Abstract
Breast cancer (BC) is a heterogeneous and multifactorial disease. The system formed by glutathione-S-transferases (GSTs) acts to protect the organism against the oxidative stress generated by xenobiotics and their active products. Glutathione transferase mu 1 (GSTM1) and glutathione transferase theta 1 (GSTT1) present null polymorphic variants by complete deletion. The absence of these enzymes may influence the susceptibility to several diseases such as BC. This study aimed to systematically review and investigate the existence of a possible correlation between the presence/absence of these genetic variants and the development of BC and their influence in chemotherapy response. The Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) protocol was used, and the searches were performed in the portal of the Virtual Health Library (VHL) and the PubMed, resulting in 21 articles. It is clear that most studies revealed a risk association between the deletion of GSTM1 and/or GSTT1 and the development and/or prognosis of BC.Moreover, it should be noted that these results of risk association were found in large part in the populations of the Americas and Europe, followed by Asians. Regarding the response to treatment, protective associations were found in the presence of GSTM1 deletion. However, due to the inconclusive results of many studies, further analysis in this area is required.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer is a heterogeneous disease that affects individuals worldwide. The International Agency for Research on Cancer (IARC) expected, for 2018, 18.1 million new cases and 9.6 million deaths by neoplasia. Female breast cancer is in the second place in the ranking of causes of death by cancer, with 11.6% of all deaths in the world, followed by prostate tumors (7.1% of cases) [1]. According to the National Cancer Institute (INCA), in Brazil, there were be about 60 thousand new cases of breast cancer for the biennium 2018–2019. Regardless of non-melanoma skin tumors, the breast cancer is the most incident in the Brazilian Southern, resulting in an estimated risk of 73.07 cases per 100,000 women [2].
Similar to other types of malignant tumors, the development of breast carcinoma is complex, environmental variables such as lifestyle and intrinsic characteristics of the patient also influence the development and evolution of this malignant disease, such as age [3, 4], hormone factors [5], menopause [6], smoking [7], exposure to ionizing radiation [8], and overweight [9]. Hence, it is important to understand these elements to determine the diagnosis and prognosis of patients.
Most cancers are associated with external risk factors. Therefore, a wide of substances are constantly interacting with the body by several routes, and they need to be eliminated securely to avoid any kind of injury to its DNA. Likewise, there is a complex system formed by phase I detoxification enzymes, components of the P450 cytochrome system [10], and by phase II detoxification enzymes, that includes the glutathione S-transferases (GSTs) [11] and the N-acetyltransferases (NATs) [12]. All this system is responsible for metabolizing environmental and xenobiotic factors that can be potentially associated with the increased carcinogenesis [11].
The GSTs represent a superfamily of cytosolic, mitochondrial and microsomal enzymes, which are involved in the metabolism of xenobiotic compounds and their reactive products, preventing the oxidative stress [11, 13]. This superfamily is divided into Alpha, Mu, Omega, Pi, Sigma, Theta and Zeta classes, according to its structural, chemical and physical characteristics. Their N-terminal residue interacts with the thiol group of the glutathione peptide in its reduced form (γ-l-glutamyl- l-cysteine-glycine, GSH), providing the conjugation of compounds to be excreted, including carcinogens, drugs and metabolism products [14, 15].
Genetic polymorphisms can lead to variations in the activity of the enzymes, resulting in combinations ranging from partial to complete deletions, which can result even in a null phenotype [16, 17]. The glutathione transferase mu 1 (GSTM1) and glutathione transferase theta 1 (GSTT1) genes belong to the mu and theta classes, respectively. They present null polymorphic variants by complete deletion also called null genotypes that result in a complete absence of the enzyme function in both cases, GSTM1- null and GSTT1- null genotypes [18].
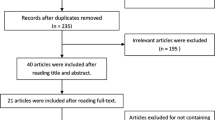
As already stated by researchers these polymorphisms are associated with the development of several diseases, such as uterine leiomyoma [19], hypertension [20], psoriasis [21], prostate carcinoma [22] and chronic myeloid leukemia [23], among others. Studies also showed that genetic polymorphisms of this system may act as predictors of susceptibility to some types of cancer, such as lung [24], colorectal [25], breast [26] and cervical [27]. However, few studies have discussed the clinical meaning of such variants, as well its correlation with the parameters that are determinant of poor prognosis. Thus, results in the correlation with breast cancer have been controversial because some of them found a significant association and others showed a risk association. Therefore, this study aimed to systematically review and evaluate studies regarding the occurrence of polymorphisms in the GST system in patients with breast cancer. The current study also aimed to investigate if there are correlation between the presence/absence of the genetic variants and the susceptibility, as well as between determinants of pathology’s prognosis (Fig. 1).
Materials and methods
Review protocol
The systematic review followed the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) protocol, adapted from Moher et al. [28].
Literature search
PubMed and the regional portal of the Virtual Health Library (VHL) were used to perform the searches (the last search was conducted on April 15th, 2020). The descriptors applied were "glutathione transferase" and "breast cancer" and "genetic polymorphism". Then, some filters were selected: Portuguese and English language, type of document (article), published between 2013 and 2020. Additionally, complete readings of literature reviews and systematic reviews with similar subjects were performed. The studies were pre-selected by reading their titles and abstracts and separated for further analysis and extraction of data.
Inclusion and exclusion criteria
Inclusion criteria:
-
1.
Case–control studies and case studies;
-
2.
The aim of the studies: study the influence of GSTT1 and/or GSTM1 polymorphisms on breast cancer;
-
3.
Studies with complete data and statistical results.
Exclusion criteria:
-
1.
Review studies, meta-analyses, case reports, comments or clinical tests;
-
2.
Other types of cancer or hematological malignancies;
-
3.
Studies containing incomplete data or duplicate data.
Biases
The possible biases of the eligible studies were analyzed according to the limitations of each study, such as sample size, statistics of the results and the parameters involved.
Results
Eligible studies
The VHL database search resulted 769 articles. After the filters’ refining 139 articles were found, and after the analysis of the titles and abstracts, 26 articles were selected. In PubMed database, 244 articles were found and, after selecting the filters, 67 articles remained, of which 35 were selected for a full reading. Other articles from literature review readings were not included, because these articles did not attend the inclusion criteria or were already included. Thus, 21 articles were included in this review.
Characteristics of the studies
From the studies included in the systematic analysis, 13 are case–control studies [25, 29,30,31,32,33,34,35,36,37,38,39,40], and 8 are cohort studies [41,42,43,44,45,46,47,48]. Most studies (28.6%) are from China, followed by Mexico (19.0%). The sample size ranged from 49 to 1109 (6747 cases). Table 1 summarizes the main results found by the included studies for GSTT1 and GSTM1 polymorphisms, and their correlation with the development of breast cancer and/or response to treatment.
Discussion
We performed a review to evaluate the association between GSTM1 and GSTT1 polymorphisms (combined or not) on breast cancer risk and response to treatment, including 6747 breast cancer patients.
Polymorphic variants of GSTs influence the effectiveness of detoxification of the cytotoxins from drugs or carcinogens, and it can increase the susceptibility to cancer development. Several studies discussed the influence of GSTT1 and GSTM1 deletion polymorphisms in some malignancies such as acute myeloid leukemia [49], lung cancer [50] and hepatocellular carcinoma [51]. However, the role of these polymorphisms is not clear towards the susceptibility to breast cancer development, as well as their correlation with the factors that determine the prognosis of this disease.
The simultaneous deletion of GSTM1 and GSTT1 has been associated with a higher risk of developing breast cancer, which increases when it is correlated with exposure to environmental factors such as pesticides, as demonstrated by Sohail et al. (2013) in a case–control study conducted in 200 Pakistani women. Besides, the authors reported an association between these GST variants and a higher risk of developing breast cancer in women who smoke or have a positive family history for the disease [32]. Another article published by the group of Garcia-Martinez and collaborators (2017) observed an association of susceptibility to the development of breast cancer in Mexican women with a deletion in GSTM1, in a case–control study with 1882 women [36]. In other studies, significant results were obtained concerning the deletion of GSTM1, suggesting that it would be associated with a higher risk of developing breast cancer [30, 31, 39, 47]. In a study performed in the population of Cyprus, it was concluded that the null variant for GSTT1 was positively associated with the development of breast cancer, in relation to the wild variant, according to a study of 2286 women [25].
Breast tumor can be divided into grades I, II and III, according to the differentiation of the carcinoma cells into well differentiated, moderately differentiated and less differentiated, respectively. This parameter is considered a significant prognostic factor, considering that the less differentiated the tumor cells are, less similar to the normal breast cells [52, 53]. Brazilian researchers showed that the deletion of GSTT1 was positively associated with increased risk of disease recurrence, as well as deleted GSTM1 was correlated with a worse prognosis of patients because a higher percentage of patients with histopathologic grade III tumors was observed in the presence of this polymorphism [41].
GSTM1 active genotype can influence breast cancer progression, preventing the evolution of the disease in non-chemotherapy patients, observed a Chinese cohort study with 714 participants [48]. On the other hand, individuals carrying the null genotype for GSTT1 and/or GSTM1 may have a better response to treatment with chemotherapy. According to an Indian cohort study, when null GSTM1 genotype was evaluated combined with the Ile/Val GSTP1 genotype (another GST family polymorphism), an association was found with the presence of a response to neoadjuvant chemotherapy [44]. It was also found an association of the null genotype of GSTM1 as a protective factor in relation to the response to chemotherapy in patients with breast cancer who had high plasma levels of glucose [29]. A similar association was described in a study conducted in 262 women in the Chinese population, and showed a better response to chemotherapy among patients with null GSTM1 genotype [42]. In contrast, Wang and Huang (2015) demonstrated that the null GSTM1 genotype was more associated with a worse response to chemotherapy and lower survival [46].
Some limitations were identified by the authors in the analyzed studies. Tulsyan et al. (2013) highlighted the fact that they did not analyze variants in the genes that regulate the detoxification phase I, as well as the sample size used [44]. Another study performed by Yuan et al. (2015) stated that the disagreements found in the literature about polymorphisms of the GSTs family may be attributed to the different ethnic groups analyzed, as to the sample universe used [41]. We agree that there should be one more exclusion criteria concerning ethnicity, but few studies would be included in the review if we limited our search based on this. It has been indicated that neither phenotypic characteristics nor self-declaration replaces genetic ancestry, therefore we cannot use ethnicity as a parameter. Also, there is the possibility of biases in some correlations, such as in the assessment of exposure to environmental factors such as pesticides, that the bias can occur because exposure is verbally reported by patients, without the application of a structured quantitative instrument [39].
Conclusions
It is clear that most studies reveal a risk association between the deletion of GSTM1 and/or GSTT1 and the development and/or prognosis of breast cancer. In addition, it should be noted that these results of risk association were found in large part in the populations of the Americas and Europe, followed by Asians. Regarding the response to treatment, protective associations were found in the presence of GSTM1 deletion. However, due to the inconclusive results of many studies, further analysis in this area is required.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer J Clin 68(6):394–424. https://doi.org/10.3322/caac.21492
INCA (2017) Estimativa 2018: incidência de câncer no Brasil. Instituto Nacional de Câncer José Alencar Gomes da Silva, Rio de Janeiro
Fredholm H, Eaker S, Frisell J, Holmberg L, Fredriksson I, Lindman H (2009) Breast cancer in young women: poor survival despite intensive treatment. PLoS ONE 4(11):e7695–e7695. https://doi.org/10.1371/journal.pone.0007695
Anders CK, Johnson R, Litton J, Phillips M, Bleyer A (2009) Breast cancer before age 40 years. Semin Oncol 36(3):237–249. https://doi.org/10.1053/j.seminoncol.2009.03.001
Yerushalmi R, Woods R, Ravdin PM, Hayes MM, Gelmon KA (2010) Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol 11(2):174–183. https://doi.org/10.1016/S1470-2045(09)70262-1
Ahmed K, Jahan P, Nadia I, Ahmed F, Abdullah Al E (2016) Assessment of menopausal symptoms among early and late menopausal midlife bangladeshi women and their impact on the quality of life. J. Menopausal Med 22(1):39–46. https://doi.org/10.6118/jmm.2016.22.1.39
Catsburg C, Miller AB, Rohan TE (2015) Active cigarette smoking and risk of breast cancer. Int J Cancer 136(9):2204–2209. https://doi.org/10.1002/ijc.29266
Boice JD Jr, Preston D, Davis FG, Monson RR (1991) Frequent chest X-ray fluoroscopy and breast cancer incidence among tuberculosis patients in Massachusetts. Radiat Res 125(2):214–222
Sebastiani F, Cortesi L, Sant M, Lucarini V, Cirilli C, De Matteis E, Marchi I, Negri R, Gallo E, Federico M (2016) Increased incidence of breast cancer in postmenopausal women with high body mass index at the modena screening program. J Breast Cancer 19(3):283–291. https://doi.org/10.4048/jbc.2016.19.3.283
McFadyen MC, Melvin WT, Murray GI (2004) Cytochrome P450 enzymes: novel options for cancer therapeutics. Mol Cancer Ther 3(3):363–371
Hayes JD, Flanagan JU, Jowsey IR (2005) Glutathione transferases. Annu Rev Pharmacol Toxicol 45:51–88. https://doi.org/10.1146/annurev.pharmtox.45.120403.095857
Windmill KF, Gaedigk A, de la Hall PM, Samaratunga H, Grant DM, McManus ME (2000) Localization of N-acetyltransferases NAT1 and NAT2 in human tissues. Toxicol Sci 54(1):19–29. https://doi.org/10.1093/toxsci/54.1.19
Daly AK, Cholerton S, Armstrong M, Idle JR (1994) Genotyping for polymorphisms in xenobiotic metabolism as a predictor of disease susceptibility. Environ Health Perspect 102(Suppl 9):55–61. https://doi.org/10.1289/ehp.94102s955
Dirr H, Reinemer P, Huber R (1994) Refined crystal structure of porcine class Pi glutathione S-transferase (pGST P1–1) at 2·1 Å resolution. J Mol Biol 243(1):72–92. https://doi.org/10.1006/jmbi.1994.1631
Oakley A (2011) Glutathione transferases: a structural perspective. Drug Metab Rev 43(2):138–151. https://doi.org/10.3109/03602532.2011.558093
Singh S (2015) Cytoprotective and regulatory functions of glutathione S-transferases in cancer cell proliferation and cell death. Cancer Chemother Pharmacol 75(1):1–15. https://doi.org/10.1007/s00280-014-2566-x
Townsend DM, Tew KD (2003) Cancer drugs, genetic variation and the glutathione-S-transferase gene family. Am J Pharmacogenomics 3(3):157–172. https://doi.org/10.2165/00129785-200303030-00002
Di Pietro G, Magno LA, Rios-Santos F (2010) Glutathione S-transferases: an overview in cancer research. Expert Opinion Drug Metab Toxicol 6(2):153–170. https://doi.org/10.1517/17425250903427980
Mostafavi SS, Ebrahimi A, Sadat SM, Davari Tanha F, Aghasadeghi MR, Bahramali G, Abbasi Ranjbar P, Sadeghifard V, Javadi F (2016) Impact of null genotypes of GSTT1 and GSTM1 with uterine leiomyoma risk in Iranian population. J Obstetr Gynaecol Res 42(4):434–439. https://doi.org/10.1111/jog.12924
Eslami S, Sahebkar A (2014) Glutathione-S-transferase M1 and T1 null genotypes are associated with hypertension risk: a systematic review and meta-analysis of 12 studies. Curr Hypertens Rep 16(6):432. https://doi.org/10.1007/s11906-014-0432-1
Hruska P, Rybecka S, Novak J, Zlamal F, Splichal Z, Slaby O, Vasku V, Bienertova-Vasku J (2017) Combinations of common polymorphisms within GSTA1 and GSTT1 as a risk factor for psoriasis in a central European population: a case-control study. J Eur Acad Dermatol Venereol:JEADV 31(10):e461–e463. https://doi.org/10.1111/jdv.14266
Malik SS, Kazmi Z, Fatima I, Shabbir R, Perveen S, Masood N (2016) Genetic polymorphism of GSTM1 and GSTT1 and risk of prostatic carcinoma—a meta-analysis of 7,281 prostate cancer cases and 9,082 healthy controls. Asian Pac J Cancer Prevention:APJCP 17(5):2629–2635
Weich N, Ferri C, Moiraghi B, Bengio R, Giere I, Pavlovsky C, Larripa IB, Fundia AF (2016) GSTM1 and GSTP1, but not GSTT1 genetic polymorphisms are associated with chronic myeloid leukemia risk and treatment response. Cancer Epidemiol 44:16–21. https://doi.org/10.1016/j.canep.2016.07.008
Yuan Z, Li J, Hu R, Jiao Y, Han Y, Weng Q (2015) Predictive assessment in pharmacogenetics of XRCC1 gene on clinical outcomes of advanced lung cancer patients treated with platinum-based chemotherapy. Sci Rep 5:16482. doi:10.1038/srep16482. https://www.nature.com/articles/srep16482#supplementary-information
Kakkoura MG, Loizidou MA, Demetriou CA, Loucaides G, Daniel M, Kyriacou K, Hadjisavvas A (2017) The synergistic effect between the Mediterranean diet and GSTP1 or NAT2 SNPs decreases breast cancer risk in Greek-Cypriot women. Eur J Nutr 56(2):545–555. https://doi.org/10.1007/s00394-015-1099-3
Krishna BM, Chaudhary S, Panda AK, Mishra DR, Mishra SK (2018) Her2 Ile655Val polymorphism and its association with breast cancer risk: an updated meta-analysis of case-control studies. Sci Rep 8(1):7427. https://doi.org/10.1038/s41598-018-25769-y
Pandey NO, Chauhan AV, Raithatha NS, Patel PK, Khandelwal R, Desai AN, Choxi Y, Kapadia RS, Jain ND (2019) Association of TLR4 and TLR9 polymorphisms and haplotypes with cervical cancer susceptibility. Sci Rep 9(1):9729. https://doi.org/10.1038/s41598-019-46077-z
Moher D, Liberati A, Tetzlaff J, Altman DG (2009) Preferred Reporting Items for Systematic Reviews and Meta-Analyses: The PRISMA Statement. PLoS Med. https://doi.org/10.1371/journal.pmed.1000097
Soto-Quintana O, Zuniga-Gonzalez GM, Ramirez-Patino R, Ramos-Silva A, Figuera LE, Carrillo-Moreno DI, Gutierrez-Hurtado IA, Puebla-Perez AM, Sanchez-Llamas B, Gallegos-Arreola MP (2015) Association of the GSTM1 null polymorphism with breast cancer in a Mexican population. Genetics Mol Res: GMR 14(4):13066–13075. https://doi.org/10.4238/2015.October.26.2
Jaramillo-Rangel G, Ortega-Martinez M, Cerda-Flores RM, Barrera-Saldana HA (2015) Polymorphisms in GSTM1, GSTT1, GSTP1, and GSTM3 genes and breast cancer risk in northeastern Mexico. Genetics Mol Res: GMR 14(2):6465–6471. https://doi.org/10.4238/2015.June.11.22
Possuelo LG, Peraça CF, Eisenhardt MF, Dotto ML, Cappelletti L, Foletto E, Valim ARdM (2013) Polymorphisms of GSTM1 and GSTT1 genes in breast cancer susceptibility: a case-control study. Rev Brasil Ginecol Obstetr 35:569–574
Sohail A, Kanwal N, Ali M, Sadia S, Masood AI, Ali F, Iqbal F, Crickmore N, Shaikh RS, Sayyed AH (2013) Effects of glutathione-S-transferase polymorphisms on the risk of breast cancer: a population-based case-control study in Pakistan. Environ Toxicol Pharmacol 35(2):143–153. https://doi.org/10.1016/j.etap.2012.11.014
Martinez-Ramirez OC, Perez-Morales R, Castro C, Flores-Diaz A, Soto-Cruz KE, Astorga-Ramos A, Gonsebatt ME, Casas L, Valdes-Flores M, Rubio J (2013) Polymorphisms of catechol estrogens metabolism pathway genes and breast cancer risk in Mexican women. Breast (Edinburgh, Scotland) 22(3):335–343. https://doi.org/10.1016/j.breast.2012.08.004
Zgheib NK, Shamseddine AA, Geryess E, Tfayli A, Bazarbachi A, Salem Z, Shamseddine A, Taher A, El-Saghir NS (2013) Genetic polymorphisms of CYP2E1, GST, and NAT2 enzymes are not associated with risk of breast cancer in a sample of Lebanese women. Mutat Res 747–748:40–47. https://doi.org/10.1016/j.mrfmmm.2013.04.004
Campos CZ, Losi Guembarovski R, de Oliveira CEC, Banin Hirata BK, Vitiello GAF, Dias FL, Hiroki CH, Watanabe MAE, Mazzuco TL (2017) Glutathione S-transferases deletions may act as prognosis and therapeutic markers in breast cancer. Clin Exp Med 18(1):27–35. https://doi.org/10.1007/s10238-017-0461-6
Garcia-Martinez A, Gamboa-Loira B, Tejero ME, Sierra-Santoyo A, Cebrian ME, Lopez-Carrillo L (2017) CYP1A1, CYP1B1, GSTM1 and GSTT1 genetic variants and breast cancer risk in Mexican women. Salud Publica Mex 59(5):540–547. https://doi.org/10.21149/8527
Kiendrebeogo IT, Zoure AA, Sorgho PA, Yonli AT, Djigma FW, Ouattara AK, Sombie HK, Tovo SF, Yelemkoure ET, Bambara AH, Sawadogo AY, Bakri Y, Simpore J (2019) Glutathione S-transferase M1 (GSTM1) and T1 (GSTT1) variants and breast cancer risk in Burkina Faso. Biomol Concepts 10(1):175. https://doi.org/10.1515/bmc-2019-0020
Al-Eitan LN, Rababa'h DM, Alghamdi MA, Khasawneh RH (2019) Association of GSTM1, GSTT1 And GSTP1 Polymorphisms With Breast Cancer Among Jordanian Women. OncoTargets Therapy 12:7757–7765. https://doi.org/10.2147/ott.s207255
Sapcharoen K, Sanguansermsri P, Yasothornsrikul S, Muisuk K, Srikummool M (2019) Gene combination of CD44 rs187116, CD133 rs2240688, NF-kappaB1 rs28362491 and GSTM1 deletion as a potential biomarker in risk prediction of breast cancer in lower Northern Thailand. Asian Pac J Cancer Preven: APJCP 20(8):2493–2502. https://doi.org/10.31557/apjcp.2019.20.8.2493
Kalacas NA, Garcia JA, Sy Ortin T, Valdez A Jr, Fellizar A, Ramos MC, Albano PM (2019) GSTM1 and GSTT1 genetic polymorphisms and breast cancer risk in selected Filipino cases. Asian Pac J Cancer Preven: APJCP 20(2):529–535. https://doi.org/10.31557/apjcp.2019.20.2.529
Yuan P, Yuan L, Xu BL, Wang CZ, Yang HZ, Li Y (2015) Predictive potential role of glutathione S-transferases polymorphisms in response to chemotherapy and breast cancer prognosis. Genetics Mol Res: GMR 14(4):16675–16681. https://doi.org/10.4238/2015.December.11.15
Wang J, Wang T, Yin GY, Yang L, Wang ZG, Bu XB (2015) Glutathione S-transferase polymorphisms influence chemotherapy response and treatment outcome in breast cancer. Genetics Mol Res: GMR 14(3):11126–11132. https://doi.org/10.4238/2015.September.22.6
Zhou L, Huang A, Zhang D, Yao J, Zhang Y, Li X (2015) Genetic variability of glutathione S-transferases influences treatment outcome of breast cancer. Tumour Biol 36(8):5925–5929. https://doi.org/10.1007/s13277-015-3266-9
Tulsyan S, Chaturvedi P, Agarwal G, Lal P, Agrawal S, Mittal RD, Mittal B (2013) Pharmacogenetic influence of GST polymorphisms on anthracycline-based chemotherapy responses and toxicity in breast cancer patients: a multi-analytical approach. Mol Diagn Ther 17(6):371–379. https://doi.org/10.1007/s40291-013-0045-4
Liu J, Luo J, Wang Y, Li L, Yang S (2014) Predictive potential role of glutathione S-transferases polymorphisms on prognosis of breast cancer. Int J Clin Exp Pathol 7(12):8935–8940
Wang X, Huang ZH (2015) Predictive potential role of glutathione S-transferase polymorphisms in the prognosis of breast cancer. Genetics Mol Res: GMR 14(3):10236–10241. https://doi.org/10.4238/2015.August.28.7
Almeida M, Soares M, Ramalhinho AC, Moutinho JF, Breitenfeld L (2019) Prognosis of hormone-dependent breast cancer seems to be influenced by KEAP1, NRF2 and GSTM1 genetic polymorphisms. Mol Biol Rep 46(3):3213–3224. https://doi.org/10.1007/s11033-019-04778-8
Li S, Lang GT, Zhang YZ, Yu KD, Shao ZM, Zhang Q (2018) Interaction between glutathione S-transferase M1-null/present polymorphism and adjuvant chemotherapy influences the survival of breast cancer. Cancer Med 7(9):4202–4207. https://doi.org/10.1002/cam4.1567
He HR, You HS, Sun JY, Hu SS, Ma Y, Dong YL, Lu J (2014) Glutathione S-transferase gene polymorphisms and susceptibility to acute myeloid leukemia: meta-analyses. Jpn J Clin Oncol 44(11):1070–1081. https://doi.org/10.1093/jjco/hyu121
Liu K, Lin X, Zhou Q, Ma T, Han L, Mao G, Chen J, Yue X, Wang H, Zhang L, Jin G, Jiang J, Zhao J, Zou B (2014) The associations between two vital GSTs genetic polymorphisms and lung cancer risk in the Chinese population: evidence from 71 studies. PLoS ONE 9(7):e102372. https://doi.org/10.1371/journal.pone.0102372
Sui C, Ma J, He X, Wang G, Ai F (2014) Interactive effect of glutathione S-transferase M1 and T1 polymorphisms on hepatocellular carcinoma. Tumour Biol 35(8):8235–8241. https://doi.org/10.1007/s13277-014-2071-1
Elston CW, Ellis IO (1991) Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 19(5):403–410. https://doi.org/10.1111/j.1365-2559.1991.tb00229.x
Rakha EA, Reis-Filho JS, Baehner F, Dabbs DJ, Decker T, Eusebi V, Fox SB, Ichihara S, Jacquemier J, Lakhani SR, Palacios J, Richardson AL, Schnitt SJ, Schmitt FC, Tan PH, Tse GM, Badve S, Ellis IO (2010) Breast cancer prognostic classification in the molecular era: the role of histological grade. Breast cancer Res: BCR 12(4):207. https://doi.org/10.1186/bcr2607
Funding
Fundação Araucária, Programa de Pesquisa Para o SUS – PPSUS, Coordenação de Aperfeiçoamento de Pessoal de Nível Superior.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict to interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pacholak, L.M., Amarante, M.K., Guembarovski, R.L. et al. Polymorphisms in GSTT1 and GSTM1 genes as possible risk factors for susceptibility to breast cancer development and their influence in chemotherapy response: a systematic review. Mol Biol Rep 47, 5495–5501 (2020). https://doi.org/10.1007/s11033-020-05555-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05555-8