Abstract
Seed is an important way to store germplasm resources but its genetic integrity will decrease during long-term preservation. So, it’s essential to update seeds according to the aging level of different species. Pearl millet [Cenchrus americanus (L.) Morrone syn., Pennisetum glaucum (L.) R. Br.] is a crucial forage grass, biofuel plant and important crops in the world bringing huge economic and ecological benefits. However, there is no report about the impact of aging on genetic integrity of its seeds. In this study, four genetic diversity indexes (the percentage of polymorphic bands, PPB; the effective number of alleles, Ne; the Nei's gene diversity index, H; the Shannon's information index, I) and 20 pairs of genomic-SSR primers were used to certify the optimal sample volume of pearl millet for molecular study and found that the best sample volume was 60. After the artificial aging test, the germination rate and four genetic diversity parameters (the number of alleles, Na; Ne; H; I) were used to evaluate the change of genetic integrity at different aging levels. The results showed that the germination rate and these four genetic diversity parameters declined with the increase of aging levels. Furthermore, when the germination rate of pearl millet seeds went down to 68.23%, a significant difference in genetic integrity was observed with unaged seeds. In conclusion, the optimal sample size of pearl millet was 60 and the critical point of germination rate to renew germplasm resources was 68.23% and these finds might contribute to the scientific study and the safe conservation of pearl millet.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The germplasm resources are the basis of genetic breeding, biotechnology and plant science. Nowadays, around 61 million germplasm resources have been collected and 90% of which are seeds. Seeds are one of the most common and cost-optimal ways to store germplasm resources for a long time [1]. The condition of seeds is monitored by genetic integrity, which refers to the difference in alleles and the frequency of genotype compared with the original population. The genetic integrity of cross-pollination plants like pearl millet is mainly affected by seeds aging, population and environment [2]. However, even if the seeds are stored suitably, they will age inevitably [3]. Seeds aging can cause not only the mutation and drift of gene but also the loss of genetic integrity, which is a big disadvantage to the conservation of germplasm resources [4]. Thus, it is necessary to make clear the impact of seeds aging on genetic integrity of plants.
Since it always takes a long time to age seeds in natural conditions, researchers often age seeds in an artificial way to accelerate the process and build germination gradient for profound research [5]. At present, there are lots of commonly used methods, for instance, hot water aging (58 ± 1 °C) [6], high temperature with high relative humidity (40 °C,100% RH) [7] and methanol solution (MS) [8, 9].
Among the molecular markers used to evaluate genetic integrity of plants, SSR (also known as short tandem repeat, STR) has high polymorphism and conservation [10]. The genomic-SSR, which is more polymorphic than expressed sequence tag SSR (EST-SSR) [11], has been widely used in numerous studies on different plant species, such as Cannabis sativa var. indica (Lam.) E. Small et Cronq. [12], Brassica rapa L. [13], Jatropha curcas L. [14] and Cuminum cyminum L. [15]. Thus, it is ideal to detect genetic integrity by using genomic-SSR to develop molecular markers.
Pearl millet (Cenchrus americanus (L.) Morrone syn., P glaucum (L.) R. Br., Poaceae family) is a staple crop in the world, which feeds one-third of the world’s population [16]. It is an annual, warm-season C4 monocot crop that widely distributes in South Asia [17], sub-Saharan Africa and Latin America [18]. Due to its characteristics of abundant biomass, high stress resistance, and excellent palatability, it is always regarded as an ideal food for poor people in the arid districts in Africa, India and other semi-arid tropical regions [19, 20]. This green fodder has a high assay in protein, iron, zinc, calcium and other minerals with low hydrocyanic acid content, which is used as a quality forage crop in South China, Korea and South America [21]. Although it has such high economic and ecological values, a few studies have been done about pearl millet, especially on genetic integrity.
When using markers to analyze the genetic integrity of plants, we can observe some significant differences with different sample sizes [22]. Theoretically, an infinite sample size could cover all genetic diversity, while many constraints like experimental expenditure, climate, seeds production and other limitations make it impossible [23]. However, a small sample size cannot cover all the genetic diversity of plants and is unrepresentative [24]. It is essential to select an optimal sample volume to reduce errors and the follow-up workload. Thus, this research has two aims, (i) to determine the optimal sample size of pearl millet for molecular study based on genomic-SSR markers; (ii) to analyze the effects of seeds aging on genetic integrity, which can help us to find the critical point of germination rate for updating pearl millet germplasm resources.
Materials and methods
Plant materials
The seeds of pearl millet cultivar “Tifleaf 3” (open-pollinated cultivar) were provided by Beijing Mammoth Seed Company (Beijing, China) and stored at 4 °C.
Plants used to determine the optimal sample volume were maintained in the growth chamber (Wenjiang, Sichuan, China) with a photoperiod of 16 h/8 h (day/night) at 26 °C/22 °C (day/night) [25] and 80% relative humidity (RH). 35 days later, 96 of the experimental samples were selected randomly and young leaves of each sample were collected to extract DNA.
Accelerated aging test and germination test
A pilot experiment was performed ahead of the accelerated aging test to confirm the most suitable conditions (for instance: temperature, humidity and aging gradient) to acquire different germination rate of seeds. The artificial accelerated aging treatment was based on the results of prior test and Delouche’s method [7]. Seeds were artificially aged by accelerated aging chamber (Top instrument, Zhejiang, China) at 45 °C, 99% RH for 4 h, 8 h, 12 h, 16 h, 20 h and 24 h. Seeds that were not disposed of the accelerated aging protocol were used as the control group.
The germination test was performed as the method recommended by the Rules of Seed Testing for Forage, Turfgrass and other Herbaceous Plant—Species and Variety Testing [26]. Three replicates of 75 seeds were sown in the petri dish (d = 9 cm, Biosharp, Hefei, China) with blotting paper wetted by distilled water and kept in the plant incubator (Wenjiang, Sichuan, China) at 26 °C /22 °C (day/night) with an illumination of 16 h /8 h (day/night). The germination rate was recorded on the 14th day (the sprout was identified as when the radicle was as long as the seed) [27]. Then, transplanted all of the seedlings to the experimental field of College of Animal Science and Technology, Sichuan Agricultural University (30° 37′ N, 103° 40′ E, Chongzhou, Sichuan, China) [28]. 25 days later, fresh leaves of each plant were collected to extract DNA.
DNA extraction
The genomic DNA of pearl millet cultivar “Tifleaf 3” was extracted from fresh leaves by a genomic DNA extraction kit (Tiangen Biochemical Technology Co., Ltd., Beijing, China) [29]. The quality and quantity of DNA were tested by 1% agarose gel electrophoresis (1% AGE) and Nano Drop 2000 spectrophotometer (Thermo Fisher, Wilmington, United States), respectively. The qualified DNA was diluted to 20 ng/μL as the template for amplification and stored at − 20 °C until required.
Genomic-SSR PCR amplification and detection
Sequence information of the 20 pairs of genomic-SSR markers was obtained from previously published literature of Wang’s research (Supplementary Table 1) [30]. The Polymerase Chain Reaction (PCR) amplification was as follows: 1.5 μL DNA template (20 ng/μL), 7.5 μL 2× Reaction Mix, 0.3 μL Golden DNA Polymerase (Tiangen Biochemical Technology Co., Ltd., Beijing, China), 0.6 μL forward and reverse primers (10 pmol/mL, synthesized by Shanghai Shenggong Biological Engineering Technology Services Ltd., Shanghai, China), then added ddH2O up to 15 μL [31]. The PCR process was as follows: 94 °C pre-denaturation 5 min; 94 °C denaturation 30 s, 57–60 °C annealing 45 s, 72 °C extension 1 min, repeated for 35 cycles, 72 °C final extension 7 min [32] and the amplified DNA products were stored at 4 °C. The amplification process was done twice to ensure there was no missing data and the PCR products were inspected on 8% polyacrylamide gel electrophoresis (8% PAGE). The process was as follows: 4 μL sample DNA (20 ng/μL), 4 μL 50 bp Marker (Tiangen Biochemical Technology Co., Ltd., Beijing, China), electrophoresis on the condition of 350 V for 90 min, silver stain 15 min and picture preservation.
Statistical analysis
The electrophoresis results were compared and revised manually. The present bands were labeled as ‘1’, while absent bands were marked as ‘0’ to establish the data matrix [33, 34] in Excel 2016 (each vertical row represented a sample).
Analysis of the detection of optimal sample amount
The data matrix of 96 samples was intercepted by vertical rows randomly to generate the submatrix, which was used to simulate the population with the sample size of 15, 30, 45, 60, 75, 90 and 95. And this process was repeated for 1000 times by Python 3.6. According to the results of the simulation process, the percentage of polymorphic bands (PPB, Eq. 1), the effective number of alleles (Ne, Eq. 2), the Nei’s gene diversity index (H, Eq. 3) [35] and the Shannon’s information index (I, Eq. 4) [36] were calculated by Popgene version 1.32 [37]. Finally, the significance level was tested by one-way analysis of variance (one-way ANOVA) by using SPSS 19. Besides, the equations of genetic diversity parameters are as below.
N is the number of polymorphic bands, M is the total number of bands, k is the number of alleles and pi is the frequency of the ith allele.
Analysis of the genetic integrity
According to the data matrix of the accelerated aging test, PPB (Eq. 1), the number of alleles (Na), H [35] (Eq. 3) and I [36] (Eq. 4) were calculated by using Popgene version 1.32 [37]. Moreover, the significant difference in each aging level was tested by SPSS 19 via Duncan’s Multiple Comparisons (DMC). Finally, the genetic similarity coefficient among different treatments was analyzed by the unweighted pair-group method with arithmetic mean (UPGMA) based on the SM similarity matrices, using the NTSYS-PC 2.10 edition software package [38].
Results
Ascertainment of optimal sample volume
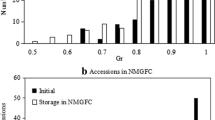
In this study, 96 individual plants were amplified by 20 pairs of genomic-SSR primers and a total of 84 bands were detected, 79 of which were polymorphic with an average of 3.95 polymorphic bands per pair of primers. We observed that with the increase in sample amount, the percentage of polymorphic bands (PPB) and the three genetic diversity indexes increased as well. The effective number of alleles (Ne) arose from 1.6554 to 1.7302 with a decreased slope and standard deviation.
The tendency of the Nei’s gene diversity index (H), the Shannon’s information index (I) and PPB was similar to Ne (Fig. 1). H was from 0.3719 to 0.4008, I was 0.5374 to 0.5780 and PPB ranged from 87.7045 to 94.0476% with the sample size varying from 15 to 95 (Supplementary Table 2). Based on the behaviors of four genetic diversity indexes, there were no distinct divergences when the sample size was 15, 30, 45, 60 while there were no significant differences when the sample size was 60, 75, 90, 95, which indicated that the minimum sample volume representing the population was 60. Thus, the optimal sample volume of pearl millet for molecular research was 60.
Relationship between four genetic indexes and sample size of pearl millet “Tifleaf 3”. a Relationship between the percentage of polymorphic bands (PPB) and sample size, b relationship between the effective number of alleles (Ne) and sample size, c relationship between the Nei's gene diversity index (H) and sample size, d relationship between the Shannon’s information index (I) and sample size. Data are expressed as means with ± SD (black bars) and different letters indicate significant differences at the 0.05 level
Effects of seeds aging on genetic integrity
Analysis of seeds aging on germination and polymorphism
After the seeds treated with 45 °C, 99% relative humidity (RH) for 4 h, 8 h, 12 h, 16 h, 20 h and 24 h, the germination rate reduced as the aging level increased (Fig. 2, Supplementary Table 3). Significant differences with control group (0 h) were observed at the germination rate of 68.00% (4 h) and 68.23% (8 h), while the differences between 4 and 8 h, 12 h and 16 h, 20 h and 24 h were not significant. In this research, seeds of 0 h (80.8%), 8 h (68.23%), 12 h (57.80%), 16 h (52.90%) and 20 h (39.10%) were selected to evaluate the genetic integrity, for 4 h (68.00%) and 24 h (40.87%) had a higher standard deviation.
The five selected groups were evaluated for genetic integrity with the sample size of 60. As for unaged materials, 187 bands were amplified by 20 pairs of primers, 186 (99.47%) of which were polymorphic. For the rest, the bands that generated by 20 pairs of primers were 138 (8 h), 115 (12 h), 139 (16 h) and 131 (20 h), respectively. The number of polymorphic bands and the percentage of polymorphic bands were 137 (99.28%, 8 h), 110 (95.65%, 12 h), 128 (92.09%, 16 h) and 119 (90.84%, 20 h), respectively. Thus, PPB deceased because of seeds aging. The results of four genetic diversity indices were presented in Table 1. Apparently, the four genetic diversity indexes of 0 h reached the highest of this study. A significant decrease with control group in Ne was observed at 8 h while in the number of alleles (Na) was 16 h. I and H showed similar behaviors with control group. Generally, with the decrease in germination rate, all of the polymorphism parameters went down, as well. However, the values of Ne, H and I in 12 h were abnormally higher than that in 8 h.
Analysis of seeds aging on genetic similarity
The genetic similarity coefficients of five selected populations were calculated by using the UPGMA approach (Supplementary Table 4). The average of genetic similarity coefficients was 0.8140 and the five treatment groups could be divided into two major groups at this point (Fig. 3). The control group belonged to one cluster while 8 h, 12 h, 16 h and 20 h belonged to another cluster.
Discussion
The age and deterioration of seeds are inevitable phenomena during storage [39]. In order to make sure the genetic integrity of seeds can be preserved completely, it is necessary to study the genetic integrity of germplasm resources. Sampling strategy is the first problem to face in genetic integrity research. It is of great importance to develop effective sampling strategies for the evaluation of genetic diversity of plant populations [40]. Mixed sampling strategy and single plant sampling strategy are two major sampling methods [41]. Mixed sampling is timesaving, but the more mixed, the greater deviation and less reliability will be. The biggest disadvantage of single plant sampling is time consuming and huge workload, but it can be used to analyze the genetic variations between or within populations, which is comprehensive and reliable [42].
In this experiment, single plant sampling method and four genetic diversity parameters (the percentage of polymorphism sites, PPB; the number of effective alleles, Ne; the Nei's gene diversity index, H and the Shannon's information index, I) were selected to determine the optimal sampling size of pearl millet for molecular research. To make results reliable, the select process was repeated for 1000 times. The results showed that when the sample size arose to 60, those four genetic diversity parameters had no significant differences with the larger sample quantity. In case of finding the minimum optimal sampling volume that could replace the population, experiments with the sample size of 50 and 55 were added in the subsequent experiments, and there was a significant difference with the sample size over 60. Therefore, 60 would be the optimal volume of samples to study genetic integrity of pearl millet. In the study of the optimal sample size of plants for molecular research, the best sample volume of Astragalus sinicus L. was 30 [22], annual ryegrass (Lolium multiflorum Lam.) was 20 [43], Psathyrostachys Huashanica Keng ex P. C. Kuo was 26 [44], and A nuda L. was 70 and 50 [45]. It means that different plants have different biological characteristics so that the optimal sampling volume is usually different. Compared with self-pollinated and asexual plants, cross-pollinated plants have high gene heterozygosity and are different from individuals within populations. Thus, they often have a smaller sample size than self-pollinated and asexual plants when doing genetic diversity analysis [46].
In this study, seeds of pearl millet were artificially aged with the optimal sample volume of 60. In the germination test, the germination was decreased as the aging level increased. Moreover, at the point of 68.32%, there was a significant decrease with the unaged group. Similarly, the values of Ne, the number of alleles (Na), H and I had the same tendency with the germination rate. However, the values of Ne, H and I in 12 h were abnormally higher than 8 h, showing irregularity. This phenomenon might be caused by the fact that the pearl millet in this study is an open-pollinated cultivar and the small number of base pairs labeled by genomic-SSR markers or the inhomogeneous distribution of markers.
Owing to the age of seeds could cause the loss of genetic integrity and the reduction of yield. It is necessary to find out the critical point of germination to make sure the germplasm resources could be updated in time. At present, the critical point of germination rate is different around the world. For example, in India, the germination rate to renew germplasm resources is 75% [47], while Britain is 70% [48]. The United States does not update germplasm resources until the germination rate reduces to 50% [49]. Moreover, in China, the standard for regeneration germplasm resources is 60% according to the government documents made by the national forage seed bank [50]. In this research, the maximum germination rate with significant differences of the control group was 68.23%, which indicated that the germplasm resources of pearl millet should be updated when the germination rate decreased to 68.23%. Although 68.23% was significantly lower than the Chinese standard, the genetic integrity of pearl millet might have undergone at a higher germination rate. So, we still need to develop better molecular markers, detail the artificially accelerated aging gradient, use more advanced methods and do further analysis of genetic diversity parameters to get the precise critical point of renewal germplasm resources of pearl millet.
References
Zhu YQ, Yan HF, Xia FS, Chen QZ, Wang MY, Mao PS (2016) The relationship between mitochondria and seed aging. Pratacult Sci 32(2):290–298
Parzies HK, Spoor W, Ennos RA (2000) Genetic diversity of barley landrace accessions (Hordeum vulgare ssp. vulgare) conserved for different lengths of time in ex situ gene banks. Heredity 84:476–486. https://doi.org/10.1046/j.1365-2540.2000.00705.x
Taiz L, Zeiger E (2015) Plant physiology, 5th edn. Science Press, Beijing
Priestley DA (1986) Seed aging: implications for seed storage and persistence in the soil. Q Rev Biol 113(3):130. https://doi.org/10.1086/415450
Rajjou L, Lovigny Y, Groot SP, Belghazi M, Job C, Job D (2008) Proteome-wide characterization of seed aging in Arabidopsis: a comparison between artificial and natural aging protocols. Plant Physiol 148(1):620–641. https://doi.org/10.1104/pp.108.123141
Bhattacharyya S, Hazra A, Sen-Mandi S (1985) Accelerated ageing of seeds in hot water: germination characteristics of aged wheat seeds. Seed Sci Technol 13(3):683–690
Delouche JC, Baskin CC (1973) Accelerated aging techniques for predicting the relative storability of seeds lots. Seed Sci Technol 1(2):427–452
Dao PT, Ram HH (1997) Prediction of seed longevity in soybean (Glycine max) based on seed-stress test. Indian J Agric Sci 67(7):322–323
Mugnisjah WQ, Nakamura S (1985) Methanol and ethanol stress for seed vigour evaluation in soybean. Seed Sci Technol 14(1):95–103
Gilmore S, Peakall R, Robertson J (2003) Short tandem repeat (STR) DNA markers are hypervariable and informative in Cannabissativa: implications for forensic investigations. Forensic Sci Int 131(1):65–74. https://doi.org/10.1016/S0379-0738(02)00397-3
Zhou SF, Wang CR, Frazier TP, Yan HD, Chen PL, Chen ZH, Huang LK, Zhang XQ, Peng Y, Ma X, Yan YH (2018) The first illumina-based de novo transcriptome analysis and molecular marker development in napier grass (Pennisetum purpureum). Mol Breed 38(7):95. https://doi.org/10.1007/s11032-018-0852-8
Soler S, Gramazio P, Figàs MR, Vilanova S, Rosa E, Llosa ER, Borràs D, Plazas M, Prohens J (2017) Genetic structure of Cannabis sativa var. indica cultivars based on genomic SSR (gSSR) markers: implications for breeding and germplasm management. Ind Crops Prod 104:171–178. https://doi.org/10.1016/j.indcrop.2017.04.043
Wang Z, Ge Y, Jing J, Han XL, Piao ZY (2014) Integrated genetic linkage map based on UGMS and gSSR markers in Brassica rapa. Sci Horticult 179:293–300. https://doi.org/10.1016/j.scienta.2014.09.046
Wen MF, Wang HY, Xia ZQ, Zou ML, Lu C, Wang WQ (2010) Developmenrt of EST-SSR and genomic-SSR markers to assess genetic diversity in Jatropha curcas L. BMC Res Notes 3(1):42. https://doi.org/10.1186/1756-0500-3-42
Bharti R, Kumar S, Parekh MJ (2018) Development of genomic simple sequence repeat (gSSR) markers in cumin and their application in diversity analyses and cross-transferability. Ind Crops Prod 111:158–164. https://doi.org/10.1016/j.indcrop.2017.10.018
Kumar S, Hash CT, Nepolean T, Satyavathi CT, Singh G, Mahendrakar MD, Yadav RS, Srivastava RK (2017) Mapping QTLs controlling flowering time and important agronomic traits in pearl millet. Front Plant Sci 8:1731. https://doi.org/10.3389/fpls.2017.01731
Khan I, Raza MA, Khalid MHB, Awan SA, Raja NI, Zhang X, Min S, Wu BC, Hassan MJ, Huang L (2019) Physiological and biochemical responses of pearl millet (Pennisetum glaucum L.) seedlings exposed to silver nitrate (AgNO3) and silver nanoparticles (AgNPs). Int J Environ Res Public Health 16(13):2261. https://doi.org/10.3390/ijerph16132261
Yadav M, Rengasamy RS, Gupta D (2019) Characterization of pearl millet (Pennisetum glaucum) waste. Carbohyd Polym 212:160–168. https://doi.org/10.1016/j.carbpol.2019.02.034
Chelpuri D, Sharma R, Durga KK, Katiyar P, Mahendrakar MD, Singh RB, Yadav RS, Gupta R, Srivastava RK (2019) Mapping quantitative trait loci (QTLs) associated with resistance to major pathotype-isolates of pearl millet downy mildew pathogen. Eur J Plant Pathol 154(4):983–994. https://doi.org/10.1007/s10658-019-01718-x
Pucher A, Høgh-Jensen H, Gondah J, Hash CT, Haussmann BIG (2014) Micronutrient density and stability in west African pearl millet—potential for biofortification. Crop Sci 54(4):1709–1720. https://doi.org/10.2135/cropsci2013.11.0744
Khairwal IS, Yadav SK, Rai NK, Upadhyaya HU, Kachhawa D, Nirwan B, Bhattacharjee R, Rajpurohit BS, Dangaria CJ, Srikant (2007) Evaluation and identification of promising pearl millet germplasm for grain and fodder traits. J SAT Agricult Res 5(1):1–6
Chen J, Lin XJ, Zhong SJ (2015) Effect of sampling strategy on identification of cultivars of Astragalus sinicus L. by SSR marker. Plant Genet Resour C 16(6):1245–1251
Marshall DR (1990) Crop genetic resources: current and emerging issues. Wiley, Chichester
Jin Y, Lu BR (2003) Sampling strategy for genetic diversity. Chin Biodivers 11(2):155–161
Yan HD, Bombarely A, Xu B, Wu BC, Frazier TP, Zhang XQ, Chen J, Chen PL, Sun M, Feng GY, Wang CR, Cui CM, Li Q, Zhao BY (2019) Autopolyploidization in switchgrass alters phenotype and flowering time via epigenetic and transcription regulation. J Exp Bot 70(20):5673–5686. https://doi.org/10.1093/jxb/erz325
Wang YR, Zhang JY, Liu WX, Liu ZP, Duan YY, Yu L, Li YR, Sun JH, Hu XW, Liu YJ (2017) Rules of seed testing for forage, turfgrass and other herbaceous plant—species and variety testing. Standards Press of China, Beijing
Gutierrez G, Cruz F, Moreno J, Gonzalez-Hernandez VA, Vazquez-Ramos JM (1993) Natural and artificial seed ageing in maize: germination and DNA synthesis. Seed Sci Res 3(4):279–285
Zhou SF, Chen J, Lai YS, Yin GH, Chen PL, Pennerman KK, Yan HD, Wu BC, Zhang H, Yi XF, Wang CR, Fu MJ, Zhang XQ, Huang LK, Ma X, Peng Y, Yan YH, Nie G, Liu L (2019) Integrative analysis of metabolome and transcriptome reveals anthocyanins biosynthesis regulation in grass species, Pennisetum purpureum. Ind Crops Prod 144:442–454. https://doi.org/10.1016/j.indcrop.2019.111470
Huang LK, Huang X, Yan HD, Yin GH, Zhang XQ, Tian Y, Zhang Y, Jiang XM, Yan YH, Ma X, Peng Y, Zhou JN, Nie G (2014) Constructing DNA fingerprinting of Hemarthria cultivars using EST-SSR and SCoT markers. Genet Resour Crop Evolut 61(6):1047–1055. https://doi.org/10.1007/s10722-014-0107-4
Wang CR, Yan HD, Li J, Zhou SF, Liu T, Zhang XQ, Huang LK (2018) Genome survey sequencing of purple elephant grass (Pennisetum purpureum Schum ‘Zise’) and identification of its SSR markers. Mol Breed 38(7):94. https://doi.org/10.1007/s11032-018-0849-3
Xie WG, Robins JG, Bushman BS (2012) A genetic linkage map of tetraploid orchardgrass (Dactylis glomerata L.) and quantitative trait loci for heading date. Genome 55(5):360–369. https://doi.org/10.1139/g2012-026
Qi X, Pittaway TS, Lindup S, Liu H, Waterman E, Padi FK, Hash CT, Zhu J, Gale MD, Devos KM (2004) An integrated genetic map and a new set of simple sequence repeat markers for pearl millet, Pennisetum glaucum. Theor Appl Genet 109(7):1485–1493. https://doi.org/10.1007/s00122-004-1765-y
Liwu Z, Rongrong C, Minhang Y, Aifen T, Jiantang X, Lihui L, Pingping F, Jianmin Q (2015) Genetic diversity and DNA fingerprinting in jute (Corchorus spp.) based on SSR markers. Crop J 3(5):416–422. https://doi.org/10.1016/j.cj.2015.05.005
Xiaoyan S, Chunzhi Z, Ying L, Shuangshuang F, Qing Y, Sanwen H (2016) SSR analysis of genetic diversity among 192 diploid potato cultivars. Horticult Plant J 2(3):163–171. https://doi.org/10.1016/j.hpj.2016.08.006
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci 70(12):187–192
Strong WL (2016) Biased richness and evenness relationships within Shannon-Wiener index values. Ecol Ind 67:703–713. https://doi.org/10.1016/j.ecolind.2016.03.043
Yeh FC, Boyle TJB (1997) Population genetic analysis of codominant and dominant markers and quantitative traits. Belg J Bot 129:157
Huang LK, Zhang Y, Zhang J, Zhang XQ, Xie WG, Jiang XM, Peng F, Yan YH, Ma X, Liu W, Peng Y, Yin GH, Li X (2014) Genetic stability and DNA fingerprinting of the Hemarthria compressa cultivar “Guangyi”. Biochem Syst Ecol 55:310–316. https://doi.org/10.1016/j.bse.2014.03.027
Villiers TA (1974) Seed aging: chromosome stability and extended viability of seeds stored fully imbided. Plant Physiol 53(6):875–878
Gargiulo R, Saubin M, Rizzuto G, West B, Fay MF, Kallow S, Trivedi C (2019) Genetic diversity in British populations of Taxus baccata L.: is the seedbank collection representative of the genetic variation in the wild? Biol Conserv 233:289–297. https://doi.org/10.1016/j.biocon.2019.01.014
Ban T, Han P, Liu X, Jin L, Wang XJ (2009) Sampling numbers in genetic diversity analysis of alfalfa using bulked DNA based on RAPD and SSR markers. Life Sci Res 13:158–162
Jennifer FC, James MA, Hamrick JL (1997) Developing a sampling strategy for Baptisia arachnifera based on allozyme diversity. Conserv Biol 11(5):1133–1139
Luo YC, Ma X, Zhang XQ (2013) Bulking strategies for genetic diversity analysis of annual ryegrass based on SSR markers. Pratacult Sci 30(3):376–385
Liu WX, Li LH, Liu WH, Zhang ZM, Wu ZH, Wang CS (2006) SSR analysis on the sampling strategy of Psathyrostachys huashanic Keng population. J Triticeae Crops 26:16–20
Yang MM, Lu P, Liu JQ, Li HY, Huang F, Li J, Wang XX, Xi LJ (2017) Study on sample size in analysis of genetic integrity of Avena nuda L. based on SSR markers. Chin J Grassl 39(3):31–37
Tan LT, Yu CM, Chen P, Wang YZ, Chen JK, Wen L, Xiong HP (2012) Effects of sample size on genetic diversity parameters for “Zhong Zhu No. 1” Group. Plant Fiber Sci China 34(4):179–183
Engels J, Ramanatha RR (1998) Regeneration of seed crops and their wild relatives. ResearchGate. https://www.researchgate.net/publication/236143293_Regeneration_of_seed_crops_and_their_wild_relatives. Accessed January 1998
Vellvé R (2013) Saving the seed: genetic diversity and European agriculture. Routledge, London
Wu YH, Wang HP, Li XX, Song JP, Zhao Q, Zhang XH, Yang WL (2020) Monitoring seed viability of different Allium vegetables in the national medium-term gene bank of vegetable germplasm resources. Plant Genet Resour C 21(3):648–654
Chen ZH, Wang ZH, Yun XJ, Li XF (2007) Viability monitoring and regeneration of seeds in the forage genebank of China. Grassl Turf 3:72–75
Acknowledgements
This research was funded by the Sichuan Province Research Grant (2016NYZ0036), the Modern Agro-industry Technology Research System (CARS-34), College Students' Innovation and Entrepreneurship Training Program and the Modern Agricultural Industry System Sichuan Forage Innovation Team (SCCXTD-2020–16).
Author information
Authors and Affiliations
Contributions
LH planned and designed the research XD and CW wrote the manuscript and performed the experiments. AZ, YS and IK revised the manuscript. RW provided partial data basis. And all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest to report.
Research involving human and animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dan, X., Wang, C., Su, Y. et al. Evaluation of genetic integrity of pearl millet seeds during aging by genomic-SSR markers. Mol Biol Rep 47, 5747–5754 (2020). https://doi.org/10.1007/s11033-020-05642-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05642-w