Abstract
Vitiligo is a common chronic depigmented skin disease characterized by melanocyte loss or dysfunction in the lesion. The pathogenesis of vitiligo has not been fully clarified. Most studies have suggested that the occurrence and progression of vitiligo are due to multiple factors and gene interactions in which noncoding RNAs contribute to an individual’s susceptibility to vitiligo. Noncoding RNAs, including microRNAs (miRNAs), are a hot topic in posttranscriptional regulatory mechanism research. miRNAs are noncoding RNAs with a length of approximately 22 nucleotides and play a negative regulatory role by binding to the 3′-UTR or 5′-UTR of the target mRNA to inhibit translation or initiate mRNA degradation. Previous studies have screened the differential expression profiles of miRNAs in the skin lesions, melanocytes, peripheral blood mononuclear cells (PBMCs) and sera of patients and mouse models with vitiligo. Moreover, several studies have focused on miRNA-25, miRNA-155 and other miRNAs involved in melanin metabolism, oxidative stress, and melanocyte proliferation and apoptosis. These miRNAs and regulatory processes further illuminate the pathogenesis of vitiligo and provide hope for the application of small molecules in the treatment of vitiligo. In this review, we summarize miRNA expression profiles in different tissues of vitiligo patients and the mechanisms by which key miRNAs mediate vitiligo development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vitiligo is a common pigment loss disease characterized by a significant decrease in melanocytes in the lesions [1, 2]. Depigmentation can occur on all parts of the body but is most commonly found on the back of the fingers, wrists, forearms, face, neck and around the genitals [3]. The theories of the pathogenesis of vitiligo are focused on autoimmunity, mental stress, oxidative stress, metabolic toxins and heredity [3,4,5]. However, the mechanism of melanocyte destruction or apoptosis has yet to be fully elucidated. An increasing number of studies suggest that the occurrence and progression of vitiligo is caused by multiple factors and is polygenic, and noncoding RNAs may play a vital role in an individual’s susceptibility to vitiligo [6].
Noncoding RNAs, including long non-coding RNA (lncRNA), microRNA (miRNA) and circular RNA (circRNA), play an important role in the regulation of gene expression. To our knowledge, there are still no studies on lncRNAs and circRNAs in the pathogenesis of vitiligo. In contrast, thus far, studies of noncoding RNA in the etiology of vitiligo have focused primarily on miRNAs.
miRNAs are a class of noncoding single-stranded RNA with a length of approximately 20–24 nucleotides [7]. During miRNA processing in the nucleus, primary miRNAs (pri-miRNAs) are first synthesized by RNA polymerase and then modified into precursor miRNAs (pre-miRNAs) by Drosha and its cofactor DGCR8. Pre-miRNAs are transported out of the nucleus through the Exportin-5/Ran-GTP complex and then cut by Dicer. miRNAs inhibit or degrade mRNA translation by binding to the 3′-UTR or 5′-UTR of the target mRNA at a posttranscriptional level [8,9,10,11]. Studies have shown that approximately 60% of human encoded protein genes are regulated by miRNAs [12] (Fig. 1).
Biogenesis of microRNA (miRNA). The initial miRNA transcript (pri-miRNA) is first processed by RNA polymerase III. Pri-miRNA has incomplete complementary double-stranded RNA regions and hairpin structures, which transformed into mature microRNA and underwent two consecutive splicing. In the nucleus, pri-miRNA is cut by Drosha enzyme to form the stem-loop structure of about 70 nt, that is, pre-miRNA. Pre-miRNA is then transported to the cytoplasm by the transporter Exportin-5 (Exp-5). Under the action of Dicer enzyme, a single chain structure of miRNA with a length of 20–24 is produced, forming a mature miRNA. Mature microRNA then binds to the RNA-induced silencing complex (RISC), mediates the silencing of post-transcriptional gene expression via mRNA deadenylating followed by degradation (partial complementarity.) or initiating mRNA cleavage activity (complete complementarity). pri-miRNAs, primary-miRNAs; pre-miRNAs, precursor miRNAs; RISC, The RNA-induced silencing complex
This review focuses on the expression profiles of miRNAs in different tissues of vitiligo patients and summarizes the latest knowledge of the molecular and cellular mechanisms of key miRNAs to pave the way for a new diagnosis and treatment for vitiligo.
miRNA expression profiles in vitiligo
miRNA differential expression profiles were obtained from skin lesion, melanocyte, peripheral blood mononuclear cells (PBMCs) and serum samples from patients with vitiligo [6, 13,14,15]. The results of the initial screen of the microarray, the PCR verification and the subsequent bioinformatics analysis provide an important basis for the in-depth study of the actions of miRNAs in the development of vitiligo (Table 1).
Skin miRNA expression profiles in vitiligo
Mansuri et al. found that there were 13 differentially expressed miRNAs in the lesion areas of nonsegmental vitiligo (NSV) when compared to the miRNAs in healthy controls. Among them, the expression levels of 12 miRNAs, including miRNA-1, miRNA-133b and miRNA-135a, in the skin lesions of NSV patients increased significantly, while the expression level of miRNA-211 decreased. Bioinformatics analysis reveals that these miRNAs have a coordinated role of oxidative stress and autoimmunity in melanocyte destruction and disease development and are critical for the development or susceptibility of NSV [6].
Another study suggested that compared to the nonpathological regions of patients with vitiligo, lesions had 29 significantly upregulated miRNAs, of which 6 miRNAs were transfected into normal human epidermal keratinocytes (NHEK), and the quantitative proteomics results showed that a melanogenesis protein, tyrosinase related protein-1 (TRP1/TYRP1), was downregulated after 6 miRNAs were transfected. Under physiological conditions, melanocytes synthesize and transfer melanosomes to peripheral keratinocytes. Further evidence suggests that the above miRNAs may act in an integrated manner to regulate the process of melanosome transfer [16].
Serum miRNA expression profile in vitiligo
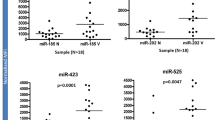
The expression of 20 miRNAs, including miRNA-146a and miRNA-191, in the sera of vitiligo mice was changed compared with that in the sera of control mice [17]. In another study from the same research group, the serum miRNA expression profiles of 10 patients with NSV and 20 healthy controls were examined; there were 31 differentially expressed miRNAs, of which 12 miRNAs had a fold-change of more than 3.0 times. Three of the miRNAs (miRNA-16, miRNA-19b and miRNA-720) seemed to be the best serum markers to differentiate NSV patients from healthy controls [18].
In one study of a Chinese population, 29 upregulated and 71 downregulated miRNAs were screened. GO analysis was used to assess the function of differentially expressed miRNA-associated predicted target genes, suggesting that the functions of the most significantly enriched target gene are focused on axon guidance, cell adhesion and cell junctions [15].
PBMC miRNA expression profile in vitiligo
miRNA microarray analysis was used to compare the miRNA expression profiles of PBMCs in the peripheral blood of patients with NSV and those of normal controls. Four miRNAs in the PBMCs of NSV patients showed significant differential expression, and the results showed that miRNA-224-3p, miRNA-2682-3p and miRNA-4712-3p expression was upregulated, and miRNA-3940-5p expression was significantly downregulated. The abovementioned miRNAs were further verified by PCR after the sample was added. The results showed that the expression of miRNA-224-3p and miRNA-4712-3p was upregulated in the PBMCs of NSV patients, and the expression of miRNA-3940-5p was downregulated, and these results are consistent with the results of the miRNA microarray data analysis. This study suggests that miRNAs may regulate T lymphocyte-mediated immune imbalance in vitiligo and that targeting these miRNAs may be a potential treatment for vitiligo [13].
Melanocyte miRNA expression profile in vitiligo
Hydrogen peroxide (H2O2) can induce oxidative stress in melanocytes and stimulate the pathological process of melanocyte injury in vitiligo [19, 20]. By differential analysis of the miRNA microarray expression profile, Zhang et al. found that 10 miRNAs in PIG1 melanocytes (a strain of normal human skin melanocytes) changed after H2O2 treatment, and 7 miRNAs were upregulated and 3 were downregulated. In PIG3V melanocytes, an immortalized melanocyte cell line from vitiligo patients, three miRNAs were upregulated and 5 were downregulated when treated with H2O2. Interestingly, after H2O2 treatment, three miRNAs (miRNA-93, miRNA-320b and miRNA-423-5p) were altered in both melanocyte types. These three miRNAs were further verified by RT-PCR. The trend between the miRNA microarray and PCR is consistent. The expression of miRNA-93 and miRNA-320b in PIG1 melanocytes decreased after H2O2 treatment. The expression of miRNA-423-5p in the two kinds of melanocytes was higher after H2O2 treatment than before treatment, and the expression level of PIG3V cells was significantly different before and after treatment. The results were consistent with the microarray results [21].
Pivotal miRNAs
miRNA-155-5p
miRNA-155-5p (miRNA-155) is considered to be a common oncogenic miRNA involved in many biological processes, such as hematopoietic cell differentiation, immune cell differentiation, inflammatory and immune response, muscle development and adipose differentiation [22,23,24]. In situ hybridization analysis showed that miRNA-155 expression increased in the epidermis of patients with vitiligo. In human primary melanocytes and keratinocytes, vitiligo-related cytokines induced the production of miRNA-155, and miRNA-155 inhibited the expression of melanogenesis-related genes and changed the expression of IFN-regulated genes in melanocytes and keratinocytes. It is suggested that miRNA-155 is involved in the pathogenesis of vitiligo [25] (Fig. 2).
Multiple miRNAs are involved in vitiligo. miRNA-155 is involved in abnormal melanin metabolism. miRNA-211, miRNA-25 and miRNA-423 regulate oxidative stress during melanocyte injury. SCF, supercoiling factor; bFGF, fibroblast growth factor 2; YAP1, Yes associated protein 1; IL-2R, interleukin 2 receptor; PPARGC1A, PPARG coactivator 1 alpha; GSTM1, Glutathione S-transferase Mu 1; TYR, tyrosinase; MITF, melanogenesis associated transcription factor; TGF-BR2, transforming growth factor beta receptor 2
Interestingly, miRNA-155 was found to be down-regulated in T cells from patients with vitiligo. Further research found that miRNA-155 agonists significantly reduced the growth rate of CD8+ T cells and promoted the proliferation of melanocytes by inducing an increase in the percentage of Treg cells. In contrast, miRNA-155 antagonists inhibited melanocyte proliferation by reducing the percentage of Treg cells. In short, miRNA-155 protects melanocyte survival by increasing the number of Treg cells and reducing the number of CD8+ T cells, which may provide new prospects for the treatment of vitiligo [26].
miRNA-211-5p
miRNA-211 positively regulates pigment production by targeting transforming growth factor beta receptor 2 (TGFBR-2). TGFBR-2 is considered to be a negative regulator of tyrosinase (TYR) and TYRP-1 [27]. Therefore, miRNA-211 plays an important role in the regulation of melanocyte biology and development. One study found that miRNA-211 acts as a metabolic switch in nonpigmented melanoma cells [28]. Another study confirmed one aspect of the above conclusion. This microarray study found that the expression of miRNA-211 was significantly decreased in the lesions of vitiligo and in the vitiligo cell line PIG3V. Of all the predicted miRNA-211 targets, analysis revealed an inverse upregulation of peroxisome proliferative activated receptor, gamma, coactivator 1 alpha (PPARGC1A), ribonucleotide reductase regulatory subunit M2 (RRM2) and TAO kinase 1 (TAOK1), which was confirmed with RNA-seq and gene expression array data derived from biopsy samples taken from patients with vitiligo (GEO accession series GSE53148) [29]. In vitro, it was further found that miRNA-211 can bind to the 3′-UTR site of PPARGC1A/PGC1-α and inhibit its expression. The increased production of reactive oxygen species in vitiligo cells may be partly due to the abnormal expression of miRNA-211 and its subsequent regulation of its target gene. miRNA-211 coated with nanoparticles can improve the oxygen consumption rate of PIG3V cells to a certain extent, so miRNA-211 and its target genes may be potential biomarkers and therapeutic targets for vitiligo [30,31,32].
miRNA-25-5p
The serum level of miRNA-25 in patients with vitiligo was increased, and this serum level was related to the activity of vitiligo. The reason may be that oxidative stress can upregulate the level of miRNA-25 in melanocytes and keratinocytes by inducing demethylation of the miRNA-25 promoter region. Similarly, the recovery of miRNA-25 promoted H2O2-induced damage to melanocytes and resulted in melanocyte dysfunction. In this process, MITF, a target gene of miRNA-25, is involved in the damage to melanocytes. Further experiments have shown that miRNA-25 can inhibit the production and secretion of stem cell factor (SCF) and basic fibroblast growth factor (bFGF) by keratinocytes and thus inhibit the paracrine protection of melanocytes under oxidative stress [33]. Therefore, it is worthwhile to study the possibility of miRNA-25 as a target of antioxidation therapy for vitiligo.
miRNA-423-5p
The expression of miRNA-423 in the immortalized human epidermal melanocytes PIG1 and the vitiligo melanocytes PIG3V was upregulated after H2O2 treatment. Therefore, miRNA-423 may be involved in the oxidative stress response of melanocytes [21]. This study further revealed that miRNA-423 can decrease the viability of human melanocytes and promote melanocyte apoptosis under the oxidative stress induced by H2O2. It was proven that miRNA-423 may be a risk factor for regulating oxidative stress damage in human melanocytes. Further studies showed that miRNA-423 mediated oxidative stress injury in human melanocytes by targeting GSTM1, which is a key antioxidant component of human melanocytes, and has significant antioxidant capacity [21, 34].
miRNA-202-3p
miRNA-202-3p may participate in the occurrence and development of vitiligo. Overexpression of miRNA-202-3p can significantly inhibit the proliferation and adhesion of human melanocytes, inhibit melanin synthesis and promote apoptosis. Silencing miRNA-202-3p can promote melanin synthesis but has no obvious effect on cell proliferation, adhesion and apoptosis. Yes1 associated transcriptional regulator (YAP1) may be a direct target gene of miRNA-202-3p, and it affects the expression of p73 and BCL2 binding component 3 (PUMA) in cell proliferation and apoptosis and affects the biological function of normal melanocytes [14].
miRNA-3940-5p
Vitiligo is an immune disorder characterized by the presence of activated T lymphocytes in the skin and the destruction of melanocytes, which is one of the main mechanisms of the pathogenesis of NSV [35,36,37]. The expression of miRNA-3940 in the PBMCs of patients with NSV was found to be downregulated. Following the construction of the miRNA-3940-inhibited T lymphoblastic leukemia cell line HuT78, interleukin 2 receptor γ (IL-2Rγ), an immune-associated gene of vitiligo, has been confirmed to be the target gene of miRNA-3940. After miRNA-3940 was inhibited, the proliferation rate of HuT78 cells was significantly upregulated. Based on these results, miRNA-3940 may be a novel miRNA that plays a potential role in the mechanism of immune imbalance in vitiligo [38].
miRNA-9-5p
The skin microenvironment is critical to normal melanocyte function [39]. Human vitiligo lesions showed a decrease in adhesion molecules, such as E-cadherin and β1 integrin [36, 40, 41]. Therefore, the reduction in the adhesion of melanocytes to the adjacent keratinocytes and the loss of melanocytes from the epidermis are key steps in the pathogenesis of vitiligo [42]. The melanocytes adhere and migrate to the edge regions to initiate the repigmentation process [43]. miRNA-9 is involved in the adhesion and migration of melanocytes during UVB-induced vitiligo repigmentation [35]. In this process, Su M et al found that UVB could increase the level of IL-10 in HaCaT cells and trigger the methyl modifications of miRNA-9 induced by the methyltransferase DNMT3A, thereby reducing the level of miRNA-9. It was suggested that miRNA-9 targeted and inhibited E-cadherin and β1 integrin in HaCaT cells and inhibited PIG1 cell migration when HaCaT cells were exposed to UVB. Therefore, once miRNA-9 is inhibited, the migration of PIG1 cells to HaCaT cells can be improved. To be more optimistic, interfering with miRNA-9 may provide an opportunity to treat vitiligo.
miRNA polymorphism
Single nucleotide polymorphism (SNP) is the most common gene variation in human genes and can affect gene expression, transcription and modification [44, 45]. It is one of the important reasons for disease susceptibility, and individual and ethnic differences in drug reactivity. Studies have shown that SNPs of miRNAs are associated with many human diseases, which can affect the phenotype or development of the disease [46,47,48].
Four SNP loci of miRNAs, including miRNA-146a rs2910164, miRNA-149 rs2292832, miRNA-196a2 rs11614913, and miRNA-499 rs3746444, were identified from 400 known human pre-miRNAs in patients with vitiligo and controls. It was found that only individuals with the miRNA-196a2 CC genotype had lower susceptibility to vitiligo than patients with the combined genotype of TT and TC. The genotypes of miRNA-196a2 rs11614913 CC combined with TT and TC were further analyzed by stratification [49]. A subsequent study showed that rs11614913 T > C can reduce the early apoptosis rate of melanocytes and has a protective effect against melanocyte apoptosis under the oxidative stress induced by H2O2. TRP-1 was proven to be the target gene of miRNA-196a2 [50].
Further study evaluated the potential association between the rs11614913 SNP in miRNA-196a2 and the serum tyrosine (Tyr) levels in 116 patients with vitiligo and 116 controls. Interestingly, individuals with a TT + TC genotype in miRNA-196a2 and high serum Tyr levels had a higher risk of vitiligo than patients with lower CC genotypes and low Tyr levels. In addition, the rs11614913 C allele enhanced its inhibitory effect on Tyr expression, and its downregulation in melanocytes successfully reduced intracellular ROS levels and apoptotic rates [51]. In summary, it suggested that miRNA-196a2 polymorphisms can regulate Tyr levels and thus affect susceptibility to vitiligo.
Conclusion
The differential expression of miRNAs that have been reported provides an opportunity for us to understand the pathogenesis of vitiligo. Regulating miRNA and its downstream signaling pathway may provide more potentially valuable options for the prevention and treatment of vitiligo.
References
Ezzedine K, Whitton M, Pinart M (2016) Interventions for vitiligo. JAMA 316(16):1708–1709. https://doi.org/10.1001/jama.2016.12399
Ezzedine K, Eleftheriadou V, Whitton M, van Geel N (2015) Vitiligo. Lancet 386(9988):74–84. https://doi.org/10.1016/S0140-6736(14)60763-7
Delmas V, Larue L (2019) Molecular and cellular basis of depigmentation in vitiligo patients. Exp Dermatol 28(6):662–666. https://doi.org/10.1111/exd.13858
Singh RK, Lee KM, Vujkovic-Cvijin I, Ucmak D, Farahnik B, Abrouk M et al (2016) The role of IL-17 in vitiligo: a review. Autoimmun Rev 15(4):397–404. https://doi.org/10.1016/j.autrev.2016.01.004
Jacquemin C, Taieb A, Boniface K, Seneschal J, Fhu A (2019) Imbalance of peripheral follicular helper T lymphocyte subsets in active vitiligo. Pigm Cell Melanoma Res 32(4):588–592. https://doi.org/10.1111/pcmr.12763
Mansuri MS, Singh M, Dwivedi M, Laddha NC, Marfatia YS, Begum R (2014) MicroRNA profiling reveals differentially expressed microRNA signatures from the skin of patients with nonsegmental vitiligo. Br J Dermatol 171(5):1263–1267. https://doi.org/10.1111/bjd.13109
Li Y, Jin X, Wang Z, Li L, Chen H, Lin X et al (2019) Systematic review of computational methods for identifying miRNA-mediated RNA-RNA crosstalk. Brief Bioinform 20(4):1193–1204. https://doi.org/10.1093/bib/bbx137
Grijalvo S, Alagia A, Jorge AF, Eritja R (2018) Covalent strategies for targeting messenger and non-coding RNAs: an updated review on siRNA, miRNA and antimiR conjugates. Genes. https://doi.org/10.3390/genes9020074
Sarkar D, Leung EY, Baguley BC, Finlay GJ, Askarian-Amiri ME (2015) Epigenetic regulation in human melanoma: past and future. Epigenetics 10(2):103–121. https://doi.org/10.1080/15592294.2014.1003746
Mione M, Bosserhoff A (2015) MicroRNAs in melanocyte and melanoma biology. Pigm Cell Melanoma Res 28(3):340–354. https://doi.org/10.1111/pcmr.12346
Lee I, Ajay SS, Yook JI, Kim HS, Hong SH, Kim NH et al (2009) New class of microRNA targets containing simultaneous 5’-UTR and 3’-UTR interaction sites. Genome Res 19(7):1175–1183. https://doi.org/10.1101/gr.089367.108
Cai Y, Yu X, Hu S, Yu J (2009) A brief review on the mechanisms of miRNA regulation. Genom Proteom Bioinform 7(4):147–154. https://doi.org/10.1016/S1672-0229(08)60044-3
Wang Y, Wang K, Liang J, Yang H, Dang N, Yang X et al (2015) Differential expression analysis of miRNA in peripheral blood mononuclear cells of patients with non-segmental vitiligo. J Dermatol 42(2):193–197. https://doi.org/10.1111/1346-8138.12725
Shang Z (2017) The study of biological effects and molecular mechanism of miR-202-3p targeting YAP1 in melanocyte [Ph.D]. Zhengzhou University
Shang Z, Li H (2017) Altered expression of four miRNA (miR-1238-3p, miR-202-3p, miR-630 and miR-766-3p) and their potential targets in peripheral blood from vitiligo patients. J Dermatol 44(10):1138–1144. https://doi.org/10.1111/1346-8138.13886
Vaish U, Kumar AA, Varshney S, Ghosh S, Sengupta S, Sood C et al (2019) Micro RNAs upregulated in citiligo skin play an important role in its aetiopathogenesis by altering TRP1 expression and keratinocyte-melanocytes cross-talk. Sci Rep 9(1):10079. https://doi.org/10.1038/s41598-019-46529-6
Shi YL, Weiland M, Lim HW, Mi QS, Zhou L (2014) Serum miRNA expression profiles change in autoimmune vitiligo in mice. Exp Dermatol 23(2):140–142. https://doi.org/10.1111/exd.12319
Shi YL, Weiland M, Li J, Hamzavi I, Henderson M, Huggins RH et al (2013) MicroRNA expression profiling identifies potential serum biomarkers for non-segmental vitiligo. Pigm Cell Melanoma Res 26(3):418–421. https://doi.org/10.1111/pcmr.12086
He Y, Li S, Zhang W, Dai W, Cui T, Wang G et al (2017) Dysregulated autophagy increased melanocyte sensitivity to H2O2-induced oxidative stress in vitiligo. Scientific reports 7:42394. https://doi.org/10.1038/srep42394
Qiu L, Song Z, Setaluri V (2014) Oxidative stress and vitiligo: the Nrf2-ARE signaling connection. J Invest Dermatol 134(8):2074–2076. https://doi.org/10.1038/jid.2014.241
Zhang Y (2011) The mechanism research of miR-423-5p regulated human melanocyte oxidative stress injury induced by hydrogen peroxide [Ph.D]. Fourth Military Medical University
Gu J, Lu Z, Ji C, Chen Y, Liu Y, Lei Z et al (2017) Melatonin inhibits proliferation and invasion via repression of miRNA-155 in glioma cells. Biomed Pharmacother 93:969–975. https://doi.org/10.1016/j.biopha.2017.07.010
Alivernini S, Gremese E, McSharry C, Tolusso B, Ferraccioli G, McInnes IB et al (2017) MicroRNA-155-at the critical interface of innate and adaptive immunity in arthritis. Front Immunol 8:1932. https://doi.org/10.3389/fimmu.2017.01932
El-Lithy GM, El-Bakly WM, Matboli M, Abd-Alkhalek HA, Masoud SI, Hamza M (2016) Prophylactic l-arginine and ibuprofen delay the development of tactile allodynia and suppress spinal miR-155 in a rat model of diabetic neuropathy. Transl Res 177:85-97 e1. https://doi.org/10.1016/j.trsl.2016.06.005
Sahmatova L, Tankov S, Prans E, Aab A, Hermann H, Reemann P et al (2016) MicroRNA-155 is dysregulated in the skin of patients with vitiligo and inhibits melanogenesis-associated genes in melanocytes and keratinocytes. Acta Dermato-Venereol 96(6):742–747. https://doi.org/10.2340/00015555-2394
Lv M, Li Z, Liu J, Lin F, Zhang Q, Li Z et al (2019) MicroRNA155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo. Mol Med Rep 20(4):3617–3624. https://doi.org/10.3892/mmr.2019.10607
Dai X, Rao C, Li H, Chen Y, Fan L, Geng H et al (2015) Regulation of pigmentation by microRNAs: MITF-dependent microRNA-211 targets TGF-beta receptor 2. Pigm Cell Melanoma Res 28(2):217–222. https://doi.org/10.1111/pcmr.12334
Mazar J, Qi F, Lee B, Marchica J, Govindarajan S, Shelley J et al (2016) MicroRNA 211 functions as a metabolic switch in human melanoma cells. Mol Cell Biol 36(7):1090–1108. https://doi.org/10.1128/MCB.00762-15
Rashighi M, Agarwal P, Richmond JM, Harris TH, Dresser K, Su MW et al (2014) CXCL10 is critical for the progression and maintenance of depigmentation in a mouse model of vitiligo. Sci Transl Med 6(223):223ra23. https://doi.org/10.1126/scitranslmed.3007811
Sahoo A, Lee B, Boniface K, Seneschal J, Sahoo SK, Seki T et al (2017) MicroRNA-211 regulates oxidative phosphorylation and energy metabolism in human vitiligo. J Invest Dermatol 137(9):1965–1974. https://doi.org/10.1016/j.jid.2017.04.025
Sahoo A, Sahoo SK, Joshi P, Lee B, Perera RJ (2019) MicroRNA-211 loss promotes metabolic vulnerability and BRAF inhibitor sensitivity in melanoma. J Invest Dermatol 139(1):167–176. https://doi.org/10.1016/j.jid.2018.06.189
Spiegelman VS, Elcheva IA, Metabo-miR (2017) miR-211 regulates mitochondrial energy metabolism in vitiligo. J Invest Dermatol 137(9):1828–1830. https://doi.org/10.1016/j.jid.2017.06.012
Shi Q, Zhang W, Guo S, Jian Z, Li S, Li K et al (2016) Oxidative stress-induced overexpression of miR-25: the mechanism underlying the degeneration of melanocytes in vitiligo. Cell Death Differ 23(3):496–508. https://doi.org/10.1038/cdd.2015.117
Lu L, Wu W, Tu Y, Yang Z, He L, Guo M (2014) Association of glutathione S-transferase M1/T1 polymorphisms with susceptibility to vitiligo. Gene 535(1):12–16. https://doi.org/10.1016/j.gene.2013.11.024
Su M, Yi H, He X, Luo L, Jiang S, Shi Y (2019) miR-9 regulates melanocytes adhesion and migration during vitiligo repigmentation induced by UVB treatment. Exp Cell Res 384(1):111615. https://doi.org/10.1016/j.yexcr.2019.111615
Reichert Faria A, Jung JE, Silva de Castro CC, de Noronha L (2017) Reduced immunohistochemical expression of adhesion molecules in vitiligo skin biopsies. Pathol Res Pract 213(3):199–204. https://doi.org/10.1016/j.prp.2016.12.019
Boniface K, Taieb A, Seneschal J (2016) New insights into immune mechanisms of vitiligo. G Ital Dermatol Venereol 151(1):44–54
Wang Y, Wang K, Dang N, Wang L, Zhang M (2016) Downregulation of miR-3940-5p promotes T-cell activity by targeting the cytokine receptor IL-2R gamma on human cutaneous T-cell lines. Immunobiology 221(12):1378–1381. https://doi.org/10.1016/j.imbio.2016.07.008
Speeckaert R, Speeckaert M, De Schepper S, van Geel N (2017) Biomarkers of disease activity in vitiligo: a systematic review. Autoimmun Rev 16(9):937–945. https://doi.org/10.1016/j.autrev.2017.07.005
Mohan GC, Silverberg JI (2015) Association of vitiligo and alopecia areata with atopic dermatitis: a systematic review and meta-analysis. JAMA Dermatol 151(5):522–528. https://doi.org/10.1001/jamadermatol.2014.3324
Bordignon M, Castellani C, Fedrigo M, Thiene G, Peserico A, Alaibac M et al (2013) Role of alpha5beta1 integrin and MIA (melanoma inhibitory activity) in the pathogenesis of vitiligo. J Dermatol Sci 71(2):142–145. https://doi.org/10.1016/j.jdermsci.2013.04.005
Lee AY (2012) Role of keratinocytes in the development of vitiligo. Ann Dermatol 24(2):115–125. https://doi.org/10.5021/ad.2012.24.2.115
Majid I, Imran S (2015) Targeted ultraviolet B phototherapy in vitiligo: a comparison between once-weekly and twice-weekly treatment regimens. Indian J Dermatol Venereol Leprol 81(6):600–605. https://doi.org/10.4103/0378-6323.168325
Jadeja SD, Mansuri MS, Singh M, Dwivedi M, Laddha NC, Begum R (2017) A case-control study on association of proteasome subunit beta 8 (PSMB8) and transporter associated with antigen processing 1 (TAP1) polymorphisms and their transcript levels in vitiligo from Gujarat. PLoS ONE 12(7):e0180958. https://doi.org/10.1371/journal.pone.0180958
Mansuri MS, Jadeja SD, Singh M, Laddha NC, Dwivedi M, Begum R (2017) The catalase gene promoter and 5′-untranslated region variants lead to altered gene expression and enzyme activity in vitiligo. Br J Dermatol 177(6):1590–1600. https://doi.org/10.1111/bjd.15681
Ahangari F, Salehi R, Salehi M, Khanahmad H (2014) A miRNA-binding site single nucleotide polymorphism in the 3’-UTR region of the NOD2 gene is associated with colorectal cancer. Med Oncol 31(9):173. https://doi.org/10.1007/s12032-014-0173-7
Jia W, Zeng L, Luo S, Bai F, Zhong R, Wu L et al (2018) Association of microRNA-423 rs6505162 C > A polymorphism with susceptibility and metastasis of colorectal carcinoma. Medicine 97(6):e9846. https://doi.org/10.1097/MD.0000000000009846
Speeckaert R, van Geel N, Vitiligo (2017) An update on pathophysiology and treatment options. Am J Clin Dermatol 18(6):733–744. https://doi.org/10.1007/s40257-017-0298-5
Huang Y (2012) Single nucleotide polymorphism and functional analysis of miR-196a2 and vitiligo susceptibility[D]. Fourth Military Medical University
Huang Y, Yi X, Jian Z, Wei C, Li S, Cai C et al (2013) A single-nucleotide polymorphism of miR-196a-2 and vitiligo: an association study and functional analysis in a Han Chinese population. Pigm Cell Melanoma Res 26(3):338–347. https://doi.org/10.1111/pcmr.12081
Cui TT, Yi XL, Zhang WG, Wei C, Zhou FB, Jian Z et al (2015) miR-196a-2 rs11614913 polymorphism is associated with vitiligo by affecting heterodimeric molecular complexes of Tyr and Tyrp1. Arch Dermatol Res 307(8):683–692. https://doi.org/10.1007/s00403-015-1563-1
Acknowledgements
Supported by grants from the National Natural Science Foundation of China (NSFC; Grant Nos. 81560502, 81960354), the National Natural Science Foundation of Yunnan Province (Grant No. 2017FB116), the Talent Project of Yunnan Province (2019HB024), 100 Talents Program of Kunming Medical University (Lechun Lyu). The authors acknowledge the editors and reviewers for their positive and constructive comments and suggestions on our study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yan, S., Shi, J., Sun, D. et al. Current insight into the roles of microRNA in vitiligo. Mol Biol Rep 47, 3211–3219 (2020). https://doi.org/10.1007/s11033-020-05336-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05336-3