Abstract
Lodging of the crop is one of the major constraints in cyclone-prone coastal rice areas. The feasibility of conventional breeding approaches for lodging resistance is limited due to high influence of structural and weather parameters. Stem lodging is the major risk for irrigated low land areas. In the present study, F2 population derived from the cross between lodging-susceptible Swarna variety and lodging-resistant advanced breeding line MTUII 110-9-1-1-1-1 was used for QTL mapping. A total of five QTLs were identified for culm diameter, culm thickness, culm strength, bending stress, and panicle length. The phenotypic response and QTLs identified from the above population elucidate that culm diameter and culm thickness play key role in lodging resistance. Gene prediction in the mapping region of culm diameter and thickness on chromosome 6 revealed that gene sets of microtubule-based movement LOC_0s06g45900 and potassium transporter LOC_0s06g45940 might be putative candidate genes responsible for lodging resistance. Near isogenic line (NIL) with wider and thick culm in Swarna background was developed using marker-assisted backcross breeding. Developed lodging resistant line in the background of widely adopted mega rice variety Swarna would enhance the rice productivity even under adverse climatic conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa L.) is an important staple food for more than half of the world population especially in South Asia. Realization of sustained yields under changing climatic conditions with shrinking resources to feed ever-increasing population is a major challenge to the rice scientists. Among the abiotic stresses, lodging of the crop at reproductive and maturity stages in cyclone-prone areas is one of the major problems hampering rice productivity. Lodging can be defined as displacement of stem from vertical position; it is either permanently or partially reversible based on extent of bending. Stem lodging is a major risk in irrigated low land ecosystem. Two types of stem lodging exist and they are the breaking and bending type. Breaking type lodging is most prevalent in coastal irrigated ecosystem where lodging occurs due to breakage of lower internodes and bending type of lodging exists in low land rice areas (Hirano et al. 2017). Lodging in rice plants not only reduces grain yield, but also deteriorates grain quality (Setter et al. 1997) and up to 40% yield loss was reported (Nishiyama 1986).
Breeding for lodging resistance using conventional methods is quite difficult as it is influenced by both structural properties of the stem and weather parameters. Crop management practices like excess nitrogen fertilizer application enhances lodging (Hashem et al. 2016) and results in reduction in structural carbohydrates of culm and also lignin deposition in secondary cell wall of lower internodes of culm (Zhang et al. 2016). Incidence of brown planthopper (BPH) and sheath blight weakens the lower internodes (Wu et al. 2012). Climate changed conditions like higher CO2 enhances the risk of stem lodging (Zhu et al. 2013).
In the early 1960s, incorporation of sd-1 gene from the Chinese variety Dee-geo-woo-gen in traditional varieties minimized the lodging risk to certain extent (Khush 1999). But still it remains a major problem even in semi-dwarf varieties because of weak culm characteristics viz. narrow culm, and elongated internodes. For instance, semi-dwarf mega rice variety Swarna is highly vulnerable to lodging by virtue of lower culm diameter (Girija Rani et al. 2017).
Although the traditional target was reduction of height, the other important traits contributing to lodging resistance are culm characteristics viz. basal internodal length, culm diameter at 4th internode, culm wall thickness, bending stress, culm strength, leaf sheath wrapping and thickness, and linear density of culm (Chang and Vergara 1972; Matsuda et al. 1983; Kashiwagi and Ishimaru 2004; Zhong et al. 2007; Zhu et al. 2016; Rao et al. 2017). Breaking type lodging resistance was improved due to increased lignin accumulation and culm diameter (Okuno et al. 2014). Silicified cells provide strength to culm by upregulation of cinnamyl alcohol dehydrogenase (CAD), a key gene responsible for lignin biosynthesis (Dorairaj and Ismail 2017). The physical strength was positively and highly significantly correlated with the total amount of potassium and silicon in culm during grain filling, and total amount of soluble sugars in culm at the full heading and milky stages (Zhang et al. 2010; Yan-Hua et al. 2011). Re-accumulation of stem non-structural carbohydrates (NSC) in the later period of grain filling related to slower senescence and is correlated with lodging resistance (Kashiwagi et al. 2006; Takashi et al. 2017). Thickened secondary cell walls with higher cellulose levels in the mature plants enhances the mechanical strength (Fan et al. 2017). Leaf sheath can delay the aging of stem and increase the strength of the stem (Kashiwagi et al. 2008). Involvement of epsistatic gene interactions for lodging resistance–related traits was reported (Girija Rani et al. 2015) and marker-assisted selection would help in fixation of favorable alleles involved in epsistatic interactions (Fethi et al. 2011). Phenotypic evaluation for lodging resistance requires visual estimates in plots of breeding lines and genetic variability is too high in early generations.
Marker-assisted breeding is one of the viable strategies to develop lodging-resistant rice varieties. Quantitative trait loci (QTL) having major effects for culm traits viz. basal culm thickness, culm length and culm strength (Mu et al. 2004), pushing resistance (Kashiwagi and Ishimaru 2004), and basal culm traits (Hu et al. 2008; Zhu et al. 2008) would be useful in selection of non-lodging lines in early generations. Effective quantitative trait loci for culm diameter, Strongculm2 (SCM2) on chromosome 6 was identified (Ookawa et al. 2010) and this QTL has role in gene regulation on vascular development and first branch of panicle to affect the yield (Terao et al. 2010). QTL for leaf sheath length in vicinity of SCM2 (Liu et al. 2011); QTLs for strong culm on chromosome 1, 5, 6, 8, and 11 (Yamamoto et al. 2013); physical strength of the upper culms (Kashiwagi 2014), Strongculm 3(SCM3), and conferring culm strength (Yano et al. 2015) were identified by earlier workers. Leaf star rice variety with superior lodging resistance possessing gold hull and internode2 (gh2) on chromosome 2 was developed by Ookawa et al. (2014). QTL preventing culm strength deterioration after grain filling BSUC11 (Kashiwagi et al. 2016) and two major effect QTLs, qCD1.1 and qCS1.1, on chromosome 1 (Yadav et al. 2017) were associated with lodging resistance. QTLs for the density of hemicellulose, cellulose, and holocellulose cell wall materials in japonica varieties contributing to increased bending stress were reported by Mulsanti et al. 2018.
The present study aimed to identify new QTLs for lodging resistance traits and to incorporate strong culm traits in highly susceptible mega rice variety Swarna using marker-assisted breeding besides adopting phenotypic techniques for lodging resistance.
Materials and methods
Selection of the parents and generation of mapping population
Swarna, a mega rice variety developed at Regional Agricultural Research Station (RARS) is being cultivated nearly in 5 million ha worldwide. Swarna is an indica variety developed by pedigree method using Vasista and Mahsuri as parents. It is a semi-dwarf erect plant type with 110 cm plant height, dark green foliage, completes its life cycle in 150 days, medium slender brown glume grain, intermediate amylose content, and is widely adopted by farmers because of its high yielding capacity under low input management. Swarna, a highly lodging-susceptible variety was used as recipient parent and an advanced breeding line MTUII 110-9-1-1-1-1 (Girija Rani et al. 2017) was used as donor for lodging resistance. Swarna was crossed with MTUII 110-9-1-1-1-1 and F1 was selfed to generate F2 population. One hundred eighty individual F2 plants were selfed to generate respective F3 families. Simultaneously, some of F1 plants were backcrossed with the recurrent parent Swarna three times followed by four generations of selfing to generate NILs of Swarna with lodging resistance (Fig. 1).
F1 and backcross F1 seedlings (25 days old) were transplanted to with spacing of 30 cm between rows and 30 cm between plants to assess maximum expression of traits, whereas F2, F3 families, and advanced backcross progenies were transplanted at 20 cm between rows and 15 cm between plants. Fertilizer application of 90:60:60 kg/ha of nitrogen, phosphorous, and potassium was practiced. Plant protection against sheath blight, BPH, and stem borer was practiced.
Phenotypic evaluation for lodging resistance
The present study was carried out at RARS, Maruteru, Andhra Pradesh, India, located in typical coastal irrigated ecosystem where cyclones are most prevalent especially during the maturity to harvesting stages. The rainfall data at reproductive stage from 2012 to 2017 was furnished in Table 1.
A total of 180 F2 individual plants and respective F3 families pertaining to Swarna/MTUII 110-9-1-1-1-1 cross were evaluated for different traits contributing to lodging resistance viz. culm diameter (mm), thickness (mm), basal internodal length at 4th internode from top (cm) at 20 days after heading and traits culm strength (score 1–9), bending stress (g stem−1) and percent of lodging just before harvesting. The culm internodes were cut transversely at 4th internode from the top at 20 days after heading with a scalpel to measure the inner and outer diameters of the internode with a vernier calipers. The averaged culm wall thickness was then calculated by the following equation:
Bending stress was measured at the time of harvesting by pushing hill at 20 cm above the ground at 45° angle using a prostrate tester (DIK 7401, Daiki Rika Kogyo Co. Ltd., Tokyo, Japan), and it was expressed in g stem−1 using the following formula as per Bhagat et al. (2011).
At the time of harvesting stage, culm strength scores and percent of lodging were recorded as per SES, IRRI 2002.
In each generation of development of NILs, confirmed positive plants using foreground markers were phenotypically evaluated for lodging resistance traits. At BC3F2 stage onwards, yield parameters like days to 50% flowering, plant height (cm), ear-bearing tillers, panicle length (cm), and grain yield per plant were measured along with lodging-related parameters for confirmed lines.
Genotyping of mapping populations using SSR markers and QTL mapping
Leaf tissue was collected from all the individual plants in all the generations. The genomic DNA was isolated as per the protocol of Zheng et al. 1995. Quality and quantity of DNA was assessed using a nanodrop eight-channel spectrophotometer (Thermo Fisher Scientific). Polymerase chain reaction was performed in 10 μL final reaction volume comprising of 1 μL of 10X Taq buffer A with 15 mM Mgcl2, 0.5 μL of 2.5 mM dNTPs, 1 μL each of 5 μM forward and reverse primers, 1 U Taq polymerase, 2.5 μL of genomic DNA (30 ng/μL), and 3 μL of sterile distilled water. Polymerase chain reactions were carried out using an Eppendorf master cycler gradient with amplification profile of 94 °C for 5 min for initial denaturation, 35 cycles of 94 °C for 30 s, 58 °C for 30 s, 72 °C for 1 min, and a final extension at 72 °C for 5 min. PCR products were subjected to a 3% high-resolution metaphor Agarose (Lonza), and gel images were captured under UV light using the syngene gel documentation system.
Statistical analysis
The means and variances of means for F1, F2 generations were computed using individual plant data and F3 generation on family basis. Identified 5 NILs were evaluated along with two parents in three replications in randomized block design, and data was recorded on 5 randomly selected plants. Statistical analysis was performed using Cropstat 7.2 version. Graphical genotyping was performed for selected NILs to assess maximum genome recovery of recurring parent Swarna using GGT 2.0 (Van Berloo 1999). A Win QTL cartographer (Batsen et al. 2005) was used to detect QTLs by using composite interval mapping with 1000 permutations.
Results
QTL mapping
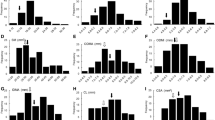
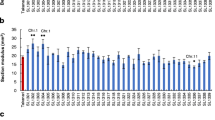
Recurrent parent Swarna as well as donor parent MTUII 110-9-1-1-1-1 were evaluated for different traits for lodging resistance and there was significant variation between the parents, F1, F2, F3, generations for most of the traits during wet season of 2013 (Table 2, Fig. 2). Parental polymorphism survey was carried out between Swarna and MTUII 110-9-1-1-1-1 using 576 SSR markers covering 12 chromosomes and identified 104 polymorphic markers were used to genotype 180 F2 individual plants and F3 families.
The QTL analysis using F2 population of Swarna/MTUII 110-9-1-1-1-1 resulted in identification of five QTLs on chromosome 2, 6, and 7 (Table 3, Fig. 3). Among them, three QTLs were identified on chromosome 6 viz. qCd6 for culm diameter, qCt6 for culm thickness, and qPl6 for panicle length. QTL for culm diameter was located in the marker interval between RM 20547 and RM 20557 with LOD score of 6.07 and phenotypic variance of 10.39% while the QTL qCt6 for culm thickness was a major QTL with a LOD score of 3.06 and phenotypic variance of 13.04%. QTL for culm strength qCs7 was mapped on chromosome 7 in the chromosomal region between RM 418 and RM 320 with LOD score value of 3.13 explaining phenotypic variance of 11.19%. A major significant QTL for panicle length (qPl6) was mapped on chromosome 6 with 3.55 LOD score and 16% phenotypic variation. Bending stress QTL (qBs2) was identified on chromosome 2 with LOD score value of 3.14 and R2 value of 2.36%. In all the detected QTLs, the donor MTUII 110-9-1-1-1-1 alleles had increasing effect. Validation of identified QTLs using linked markers among F3 families of Swarna and MTUII 110-9-1-1-1-1 cross revealed co-segregation for respective lodging-related traits. Out of five QTLs, culm diameter, culm thickness-linked markers RM 20557 and RM 5509 were found to be co-segregated with lodging resistance–related traits in another mapping population of Indra/BPT 2270 at F2 generation and it indicated that RM 20557 and RM 5509 markers can be used to transfer wider and thick culm traits though marker-assisted breeding.
Marker-assisted backcross breeding
Culm diameter and culm thickness QTL-linked markers RM 20557 at 27.45 Mb and RM 5509 at 27.82 Mb on chromosome 6 were identified as foreground markers for traits conferring lodging resistance in new donor MTUII 110-9-1-1-1-1. Polymorphic markers RM 30 located at 26.86 Mb and RM 340 at 28.49 Mb were selected as recombinant selection markers. Based on the phenotyping and genotyping, positive plants were selected in each generation and were utilized in developing NILs of Swarna (Table 4). Three positive F1 plants for foreground markers and non-lodging traits were used for generation of BC1F1. Natural screening for lodging at BC1F1 generation with the occurrence of Neelam cyclone during wet season of 2013 was depicted in Fig. 4. Out of 32 BC1F1 plant genotyped, 6 positive plants with lodging resistance were selected for generation of BC2F1. To generate BC3F1s, eight plants conferring positive for foreground markers RM 20557 and RM 5509 besides lodging resistance traits were selected out of 329 BC2F1 plants genotyped. At BC3F1 generation, 51 positive plants confirmed for strong culm traits were selfed to study BC3F2 generation. Grain yield (> 15 g/plant), wider culm diameter (> 7.0 mm), culm thickness (> 1.5 mm), stronger culm (scores 1 or 3), basal internodal length (< 5 cm), bending stress (> 40 g stem−1), and percent of lodging (< 25%) with flowering duration matching with recurring parents were used as selection criteria to select advanced breeding lines with high yield potential from BC3F2 generation onwards besides genotypic confirmation. This resulted in selection of 99 advanced progenies with Swarna characters out of 3672 plants of BC2F2 genotyped. At BC3F3 generation, 65 plants selected by genotypic and phenotypic selection of 1980 plants pertaining to 99 families. Five NILs were selected out of 65 progenies at BC3F4 generation. Based on the phenotyping, foreground, recombinant, and background selections, five NILs of Swarna with lodging resistance were generated. Results of yield evaluation of NILs over three seasons were furnished in Table 5 and three seasons mean for each trait was compared with pooled analysis of variance of three seasons. There is significant variation for the all traits except days to 50% flowering and ear-bearing tillers. Among the five NILs, NIL 2, NIL 1, and NIL 3 expressed yield increase of 51.21%, 49.63%, and 45.56% over recurrent parent Swarna with lodging-resistant traits. Out of the above three high-yielding NILs, NIL 1 was found to be the best with wider (6.37 mm) and thick (1.29 mm) culm, increased panicle length (27.47 cm), bending stress (46.19 g/stem), desirable mean culm strength score of 2.67 and minimum percentage of lodging (1.01%). Basal internodal length of NIL1 is on par with Swarna. Background selection was carried out for five NILs using 73 markers covering 12 chromosomes to assess maximum recovery of Swarna. Results of graphical genotyping revealed that NIL1 expressed maximum recovery 94.1% of Swarna (Table 6). Developed NIL1 has distinguishable characters such as wider and thick culm, long panicles with 3–5 days earlier in duration than recurrent parent Swarna and quality parameters are on par with Swarna (Fig. 5, Table 7).
Discussion
Two QTLs mapped for culm diameter (qCd6) and culm thickness (qCt6) were in congruent with previously reported Strong culm2 (SCM2) linked to culm diameter possessing pleiotropic effect with APO (Apparent panicle organization) by Ookawa et al. (2010) indicating donor MTUII 110-9-1-1-1-1 possesses alleles of strong culm and therefore the linked markers RM 20557, RM 5509 can be utilized for marker-assisted breeding for the development of lodging-resistant lines. The QTL qPl6 for panicle length identified in the chromosomal region between RM 30 and RM 340 on chromosome 6 was very close to the QTL related to panicle length (qPl6) identified by Zhu et al. 2008 in the RIL population of the cross between Lemont (japonica) and Teqing (indica) and Zhang et al. 2015 using backcross population of Nipponbare (japonica) and WS 3(indica).
The QTL for bending stress qBs2 identified at 4.3 Mb on chromosome 2 is within vicinity of strong culm 4 (SCM4) linked marker RM 3703 at 3. 86 Mb reported by Yano et al. 2015. Ookawa et al. 2016 identified a QTL for cortical fiber tissue conferring lodging resistance on chromosome 7 at 18.4 Mb which was co-localized with the QTL qCs7 for culm strength in the current study.
The results indicated that QTLs for culm diameter, culm thickness, culm strength and bending stress confers lodging resistance in new donor MTUII 110–9–1-1-1-1. Further it can be inferred that there was strong association between panicle length and lodging resistance traits which was in confirmation with Ookawa et al. 2010 who explained pleiotropic effect of SCM2 with APO (Apparent panicle organization) on chromosome 6. Culm thickness is positively associated with culm diameter and culm strength. Per cent of lodging has positive correlation with culm strength as per Girija Rani and Satyanarayana 2018.
According to the rice genome annotation database (http://rice.plantbiology.msu.edu/), gene sets of micro tubule based movement LOC_0s06g45900 and potassium transporter LOC_0s06g45940 predicted in mapping region of culm diameter, culm thickness on chromosome 6. Transverse orientation of microtubules promotes cell elongation and longitudinal orientation restricts cell elongation which inturn contributes to lodging resistance. Microtuble orientation signals environmental response for strong culms (Nick 2012; Nick and Opatrny 2014). Role of microtubule in controlling cell wall properties in rice was reported by Zhang et al. 2010.
Potassium transport gene maintains cell turgidity resulting in strong culms (Wang and Wu 2015). In rice potassium culm content is directly correlated with culm mechanical strength because potassium directly associates with lignification of sclerenchyma cell and vascular bundles (De Datta and Mikkelsen 1985; Zhang et al. 2010).This indicates that microtublular movement and potassium transporter genes might be putative candidate genes responsible for strong culm traits in the mapping population.
Marker-assisted backcross breeding
Out of five QTLs, two QTLs for culm diameter and culm thickness were found to be major effect QTLs for incorporation of strong culm traits in Swarna. These markers were used as foreground markers for incorporation wider and thick culms in Swarna. Use of recombinant markers RM 30 and RM 340 limited introgression of undesirable alleles of donor MTUII 110-9-1-1-1 aided in successful introgression of genomic region of 1.86 Mb of strong culm traits in Swarna. In each generation of development of NILs, phenotypic confirmation of positive plants of RM 20557 and RM 5509 for lodging resistance related traits and use of selection criteria for selection of non-lodging lines with Swarna characters helped in identification of better NILs. Background selection resulted in identification of best NIL of Swarna (indica) with non-lodging traits out of 5 NILs. Earlier workers, Kashiwagi et al. (2006) identified the locus responsible for pushing resistance of the lower part of the rice plant (prl5) in backcross inbred lines developed from a cross between Nipponbare (japonica) and Kasalath (indica) and developed NILs of Kasalth were characterized (Kashiwagi et al. 2008) for lodging resistance. Incorporation of SCM2 conferring for wider culm (Ookawa et al. 2010), SCM2+SCM3 pyramided for wider and strong culms (Yano et al. 2015) for the developments of NILs of Koshihikari (japonica) for lodging resistance.
Results of yield evaluation of NILs over 3 seasons revealed that developed best NIL1 has wider and thicker culms conferring lodging resistance with minimizing linkage drag of donor alleles. Thus molecular breeding of Swarna for lodging resistance using MTU II 110-9-1-1-1-1 as donor resulted in development of NIL with higher yield than Swarna under lodging prone conditions.
Conclusion
Identification QTLs linked to lodging resistance in new donor MTU II 110-9-1-1-1-1 helped in adoption of marker-assisted breeding for precise transmission of lodging resistance loci into mega rice variety Swarna (indica). Gene sets pertaining to microtubules movement and potassium transporter might be putative candidate genes responsible for strong culm traits (wider and thicker culms) in the developed mapping population. Adoption of marker assisted backcross breeding besides phenotypic selection for lodging resistance with higher yield resulted in NIL with non-lodging trait of widely grown rice variety Swarna. Identified donor MTUII 110-9-1-1-1-1 can be useful in future breeding programs, generated NIL of Swarna with lodging resistance can be released as variety after thorough multi-environment testing. Further, this material can be used as genetic stocks for future breeding programs.
References
Batsen CJ, Weir BS, Zeng ZB (2005) QTL cartographer version 2.5. Department of Statistics, North Carolina state University, USA
Bhagat KP, Sairam RK, Deshmukh PS, Kushwaha SR (2011) Biochemical analysis of stem in lodging tolerant and susceptible wheat (Triticum aestivum L.) genotypes under normal and late sown conditions. Indian J Plant Physiol 16(1):68–74
Chang TT, Vergara BS (1972) Ecological and genetic information on adaptability and yielding ability in tropical varieties. In: International Rice Research Institute (ed) Rice breeding. International Rice Research Institute, Manila, p 431
Fan C, Li Y, Hu Z, Hu H, Wang G, Li A, Wang Y, Tu Y, Xia T, Peng L, Feng S (2017) Ectopic expression of a novel OsExtensin-like gene consistently enhances plant lodging resistance by regulating cell elongation and cell wall thickening in rice. Plant Biotechnol J 16:254–263. https://doi.org/10.1111/pbi.12766
De Datta SK, Mikkelsen DS (1985) Potassium nutrition of rice. In: Munson RD (ed) Potassium in agriculture. ASA, Madison, pp 665–669
Dorairaj D, Ismail M (2017) Distribution of silicified microstructures, regulation of cinnamyl alcohol dehydrogenase and lodging resistance in silicon and paclobutrazol mediated Oryza sativa. Front Physiol 8. https://doi.org/10.3389/fphys.2017.00491
Fethi B, Hanbary C, Mohamed EG (2011) Genetic adaptability of inheritance of resistance to biotic and abiotic stress level on crop: role of epistasis. Afr J Biotechnol 10(86):19913–19917
Girija Rani M, Satyanarayana PV, Lal Ahmed M, Ashok Rani Y, Srinivasa rao V (2015) Gene action of elite rice lines for yield and lodging resistance related traits. Res Crops 16(4):689–697
Girija Rani M, Satyanarayana PV, Lal ahmed M, Ashok Rani Y, Srinivasa rao V (2017) Combining ability studies of lodging susceptible vs resistant genotypes of rice. Green Farming 8(2):254–259
Girija Rani M, Satyanarayana PV (2018) Selection of lodging resistant lines in early generations using linked molecular markers and phenotypic traits in rice. Int J Curr Microbiol App Sci 7(01):1638–1650
Hashem M, Naeem ES, Metwally TF, El Sharkawi HM (2016) Enhancement of lodging resistance and productivity of rice using growth regulators at different nitrogen levels. J Plant Breed Crop Sci 8(3):34–44
Hirano K, Odonio RL, Matsuoha M (2017) Engineering the lodging resistance mechanism of post-green revolution rice to meet future demands. Proc Japn Acad ser B Phys Biol Sci 93(4):220–233
Hu J, Fujimoto K, Guo LB, Zeng DL, Zhang GH, Dong GJ, Wang XH, Zhu LH, Qian Q (2008) QTL analysis of lodging resistance force and lodging resistance related traits in rice. Chin J Rice Sci 22(2):211–214
Kashiwagi T, Ishimaru K (2004) Identification, functional analysis of a locus for improvement of lodging resistance in rice. Plant Physiol 134:676–683
Kashiwagi T, Madoka Y, Hirotsu N, Ishimaru K (2006) Loucs prl5 improves lodging resistance of rice by delaying senescence and increasing carbohydrate re-accumulation. Plant Physiol Biochem 44:152–157
Kashiwagi T, Togawa E, Hirotsu N, Ishimaru K (2008) Improvement of lodging resistance with QTLs for stem diameter in rice (Oryza sativa L.). Theor Appl Genet 117:749–757
Kashiwagi T (2014) Identification of quantitative trait loci for resistance to bending-type lodging in rice (Oryza sativa L.). Euphytica 198(3):353–367
Kashiwagi T, Munakata J, Ishimaru K (2016) Functional analysis of the lodging resistance QTL BSUC11 on morphological and chemical characteristics in upper culms of rice. Euphytica 210:233–243. https://doi.org/10.1007/s10681-016-1707-1
Khush GS (1999) Green revolution: preparing for the 21st century. Genome 42:646–655
Liu H, Rao Y, Yang Y, Leng Y, Huang L, Zhang G, Hu J, Guo L, Gao Z, Zhu L, Dong G, Liu J, Yan M, Qian Q, Zeng D (2011) Genetic analysis of traits related to leaf sheath in rice (Oryza Sativa L.). Rice Genomics Genet 2(3):21–30
Matsuda T, Kawahara H, Chonan N (1983) Histological studies on breaking resistance of lower internodes in rice culm IV the rules of each tissue of internode and leaf sheath in breaking resistance. Proc Crop Sci Soc Jpn 52:355–361
Mu P, Zi-chao L, Chun-ping L, Hong-liang Z, Xiang-kun W (2004) QTL analysis for lodging resistance in rice using a DH population under lowland and upland cultural conditions proceedings of the 4th International Crop Science Congress 26 September - 1 October 2004, Brisbane, Australia
Mulsanti IW, Yamamoto T, Ueda T, Samadi AF, Kamahora E, Rumanti IA, Thanh V, Adachi S, Suzuki S, Kanekatsu M, HirasawaT T, Ookawa T (2018) Finding the superior allele of japonica-type for increasing stem lodging resistance in indica rice varieties using chromosome segment substitution lines. Rice. https://doi.org/10.1186/s12284-018-0216-3
Nick P (2012) Microtubles and tax payer. Protoplasma 249(suppl 2):S81–S94
Nick P, Opatrny Z (eds.) (2014) Why to spend tax money on plant microtubules? Applied Plant Cell Biology, Plant Cell Monographs 22, https://doi.org/10.1007/978-3-642-41787-0_2
Nishiyama I (1986) Lodging of rice plants and counter measures. Plant Growth Regulators in Agriculture 34:150–163
Ookawa T, Hobo T, Yano M, Murate K, Ando T, Miure H, Asno K, Ochiai Y, Ikeda M, Nishitani R, Ebistani T, Ozaki I, Angeles ER, Hirasana T, Matsuoka M (2010) New approach for rice improvement using a pleiotropic QTL gene for lodging resistance and yield. Nat Commun 1:132
Ookawa T, Inoue K, Matsuoka M, Ebitani T, Takarada T, Yamamoto T, Ueda T, Yokoyama T, Nakaba SF, Funada R, Kato H, Kanekatsu M, Toyota K, Motobayashi T, Vazirzanjani M, Tojo S, Hirasawa T (2014) Increased lodging resistance in long-culm, low-lignin gh2 rice for improved feed and bioenergy production. Chin J Sci Rep 4:6567
Ookawa T, Aoba R, Yamamoto T, Ueda T, Takai T, Fukuoka S, Tsuyu Ando T, Adachi S, Matsuoka M, Ebitani T, Kato Y, Mulsanti IW, Kishii M, Reynolds M, Piñera F, Kotake T, Kawasaki S, Motobayashi T, Hirasawa T (2016) Precise estimation of genomic regions controlling lodging resistance using a set of reciprocal chromosome segment substitution lines in rice. Sci Rep 6:30572 doi:1038/sr rep 30572
Okuno A, Hirano K, Asano K, Takase W, Masuda R, Morinaka Y, Ueguchitanaka M, Kitano H (2014) New approach to increasing rice lodging resistance and biomass yield through the use of high gibberellin producing varieties. PLoS One 9(2):e86870
Rao AN, Sreekanth B, Madhav SM, Reddy NS (2017) Studies on physical and mechanical properties of rice stem for lodging tolerance at reproductive phase (Oryza sativa L.). Int J Pure App Biosci 5(3):407–414
SES, IRRI (2002) Standard evaluation system for rice. International Rice Research Institute, Los Banos
Setter TL, Laureles EV, Mazaredo AM (1997) Lodging reduces yield of rice by self-shading and reductions in canopy photosynthesis. Field Crop Res 49:95–106
Terao T, Nagata K, Morino K, Hirose T (2010) A gene controlling the number of primary rachis branches also controls the vascular bundle formation and hence is responsible to increase the harvest index and grain yield in rice. Theor Appl Gen 120(5):875–893
Takashi ST, Tanaka YK, Hiroyuki H, Tatsuya I (2017) Rice (Oryza sativa L.) cultivars for whole crop silage having traits of higher non-structural carbohydrate accumulation in stems and lodging resistance under late direct seeding cultivation on well-drained paddy field in the Kansai region, Japan. Jpn J Crop Sci 86(3):229–235
Van Berloo R (1999) GGT: software for the display of graphical genotypes. J Hered 90(2):328–329
Wang Y, Wu W (2015) Genetic approaches for crop potassium acquisition and use efficiency. Curr Opin Plant Biol 25:46–52
Wu W, Huang J, Cui K, Nie L, Wang Q, Yang F, Shah F (2012) Sheath blight reduces stem breaking resistance and increases lodging susceptibility of rice plants. Field Crop Res 128:101–108
Yadav S, Singh UM, Naik SM, Venkateshwarlu C, Ramayya PJ, Anitha RK, Nitika S, Arvind K (2017) Molecular mapping of QTLs associated with lodging resistance in dry direct-seeded rice (Oryza sativa L.). Front Plant Sci 8. https://doi.org/10.3389/fpls.2017.01431
Yamamoto K, Yamamoto T, Sugiyama C, Hirasawa T, Ookawa T (2013) Estimation of the locus for strong culm traits, using reciprocal chromosome segment substitution lines derived from the cross between rice varieties, koshihikari and takanari. Proceedings of 7th International Rice Genetics Symposium 5–8 November 2013, Manila, Philippines, 435
Yan-Hua Y, Zhen Z, Ya-Dong Z, Tao C, Qing-Yong Z, Li-Hui C-LW (2011) Changes of stem biochemical components in different growth stages of rice and their relationship with lodging resistance. Pl Physiol J 47(12):1181–1187
Yano K, Ookawa T, Taiichiro A, Aya K, Ochiai Y, Hirasawa T, Ebitani T, Takarada T, Yano M, Yamamoto T, Fukuoka S, Wu J, Ando T, Ordonio RL, Hirano K, Matsuoka M (2015) Isolation of a novel lodging resistance QTL gene involved in strigolactone signaling and its pyramiding with a QTL gene involved in another mechanism. Mol Plant 8(2):303–315
Zhang F, Zin Z, Ma G, Shang W, Liu H, Xu M, Liu Y (2010) Dynamics between lodging resistance and chemical contents in japonica rice during grain filling. Rice Sci 17(4):311–318
Zhang L, Wang J, Wang J, Wang L, Ma B, Zeng L, Qi Y, Li Q, He Z (2015) Quantitative trait locus analysis and fine mapping of the qPL6 locus for panicle length in rice. Theor Appl Genet 128:1151–1161
Zhang W, Wu L, Ding Y, Weng F, Wu X, Li G, Liu Z, Tang S, Ding C, Wang S (2016) Top-dressing nitrogen fertilizer rate contributes to decrease culm physical strength by reducing structural carbohydrate content in japonica rice. J Integr Agric 15(5):992–1004
Zheng K, Subudhi PK, Domingo J, Magantay G, Huang N (1995) Rapid DNA isolation for marker assisted selection in rice breeding. Rice Genet News Lett 12:255–258
Zhong Z, Sui G, Hua Z, Li Q, Hao X, Zhu B, Yao J, Su Y (2007) Analysis of lodging resistance of Tiyou 418 and Liaoyou 5218 with higher stems and bigger panicles. Chin Agric Sci Bull 23(8):141
Zhu LH, Zhong DB, Xu JL, Yu SB, Li ZK (2008) Differential expression of lodging resistance related QTLs in rice (Oryza sativa L.). Plant Sci 175:898–905
Zhu CW, Cheng WG, Sakai H, Oikawa S, Laza R, Usui Y, Hasegawa DT (2013) Effects of elevated [CO2] on stem and root lodging among rice. Cultivars Chin Sci Bull 58:1787–1794
Zhu G, Li G, Wang D, Yuan S, Wang F (2016) Changes in the lodging-related traits along with rice genetic improvement in China. PLoS One 11(7):e0160104. https://doi.org/10.1371/journal.pone.0160104
Funding
We acknowledge Rastriya Krishi Vikas Yojana (RKVY), Government of India, and Acharya N G Ranga Agricultural University for providing funds to carry out above research work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Merugumala, G.R., P.V, S., Narne, C. et al. Molecular breeding of “Swarna,” a mega rice variety for lodging resistance. Mol Breeding 39, 55 (2019). https://doi.org/10.1007/s11032-019-0961-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-019-0961-z