Abstract
The brown planthopper (Nilaparvata lugens Stål, BPH) is the most destructive pest of rice (Oryza sativa L.). Utilizing resistant rice cultivars that harbor resistance gene/s is an effective strategy for integrated pest management. Due to the co-evolution of BPH and rice, a single resistance gene may fail because of changes in the virulent BPH population. Thus, it is urgent to explore and map novel BPH resistance genes in rice germplasm. Previously, an indica landrace from India, Paedai kalibungga (PK), demonstrated high resistance to BPH in both in Wuhan and Fuzhou, China. To map BPH resistance genes from PK, a BC1F2:3 population derived from crosses of PK and a susceptible parent, Zhenshan 97 (ZS97), was developed and evaluated for BPH resistance. A novel BPH resistance locus, BPH39, was mapped on the short arm of rice chromosome 6 using next-generation sequencing-based bulked segregant analysis (BSA-seq). BPH39 was validated using flanking markers within the locus. Furthermore, near-isogenic lines carrying BPH39 (NIL-BPH39) were developed in the ZS97 background. NIL-BPH39 exhibited the physiological mechanisms of antibiosis and preference toward BPH. BPH39 was finally delimited to an interval of 84 Kb ranging from 1.07 to 1.15 Mb. Six candidate genes were identified in this region. Two of them (LOC_Os06g02930 and LOC_Os06g03030) encode proteins with a similar short consensus repeat (SCR) domain, which displayed many variations leading to amino acid substitutions and showed higher expression levels in NIL-BPH39. Thus, these two genes are considered reliable candidate genes for BPH39. Additionally, transcriptome sequencing, DEGs analysis, and gene RT-qPCR verification preliminary revealed that BPH39 may be involved in the jasmonic acid (JA) signaling pathway, thus mediating the molecular mechanism of BPH resistance. This work will facilitate map-based cloning and marker-assisted selection for the locus in breeding programs targeting BPH resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa) is one of the most crucial food crops globally, and the brown planthopper (Nilaparvata lugens Stål, BPH) is the most destructive pest in rice-growing areas. Given the escalating severity and frequency of pest outbreaks over the past few decades, these have posed a substantial threat to rice production (Du et al. 2020). Global warming exacerbates the population growth and metabolic rates of pests, while environmental deterioration may reduce the number of natural pest enemies, foreseeably leading to heightened crop losses (Deutsch et al. 2018). Controlling pests is a key priority for ensuring rice yield (Crist et al. 2017). Presently, chemical insecticides predominantly serve as the means to control BPH. However, the abuse of pesticides not only escalates farmers' production and labor costs but also causes irreversible harm on the environment. Consequently, the use of such pesticides will decline in many developing countries. The adoption of resistant rice cultivars harboring resistance genes proves to be an effective strategy for integrated pest management. Breeding insect-resistant varieties aligns with the modern trend in rice breeding and stands as a key objective within the realm of green super rice (Zhang 2007). Due to the co-evolution of BPH and rice, a single resistance gene may falter owing to alterations in the BPH virulent population. It is imperative to explore and map novel BPH resistance genes in rice germplasm.
The inheritance of resistance to BPH in rice has been a focal point of research. Since the identification of BPH resistance genes BPH1 and bph2 in 1971 (Athwal et al. 1971), more than 40 genes have been mapped in cultivated and wild rice. The majority of these genes exhibit dominant or partially dominant, with only a few being recessive, including bph2, bph4, bph5 and bph29. BPH resistance genes are typically preferentially located in gene clusters on rice chromosomes 3, 4, 6 and 12 (Cheng et al. 2013b). For example, 11 genes and/or QTLs (BPH30, BPH33, BPH12, BPH15, BPH3/17, BPH20 (t), bph22 (t), qBPH4, qBPH4.2, qBPH4.3, qBPH4.4) cluster within a 0.91–8.9 Mb region between markers H99 and B44 on the short arm of chromosome 4 (4S) (Rahman et al. 2009; HOU et al. 2011; Qiu et al. 2012; Lv et al. 2014; Hu et al. 2015a, b, 2018; Liu et al. 2015; Mohanty et al. 2017; Wang et al. 2018). Eight genes (BPH1, bph2, bph7, BPH9, BPH10, BPH18, BPH21 and BPH26) cluster within a 5 Mb region on the long arm of chromosome 12 (12L) (Ishii et al. 1994; Sun et al. 2006; Cha et al. 2008; Rahman et al. 2009; Tamura et al. 2014; Qiu et al. 2014; Ji et al. 2016; Zhao et al. 2016). These genes may represent alleles or distinct genes, but there is insufficient evidence to differentiate between them.
With the rapid and in-depth advancement of rice functional genomics, map-based cloning has been employed to sequentially clone a range of BPH resistance genes. To date, 17 resistance genes have been cloned in rice, including genes such as BPH1, bph2, BPH3, BPH6, bph7, BPH9, BPH10, BPH14, BPH15, BPH18, BPH21, BPH26, bph29, BPH30, BPH32, BPH37 and BPH40 (Du et al. 2009; Cheng et al. 2013a; Tamura et al. 2014; Liu et al. 2015; Wang et al. 2015; Ji et al. 2016; Zhao et al. 2016; Ren et al. 2016; Guo et al. 2018; Shi et al. 2021; Zhou et al. 2021). BPH14 was the first BPH resistance gene cloned on chromosome 3L from O. officinalis. BPH14 encodes a member of the NLR family, known as the CC-NB-LRR (coiled-coil, nucleotide-binding, and leucine-rich repeat) protein, often observed in disease resistance proteins (Du et al. 2009). Previous research revealed that the CC and NB domains of BPH14 protein can interact with WRKY transcription factors (WRKY46 and WRKY72) to activate defense signals, thereby resisting feeding by BPH (Hu et al. 2017). Recent research has found that Bph14-mediated hypersensitive response (HPR) is achieved through the direct binding of BISP and BPH14 to the selective autophagy cargo receptor OsNBR1, which delivers BISP to OsATG8 for degradation. In susceptible plants, BISP directly targets OsRLCK185 to suppress basal defenses (Guo et al. 2023). Eight genes on chromosome 12L also encode NLR proteins. They were identified as alleles of each other and divided into four allelotypes, showing resistance to different BPH biotypes. This reveals that allelic variation of insect resistance genes is an essential strategy for rice to cope with pest virulent variation (Zhao et al. 2016). BPH32, located on chromosome 6S, encodes a specifically conserved short repeat (SCR) domain protein. BPH32 is highly expressed in leaf sheaths and may inhibit BPH feeding (Ren et al. 2016). Recently, a new broad-spectrum resistance gene, BPH6, was successfully cloned in the indica rice Swarnalata. BPH6 encodes an atypical LRR protein localized to the exocyst, where it interacts with the exocyst subunit EXO70E1 to regulate rice cell secretion and maintain cell wall integrity. It was found that OsEXO70H3 interacted with SAMSL, increasing the delivery of SAMSL outside the cells, leading to lignin deposition in cell walls, thus hindering BPH feeding (Guo et al. 2018; Wu et al. 2022). BPH30, located on chromosome 4S, belongs to a novel gene family with two leucine-rich domains. It is strongly expressed in sclerenchyma cells, enhancing cellulose and hemicellulose synthesis, making the cell walls stiffer and fortifying the sclerenchyma. BPH40 is another member of the family, and the resistance mechanism is similar to that of BPH30, related to cell wall fortification in sclerenchyma to prevent BPH stylets from reaching the phloem to feed (Shi et al. 2021). Zhou et al. used GWAS studies to identify a previously unidentified gene, BPH37, containing the CC and NB domains but lacking the LRR domain, exhibiting high resistance against BPH herbivory (Zhou et al. 2021).
Previously, approximately 300 cultivar accessions were introduced by the International Rice Research Institute (IRRI), potentially possessing resistance to multiple BPH biotypes. Following the evaluation of BPH resistance at the seedling stage in these cultivars, more than 190 cultivars exhibiting moderate resistance and above were screened (Hu et al. 2016). Among them, an indica landrace from India, named Paedai kalibungga (PK), demonstrated high resistance to BPH in Fuzhou, China. To map BPH resistance genes in PK, a BC1F2:3 population resulting from crosses of PK and a susceptible parent, Zhenshan 97 (ZS97), was developed and assessed for BPH resistance. Our study focused on (1) primarily mapping BPH resistance genes using mixed-pool sequencing (BSA-seq); (2) fine mapping the locus to a region smaller than 100 Kb through progeny testing; (3) analyzing candidate genes through comparative sequencing and gene expression studies; (4) conducting transcriptome analysis and initial verification of differential gene expression to elucidate the molecular mechanisms of BPH39-mediated resistance to brown planthoppers.
Materials and methods
Plant materials and mapping population
Paedai kalibungga (abbreviated as PK, IRGC 38545) and PTB33, both of which originating from India, are well supplied by the International Rice Research Institute (IRRI). PTB33 (carrying BPH32) was used as a resistant control for sequencing the BPH32 gene and BPH evaluation. Two Chinese cultivars, TN1 and Zhenshan 97 (abbreviated as ZS97), were used as susceptible controls.
The construction of the primary mapping population consists of the following steps: The susceptible parent, ZS97, was continuously crossed with the resistant parents, PK and F1, to obtain 16 BC1F1 plants. After evaluating BPH resistance, five resistant plants were selfed to obtain BC1F2. Ninety-two BC1F2 individuals were planted and selfed, subsequently, the corresponding 92 seeds were harvested, forming the primary mapping population. The secondary or fine mapping population was the BC3F2 (near-isogenic lines, NILs), developed by marker-assisted backcross breeding (MABC).
BPH materials
BPH insects were first captured in paddy fields in Fuzhou, China, and reared and maintained on a susceptible indica rice variety TN1 in a walk-in artificial climate light greenhouse in our lab. The temperature of the greenhouse was controlled at 28 ± 2 °C and the humidity was 75 ± 5%.
Molecular marker design
InDel markers were developed based on the sequence difference between PK and ZS97, and the genomic variations of 4,726 cultivars in the database of RiceVarMap (http://ricevarmap.ncpgr.cn/v2/) described by Zhao et al.35 A segment of 3–8 bp insertion or deletion between the two parental varieties was found in sequence data and the website, and then specific primers were designed at a distance of 50–60 bp on either side of this segment, so that the size of the amplified fragment containing the deletion target was about 100 bp. The molecular markers required for mapping BPH39 are listed in Table S1.
BSA-seq
The bulked segregant analysis method using next-generation sequencing (BSA-seq) was described by Hu et al. (2018).
Construction of a high-resolution physical map and fine mapping of locus
Approximately 5,000 BC3F2 seeds with a heterozygous genotype at the target gene locus were sown in multiple 96-well plates, The PCR plate containing the seeds, which had holes at the bottom, was then placed in the germination box, and a small amount of water was added before the box was tightly covered. About one week after germination, almost 4,500 surviving seedlings were selected, and each seedling was fixed and sampled sequentially to extract rice leaf DNA. According to the results of preliminary mapping, all seedlings were analyzed for genotypes using closely linked markers on both sides of the target gene, and individuals with recombination exchange between markers on both sides (one side was marked as homozygous ZS97 genotype B or homozygous PK genotype A, and the other side was marked as heterozygous genotype H) were selected. After transplanting all recombination exchange individuals into the field for planting, mature plants were harvested and selfed for subsequent genotype and phenotype analysis. Uniformly distributed markers with polymorphisms between the two parents were developed in the target locus region, and marker genotype analysis was performed on all recombinants to construct a physical map of the target locus based on the physical location of the marker on the chromosome. Two to three individuals of each recombination breakpoint type were selected for progeny testing (genotype and phenotype analysis). Based on the results of the progeny tests of recombinants, we can more accurately delimit the mapping interval of target genes.
Candidate gene prediction
Candidate gene prediction was undertaken based on rice genome annotation databases, namely the Rice Genome Annotation Project Database at (MSU) (http://rice.uga.edu/cgi-bin/gbrowse/rice/). By combining protein structure and function data from gene annotations, possible candidate genes can be predicted in advance. At the same time, one or two candidate genes were determined by the sequence alignment of the two parents in the fine-mapping region of the locus.
RNA isolation and RT-qPCR analysis
For the two-leaf and one-heart seedling stage, three biological replicates were used for all RNA experiments. Total RNAs were isolated from rice leaf sheaths using the TRIzol reagent (Takara). First-strand cDNA was synthesized according to the manufacturer’s instructions (All-in-One First-Strand Synthesis MasterMix, Lablead, F0202-100 T). RT-qPCRs were performed using a CFX Connect Real-Time System (Bio-Rad) and 2 × Taq PCR StarMix with Loading Dye (GenStar, A002-100–1), following the manufacturer’s instructions and using the primers listed in Table S2.
RNA-seq analysis
The NIL-BPH39 and ZS97 plants were grown to two leaves and one heart, and each plant was infested with 10 third-instar BPH nymphs. All treatments had three biological replicates, and the samples were collected from the infested (72 h) and control (0 h) plants simultaneously (Guo et al. 2018). Total RNAs were isolated using the same method as above. The cDNA libraries were checked using Qubit 2.0 and Agilent 2100 before they were sequenced by Illumina HiSeq2500/x (https://doi.org/10.21203/rs.3.rs-634003/v1). Transcriptome profiling was carried out by Wuhan Metware Biotechnology Co., Ltd. Wuhan, China (http://www.metware.cn/; https://doi.org/10.1016/j.molp.2020.06.005). All raw data were deposited into the National Center for Biotechnology Information under accession number (PRJNA1006526) and were publicly accessible at https://www.ncbi.nlm.nih.gov/. Differential expression analysis of four experimental groups, GO functional annotation, and KEGG pathway enrichment analysis were performed based on the differentially expressed genes.
Evaluation of BPH resistance
The improved methods of seedling bulk testing and seedling death rate were performed to evaluate the BPH resistance. When rice seedlings grow to the stage of two leaves and one heart, they are transferred to a black box filled with rice liquid medium. Eight plants were retained in each row (one variety in each row), and the process was repeated three times. The boxes were placed in a gauze net, and 2 days after transplanting, 8–10 s-instar nymphs were infested per seedling. When the dead seedling rate of the susceptible control was greater than 90%, the resistance level of a seedling was indicated by a resistance score (RS = 1, 3, 5, 7 or 9) and the resistance level of a line was indicated by the average RS of 10 seedlings. For the seedling death rate test, the resistance level of each line was evaluated based on the dead seedling rate of each row as follows: Mortality 0–1.0%, immunization; 1.1–10.0%, high resistance; 10.1–30.0%, resistance; 30.1–50.0%, moderate resistance; 50.1–70.0%, moderate susceptibility; 70.1–100%, susceptibility.
In addition, the amount, weight and area of honeydew excreted by BPH were measured. Honeydew weight and area measurements were performed using the method described by Pathak et al. (1982) with slight modifications (Hu et al. 2018). Fifteen replicates were tested.
Results
Phenotypic distribution of the genetic population
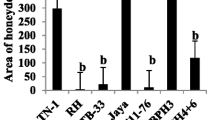
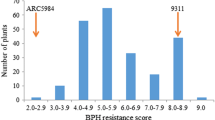
We evaluated the BPH resistance of a BC1F2:3 population comprising 92 families derived from crosses between ZS and PK using three experimental methods: seedling resistance, and the weight and area of honeydew excreted by BPH. The seedling bulk test was performed twice, in September 2017 and April 2018, respectively. The results showed that the mean seedling resistance score (RS) of the resistant parent PK was 2.0, indicating resistance, while the RS of the susceptible parent ZS was 8.9, indicating high susceptibility. The RS of 92 families showed a continuous distribution between 2.0 and 9.0, the results in 2017 had two obvious peaks at 4.0–4.9 and 6.0–6.9, and the results in 2018 had only one obvious peak at 6.0–6.9. According to previous studies (Qiu et al. 2012, 2014), we defined an RS of 0–6.9 as resistance and 7.0–9.0 as susceptibility. After statistical analysis, the resistance ratio was 62:30 in 2017 and 67:25 in 2018 (Fig. 1a, b). The results of the chi-square test showed that all observations met the Mendelian genetic segregation ratio of at 3:1 for single genes (χ2 = 2.84 and 0.23 < χ20.05, 1 = 3.84 in 2017 and 2018, respectively). The honeydew weight and area were continuously distributed between 1.3–11.9 mg and 0.6–12.0 cm2, respectively. The honeydew weight and area of the parent ZS (8.7 mg, 11.6 cm2) were significantly higher than those of PK (2.5 mg, 1.9 cm2), respectively (Fig. 1c, d). The above results indicated that the resistance of the resistant parent PK might be controlled by a dominant gene.
Frequency distribution of BPH resistance in BC1F2:3 population of ZS/PK. a and b, seedling resistance to BPH in 2017 and 2018, respectively; c and d, Honeydew weight and area secreted by BPH, respectively. PK (Paedai kalibungga) and ZS (Zhenshan 97) denote the phenotypic values of the resistant and susceptible parents, respectively. e, Mapping of BPH39 using the method of BSA-seq based on the BC1F2:3 populations from ZS/PK. AF, allele frequency difference of individual marker between the two opposite DNA pools. The arrow indicates the peak where BPH39 was located
Previous studies have shown that the RS is positively correlated with honeydew weight and area, that is, the lower the RS, the smaller the honeydew weight and area of BPH, and vice versa. In this study, there was a highly significant positive correlation between RS in 2017 and 2018 (correlation coefficient 0.88), as well as a highly significant positive correlation between honeydew area and honeydew weight amount (correlation coefficient 0.85). However, there was a small yet significant positive correlation between seedling resistance and honeydew weight or area (Table S3).
Preliminary mapping of BPH39 by BSA-seq
In order to map the BPH resistance locus in the parent PK, we selected 15 extremely resistant and 15 extremely susceptible lines to construct resistance and susceptible pools, respectively, in the BC1F2:3 population based on the RS data. Then we performed an SNP frequency difference analysis covering the 12 chromosomes of rice using BSA-seq and found a maximum peak in the interval of 0–5 Mb on the short arm of chromosome 6 (6S). This indicating that there may be a major gene in this region, which was tentatively named BPH39 in accordance with the naming rules for BPH resistance loci (Fig. 1e).
Verification of effect for BPH39
BPH39 was preliminarily mapped to the interval of 0–5 Mb on chromosome 6S by BSA-seq. To verify the QTL, we developed 8 molecular markers in this region and adjacent regions (0.93–19.51 Mb) (Table S1). These eight markers were then used to genotype 92 lines in the F2:3 population. Combined with the phenotype of seedling resistance, a t-test was performed to group the genotypes A and B using these markers, and the confidence interval of BPH39 was verified based on the significant difference results. A genotype with the same marker pattern was classified as homozygous dominant (genotype A), homozygous recessive (genotype B), or heterozygous (genotype H). The RS of genotype A was compared with the RS of genotype B by the t test. We can see that the RS of genotype A is significantly different from that of genotype B except for the two molecular markers I828 and RM541 (P > 0.05). The smallest P-value at marker I992 is 2.48 × 10–7. As the physical location of a marker gets closer to I992, the P-value decreases, whereas it increases when further away (Fig. 2). The above results indicate that there is indeed a major resistance locus near marker I992 on chromosome 6S, and BPH39 was preliminary located between the markers I369 and I392.
Marker linkage analysis of marker genotype and phenotype in BPH39 region on chromosome 6S. a-h, the BPH39 genotypes and phenotypes identified according to its linkage markers I369, I992, RM190, I532, RM204, I219, I928 and RM451. A, PK genotype; B, ZS genotype; H, heterozygote genotype; P-values are shown for the comparison between A and B
Development and evaluation of a near-isogenic line carrying BPH39
To facilitate the precise mapping of this locus and to establish a marker-assisted selection (MAS) framework for its use in insect resistance breeding, we successfully generated a near-isogenic line (NIL) of BPH39 within the genetic background of Zhenshan 97. We first selected four resistant individuals with a homozygous PK genotype (A) on both sides of BPH39 in the BC1F3 population and backcrossed them with ZS97 for two consecutive generations using MAS. They were then selfed for one generation, and finally, the BC3F2 population with segregation at the BPH39 locus were developed. Using MAS, we obtained three individuals with homozygous PK genotypes and three with homozygous ZS97 genotypes, corresponding to the near-isogenic line carrying BPH39 (NIL-BPH39) and the near-isogenic line not containing BPH39 (ZS97), respectively.
We detected the genetic background of NIL-BPH39, using 48 polymorphic markers distributed on 12 rice chromosomes. The results showed that there were 1 and 3 marker genotypes in the two independent near isogenic lines of BPH39, B39-1 and B39-2, respectively, which were different from the recurrent parent ZS97, and the background recovery rates were 97.9% and 93.8%, respectively (Figure S1, S2). B39-1 was used in this study. The above results indicate that the genetic background of the near-isogenic lines developed in this experiment is clean enough to be used for fine mapping and effect evaluation of the BPH resistance QTL.
BPH resistance was evaluated on the NILs. The results showed that the resistant NIL-BPH39 was still alive, while the susceptible ZS97 was dead (Fig. 3a). The honeydew area and weight gain of BPH on ZS97 were significantly greater than those of NIL-BPH39 (Fig. 3b-e). We monitored the survival rate of BPH nymphs continuously over seven days. The number of nymphs steadily decreased on the plants, but it declined quickly in the NIL-BPH39. After seven days of infestation, the BPH survival rate for NIL-BPH39 was only 50%, compared to 82% for ZS97 (Fig. 3f).
Phenotypic of BPH resistance of near isogenic line carrying BPH39 (NIL-BPH39). a, the photo of seedling resistance of NIL-BPH39 and ZS97 at the time of 12 days after BPH infestation; b, Color photos of honeydew secretion of BPH after ninhydrin staining. The honeydew is shown in purple red. c, Honeydew area of BPH. d, Honeydew wight of BPH. e, Weight gain of BPH. f, Survival rate of BPH. g, Preference of settled BPH. ZS97 and NIL-BPH39 denote NIL that do not carry and carry the homozygous BPH39 locus, respectively. * represents a significant difference at the 0.05 level
Furthermore, we investigated the number of settled BPHs on NIL-BPH39 and ZS97. The results showed that the number of BPH on NIL-BPH39 was significantly less than that on ZS97 after 6 h of infestation. This difference became most pronounced 48 h after infestation, with an average of only 15 BPHs on NIL-BPH39 and 33 BPHs on ZS97 (Fig. 3g). Taken together, these results indicate that BPH39 has preferential and antibiotic effects on the insects.
Fine mapping of BPH39
To fine-map the locus, we planted 4,500 individuals of BC3F2 that were heterozygous at both flanking markers I369 and I992. Marker genotype analysis was performed on 4,500 individuals, and a total of 26 recombinants were screened with exchange at markers on both sides. At the same time, a total of eight additional polymorphic markers were developed and evenly distributed in this region (Table S1), and then a high-resolution physical map of BPH39 was constructed based on the physical location of these markers on chromosome 6S (Fig. 4a).
Fine mapping of BPH39. a Primary physical map of part of chromosome 6S containing BPH39. b Physical map of marker interval I369-I992 harboring BPH39. The numbers in brackets show the times of recombination occurred between the marker loci and BPH39 among 4,500 individuals. c Graphical genotypes and resistance phenotypes of the recombinants. Black, white and grey bars denote PK, ZS and recombined segments, respectively. HR, highly resistant; MR, moderately resistant; S, susceptible. d Annotated genes within the final 85 kb interval based on Nipponbare reference from MSU database
After genotyping the 26 recombinants using eight markers, various exchange types were classified according to the recombination breakpoints at the markers. We selected two to three recombinants for each exchange type and conducted progeny testing based on BPH resistance. Only the recombinants R32-8, R32-2, and R32-19 did not show recombination in the region between markers I7494 and I1540. The left and right boundaries of the locus were determined by the recombinants R32-41, R32-18, R32-45, and R32-19, R32-9, R32-6, R32-44, respectively. The recombinants with resistant homozygous or heterozygous genotypes at marker I1432 were resistant to BPH, while the plants with susceptible homozygous genotypes were susceptible. This indicated that the marker genotype of I1432 and the phenotype co-segregated. Taken together with the above analysis of recombinants’ genotypes and progeny testing, BPH39 was finally fine-mapped within markers I7494 and I1540, corresponding to an interval size of 84 Kb based on the Nipponbare reference genome (Fig. 4b, c).
Candidates of BPH39
Six genes were identified in the 84 Kb region according to the Rice Genome Annotation Project Database (MSU). LOC_Os06g02930 and LOC_Os06g03030 encode a conserved hypothetical protein. LOC_Os06g02960 encodes a DUF594 family protein of unknown function. LOC_Os06g02980 encodes a subunit of the plasma membrane H + -ATPase involved in nutrient transport. LOC_Os06g03050 encodes a DUF594 protein containing an unknown functional domain. LOC_Os06g03080 encodes an aspartic protease (Fig. 4d). We sequenced those genes of the two cultivars, NIL-BPH39 and ZS97, and aligned them with the reference genome Nipponbare (NPB), resulting in multiple variations between NIL-BPH39 and ZS97. Among them, two genes, LOC_Os06g02930 and LOC_Os06g03030, exhibited the most variations—11 SNPs and one insertion, and 5 SNPs and one deletion, respectively—resulting in substitutions and alterations of the encoded amino acids. A base substitution led to premature termination of amino acid translation at 643 bp in LOC_Os06g02930 in NIL-BPH39 (Table 1). The proteins of other candidate genes had only a few amino acid substitutions and insertions or deletions (Table S4). We analyzed the gene expression levels of candidate genes in near-isogenic lines before and after BPH infestation. The results showed that the expression levels of only LOC_Os06g02930 and LOC_Os06g03030 in the resistant line were significantly higher than those in the susceptible line (Fig. 5). Based on the above results, we believe that LOC_Os06g02930 and LOC_Os06g03030 may be reliable candidate genes for BPH39.
Expression analysis of BPH39 candidate genes in the two NILs with contrary BPH39 genotypes. The histogram and the left ordinate axis represent the relative expression of the genes detected by RT-qPCR. SM, RM represent ZS97 and NIL-BPH39 with no infestation. ST and RT indicate ZS97 and NIL-BPH39 at 48 h after BPH infestation. The expression levels of all genes in SM are set to the value of ‘1’. Error bar indicates standard deviation. The letter a, b, c and d on the error bar indicates significant difference in the process of multiple comparison and analysis of variance, in which the same letters indicate that there is no significant difference at 0.05 level, while different letters indicate significant differences. The line chart and the right ordinate axis represent the FPKM value of the gene expression detected by RNA-seq. The orange shaded part represents the standard error
RNA sequencing and analysis of DEGs
In order to explore which genes were involved in the resistance to BPH, we performed transcriptome sequencing and differentially expressed gene (DEG) analysis of NIL-BPH39 and ZS97 before and after BPH infestation. The analysis revealed a total of 51 differentially expressed genes that were consistently present across all four treatment groups, with 17 of them exhibiting upregulation. These identified genes potentially play a crucial role in the BPH39-mediated insect resistance response (Fig. 6a). KEGG enrichment analysis showed that DEGs between RT (NIL-BPH39 after BPH infestation) and ST (ZS97 after BPH infestation) were mainly concentrated in the phenylpropanoid biosynthesis and diterpenoid biosynthesis pathways (Fig. 6b). The Gene Ontology (GO) enrichment analysis showed that DEGs between RT and ST were mainly concentrated in the terms 'oxidation–reduction', 'response to stress and stimulus', 'defense response', and 'programmed cell death' in the biological process category (Fig. 6c).
DEGs analysis by RNA-seq in the two NILs with contrary BPH39 genotypes. a, Venn diagram of DEGs in the comparison groups. b and c, DEGs of KEGG and GO enrichment results in comparison group of RT-ST, respectively. d and e, DEGs of heat maps of JA-related genes and transcript factors (TFs), respectively. SM, RM, ST and RT, see legends in Fig. 5
Numerous studies have shown the involvement of the JA, ETH, ABA, and SA plant hormone signaling pathways in the anti-insect response mechanism of BPH feeding, as well as their regulatory role in conferring resistance to BPH. It was found that 10 of the differentially expressed genes screened were significantly upregulated during BPH feeding induction (Fig. 6d), which also indicated that the plant hormones JA, ETH, ABA, and SA were involved in anti-insect molecular responses in rice-brown planthopper interactions. As can be seen from Fig. 6e, transcription factors play an important role in plant responses to biological stresses such as insect resistance and disease resistance. The expression levels of OsWRKY50, OsWRKY81, OsWRKY65, OsWRKY52, OsMYB63, OsMYB2 and OsbHLH6 were significantly up-regulated in NIL-BPH39 after BPH feeding.
Quantitative RT-qPCR verification
To verify the expression levels of insect-resistant genes before and after BPH infestation in NIL-BPH39 and ZS97 using RNA-Seq, we selected several JA-related genes for RT-qPCR verification. The expression trends of OsbHLH6, OsJAZ10, OsJAZ13 and OsWRKY81 were completely consistent with RNA-Seq data, with all being significantly upregulated in NIL-BPH39 (Fig. 7). The trends for the other seven genes were similar to the RNA-Seq results, and all related genes, except OsLOX8 in the sensitive lines, were significantly upregulated after BPH infestation. Among them, OsPR10, OsLOX12 and Bphi008a were up-regulated hundreds of times after BPH infestation, suggesting that they were strongly induced by BPH feeding. Other genes were also upregulated by tens of times and similarly induced by BPH.
Discussion
The discovery, identification and cloning of BPH resistance genes is the most basic and important work of rice insect resistance genetics and breeding, and it is also an important component of rice functional genome research. An important finding of this study was the identification of a novel BPH resistance locus, BPH39, in an Indian rice line and its fine mapping to an 84 Kb region within markers I7494 and I1540 on chromosome 6S. According to the reported localization information of the BPH resistance genes, we found that at least five other genes were mapped on chromosome 6S, which were BPH3, bph4, BPH32, bph29, BPH25 and BPH37. A recessive gene, bph4, was mapped near this interval between SSR markers RM589 and MR586 (Jairin et al. 2010). BPH32 was derived from a resistant variety, Ptb33 (an Indian variety bred at Pattambi (Ptb) in Kerala state), which was mapped between markers RM19291 and RM8072. This locus was later cloned and its exact location was delimited to be between markers RM19291 and I32-4 (Ren et al. 2016). bph29, (previously named bph20 (t)), a gene derived from Oryza rufipogon Griff, was initially found to be localized between markers BYL7 and BYL8, and subsequently, the gene was finely mapped within the 24 Kb region between BYL8 and BID2, and was finally cloned (Yang et al. 2012; Wang et al. 2015). Myint et al. (2012) mapped gene BPH25 in a highly resistant line, ADR52, from India and found it spanned a relatively large interval between markers S00310 and RM8101, encompassing 1.7 Mb. This interval also includes the aforementioned genes: bph29, BPH3, BPH32, BPH39 and bph4 (Fig. 8). Combining the above information, we speculate that BPH3 and BPH32 are likely to be the same gene because they are both mapped in the same insect-resistant line Ptb33, by two different research teams. BPH39 is located between bph29 and bph4. In addition, BPH39 is a dominant locus, while bph29 and bph4 are both recessive genes. The localization interval of BPH39 in this study adjoined that of BPH32, so we first had to figure out the relationship between the two genes.
In order to understand the relationship between BPH39 and BPH32, we sequenced the BPH32 gene in NIL-BPH39 and ZS97, and found that Ptb33 (BPH32) and NIL-BPH39 had a total of 23 variations in the BPH32 gene, which also resulted in amino acid sequence changes (Table S5). In addition, it is also the most important evidence that during the fine mapping of BPH39, 12 recombinants displayed a genotype at marker I324 (located within the BPH32 gene) that was opposite to the phenotype, that is, recombination occurred, indicating that BPH39 is likely not allelic to BPH32 (Fig. 4). Taken together, these results preliminarily indicate that BPH39 is different from BPH32, but subsequent transgenic trials are needed to complete the cloning of BPH39 before the relationship between them can be thoroughly clarified.
Six candidate genes were identified in the 84 Kb region containing BPH39. Among them, two genes, LOC_Os06g02930 and LOC_Os06g03030, had the most variations resulting in the substitution and insertion of encoded amino acids between NIL-BPH39 and ZS97 (Table 1). Furthermore, the expression levels of these two genes in resistant line were significantly higher than those in susceptible line (Fig. 5). They all encode similar conserved hypothetical proteins with short consensus repeat (SCR) domain, which are very similar to the cloned BPH resistance gene BPH32 (Ren et al. 2016). Based on the above results, we believe that LOC_Os06g02930 and LOC_Os06g03030 are reliable candidate genes for BPH39. These two genes will subsequently be used for genetic transformation verification.
Hormones play an important role in plants' responses to biotic and abiotic stresses. Salicylic acid (SA), jasmonic acid (JA) and ethylene (ETH) are common defense hormones. JA, in particular, plays a major role in the response to mechanical damage caused by insects. It has been reported that several rice genes in the JA signaling pathway are involved in the defense response against BPH. In the Bph6 near-isogenic line, the active JA substance, JA-ILE, was induced by insect pests and was found to be significantly higher than in the susceptible variety 9311, indicating that JA played an important role in BPH6-mediated resistance (Guo et al. 2018). Lipoxygenase (LOX) and allene oxide synthase (AOS) are the first and second key enzymes in JA biosynthesis, respectively. Studies have shown that OsAOS1, OsAOS2, OsLOX1 and OsLOX9 are involved in herbivorous-induced JA synthesis, thereby mediating plant resistance to insects (Wang et al. 2008; Ma et al. 2020; Zeng et al. 2021). In this study, transcriptome and RT-qPCR confirmed that two LOX-related genes, OsLOX8 and OsLOX12, were strongly induced by BPH in the resistant line NIL-BPH39. Jasmonate carboxymethyltransferase gene OsJMT1 plays a regulatory role in rice defense against BPH by altering the levels of JA and its related metabolites (Qi et al. 2016). Our study also found that the expression of OsJMT2 (homologous gene of OsJMT1) was strongly induced in NIL-BPH39. In addition, two JAZ protein genes, OsJAZ10 and OsJAZ13, as well as the JA-induced transcription factors OsbHLH6 and OsWRKY81, were significantly upregulated in NIL-BPH39 (Fig. 7). A series of BPH resistance genes involved in the JA signaling and synthesis pathway have been identified, indicating that JA plays an important role in mediated rice resistance to BPH (Zhou et al. 2023). Interestingly, a previously identified BPH resistance gene Bphi008a (Hu et al. 2011), is also significantly upregulated in the NIL-BPH39, suggesting that they may be related in some way. In conclusion, these preliminary results suggest that Bph39 may be involved in the JA signaling pathway, thereby mediating the defense response of BPH.
Conclusions
A novel BPH resistance gene BPH39 was finely mapped within an interval of 84 Kb on rice chromosome 6S. Two candidate genes for BPH39, LOC_Os06g02930 and LOC_Os06g03030, were identified by parental comparative sequencing and gene expression analysis. Transcriptome sequencing, DEG analysis, and gene RT-qPCR verification preliminarily revealed that BPH39 may be involved in the JA signaling pathway, thus mediating the molecular mechanism of BPH resistance. This work will facilitate map-based cloning and marker-assisted selection of the locus in BPH resistant breeding.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Athwal DS, Pathak MD, Bacalangco EH, Pura CD (1971) Genetics of resistance to brown planthoppers and green leafhoppers in Oryza sativa L.1. Crop Sci 11:747–750. https://doi.org/10.2135/cropsci1971.0011183X001100050043x
Cha Y-S, Ji H, Yun D-W et al (2008) Fine Mapping of the Rice Bph1 Gene, which Confers Resistance to the Brown Planthopper (Nilaparvata lugens Stal), and Development of STS Markers for Marker-assisted Selection. Genome 26:146–151. https://doi.org/10.1016/S1016-8478(23)13976-8
Cheng X, Wu Y, Guo J et al (2013a) A rice lectin receptor-like kinase that is involved in innate immune responses also contributes to seed germination. Plant J 76:687–698. https://doi.org/10.1111/tpj.12328
Cheng X, Zhu L, He G (2013b) Towards Understanding of Molecular Interactions between Rice and the Brown Planthopper. Mol Plant 6:621–634. https://doi.org/10.1093/mp/sst030
Crist E, Mora C, Engelman R (2017) The interaction of human population, food production, and biodiversity protection. Science 356:260–264. https://doi.org/10.1126/science.aal2011
Deutsch CA, Tewksbury JJ, Tigchelaar M et al (2018) Increase in crop losses to insect pests in a warming climate. Science 361:916–919. https://doi.org/10.1126/science.aat3466
Du B, Zhang W, Liu B et al (2009) Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Nucleic Acids Res 106:22163–22168. https://doi.org/10.1073/pnas.0912139106
Du B, Chen R, Guo J, He G (2020) Current understanding of the genomic, genetic, and molecular control of insect resistance in rice. Mol Breeding 40:24. https://doi.org/10.1007/s11032-020-1103-3
Guo J, Xu C, Wu D et al (2018) Bph6 encodes an exocyst-localized protein and confers broad resistance to planthoppers in rice. Nat Genet 50:297–306. https://doi.org/10.1038/s41588-018-0039-6
Guo J, Wang H, Guan W et al (2023) A tripartite rheostat controls self-regulated host plant resistance to insects. Nature 618:799–807. https://doi.org/10.1038/s41586-023-06197-z
Hou L, Yu P, Xu Q et al (2011) Genetic Analysis and Preliminary Mapping of Two Recessive Resistance Genes to Brown Planthopper, Nilaparvata lugens Stål in Rice. J Integr Plant Biol 18:238–242. https://doi.org/10.1016/S1672-6308(11)60033-4
Hu J, Zhou J, Peng X et al (2011) The Bphi008a gene interacts with the ethylene pathway and transcriptionally regulates MAPK genes in the response of rice to brown planthopper feeding. J Exp Bot 156:856–872. https://doi.org/10.1104/pp.111.174334
Hu J, Xiao C, Cheng M et al (2015a) A new finely mapped Oryza australiensis-derived QTL in rice confers resistance to brown planthopper. Gene 561:132–137
Hu J, Xiao C, Cheng M et al (2015b) Fine mapping and pyramiding of brown planthopper resistance genes QBph3 and QBph4 in an introgression line from wild rice O. officinalis. Mol Breeding 35:3. https://doi.org/10.1007/s11032-015-0228-2
Hu J, Xiao C, He Y (2016) Recent progress on the genetics and molecular breeding of brown planthopper resistance in rice. Rice 9:30. https://doi.org/10.1186/s12284-016-0099-0
Hu L, Wu Y, Wu D et al (2017) The coiled-coil and nucleotide binding domains of brown planthopper resistance14 function in signaling and resistance against planthopper in rice. Euphytica 29:3157–3185. https://doi.org/10.1105/tpc.17.00263
Hu J, Chang X, Zou L et al (2018) Identification and fine mapping of Bph33, a new brown planthopper resistance gene in rice (Oryza sativa L.). Rice 11:55. https://doi.org/10.1186/s12284-018-0249-7
Ishii T, Brar DS, Multani DS, Khush GS (1994) Molecular tagging of genes for brown planthopper resistance and earliness introgressed from Oryza australiensis into cultivated rice, O. sativa. Euphytica 37:217–221. https://doi.org/10.1139/g94-030
Jairin J, Sansen K, Wongboon W, Kothcharerk J (2010) Detection of a brown planthopper resistance gene bph4 at the same chromosomal position of Bph3 using two different genetic backgrounds of rice. Breeding Sci 60:71–75. https://doi.org/10.1270/jsbbs.60.71
Ji H, Kim S-R, Kim Y-H et al (2016) Map-based cloning and characterization of the BPH18 gene from wild rice conferring resistance to brown planthopper (BPH) insect pest. Sci Rep-uk 6:34376. https://doi.org/10.1038/srep34376
Liu Y, Wu H, Chen H et al (2015) A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice. Nat Biotechnol 33:301–305. https://doi.org/10.1038/nbt.3069
Lv W, Du B, Shangguan X et al (2014) BAC and RNA sequencing reveal the brown planthopper resistance gene BPH15 in a recombination cold spot that mediates a unique defense mechanism. BMC Genom 15:674. https://doi.org/10.1186/1471-2164-15-674
Ma F, Yang X, Shi Z, Miao X (2020) Novel crosstalk between ethylene- and jasmonic acid-pathway responses to a piercing–sucking insect in rice. New Phytol 225:474–487. https://doi.org/10.1111/nph.16111
Mohanty SK, Panda RS, Mohapatra SL et al (2017) Identification of novel quantitative trait loci associated with brown planthopper resistance in the rice landrace Salkathi. Euphytica 213:1–15. https://doi.org/10.1007/s10681-017-1835-2
Myint KKM, Fujita D, Matsumura M et al (2012) Mapping and pyramiding of two major genes for resistance to the brown planthopper (Nilaparvata lugens [Stål]) in the rice cultivar ADR52. Theor Appl Genet 124:495–504. https://doi.org/10.1007/s00122-011-1723-4
Pathak PK, Saxena RC, Heinrichs EA (1982) Parafilm sachet for measuring honeydew excretion by nilaparvata lugens on Rice1. J Econ Entomol 75:194–195. https://doi.org/10.1093/jee/75.2.194
Qi J, Li J, Han X et al (2016) Jasmonic acid carboxyl methyltransferase regulates development and herbivory-induced defense response in rice. J Integr Plant Biol 58:564–576. https://doi.org/10.1111/jipb.12436
Qiu Y, Guo J, Jing S et al (2012) Development and characterization of japonica rice lines carrying the brown planthopper-resistance genes BPH12 and BPH6. Theor Appl Genet 124:485–494. https://doi.org/10.1007/s00122-011-1722-5
Qiu Y, Guo J, Jing S et al (2014) Fine mapping of the rice brown planthopper resistance gene BPH7 and characterization of its resistance in the 93–11 background. Euphytica 198:369–379. https://doi.org/10.1007/s10681-014-1112-6
Rahman ML, Jiang W, Chu SH et al (2009) High-resolution mapping of two rice brown planthopper resistance genes, Bph20(t) and Bph21(t), originating from Oryza minuta. J Integr Plant Biol 119:1237–1246. https://doi.org/10.1007/s00122-009-1125-z
Ren J, Gao F, Wu X et al (2016) Bph32, a novel gene encoding an unknown SCR domain-containing protein, confers resistance against the brown planthopper in rice. Sci Rep-uk 6:37645. https://doi.org/10.1038/srep37645
Shi S, Wang H, Nie L et al (2021) Bph30 confers resistance to brown planthopper by fortifying sclerenchyma in rice leaf sheaths. Mol Plant 14:1714–1732. https://doi.org/10.1016/j.molp.2021.07.004
Sun L-H, Wang C-M, Su C-C et al (2006) Mapping and marker-assisted selection of a brown planthopper resistance gene bph2 in rice (Oryza sativa L.). J Integr Plant Biol 33:717–723. https://doi.org/10.1016/S0379-4172(06)60104-2
Tamura Y, Hattori M, Yoshioka H et al (2014) Map-based cloning and characterization of a brown planthopper resistance gene BPH26 from oryza sativa L. ssp. indica cultivar ADR52. Sci Rep-uk 4:5872. https://doi.org/10.1038/srep05872
Wang R, Shen W, Liu L et al (2008) A novel lipoxygenase gene from developing rice seeds confers dual position specificity and responds to wounding and insect attack. Plant Mol Biol 66:401–414. https://doi.org/10.1007/s11103-007-9278-0
Wang Y, Cao L, Zhang Y et al (2015) Map-based cloning and characterization ofBPH29, a B3 domain-containing recessive gene conferring brown planthopper resistance in rice. J Exp Bot 66:6035–6045. https://doi.org/10.1093/jxb/erv318
Wang H, Shi S, Guo Q et al (2018) High-resolution mapping of a gene conferring strong antibiosis to brown planthopper and developing resistant near-isogenic lines in 9311 background. Mol Breeding 38:1–10. https://doi.org/10.1007/s11032-018-0859-1
Wu D, Guo J, Zhang Q et al (2022) Necessity of rice resistance to planthoppers for OsEXO70H3 regulating SAMSL excretion and lignin deposition in cell walls. New Phytol 234:1031–1046. https://doi.org/10.1111/nph.18012
Yang L, Li RB, Li YR et al (2012) Genetic mapping of bph20(t) and bph21(t) loci conferring brown planthopper resistance to Nilaparvata lugens Stål in rice (Oryza sativa L.). Euphytica 183:161–171. https://doi.org/10.1007/s10681-011-0437-7
Zeng J, Zhang T, Huangfu J et al (2021) Both allene oxide synthases genes are involved in the biosynthesis of herbivore-induced jasmonic acid and herbivore resistance in rice. J Integr Plant Biol 10:442. https://doi.org/10.3390/plants10030442
Zhang Q (2007) Strategies for developing green super rice. Crop Sci 104:16402–16409. https://doi.org/10.1073/pnas.0708013104
Zhao Y, Huang J, Wang Z et al (2016) Allelic diversity in an NLR gene BPH9 enables rice to combat planthopper variation. P Natl Acad Sci USA 113:12850–12855. https://doi.org/10.1073/pnas.1614862113
Zhou C, Zhang Q, Chen Y et al (2021) Balancing selection and wild gene pool contribute to resistance in global rice germplasm against planthopper. J Integr Plant Biol 63:1695–1711. https://doi.org/10.1111/jipb.13157
Zhou S, Gao Q, Chen M et al (2023) Silencing a dehydration-responsive element-binding gene enhances the resistance of plants to a phloem-feeding herbivore. Plant Cell Environ 46:3090–3101. https://doi.org/10.1111/pce.14569
Funding
This work was supported in part by the National Natural Science Foundation of China (32025031, U1905203), Fujian Provincial Natural Science Foundation (2021J01131016), National Key Research and Development Program of China (2023YFF1000500), Fujian Agriculture and Forestry University School Innovation Fund (KFA20006A), and The Joint Open Competitive Project of the Yazhou Bay Seed Laboratory and China National Seed Company Limited (B23YQ1514, B23CQ15EP).
Author information
Authors and Affiliations
Contributions
Yangdong Ye and Yanan Wang performed experiments and analyzed the data, Ling Zou, Fangming Zhang, Shangye Xiong, Baohui Liang, Cheng Chen and Zhihong Zhu collected plant materials and performed parts of experiments, Shuai Zhang, Xiaoqing Wu, and Weiren Wu participated in handling figures and tables, Jie Hu, Jianguo Wu and Shuai Zhang, the co-correspondence authors, designed this research and wrote the manuscript. All authors have approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
We declare that these experiments comply with the ethical standards in China.
Conflict of interests
No conflict of interest exits in the submission of this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ye, Y., Wang, Y., Zou, L. et al. Identification and candidate analysis of a new brown planthopper resistance locus in an Indian landrace of rice, paedai kalibungga. Mol Breeding 44, 45 (2024). https://doi.org/10.1007/s11032-024-01485-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-024-01485-6