factors like eIF3 and eIF2 [1]. RPS3 plays a role in DNA damage repair due to the presence of its amino acid sequence that is identical to an apurinic/apyrimidinic endonuclease II [2]. In addition, RPS3 has been shown to have other extraribosomal functions including control of apoptosis, survival, transcription and tumorigenesis [3]. RPS3 is reported to be associated with p53 and MDM2 where its knockdown decreases the high level of p53 when the HEK 293 cells are under oxidative stress [4]. Furthermore, RPS3 silencing promotes ribosomal stress which impairs ribosomal biogenesis and by its turn induces the activation of c-Myc and p38 leading to p53 induction and G1 cell cycle arrest in HT1080 fibrosarcoma cells [5].
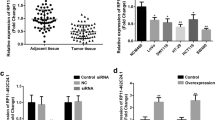
The level of mRNA corresponding to RPS3 is shown to be higher in colorectal tumor cells and adenomatous polyps, compared to normal colon cell mucosa [6]. Colon adenomatous polyps are the precursors of adenocarcinoma and colon cancer. This may indicate that RPS3 may be playing a role in the early stage of colon cancer formation. Colon adenocarcinoma cells Caco-2 has similar level of expression of RPS3 mRNA compared to the colon tumor cells, but higher than normal mucosa cells [6].
Colon cancer is a major cause of death in the world accounting for around 9% of cancer incidence [7,8,9]. It usually starts as small adenomatous colon polyps, small benign clump of cells that if left untreated may develop to become adenocarcinomas. Environmental and genetic factors play a significant role in the conversion of adenomatous polyps to carcinoma by building up mutations in certain genes that are responsible for the shifting from a normal proliferating epithelium to a rapid proliferating colon mucosa. The benign adenomatous polyps formed can become life threatening, invasive carcinomas in around 10 years [10].
The metabolism of cancer cells, including colon cancer, is altered compared to normal cells. Instead of cellular respiration, cancer cells depend on another metabolic pathway to produce energy through aerobic glycolysis even in the presence of oxygen, a process known as the Warburg effect [11, 12]. During this process, cancer cells consume large quantity of glucose and produce a high level of lactic acid due to the action of lactate dehydrogenase (LDH), an enzyme which converts pyruvate to lactic acid in glycolysis. Even with the high level of lactic acid produced, cancer cells can survive because they become resistant to the microenvironmental acidosis [11,12,13]. The Warburg effect inhibition is known to be dependent, in part, on the p53 tumor suppressor protein status in the cells [14], where LDH-A is identified as a potential cancer therapeutic target that works through both p53-dependent and independent pathways [15].
In this study, we show that RPS3 knockdown (KD) increases p53 level in colon cancer cells which correlates with an increase in apoptosis while decreasing LDH level and activity. Colon cancer cell proliferation, migration and invasion are decreased upon RPS3 KD with no significant effect on normal colon cells.
Materials and methods
Cell culture and transfection
Human colon adenocarcinoma cells Caco-2 and normal colon mucosa cells NCM-460 were used. The latter were a kind gift from Dr. Marwan El Sabban from the American University of Beirut. Caco-2 cells were grown in Dulbecco’s Modified Eagle’s Medium (Gibco) and NCM-460 cells were grown in M3: base F medium (InCell). Both media were supplemented with 1% l-glutamine, 1% penicillin/streptomycin, and 10% fetal bovine serum. Cells were incubated at 37 ºC and a 5% CO2 humid environment. 2.0 × 104 cells were plated in a 6-well plate and transfected with siRNA (Qiagen) against RPS3 (siRPS3) alone, p53 (sip53) alone, or both (siRPS3/p53) using hiperfect transfection reagent (Qiagen), according to manufacturer’s recommendations. Transfection with hiperfect alone, in the absence of siRNA, was used as a negative control.
Western blot analysis
Proteins were lysed at 48 h after transfection and separated using 15% sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis then transferred to a PVDF membrane that was blocked with 5% BSA. Primary anti-RPS3 (Abcam, Cat # 1,811,992), anti-p53 (Abcam, Cat # ab9484), anti-LDH (Abcam, Cat # ab134187), anti-GAPDH (Abcam, Cat # 9484) and anti-Actin (Abcam, Cat # ab8229) antibodies were diluted in 5% BSA (1/1000 in ratio), then incubated with the membrane overnight at 4 ºC. The membrane was washed, incubated with the appropriate secondary antibodies for 1 h then washed and incubated with ECL substrate (GE Healthcare). The bands were visualized and quantified using the Bio-Rad© Chemidoc system and normalized to GAPDH or actin controls.
WST-1 proliferation assay
Using a 96 well microtiter plate, 3.0 × 105 of cells were plated in separate wells in triplicates in 100 µl of medium and subjected to siRPS3 transfection. 24, 48, 72, 96 and 120 h post-transfection, 10 µl of WST1 reagent (Abcam) was added to each well, incubated for 3 h and the absorbance was measured at 450 nm.
Migration and invasion assays
24 h after transfection, 106 cells/ml were prepared in chemoattractant-free media and plated in inserts in 24 well plates using the QCM chemotaxis cell migration assay (EMD Millipore) and QCM collagen cell invasion assay (EMD Millipore), following the manufacturer’s recommendations. Cells were incubated for 24 h and 72 h to detect migration and invasion, respectively at 37 °C with 5% CO2. Cells that have migrated or invaded into the wells were then fixed, stained and extracted. 100 µl of the stained mixture was transferred to a 96-well plate and the absorbance was measured at 560 nm.
Apoptosis assay
2.5 × 105 cells were collected 48 h after siRPS3 transfection and washed with phosphate buffer saline (PBS) buffer. Apoptosis was detected by staining the cells with Annexin V and propidium iodide (PI) using the Annexin V/PI apoptosis kit (Abcam), according to the manufacturer’s recommendations. Quantification of apoptosis and survival was done by flow cytometry.
LDH activity assay
The LDH cytotoxicity assay kit (Abcam) was used according to the manufacturer’s recommendations. Various NADH standard dilutions were prepared to obtain concentrations ranging between 0 and 12.5 nmol. 1–2 × 106 of cells were harvested at 48 h after transfection, washed with cold PBS and homogenized on ice with cold assay buffer followed by centrifugation. The supernatant was then collected and kept on ice. 50 µl of the standard, positive control and sample dilutions were added in a 96-well plate. Then, 50 µl of the reaction mix (2 µl of substrate mix with 8 µl of LDH assay buffer) was added to the wells and mixed. The absorbance was measured at room temperature using a microplate reader at 450 nm every 2 min for 10 min. The NADH standard curve was used to calculate the LDH activity.
Statistical analysis
All experiments were performed in triplicate for at least three independent times and expressed by mean ± standard deviation. All statistical analyses were based on the student’s t-test distribution and the significance level was set at p value < 0.05.
Results
RPS3 knockdown significantly decreases proliferation and metastatic potential in Caco-2 cells, with no effect on NCM-460 cells
We measured the level of RPS3 expression in Caco-2 and NCM-460 cells by western blot and showed high expression of the protein in both cell lines. We then successfully targeted RPS3 by siRNA as shown by western blot analysis and studied the effect of this KD on various cell behaviors. GAPDH antibody was used as a loading control (Fig. 1a).
RPS3 downregulation dampens the proliferation, invasion and migration of Caco-2 cells (a). Western Blot Analysis of the expression of RPS3 upon RPS3 KD in Caco-2 (left) and NCM-460 (right) cells (b, c). WST-1 Assay for Caco2 (b) and NCM-460 cells (c) was performed for five consecutive days’ post-transfection to measure the rate of cellular proliferation (d, e). Quantitation of cell migration (d) and matrigel invasion (e) upon RPS3 KD in Caco-2 cells. All experiments were done in triplicate and repeated at least three independent times. *p = 0.04, bars denote standard error (SEM)
To evaluate the effect of this transfection on proliferation, the WST-1 cell proliferation assay was used. Caco-2 cells showed higher proliferation rates (OD above 0.2) compared to the normal NCM-460 cell line (OD below 0.2). We observed a statistically significant reduction in the proliferation rate of Caco-2 cells in siRPS3-transfected cells compared to control cells, mainly at 48 and 120 (p = 0.04) hours with a 1.29 and 1.84 fold decrease, respectively (Fig. 1b). For NCM-460 cells, there was no significant difference between siRPS3-transfected and control cells (Fig. 1c). These results might indicate that RPS3 is involved in the proliferative mechanisms specific for cancer cells.
In addition, RPS3 KD in Caco-2 cells was able to significantly impair both the migration and invasion ability of those cells compared to the control. siRPS3-transfected cells presented a statistically significant twofold decrease in their migration ability compared to the control cells, with a p-value of 0.04 (Fig. 1d). Similarly, siRPS3-transfected cells that have invaded through the Matrigel-coated insert significantly decreased in comparison with the control cells with a p-value of 0.04 (Fig. 1e). In fact, RPS3 KD decreased invasion at 48 h by 1.175-fold (data not shown) but the highest and most significant difference was observed at 72 h (2.45-fold decrease; Fig. 1e).
RPS3 knockdown significantly increases apoptotic levels selectively in Caco-2 cells
Evasion of apoptosis is one of hallmarks of cancer. In order to study the effect of RPS3 KD on this process, cells were stained with Annexin V and PI and apoptotic versus viability levels were measured by flow cytometry. The results showed that KD of RPS3 in Caco-2 cells significantly decreased cell viability from an average of 94.5% in control cells to 80.1% in siRPS3-transfected cells (p < 0.05). The late apoptosis level significantly increased from an average of 3.1% in control cells to 13% in siRPS3-transfected cells (p < 0.05; Fig. 2a, c). However, the KD of RPS3 in NCM-460 cells did not lead to noticeable change in the viability or apoptosis level between the control and the siRPS3-transfected cells (Fig. 2b, d). These results indicate that targeting RPS3 might hinder viability and induce apoptosis specifically in cancer cells with no major effect on normal cell mucosa.
Apoptosis Assay of Caco-2 and NCM-460 cells upon RPS3 KD (a, b). Representative figures of the flow cytometry analysis of apoptosis levels as measured by Annexin V/PI staining in Caco-2 (a) and NCM-460 (b) cells upon RPS3 KD (c, d). The Percentage of late apoptotic cells (left panel) and viability Level (right panel) of Caco-2 (c) and NCM-460 (d) cells upon RPS3 KD. All Experiments were done in triplicate and repeated at least three independent times. *p < 0.05, bars denote SEM
RPS3 knockdown increases p53 level while decreasing LDH protein level and activity selectively in Caco-2 cells
To understand the molecular pathways involved in the effects of RPS3 KD on colon cancer cells, we measured the level of p53 and LDH. The KD of RPS3 by around 3.9‐folds in Caco-2 cells led to around 1.95‐folds increase in p53 level and around 3‐folds decrease in the level of LDH compared to the control cells as verified by western blot analysis (Fig. 3a). However, the KD of RPS3 by around 3.23‐folds in NCM-460 did not lead to a noticeable change in the p53 or LDH levels between the control and the siRPS3-transfected NCM-460 cells (Fig. 3b). GAPDH and actin were used as loading controls and to quantitate the relative band intensities between the siRPS3-transfected and control cells.
RPS3 downregulation increases p53 level while decreasing LDH level and activity in Caco-2 cells but not in NCM-460 cells (a, b). Western Blot Analysis of the expression of RPS3, p53 and LDH levels upon RPS3 KD in Caco-2 (a) and NCM-460 (b) cells at 48 h post-transfection. Quantitation of the relative intensity of the bands is provided below each panel after normalization to either GAPDH (a) or actin (b) and was compiled from three different experiments (c, d). The LDH activity measured in Caco-2 (c) and NCM-460 (d) cells upon RPS3 KD. All experiments were done in triplicate and repeated at least three independent times. *p < 0.05, bars denote SEM
Since we observed a decrease in the level of LDH protein upon RPS3 KD in Caco-2 cells, we next looked at the level of LDH activity using LDH assay kit. The results showed that the initial LDH activity was higher in Caco-2 cells reaching 212.68 mU/ml (T = 10 min) compared to NCM-460 cells with an LDH activity of around 113.24 mU/ml (T = 10 min; Fig. 3c, d). Upon RPS3 KD, we had a significant decrease in the LDH activity level in siRPS3-transfected Caco-2 cells compared to the control cells at T = 2, 4, 6, 8 and 10 min with an average of 27% fold decrease and p < 0.05 (Fig. 1c). No significant difference in LDH activity was observed in NCM-460 cells upon RPS3 KD (Fig. 1d). Thus, the effect of RPS3 KD on p53 and LDH seems to be cell-type dependent, selectively affecting cancer cells.
p53 knockdown has no effect on RPS3 and LDH levels while the Double knockdown of RPS3 and p53 compensates for the decrease in LDH level
In order to understand the interplay among the different proteins tested, we targeted p53 by siRNA in both cell lines and measured the level of RPS3 and LDH. The KD of p53 had no significant effect on the level of the two proteins in both cells compared to control cells, although p53 levels were decreased by around 3.5‐folds in Caco-2 cells and 3‐folds in NCM-460 cells (Fig. 4a, b). This demonstrated that in Caco-2 cells, RPS3 KD affected p53, but p53 KD had no effect on RPS3 or LDH level.
Representative figures of western blot analysis of RPS3, p53 and LDH levels. The expression was analysed upon p53 KD in Caco-2 (a) and NCM-460 (b) cells at 48 h post-transfection. The level of expression was also analysed upon the concurrent KD of both RPS3 and p53 in Caco-2 (c) and NCM-460 (d) cells at 48 h post-transfection. Quantitation of the relative intensity of the bands is provided below each panel after normalization to either GAPDH (a) or actin (b–d) and was compiled from three different experiments
To further elucidate this interaction, we aimed at concurrently targeting both RPS3 and p53 by siRNA transfections and measuring the level of LDH. Western blot analysis showed that double KD of RPS3 (around threefold decrease in both cell lines) and p53 (around 2.75-fold decrease in Caco-2 cells and 2.6 in NCM-460 cells) did not lead to a significant fold change in LDH between the control and the transfected cells in either cell lines (Fig. 4c, d). These observations suggested that the increased level of p53 upon RPS3 KD might be responsible for the decrease in LDH level in Caco-2 cells.
Discussion
Although RPS3 has a known role as an endonuclease involved in DNA repair function [2], its role in tumorigenesis is still poorly understood. In human tumors, RPS3 has been reported to be highly expressed in colorectal cancer [6], melanoma [16] and hepatocellular carcinoma [17]. In the present study, we aimed at understanding the role played by RPS3 in colon cancer compared to normal colon mucosa cells through transcriptionally targeting the protein by siRNA transfection and studying the effect of this downregulation on cellular behaviors.
We show that RPS3 KD significantly dampened proliferation, viability, migration and invasion while increasing apoptosis in Caco-2 cells compared to control cells, with no significant effect on NCM-460 cells. Noteworthy, the proliferation rate of NCM-460 cells was slower than that of Caco-2 cells which was predictable since NCM-460 is assumed to be a normal cell line. Previous studies reported the involvement of RPS3 in modulating cellular behaviors leading to tumorigenesis. RPS3 KD inhibited melanoma [18], acute lymphoblastic leukemia [19] and hepatocellular carcinoma [20] cell growth in vitro and in xenograft mouse models. In NIH-3T3 fibroblasts, the inhibition of enhanced RPS3 expression induced apoptosis and reversed cell transformation [21]. In contrast, in irradiated glioblastoma cells, RPS3 was reported as a pro-apoptotic protein via the interaction with DNA damage-inducible transcript 3 (DDIT3) [22]. In Addition, silencing of RPS3 inhibited invasion and migration in osteosarcoma [23] and hepatocellular carcinoma cells [20] while RPS3 expression suppressed migration and invasion in fibrosarcoma through interaction with the tumor suppressor nm21 by blocking the ERK pathway and MMP9 [24]. These data imply that RPS3 effect on cancer growth and progression depends on the tumor type.
One of the major pathways related to induction of apoptosis and RPs alteration is p53, known to be activated upon ribosomal stress leading to a disruption in protein synthesis. In the absence of molecular and physiological stresses, the MDM2 protein binds p53 targeting the latter for degradation by ubiquitination [25]. RPS3 can modulate the MDM2-p53 pathway through the interaction with both proteins. In HT1080, the knockdown of RPS3 led to ribosomal stress and the activation of two cellular pathways, the ASK1/p38 and the c-Myc, which induced an increase in the level of p53 and cell cycle arrest at G1 [5]. Similarly, in this work, RPS3 KD in Caco-2 cells led to an increase in p53 level. However, no notable change in NCM-460 was observed, proving that the effect of RPS3 on the level of p53 is colon cancer-cell specific. We suggest that the observed increase in apoptosis could be linked to the increase in p53 level upon RPS3 KD, but this should be further elucidated in the future. It is important to note that p53 is mutated in Caco-2 cells [26] while NCM-460 cells carry a wild-type p53 allele [27] which might also be the factor that is contributing to the different results we observed in the two cell lines. In order to elaborate more on the interaction between RPS3 and p53, more work will be conducted to reveal the level of p53 mRNA, the stability of p53 and the level of MDM-2 in both cells upon RPS3 KD.
The upregulation of p53 correlated with a significant decrease in the protein level and the activity of LDH, with no significant effect on normal cells. The basal LDH activity level of Caco-2 cells was higher than that of NCM-460 cells, consistent with the fact that cancer cells have an increased level of LDH activity due to the increase in Warburg effect [12]. The KD of p53 had no significant effect on the level of RPS3 and LDH in both cells, indicating that p53 is probably a downstream target of RPS3. However, double KD of RPS3 and p53 in both cells did not lead to a significant fold change in LDH between the control and the transfected cells. This supports the possibility that the increased level of p53 upon RPS3 KD in Caco-2 cells is responsible, at least in part, for the decrease in the level of LDH. In the future, we plan to further elucidate the effect of this KD on cellular behavior and to find the molecular pathways involved since previous studies have proved that an increase in the level of p53 in the cell can affect cancer cell metabolism, including an inhibition of the Warburg effect and the level of LDH [28, 29]. It is noteworthy to mention that the LDH enzyme plays a vital role in cancer cell survival, where silencing of LDHA in cancer cells can decrease proliferation and tumorigenic potential in hypoxic condition as well as increase in mitochondrial oxidative phosphorylation [30]. Downregulation of LDHA led to oxidative stress and decreased ATP production leading to an increase in cell death in lymphoma and pancreatic cancer [31]. Targeting LDHA may be a great therapeutic option in treatment of various cancer types [32], including breast [33], pancreatic [34], glioblastoma [35] and colorectal cancer [36]. Thus, the decrease in Caco-2 cellular growth upon RPS3 KD might be in part, linked to the inhibition of the Warburg effect by the decrease in the level and activity of the LDH enzyme.
This study shows an important interplay between RPS3, p53 and LDH selectively in colon cancer cells. This effect was associated with a decrease in proliferation, survival, migration and invasion while increasing apoptotic levels. Interestingly, NCM-460 cells were not affected by the downregulation of RPS3. These observations suggest that RPS3 is specifically affecting cancer cell growth and progression with minimal effects on normal cells making it a possible selective target for colon cancer therapy.
References
Henras AK, Soudet J, Gerus M, Lebaron S, Caizergues-Ferrer M, Mougin A, Henry Y (2008) The post-transcriptional steps of eukaryotic ribosome biogenesis. Cell Mol Life Sci 65(15):2334–2359. https://doi.org/10.1007/s00018-008-8027-0
Kim J, Chubatsu LS, Admon A, Stahl J, Fellous R, Linn S (1995) Implication of mammalian ribosomal protein S3 in the processing of DNA damage. J Biol Chem 270(23):13620–13629
Zhou X, Liao WJ, Liao JM, Liao P, Lu H (2015) Ribosomal proteins: functions beyond the ribosome. J Mol Cell Biol 7(2):92–104. https://doi.org/10.1093/jmcb/mjv014
Yadavilli S, Mayo LD, Higgins M, Lain S, Hegde V, Deutsch WA (2009) Ribosomal protein S3: a multi-functional protein that interacts with both p53 and MDM2 through its KH domain. DNA Repair 8(10):1215–1224. https://doi.org/10.1016/j.dnarep.2009.07.003
Kim HD, Kim TS, Kim J (2011) Aberrant ribosome biogenesis activates c-Myc and ASK1 pathways resulting in p53-dependent G1 arrest. Oncogene 30(30):3317–3327. https://doi.org/10.1038/onc.2011.47
Pogue-Geile K, Geiser JR, Shu M, Miller C, Wool IG, Meisler AI, Pipas JM (1991) Ribosomal protein genes are overexpressed in colorectal cancer: isolation of a cDNA clone encoding the human S3 ribosomal protein. Mol Cell Biol 11(8):3842–3849
Boyle P, Ferlay J (2005) Mortality and survival in breast and colorectal cancer. Nat Clin Pract Oncol 2(9):424–425. https://doi.org/10.1038/ncponc0288
International Agency for Research on Cancer Washington (2002) World Health Organization cancer incidence in five continents. The World Health Organization and the International Agency for Research on Cancer, Lyon
World Cancer Research Fund (2007) Food, Nutrition, Physical Activity, and the Prevention of Cancer: A Global Perspective. American Institute for Cancer Research, Washington
De Rosa M, Pace U, Rega D, Costabile V, Duraturo F, Izzo P, Delrio P (2015) Genetics, diagnosis and management of colorectal cancer (Review). Oncol Rep 34(3):1087–1096. https://doi.org/10.3892/or.2015.4108
DeBerardinis RJ, Lum JJ, Hatzivassiliou G, Thompson CB (2008) The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab 7(1):11–20. https://doi.org/10.1016/j.cmet.2007.10.002
Icard P, Shulman S, Farhat D, Steyaert JM, Alifano M, Lincet H (2018) How the Warburg effect supports aggressiveness and drug resistance of cancer cells? Drug Resist Updat 38:1–11. https://doi.org/10.1016/j.drup.2018.03.001
Hsu PP, Sabatini DM (2008) Cancer cell metabolism: Warburg and beyond. Cell 134(5):703–707. https://doi.org/10.1016/j.cell.2008.08.021
Rajeshkumar NV, Dutta P, Yabuuchi S, de Wilde RF, Martinez GV, Le A, Kamphorst JJ, Rabinowitz JD, Jain SK, Hidalgo M, Dang CV, Gillies RJ, Maitra A (2015) Therapeutic targeting of the Warburg effect in pancreatic cancer relies on an absence of p53 function. Cancer Res 75(16):3355–3364. https://doi.org/10.1158/0008-5472.CAN-15-0108
Allison SJ, Knight JR, Granchi C, Rani R, Minutolo F, Milner J, Phillips RM (2014) Identification of LDH-A as a therapeutic target for cancer cell killing via (i) p53/NAD(H)-dependent and (ii) p53-independent pathways. Oncogenesis 3:e102. https://doi.org/10.1038/oncsis.2014.16
Ishiguro T, Nakajima M, Naito M, Muto T, Tsuruo T (1996) Identification of genes differentially expressed in B16 murine melanoma sublines with different metastatic potentials. Cancer Res 56(4):875–879
Shuda M, Kondoh N, Tanaka K, Ryo A, Wakatsuki T, Hada A, Goseki N, Igari T, Hatsuse K, Aihara T, Horiuchi S, Shichita M, Yamamoto N, Yamamoto M (2000) Enhanced expression of translation factor mRNAs in hepatocellular carcinoma. Anticancer Res 20(4):2489–2494
Tian Y, Qin L, Qiu H, Shi D, Sun R, Li W, Liu T, Wang J, Xu T, Guo W, Kang T, Huang W, Wang G, Deng W (2015) RPS3 regulates melanoma cell growth and apoptosis by targeting Cyto C/Ca2+/MICU1 dependent mitochondrial signaling. Oncotarget 6(30):29614–29625. https://doi.org/10.18632/oncotarget.4868
Hua W, Yue L, Dingbo S (2016) Over-expression of RPS3 promotes acute lymphoblastic leukemia growth and progress by down-regulating COX-2 through NF-κb pathway. Blood 128(22):3927
Zhao L, Cao J, Hu K, Wang P, Li G, He X, Tong T, Han L (2018) RNA-binding protein RPS3 contributes to hepatocarcinogenesis by post-transcriptionally up-regulating SIRT1. Nucleic Acids Res. https://doi.org/10.1093/nar/gky1209
Naora H, Takai I, Adachi M, Naora H (1998) Altered cellular responses by varying expression of a ribosomal protein gene: sequential coordination of enhancement and suppression of ribosomal protein S3a gene expression induces apoptosis. J Cell Biol 141(3):741–753
Kim W, Youn H, Lee S, Kim E, Kim D, Sub Lee J, Lee JM, Youn B (2018) RNF138-mediated ubiquitination of rpS3 is required for resistance of glioblastoma cells to radiation-induced apoptosis. Exp Mol Med 50(1):e434. https://doi.org/10.1038/emm.2017.247
Nagao-Kitamoto H, Setoguchi T, Kitamoto S, Nakamura S, Tsuru A, Nagata M, Nagano S, Ishidou Y, Yokouchi M, Kitajima S, Yoshioka T, Maeda S, Yonezawa S, Komiya S (2015) Ribosomal protein S3 regulates GLI2-mediated osteosarcoma invasion. Cancer Lett 356:855–861. https://doi.org/10.1016/j.canlet.2014.10.042
Kim SH, Kim J (2006) Reduction of invasion in human fibrosarcoma cells by ribosomal protein S3 in conjunction with Nm23-H1 and ERK. Biochim Biophys Acta 1763(8):823–832. https://doi.org/10.1016/j.bbamcr.2006.03.011
Momand J, Wu HH, Dasgupta G (2000) MDM2—master regulator of the p53 tumor suppressor protein. Gene 242(1–2):15–29
Liu Y, Bodmer WF (2006) Analysis of P53 mutations and their expression in 56 colorectal cancer cell lines. Proc Natl Acad Sci USA 103(4):976–981. https://doi.org/10.1073/pnas.0510146103
Kim H, Kokkotou E, Na X, Rhee SH, Moyer MP, Pothoulakis C, Lamont JT (2005) Clostridium difficile toxin A-induced colonocyte apoptosis involves p53-dependent p21(WAF1/CIP1) induction via p38 mitogen-activated protein kinase. Gastroenterology 129(6):1875–1888. https://doi.org/10.1053/j.gastro.2005.09.011
Vousden KH, Ryan KM (2009) p53 and metabolism. Nat Rev Cancer 9(10):691–700. https://doi.org/10.1038/nrc2715
Simabuco FM, Morale MG, Pavan ICB, Morelli AP, Silva FR, Tamura RE (2018) p53 and metabolism: from mechanism to therapeutics. Oncotarget 9(34):23780–23823. https://doi.org/10.18632/oncotarget.25267
Fantin VR, St-Pierre J, Leder P (2006) Attenuation of LDH-A expression uncovers a link between glycolysis, mitochondrial physiology, and tumor maintenance. Cancer Cell 9(6):425–434. https://doi.org/10.1016/j.ccr.2006.04.023
Le A, Cooper CR, Gouw AM, Dinavahi R, Maitra A, Deck LM, Royer RE, Vander Jagt DL, Semenza GL, Dang CV (2010) Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc Natl Acad Sci USA 107(5):2037–2042. https://doi.org/10.1073/pnas.0914433107
Miao P, Sheng S, Sun X, Liu J, Huang G (2013) Lactate dehydrogenase A in cancer: a promising target for diagnosis and therapy. IUBMB Life 65(11):904–910. https://doi.org/10.1002/iub.1216
Zhou M, Zhao Y, Ding Y, Liu H, Liu Z, Fodstad O, Riker AI, Kamarajugadda S, Lu J, Owen LB, Ledoux SP, Tan M (2010) Warburg effect in chemosensitivity: targeting lactate dehydrogenase-A re-sensitizes taxol-resistant cancer cells to taxol. Mol cancer 9:33. https://doi.org/10.1186/1476-4598-9-33
Mohammad GH, Olde Damink SW, Malago M, Dhar DK, Pereira SP (2016) Pyruvate kinase M2 and lactate dehydrogenase A are overexpressed in pancreatic cancer and correlate with poor outcome. PLoS ONE 11(3):e0151635. https://doi.org/10.1371/journal.pone.0151635
Li J, Zhu S, Tong J, Hao H, Yang J, Liu Z, Wang Y (2016) Suppression of lactate dehydrogenase A compromises tumor progression by downregulation of the Warburg effect in glioblastoma. NeuroReport 27(2):110–115. https://doi.org/10.1097/WNR.0000000000000506
Wang J, Wang H, Liu A, Fang C, Hao J, Wang Z (2015) Lactate dehydrogenase A negatively regulated by miRNAs promotes aerobic glycolysis and is increased in colorectal cancer. Oncotarget 6(23):19456–19468. https://doi.org/10.18632/oncotarget.3318
Acknowledgements
This work was supported by the Lebanese National Council for Scientific Research (Grant # CNRS020614). We would also like to acknowledge Dr. Samer Bazzi for performing the flow cytometry experiments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Alam, E., Maaliki, L. & Nasr, Z. Ribosomal protein S3 selectively affects colon cancer growth by modulating the levels of p53 and lactate dehydrogenase. Mol Biol Rep 47, 6083–6090 (2020). https://doi.org/10.1007/s11033-020-05683-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05683-1