Abstract
Lodging is one of the major problems in rice production. In this study, we identified quantitative trait loci (QTLs) for lodging resistance traits using recombinant inbred lines (RILs) derived from a cross between two high yielding and superior lodging resistant rice cultivars, Tachisugata and Hokuriku 193. A total of 19 QTLs were detected on chromosomes (Chrs.) 2, 4, 6, 8, 9 and 10 in the F6 and F7 populations. In particular, one stable QTL for the section modulus (SM) and the outer diameter of the major axis (ODMA) was detected on Chr. 2 in both the F6 and F7 populations, and the positive allele was derived from Tachisugata. Correlation analysis showed that the SM and ODMA were positively correlated with the bending moment at breaking, the primary factor for the breaking type lodging resistance in rice. In addition, two QTLs for the spikelet number of the secondary branch and the spikelet number per panicle were also detected on Chr. 2, which overlapped with the stable QTL for the traits related to lodging resistance. These results indicated that the stable QTL might have combined effects on the traits associated with lodging resistance and grain yield. The novel QTL would be a promising target in marker-assisted selection to improve lodging resistance and grain yield in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lodging in cereal crops is a major problem that results in decreased grain yield and deteriorated grain quality (Braumann et al. 2018). Since the 1960s, the lodging resistance of rice has been significantly improved by introducing the semi-dwarf gene (sd-1), known as the “Green Revolution Gene”. This improvement of lodging resistance is ascribed to the shortened plant height partly regulated by gibberellin (GA) deficiency or insensitivity. However, GA deficiency or insensitivity may have a negative influence on the culm strength by reducing the culm diameter and decreasing the lignin content (Okuno et al. 2014). In addition, the semi-dwarf plant itself has a bottleneck of the yield ceiling due to the limitation of source and sink capacities (Yano et al. 2015). To meet future food demands, the strategy of rice breeding should be aimed at enhancing taller plant architecture with a strong lodging resistance to develop new high yielding rice cultivars.
Lodging resistance is a complex quantitative trait, which is affected by many factors, such as the culm morphology, the culm quality, and the external environmental conditions. In particular, the culm diameter as a factor that influences the section modulus is highly correlated with the bending moment at breaking and regulates the lodging resistance of the rice plant (Ookawa et al. 2016). To date, many quantitative trait loci (QTLs) for the traits relating to lodging resistance have been identified, such as prl5 (Kashiwagi et al. 2006), qPRF-8 (Hu et al. 2008), lrt5 (Ishimaru et al. 2008), BSUC11 (Kashiwagi et al. 2016), SCM1 and SCM2 (Ookawa et al. 2010a), and SCM3 and SCM4 (Yano et al. 2015). Some QTLs and their responsible genes that simultaneously control lodging resistance and grain yield have been identified. Two pleiotropic QTL genes, SCM2 and SCM3 (Ookawa et al. 2010a; Yano et al. 2015), which can both increase the culm strength and spikelet number per panicle without causing panicle deformities, have been identified using chromosome segment substitution lines (CSSLs) derived from Sasanishiki and Habataki and backcross inbred lines (BILs) derived from Chugoku117 and Koshihikari, respectively. This finding indicates that it is possible to utilize many kinds of QTLs and their combinations affecting both lodging resistance and grain yield from natural variation in rice. It is necessary to examine the superior alleles from natural variation in rice.
Two rice cultivars, Tachisugata (TS) and Hokuriku 193 (H193), which were developed by the National Agriculture and Food Research Organization (NARO), Japan, both have high grain yield and strong lodging resistance due to their large panicles and culms (Goto et al. 2009; Ohta et al. 2010). In particular, TS is a high biomass rice cultivar for the use of whole crop silage and has superior lodging resistance designated “Tachi”, while H193 is a typical high yielding indica type rice cultivar. TS can be expected to be a promising genetic donor to further improve the lodging resistance to develop new high yielding indica type rice cultivars. In a previous study (Matsubara et al. 2016), the underlying QTLs associated with biomass yield traits, including culm length, panicle number, spikelet number per panicle, and 1000-grain weight, have been identified using the recombinant inbred lines (RILs) derived from a cross between TS and H193. However, the genetic mechanisms of strong lodging resistance of these two cultivars still remain to be elucidated. In this study, we carried out QTL analysis for the lodging resistance using the same RILs of TS/H193, which were developed by Matsubara et al. (2016) and discovered meaningful information for future rice improvement.
Materials and methods
Plant materials and field management
Two high yielding rice cultivars, TS and H193, were used to develop the RILs populations as described by Matsubara et al. (2016). In 2013, TS, H193, and 93 RILs of the F6 population were cultivated in the paddy field at the National Institute of Agrobiological Science (NIAS) in Tsukuba, Japan. Seedlings were transplanted into the paddy field at a density of one plant per hill, and the planting density was 18.5 hills m−2 with a spacing of 18 cm × 30 cm (40 plants for each line). The fertilizer was applied as the basal dressing with compost (~ 5 kg m−2) and chemical fertilizer at a rate of 8 g N m−2, 8.9 g P m−2, and 4.3 g K m−2. No additional fertilizer was applied at the later stage. In 2017, TS, H193, and 92 RILs of the F7 population were cultivated in the paddy field of the farm of the Tokyo University of Agriculture and Technology in alluvial soil from the Tama River during the rice growing seasons. Seeds were sown in nursery boxes on May 8, 2017. Seedlings were transplanted into the paddy field at a density of one plant per hill on May 25, 2017. The planting density was 22.2 hill m−2 with a spacing of 15 cm × 30 cm (40 plants for each line). The fertilizer was applied as the basal dressing with compound fertilizer at a rate of 5 g N m−2, 6 g P m−2, and 6 g K m−2. The weeds were controlled using herbicide and hand weeding when necessary. Other field management techniques were the same as those used in conventional management.
Measurement of the traits related to lodging resistance
At 14 days after heading, six main culms per RIL and parent were sampled to evaluate the physical parameters related to lodging resistance. After the measurements of culm length (CL) and the length of the basal internode (LI), the bending moment at breaking (BM) and Young’s modulus (YM) were measured at a distance of 4 cm from the supporting points using the load testing machine Tensilon (A&D, RTG-1210, Tokyo, Japan). The central section of the basal internode was sectioned by hand, and the inner and outer diameter was measured using a sliding Vernier caliper. The section modulus (SM) was calculated by π(a13b1 − a23b2)/32a1; the secondary moment of inertia (SMI) was calculated by π(a13b1 − a23b2)/64, and the cross-sectional area was calculated by π(a1b1 − a2b2)/4, where a1 is the outer diameter of the minor axis (ODMI), b1 is the outer diameter of the major axis (ODMA), a2 is the inner diameter of the minor axis (IDMI), and b2 is the inner diameter of the major axis (IDMA).
Measurement of yield traits
At the maturity stage, six plants of the middle individuals were sampled to count the panicle number (PN) and evaluate the grain yield (GY) and 1000-grain weight (1000-GW). The three largest panicles of the main culm from these plants were used to investigate the primary branch number (PBN), secondary branch number (SBN), spikelet number of primary branch (SNPB), spikelet number of secondary branch (SNSB), spikelet number per panicle (SN), grain length (GL), and grain width (GW).
QTL analysis and sequence analysis
A total of 175 SNP markers were used to construct the genetic linkage map covering all 12 chromosomes as described by Matsubara et al. (2016). Composite Interval Mapping (CIM) was performed using WinQTL Cartographer 2.5 (Wang et al. 2006) to detect the QTLs associated with lodging resistance and yield traits. The QTL regions were determined using forward and backward regression with the standard model (Model 6). The significant logarithm of odds (LOD) value threshold for each trait was determined following the 5% permutation test with 1000 replicates. The putative genes on the QTL region were identified based on the RAP-DB database (http://rapdb.dna.affrc.go.jp/). The sequences analysis of the QTL region was based on the TASUKE database (http://ricegenomes.dna.affrc.go.jp/).
Statistical analysis
The significant differences in the traits evaluated between TS and H193 were determined using a T-test. Phenotypic correlations among the traits associated with lodging were evaluated using Pearson’s correlation.
Results
Difference in the traits evaluated between TS and H193
The traits related to bending type lodging, breaking type lodging, and culm morphology were investigated in Tsukuba and Fuchu (Table 1). H193 showed significant higher values of the BM, SM, ODMI, and CSA compared to TS, but no significant differences in the BS, ODMA, IDMA, IDMI, and LI were detected between TS and H193 in Tsukuba. However, the mean values of the BM, BS, and CL of TS were higher than those of H193 in Fuchu. These results indicated that, although TS and H193 showed different phenotypes in Tsukuba and Fuchu, the higher lodging resistance of the rice was ascribed to the larger section modulus, culm diameter, and cross-sectional area.
Differences in the yield traits between TS and H193 in 2017 were also compared (Supplemental Table 1). The results showed that TS had more SNPB, a higher 1000-GW, and larger grain size compared to H193. However, the yield of H193 was higher than that of TS due to the increase in the PN and SNSB.
Phenotypic variance of the RILs
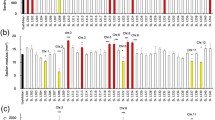
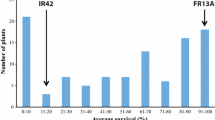
The traits related to lodging resistance and grain yield of the RILs in the F6 and F7 populations mostly showed the normal distribution with the obvious transgressive segregation (Fig. 1 and Supplemental Fig. 1).
Frequency distribution of the traits related to lodging resistance in the F6 and F7 populations. A–C The frequency distribution of the traits of RILs in the F6 population; D–I the frequency distribution of the traits of RILs in the F7 population. The abbreviations for SM, ODMA, ODMI, IDMA, CL, and CSA are the same as the Table 1. The black and white arrowheads indicate the mean values of TS and H193, respectively
To illustrate the relationships between the traits associated with lodging resistance, the correlation coefficients between the traits of the RILs in the F6 and F7 populations were analyzed using Pearson’s method (Table 2). The BM and SM were positively correlated with the CSA, ODMA, ODMI, IDMA, IDMI, CL, YM, SMI, and FR (p < 0.001), while the BS showed a significantly negative correlation with the SM, CSA, ODMA, ODMI, IDMA, IDMI, LI, CL, SMI, and FR (p < 0.001). These results indicated that the traits related to the culm diameter (SM, CSA, ODMA, ODMI, IDMA, and IDMI) were advantageous to improve the lodging resistance of rice.
QTL detection in the F6 population
A total of five QTLs controlling the SM, ODMA, and ODMI were detected in the F6 population (Table 3). Two QTLs for the SM were detected on Chrs. 2 and 8, explaining 10.0% and 12.5% of the phenotypic variance, respectively. Two QTLs controlling the ODMA were detected on Chrs. 2 and 4, explaining 11.1% and 11.6% of the phenotypic variance, respectively. One QTL affecting the ODMI was detected on Chr. 8, explaining 11.2% of the phenotypic variance. The positive alleles of the QTL for the SM on Chr. 2 and the QTLs for the ODMA on Chrs. 2 and 4 were derived from the parent TS, while the positive alleles of QTL for the SM and ODMI on Chr. 8 were derived from the parent H193.
QTL detection in the F7 population
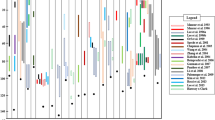
The QTL for the traits related to lodging resistance and grain yield were detected in the F7 population (Table 3; Fig. 2 and Supplemental Table 2). One QTL for the SM and one QTL for the ODMA were detected on Chr. 2 near the Marker aa02001611, explaining 13.1% and 15.6% of the phenotypic variance, respectively, and the positive alleles of these two QTLs derived from the TS. Two QTLs affecting the ODMI were detected on Chr. 2, which individually explained 13.0% and 12.4% of the phenotypic variance, respectively. One QTL for the IDMA was detected near the Marker P0815_2 on Chr. 2, explaining 13.3% of the phenotypic variance. One QTL affecting the CL was detected on Chr. 10, and it explained 52.5% of the phenotypic variance. In addition, one QTL controlling the CSA was detected on Chr. 2 with a phenotypic variance of 17.3% (Table 3).
QTLs location for the traits related to lodging resistance and grain yield in the F6 and F7 populations. The QTLs within a dotted box refer to the identification in F6 population relative to that in F7 population. SM section modulus, ODMA outer diameter of major axis of basal internode, ODMI outer diameter of minor axis of basal internode, IDMA inner diameter of major axis of basal internode, CL culm length, CSA cross sectional area, SNSB spikelet number of secondary branch, SN spikelet number per panicle, PN panicle number, and GW grain width
For the traits related to grain yield, two QTLs for the SNSB were detected on Chr. 2 near the markers aa02001611 and AD02013288, and these two QTLs explained 13.0% and 12.3% of the phenotypic variance, respectively. The positive alleles of these two QTLs were derived from TS with the additive effect 15.5 and 15.1, respectively. One QTL for the SN was detected on Chr. 2 with 19.8% of the phenotypic variance. Two QTLs controlling the PN were detected on Chr. 2 and Chr. 6, and these two QTLs explained 10.9% and 15.4% of the phenotypic variance, respectively. Two QTLs affecting the GW were detected on Chr. 2 near the markers aa02000049 and ac02000844, respectively. These two QTLs explained 13.8% and 32.2% of the phenotypic variance, and the positive alleles of the QTLs for the GW were derived from H193 (Supplemental Table 2).
Combined effects of the QTLs
A total of five QTLs in the F6 population and 14 QTLs in the F7 population were detected in Tsukuba and Fuchu, Japan, respectively. Among these QTLs, one locus with combined effects on controlling the traits related to lodging resistance and grain yield had been identified. The QTL near the markers aa02001611 on Chr. 2 simultaneously dominating the SM and ODMA was detected in both the F6 and F7 populations, and this locus was also proven to control the SN, ODMI, CSA, IDMA, and SNS in the F7 populations; the positive allele of this QTL was derived from TS (Fig. 2).
Putative genes
In this study, using RILs, one locus controlling the SM and ODMA was identified on Chr. 2 in the F6 and F7 populations, and this locus may also control the SN, ODMI, CSA, IDMA, and SNS. Thus, the putative genes on this region were investigated based on the Rice Annotation Project Database (RAP-DB, http://rapdb.dna.affrc.go.jp/). Six genes that are responsible for cytokinin signaling, NAC transcription factor expression, GA signaling, cell cycle regulation and other biological processes within the stable QTL were identified (Table 4). These putative genes are associated with different types of plant growth and development that may influence the culm morphology and yield traits, which have been reported in the previous studies.
Discussion
In this study, we used the RILs derived from a cross between two high yielding cultivars TS and H193 to map QTLs for the traits related to lodging resistance. TS and H193 not only have high grain yield but also have strong resistance to lodging due to their large culms (Matsubara et al. 2016).
Two different types of culm lodging have been observed in rice: bending type lodging and breaking type lodging (Hirano et al. 2017). In this study, the differences in the traits relating to the breaking type lodging were found between the parent TS and H193, while no difference in the bending type lodging-associated traits (YM, SMI, and FR) was observed (Table 1). Thus, we mainly focused on the traits related to breaking type lodging. The bending moment at breaking (BM), an indicator of culm strength, is typically used to evaluate the breaking type lodging resistance of rice (Mulsanti et al. 2018; Ookawa et al. 2010b). The BM is influenced by the SM and BS, and the SM can be calculated by the culm diameter and culm wall thickness (Berry et al. 2007; Ookawa et al. 2010a). The correlation analysis between the traits related to the lodging resistance showed that the BM was highly positively correlated with the SM, CSA, ODMA, ODMI, IDMA, and IDMI (Table 2), indicating that increasing the culm diameter could be an effective strategy to improve lodging resistance in rice, which was consistent with a previous study (Kashiwagi et al. 2008). In addition, both TS and H193 have larger culm diameters compared with other Japanese cultivars investigated in the previous studies (Ookawa et al. 2016; Ookawa and Ishihara 1992). We hypothesized that these two cultivars have superior strong culm genes.
In this study, the allele from TS had a positive effect on the SM, ODMA, ODMI, IDMA, and CSA. This region was different from the QTL corresponding to the SM, which was detected on Chr. 2 in our previous study (Yano et al. 2015). Culm diameter is a major factor associated with the SM, ODMA, ODMI, IDMA, and CSA. One QTL for culm diameter (qCD2.1) was detected on Chr. 2 in the dry direct seeded rice (Yadav et al. 2017), which was identified to the similar region in the QTL detected in this study. One QTL for the ODMA was detected on Chr. 4 in the F6 population, consistent with the QTL for the culm diameter detected near the marker RM4835 using a genome-wide association study (Sowadan et al. 2018). One QTL affecting the SM and ODMI was mapped on Chr. 8 in the F6 population, and the positive allele was derived from H193. Few QTLs for the culm diameter and SM have been identified on Chr. 8, and to our knowledge, this QTL would be novel. One QTL for ODMI and one QTL for CL were detected on Chrs. 9 and 10 in the F7 population, respectively. The QTL for the CL determined on Chr. 10 was not identified in the previous study using the same set of RILs used in our study (Matsubara et al. 2016). Together with the inconsistent results between the two experimental sites of Tsukuba and Fuchu, this finding indicates that the QTLs for lodging resistance might also be influenced by environmental factors (Weinig and Schmitt 2004). In contrast, we repeatedly identified QTLs on the same region of Chr. 2 in both the F6 and F7 RILs. The TS allele always has positive effects on the SM and several characteristics relating to the culm diameter, indicating that this region helps to improve lodging resistance by manipulating the culm diameter.
As for the yield traits, the QTLs for the SNSB, SN, PN, and GW were detected on Chrs. 2 and 6 in the F7 population (Supplemental Table 2). Interestingly, the QTLs for the SNSB, SN, and PN on Chr. 2 had overlapped segments with the QTL for the traits related to lodging resistance (Fig. 2). However, the TS alleles of the QTLs on Chr. 2 had positive effects on the SNSB and SN but negative effects on the PN, indicating a tradeoff between the spikelet number and panicle number. The QTLs for the number of panicles per plant (Marri et al. 2005), tiller number (Miyamoto et al. 2004), and panicle number (Yuan et al. 2003) were also detected on Chr. 2 in previous studies, and the QTL regions were consistent with this study. This result further proved that the TS allele of the QTL on Chr. 2 was responsible for improving lodging resistance and spikelet number. Two QTLs for the GW at the different regions were detected on Chr. 2, and the positive alleles were derived from H193, although H193 had a smaller grain size than TS. One QTL for the PN was detected on Chr. 6, consistent with a QTL for the tiller number at the maturity stage (qTN2-6-1) detected using RILs derived from Zhenshan 97 and Minghui 63 (Cui et al. 2004). From these results, we hypothesize that the allele of QTL on Chr. 2 from TS could also increase the spikelet number.
The stable QTL on Chr. 2 might have combined effects on lodging resistance and grain yield as described above. First, we compared the sequence differences of the parental varieties using the TASUKE database. Interestingly, the QTL region detected on Chr. 2 of TS seemed to be the japonica type but differed slightly from Nipponbare, while H193 was the indica type. We analyzed the putative genes within the stable QTL (Table 4). Six genes associated with cytokinin signaling (Hirose et al. 2007), NAC transcription factor (Kikuchi et al. 2000), fruit weight-like protein (Xu et al. 2013), F-box protein (Gomi et al. 2004), D-box protein (Ogawa et al. 2011, 2012), and CO-like protein containing two B-box zinc finger domains and one CCT domain (Lee et al. 2010) had been selected as the putative genes in this region due to their functions in plant development. In addition, the stable QTL in this study might also correspond with a mutant (bc14) conferring inferior culm strength and abnormal development (Zhang et al. 2011), OsCTR2 controlling tiller number (Wang et al. 2013), and OsABC1-2 simultaneously regulating tiller number, plant architecture, and grain size (Gao et al. 2012). These results demonstrate that many genes regulating plant growth and development have been identified within the stable QTL region on Chr. 2 in this study, and some of these genes might be involved in regulating lodging resistance and grain yield in rice. Further polymorphism analysis of the putative genes showed that all these genes in TS had the same haplotypes as those in Nipponbare (Supplemental Table 3), suggesting that we can further improve the lodging resistance of indica type rice cultivars by the combinations of these superior alleles from japonica type cultivars.
As described above, H193 might also have superior strong culm genes. However, we failed to detect any H193 alleles of the QTLs related to lodging resistance. We compared the genetic background of TS and H193. Interestingly, these two cultivars both have the genetic background from Milyang42 and Milyang25 (Supplemental Fig. 2A). In a previous study (Ashikari et al. 2005; Takai et al. 2014; Ookawa et al. 2016), two rice cultivars Takanari and Habataki, which were derived from the same parents Milyang 42 and Milyang 25, both carried the same type of Gn1a and SCM2/APO1 alleles that enhance lodging resistance and/or yield traits. Further genotypic analysis showed that Gn1a in TS and H193 were the same as that in Takanari and Habataki, but SCM2/APO1 in TS showed a different genotype from that of Takanari, Habataki, and H193 (Supplemental Fig. 2B). These results suggest that the H193 alleles of Gn1a and SCM2/APO1 might also have positive effects on lodging resistance and yield traits. Further study will be focused on the genetic mechanisms in H193 using different populations.
In this study, we identified one stable QTL associated with lodging resistance traits on Chr. 2. However, it is difficult to determine whether this QTL consists of a major effect QTL or many minor effect QTLs (Yano and Sasaki 1997). Further study will be focused on fine mapping using high resolution mapping populations, such as CSSLs and NILs. Dissection of this QTL may be utilized in marker-assisted selection to develop new varieties with strong lodging resistance and high grain yield.
References
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, Angeles ER, Qian Q, Kitano H, Matsuoka M (2005) Cytokinin oxidase regulates rice grain production. Science 309(5735):741–745
Berry PM, Sylvester-Bradley R, Berry S (2007) Ideotype design for lodging-resistant wheat. Euphytica 154(1–2):165–179
Braumann I, Urban W, Preuáẞ A, Dockter C, Zakhrabekova S, Hansson M (2018) Semi-dwarf barley (Hordeum vulgare L.) brh2 and ari-l mutants are deficient in a U-box E3 ubiquitin ligase. Plant Growth Regul 86(2):223–234
Cui K, Peng S, Ying Y, Yu S, Xu C (2004) Molecular dissection of the relationships among tiller number, plant height and heading date in rice. Plant Prod Sci 7(3):309–318
Gao Q, Yang Z, Zhou Y, Yin Z, Qiu J, Liang G, Xu C (2012) Characterization of an Abc1 kinase family gene OsABC1-2 conferring enhanced tolerance to dark-induced stress in rice. Gene 498(2):155–163
Gomi K, Sasaki A, Itoh H, Ueguchi-Tanaka M, Ashikari M, Kitano H, Matsuoka M (2004) GID2, an F-box subunit of the SCF E3 complex, specifically interacts with phosphorylated SLR1 protein and regulates the gibberellin-dependent degradation of SLR1 in rice. Plant J 37(4):626–634
Goto A, Sasahara H, Shigemune A, Miura K (2009) Hokuriku 193: a new high-yielding indica rice cultivar bred in Japan. JARQ 43(1):13–18
Hirano K, Ordonio RL, Matsuoka M (2017) Engineering the lodging resistance mechanism of post-green revolution rice to meet future demands. Proc Jpn Acad Ser B 93(4):220–233
Hirose N, Makita N, Kojima M, Kamada-Nobusada T, Sakakibara H (2007) Overexpression of a type-A response regulator alters rice morphology and cytokinin metabolism. Plant Cell Physiol 48(3):523–539
Hu J, Fujimoto K, Guo LB, Zeng DL, Zhang GH, Dong GJ, Wang XH, Zhu LH, Qian Q (2008) QTL analysis of lodging resistance force and lodging resistance-related traits in rice. Chinese J Rice Sci 22(2):211–214
Ishimaru K, Togawa E, Ookawa T, Kashiwagi T, Madoka Y, Hirotsu N (2008) New target for rice lodging resistance and its effect in a typhoon. Planta 227(3):601–609
Kashiwagi T, Madoka Y, Hirotsu N, Ishimaru K (2006) Locus prl5 improves lodging resistance of rice by delaying senescence and increasing carbohydrate reaccumulation. Plant Physiol Biochem 44(2–3):152–157
Kashiwagi T, Togawa E, Hirotsu N, Ishimaru K (2008) Improvement of lodging resistance with QTLs for stem diameter in rice (Oryza sativa L.). Theor Appl Genet 117(5):749–757
Kashiwagi T, Munakata J, Ishimaru K (2016) Functional analysis of the lodging resistance QTL BSUC11 on morphological and chemical characteristics in upper culms of rice. Euphytica 210(2):233–243
Kikuchi K, Ueguchi-Tanaka M, Yoshida K, Nagato Y, Matsusoka M, Hirano H-Y (2000) Molecular analysis of the NAC gene family in rice. Mol Gen Genet 262(6):1047–1051
Lee YS, Jeong DH, Lee DY, Yi J, Ryu CH, Kim SL, Jeong HJ, Choi SC, Jin P, Yang J, Cho LH, Choi H, An G (2010) OsCOL4 is a constitutive flowering repressor upstream of Ehd1 and downstream of OsphyB. Plant J 63(1):18–30
Marri PR, Sarla N, Reddy LV, Siddiq EA (2005) Identification and mapping of yield and yield related QTLs from an Indian accession of Oryza rufipogon. BMC Genet 6:33
Matsubara K, Yamamoto E, Kobayashi N, Ishii T, Tanaka J, Tsunematsu H, Yoshinaga S, Matsumura O, Yonemaru J, Mizobuchi R, Yamamoto T, Kato H, Yano M (2016) Improvement of rice biomass yield through QTL-based selection. PLoS ONE 11(3):e0151830
Miyamoto N, Goto Y, Matsui M, Ukai Y, Morita M, Nemoto K (2004) Quantitative trait loci for phyllochron and tillering in rice. Theor Appl Genet 109(4):700–706
Mulsanti IW, Yamamoto T, Ueda T, Samadi AF, Kamahora E, Rumanti IA, Thanh VC, Adachi S, Suzuki S, Kanekatsu M, Hirasawa T, Ookawa T (2018) Finding the superior allele of japonica-type for increasing stem lodging resistance in indica rice varieties using chromosome segment substitution lines. Rice (N Y) 11(1):25
Ogawa D, Abe K, Miyao A, Kojima M, Sakakibara H, Mizutani M, Morita H, Toda Y, Hobo T, Sato Y, Hattori T, Hirochika H, Takeda S (2011) RSS1 regulates the cell cycle and maintains meristematic activity under stress conditions in rice. Nat Commun 2:278
Ogawa D, Morita H, Hattori T, Takeda S (2012) Molecular characterization of the rice protein RSS1 required for meristematic activity under stressful conditions. Plant Physiol Biochem 61:54–60
Ohta H, Nemoto H, Ando I, Kato H, Sato H, Hirabayashi H (2010) “Tachisugata”, a new rice cultivar for whole crop silage use. Bull Natl Inst Crop Sci 11:67–84
Okuno A, Hirano K, Asano K, Takase W, Masuda R, Morinaka Y, Ueguchi-Tanaka M, Kitano H, Matsuoka M (2014) New approach to increasing rice lodging resistance and biomass yield through the use of high gibberellin producing varieties. PloS one 9(2):e86870
Ookawa T, Ishihara K (1992) Varietal difference of physical characteristics of the culm related to lodging resistance in paddy rice. Jpn J Crop Sci 61(3):419–425
Ookawa T, Hobo T, Yano M, Murata K, Ando T, Miura H, Asano K, Ochiai Y, Ikeda M, Nishitani R, Ebitani T, Ozaki H, Angeles ER, Hirasawa T, Matsuoka M (2010a) New approach for rice improvement using a pleiotropic QTL gene for lodging resistance and yield. Nat Commun 1:132
Ookawa T, Yasuda K, Kato H, Sakai M, Seto M, Sunaga K, Motobayashi T, Tojo S, Hirasawa T (2010b) Biomass production and lodging resistance in’Leaf Star’, a new long-culm rice forage cultivar. Plant Prod Sci 13(1):58–66
Ookawa T, Aoba R, Yamamoto T, Ueda T, Takai T, Fukuoka S, Ando T, Adachi S, Matsuoka M, Ebitani T, Kato Y, Mulsanti IW, Kishii M, Reynolds M, Pinera F, Kotake T, Kawasaki S, Motobayashi T, Hirasawa T (2016) Precise estimation of genomic regions controlling lodging resistance using a set of reciprocal chromosome segment substitution lines in rice. Sci Rep 6:30572
Sowadan O, Li D, Zhang Y, Zhu S, Hu X, Bhanbhro LB, Edzesi WM, Dang X, Hong D (2018) Mining of favorable alleles for lodging resistance traits in rice (Oryza sativa) through association mapping. Planta. https://doi.org/10.1007/s00425-018-2885-y
Takai T, Ikka T, Kondo K, Nonoue Y, Ono N, Arai-Sanoh Y, Yoshinaga S, Nakano H, Yano M, Kondo M (2014) Genetic mechanisms underlying yield potential in the rice high-yielding cultivar Takanari, based on reciprocal chromosome segment substitution lines. BMC Plant Biol 14(1):295
Wang S, Basten C, Zeng Z (2006) Windows QTL Cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh
Wang Q, Zhang W, Yin Z, Wen CK (2013) Rice CONSTITUTIVE TRIPLE-RESPONSE2 is involved in the ethylene-receptor signalling and regulation of various aspects of rice growth and development. J Exp Bot 64(16):4863–4875
Weinig C, Schmitt J (2004) Environmental effects on the expression of quantitative trait loci and implications for phenotypic evolution. AIBS Bull 54(7):627–635
Xu J, Xiong W, Cao B, Yan T, Luo T, Fan T, Luo M (2013) Molecular characterization and functional analysis of “fruit-weight 2.2-like” gene family in rice. Planta 238(4):643–655
Yadav S, Singh UM, Naik SM, Venkateshwarlu C, Ramayya PJ, Raman KA, Sandhu N, Kumar A (2017) Molecular mapping of QTLs associated with lodging resistance in dry direct-seeded rice (Oryza sativa L.). Front Plant Sci 8:1431
Yano M, Sasaki T (1997) Genetic and molecular dissection of quantitative traits in rice. Plant Mol Biol 35(1–2):145–153
Yano K, Ookawa T, Aya K, Ochiai Y, Hirasawa T, Ebitani T, Takarada T, Yano M, Yamamoto T, Fukuoka S, Wu J, Ando T, Ordonio RL, Hirano K, Matsuoka M (2015) Isolation of a novel lodging resistance QTL gene involved in strigolactone signaling and its pyramiding with a QTL gene involved in another mechanism. Mol Plant 8(2):303–314
Yuan AP, Cao LY, Zhuang JY, Li RZ, Zheng KL, Zhu J, Cheng SH (2003) Analysis of additive and AE interaction effects of QTLs controlling plant height, heading date and panicle number in rice (Oryza sativa L.). Acta Genet Sin 30(10):899–906
Zhang B, Liu X, Qian Q, Liu L, Dong G, Xiong G, Zeng D, Zhou Y (2011) Golgi nucleotide sugar transporter modulates cell wall biosynthesis and plant growth in rice. Proc Natl Acad Sci USA 108(12):5110–5115
Acknowledgements
The authors gratefully acknowledge the financial support that was provided by China Scholarship Council (Grant No. 201606910002); a Grant-in-Aid for Scientific Research (B) by the Japan Society for the Promotion of Science (Grant No. 15H04442).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, M., Yamamoto, E., Yamamoto, T. et al. Mapping of QTLs associated with lodging resistance in rice (Oryza sativa L.) using the recombinant inbred lines derived from two high yielding cultivars, Tachisugata and Hokuriku 193. Plant Growth Regul 87, 267–276 (2019). https://doi.org/10.1007/s10725-018-0468-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-018-0468-3