Abstract
Soybean is a primary source of vegetable oil, accounting for 53 % of the total vegetable oil consumption in the USA in 2013. Soybean oil with high oleic acid and low linolenic acid content is desired, because it not only improves the oxidative stability of the oil, but also reduces the amount of undesirable trans fat production by the hydrogenation processing. The mutant FAD2-1A and FAD2-1B alleles contribute to an elevated oleic acid level, while mutations in FAD3A and FAD3C lead to a reduction in linolenic acid levels of soybean seed oil. Although single nucleotide polymorphism (SNP) SimpleProbe assays have been developed for these genes, breeder-friendly and high-throughput marker systems are needed for large-scale selections in soybean breeding programs. TaqMan or Kompetitive Allele Specific PCR assays were successfully developed to detect and discriminate mutant alleles of FAD2-1A (17D and PI 603452 sources) and FAD2-1B (PI 283327 source) for the high oleic acid trait, as well as those of FAD3A (CX1512-44 and C1640 sources) and FAD3C (CX1512-44 source) for low linolenic acid content soybean. The accuracy of the assays was validated using multiple populations as well as a panel of diverse soybean lines, and a perfect association was observed between the SNP alleles and their corresponding fatty acid phenotypes. These marker assays have been proven to be robust and reliable for detecting mutant and wild-type alleles and thus will assist forward and backcrossing breeding of soybean lines with desired oil profiles in an accurate and efficient manner.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
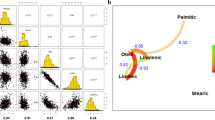
Soybean [Glycine max (L.) Merr.] is one of the most important vegetable oil providers, representing 53 % of the total vegetable oil consumption in the USA in 2013 (http://www.soystats.com/2014). Commercial soybean oil typically contains 11 % palmitic acid (16:0), 4 % stearic acid (18:0), 25 % oleic acid (18:1), 52 % linoleic acid (18:2), and 7 % linolenic acid (18:3) (Fehr 2007). Despite the nutritional benefits of unsaturated fatty acids, the high levels of linoleic and linolenic acids in soybean oil result in low oxidative stability and rapid rancidity, which would significantly reduce food storage time (Warner and Fehr 2008). Thus, soybean oil has partially been hydrogenated to reduce the amount of unstable polyunsaturated fatty acids for industrial and food applications. This hydrogenation process introduces undesirable trans fats, which are believed to be linked to many health concerns, such as risks of obesity, heart diseases, and high levels of cholesterol (Gebauer et al. 2007; Mozaffarian et al. 2009; Katan 1998). The US Food and Drug Administration recently released a final determination indicating that the health risks are associated with the consumption of trans fat. Therefore, soybean oil with high oleic and low linolenic acids is desired because the monounsaturated fatty acids not only provide higher oxidative stability to increase the oil shelf life, but also eliminate the need for trans fat production during food processing.
The molecular genetic basis of high oleic acid soybeans has been found to be the result of mutations in two microsomal delta 12 fatty acid desaturase two enzymes, FAD2-1A (Glyma10g42470) and FAD2-1B (Glyma20g24530), which are responsible for the step of conversion of oleic acid (18:1) precursors to linoleic acid (18:2) precursors in the soybean seed lipid biosynthesis pathway (Schlueter et al. 2007; Pham et al. 2010). FAD2-1A and FAD2-1B are mainly expressed in developing seeds and thus were considered as candidate genes in controlling oleic acid levels in soybean seeds (Pham et al. 2010). Therefore, they have been selected as target genes for improving the oleic acid in breeding programs (Tang et al. 2005; Pham et al. 2010). Mutant alleles of FAD2-1 have been identified from different sources with elevated oleic acid levels (Table 1) (Pham et al. 2010, 2011). In addition, EMS treatment introduced a missense G350A mutation in the coding sequence of FAD2-1A gene in line 17D, resulting in an amino acid change (Dierking and Bilyeu 2009). This missense mutation led to the increase in oleic acid content in seed oil up to 37 % of the total. PI 603452 that has the similar amount of oleic acid content in the seed oil as 17D was also identified by screening the USDA Soybean Germplasm Collection (Pham et al. 2011). A single nucleotide deletion was found in the exon of FAD2-1A gene from PI 603452, which introduced a frame shift of the translation and resulted in a truncated, non-functional protein. Moreover, PI 283327 and PI 210179 exhibit elevated oleic acid level by 10 % and both accessions were shown to possess a mutant allele of FAD2-1B (Pham et al. 2010). These two lines have eight single nucleotide polymorphisms (SNPs) at second exon of FAD2-1B, compared to the reference genome of Williams 82, which resulted in three amino acid changes: S86F, M126V, and P137R. Sequence and association analyses using recombinant inbred lines (RILs) of FAD2-1B mutant suggested that the P137R mutation has a disruptive effect on the enzyme function and that the mutant alleles were associated with the elevated oleic acid content (Pham et al. 2010).
Three loci are associated with linolenic acid content in soybean seed oil, designated as fan1, fan2, and fan3 (Fehr et al. 1992; Rennie and Tanner 1991; Fehr and Hammond 2000). Three versions of omega-3 fatty acid desaturase (FAD3) underlie the three fan loci and are responsible for the conversion of linoleate to linolenate: FAD3A (Glyma14g37350/fan1), FAD3B (Glyma02g39230/fan3), and FAD3C (Glyma18g06950/fan2) (Bilyeu et al. 2003, 2011; Pham et al. 2014). The mutant alleles of FAD3 have been identified from several sources with the reduced linolenic acid content (Table 1) (Chappell and Bilyeu 2006; Bilyeu et al. 2005, 2006). The combination of mutations at all three FAD3 loci significantly reduces the linolenic acid level in seed oil to 1 % in newly developed soybean lines (Bilyeu et al. 2006, 2011). C1640 was the first reported low linolenic acid mutant soybean line with an average of 3.4 % linolenic acid, while the level of linolenic acid is about 7–10 % in wild-type lines (Wilcox and Cavins 1985; Chappell and Bilyeu 2006). A mutation of G798A at the FAD3A locus in C1640 results in a premature stop codon and thus a non-functional enzyme. Genetic studies demonstrated that the mutation in the FAD3A gene in C1640 is associated with the reduced linolenic acid content (Chappell and Bilyeu 2006). The seeds of another low linolenic acid soybean line, CX1512-44, contained about 3 % linolenic acid (Bilyeu et al. 2005, 2011). The characterization of three FAD3 genes from CX1512-44 and association analysis revealed that novel mutations in FAD3A and FAD3C contributed to the reduced linolenic acid content of this soybean line. A mutation of G810A was identified in FAD3A at the first nucleotide following the sixth exon, resulting in misspliced mRNA and failure to produce the authentic amino acid sequence of the protein. FAD3C in CX1512-44 has a SNP of G383A that led to the amino acid change of G128E in a highly conserved region.
Efforts have been made to develop molecular markers for all of the above loci to facilitate the breeding of high oleic acid and low linolenic acid soybean lines. SimpleProbe SNP assays were developed for FAD2-1A (17D) (Pham et al. 2012), FAD2-1B (PI 283327) (Pham et al. 2010), FAD3A (CX1512-44 and C1640), and FAD3C (CX1512-44) (Bilyeu et al. 2011) to select these genes. Cleaved amplified polymorphic sequences (CAPS) were also used to distinguish the mutant and wild-type FAD3A alleles from C1640 (Chappell and Bilyeu 2006). However, these assays require DNA with higher quantity and quality, and longer processing time compared to the TaqMan SNP assays (Chantarangsu et al. 2007; Houghton and Cockerill 2006). Therefore, the goals of this study were to (1) develop breeder-friendly, robust TaqMan (Life Technologies, Carlsbad, CA, USA) or Kompetitive Allele Specific PCR (KASP) (LGC, Petaluma, CA, USA) assays to detect SNPs associated with high oleic acid and low linolenic acid traits for high-throughput marker-assisted selection, and (2) validate these SNPs across diverse soybean germplasms.
Materials and methods
Populations, plant materials, and DNA extraction for genotyping
Two BC3F2 populations obtained from backcrossing programs were used for the validation of FAD2-1 markers. Parent 1 was G00-3213 or G00-3880, which are high-yielding lines developed at the University of Georgia, and parent 2 was progeny selections from the cross of S06-4649RR × (17D × S07-14788), which carried a high oleic acid trait developed at University of Missouri, Portageville, MO (Pham et al. 2010). 17D is a soybean line developed by mutagenesis with 35 % oleic acid (Dierking and Bilyeu 2009) and possesses the FAD2-1A mutant. S07-14788 was selected from an F6 RIL of Jake × PI 283327 developed at the Delta Research Center at Portageville, MO. BC3F2 seeds were planted at Plant Science Farm in Athens, GA, and leaf samples were collected from each individual plant for DNA extraction. At harvest, five seeds from each plant were used for fatty acid analysis.
The materials used to validate the FAD3A (C1640 source) marker were two BC3F2 populations derived from the cross of G00-3213 × [Benning low lin/low palm] and G00-3880 × [Benning low lin/low palm]. Benning low lin/low palm is a near-isogenic line (NIL) carrying reduced palmitic acid (N87-2212-4 source) and linolenic acid (C1640 source) traits, which was developed at the University of Georgia. The BC3F2 seeds were planted at Plant Science Farm in Athens, GA, and leaf samples were collected from each individual plant for DNA extraction. Five seeds from each plant were used for fatty acid profiling.
Two BC1F2 populations were used for the validation of FAD3A and FAD3C from the CX1512-44 source. The recurrent parents were G00-3213HO or G00-3880HO with the high oleic trait. The other parent was TN10-5002 developed at the University of Tennessee (Knoxville, TN, USA), which has the low linolenic acid trait from the CX1512-44 source. BC1F2 seeds were harvested from the winter nursery in Puerto Rico, and 60 seeds from each of three BC1F2 plants for each population were chipped with a razor blade into two pieces: ¼ seed for fatty acid profile analysis and ¾ seed with the embryo for planting in the greenhouse for DNA sampling. Of 180 chipped seeds planted for each BC1F2 population, 123 seeds from the G00-3880HO population and 150 seeds from the G00-3213HO population germinated and leaf tissue were collected from these plants for DNA extraction.
Leaf samples collected from greenhouse-grown seedlings or field-grown plants were arranged into 96-well plates. After sampling, the leaf samples were immediately dried at 55 °C for 24 h and were ground using GenoGrinder (SPEX SamplePrep, Metuchen, NJ, USA) with a single BB (Daisy, Rogers, AR, USA) in each well at 1600 rpm for 2 min. One hundred and twenty soybean germplasms, including 33 North American soybean ancestral lines defined by Gizlice et al. (1994) and 87 plant introductions (PI) and soybean cultivars, were also selected for validation of the SNP marker assays (Table S1). Fifteen seeds were planted in greenhouse for each line in this panel. At least 10 leaves were pooled in 50 ml conical tube (VWR International, Radnor, PA, USA) with one from each plant. Samples were freeze-dried for 48 h and then ground using GenoGrinder with 10 BBs at 1600 rpm for 2 min. DNA was extracted as previously described (Keim et al. 1988).
Development of TaqMan and KASP assays
TaqMan assay primers and probes were designed using Primer Express 3.0.1 software to detect mutant SNPs in FAD2-1A (17D), FAD2-1B (PI 283327), FAD3A (C1640), FAD3A (CX1512-44), and FAD3C (CX1512-44) based on the sequences published (Dierking and Bilyeu 2009; Pham et al. 2010; Bilyeu et al. 2005; Chappell and Bilyeu 2006). Each assay consists of one primer pair to amplify amplicons ranging from 141 to 167 bp, and two SNP-containing probes labeled with VIC (wild type) and FAM (mutant) dyes, respectively.
PCRs were conducted in a 4-µL reaction system with 2 µL template DNA (5–10 ng/µL), 1.8 µL of 2× TaqMan Universal master mix (Life Technologies, Carlsbad, CA, USA), and 0.2 µL of 5× assay mix (final concentration of 0.255 µM for each primer and 0.05 µM for each probe). PCR conditions for TaqMan marker assays were as follows: 95 °C for 10 min, followed by ten cycles of touch down PCR from 68 to 60 °C with 0.8 °C decrease per cycle, then followed by 30 cycles of 92 °C for 15 s and 60 °C for 1 min.
An KASP assay was developed to detect the previously reported SNP in FAD1-2A (PI 603452) (Pham et al. 2011). Primer sequences are listed in Table 2. The KASP assays were performed with 4-µL reaction system including 1.894 µL low rox KASP master mix (KBiosciences, Herts England), 0.106 µL of primer mix (0.318 µM of each forward primer at final concentration), and 2 µL of genomic DNA (10–25 ng/µl). The PCR condition for KASP assay was 94 °C for 15 min, followed by 10 cycles of touch down PCR from 68 to 60 °C with 0.8 °C decrease per cycle, then followed by 30 cycles of 94 °C for 20 s and 57 °C for 1 min.
The PCR plates for both TaqMan and KASP assays were read with Tecan M1000 Pro Infinite Reader (TecanGroup Ltd, Morrisville, NC, USA), and the reader files were imported into Kluster Caller software (LGC, Petaluma, CA, USA) for visualization of clusters and SNP allele calling.
Fatty acid profile and association analysis
The fatty acid profiles of the validation populations, soybean ancestral lines, and cultivars were performed at the USDA Molecular Genetics Laboratory at the University of Missouri–Columbia using five whole seeds or the ¼ seed chips depending on populations. The fatty acid profile was analyzed by following the methods as previously described (Beuselinck et al. 2006). The results were presented as the relative percentage of each fatty acid class in the whole extracted oil fraction for each sample. The fatty acid profiles of 81 plant introductions (PIs) included in the diverse panel were acquired from the USDA Germplasm Resources Information Network (http://www.ars-grin.gov).
Association analysis of genotypes with oleic and linolenic acid contents was performed using single-factor analysis of variance in SAS 9.3 (SAS Institute, 2013).
Results and discussion
To improve the marker assays for FAD2-1A (17D source) and FAD2-1B (PI 283327 source), we successfully developed robust TaqMan assays for both genes based on the SNPs reported by Dierking and Bilyeu (2009) and Pham et al. (2010) and the markers were encoded as GSM257 and GSM256, respectively (Table 2). To validate these marker assays, we performed the association analysis of oleic acid content and FAD2-1A and FAD2-1B marker genotypes using two BC3F2 populations. These populations segregated for mutant FAD2-1A and FAD2-1B alleles derived from 17D and PI 283327, respectively. A clear separation of three genotype clusters with these two markers was observed (Fig. 1a). The seeds with only one FAD2-1 mutation showed only marginal increase in oleic acid content, though the elevation is statistically significant (Table 3), which is consistent with the previous study (Pham et al. 2010). All of the seeds with combined mutations of both FAD2-1A and FAD2-1B exhibited a significant increase in oleic acid content with an average of 77.1 %. Statistical analysis with general linear models (GLM, SAS 9.3) showed a significant correlation between the seed oleic acid level and the interaction of the two SNP markers (R 2 = 0.77, P < 0.001). Because the plants were selected based on the marker genotypes for fatty acid analysis, not all combinations were included. The heterozygotes showed an intermediate level of seed oleic acid content compared to homozygous wild type and mutant (Table 3), indicating that both mutations behave as incompletely dominant and dosage-dependent alleles.
Graphs of five TaqMan marker assays. a Representative SNP graphs of two TaqMan assays of GSM257 (17D source) and GSM256 (PI 283327 source) (samples from one 96-well DNA plate) for the BC3F2 population of G00-3213 × [S06-4649RR × (17D × S07-14788)]. b Representative SNP graphs of two TaqMan assays of GSM254 and GSM255 (CX1512-44) (samples from one 96-well DNA plate) for the BC1F2 population of G00-3213 × TN10-5002. c Representative SNP graph of the TaqMan assay of GSM329 (C1640 source) (samples from one 96-well DNA plate) for the BC3F2 population of G00-3213 x [Benning low lin/low palm]. MUT mutant homozygous genotypes, HET heterozygous genotypes, WT wild-type homozygous genotypes, NTC no template controls
Similarly, three robust TaqMan marker assays were developed for FAD3A (C1640 and CX1512-44) and FAD3C (CX1512-44) based on the SNPs reported previously (Chappell and Bilyeu 2006; Bilyeu et al. 2005) for the selection of low linolenic acid content soybean, and they were designated as GSM329, GSM254, and GSM255, respectively (Table 2). To validate the marker assays of GSM329, we used two BC3F2 populations derived from G00-3213 × [Benning low lin/low palm] and G00-3880 × [Benning low lin/low palm]. DNA samples for genotyping were collected from a combined total of 248 field plants from two populations, and the fatty acid profiles were obtained from a composite of five seeds from each plant. The TaqMan assay of GSM329 showed a clear separation and tight clusters of three genotypes (Fig. 1c). The seed linolenic acid content showed a significant reduction to 6.8 % in heterozygous plants compared to wild type, and a further decrease to the average of 4.8 % was observed in the mutant genotypes (Table 4). A strong association was observed between the seed linolenic acid content and FAD3A (C1640) genotype with R 2 = 0.81 (P < 0.001).
Furthermore, we performed association analysis of TaqMan marker GSM254 and GSM255 with two BC1F2 segregation populations of G00-3213HO × TN10-5002 and G00-3880HO × TN10-5002 for low linolenic acid content. The TaqMan marker assays performed well with a clear separation of three clusters (Fig. 1b). The plants possessing wild-type alleles for both FAD3A and FAD3C showed 5.8 % linolenic acid content; in contrast, the double mutant plants exhibited a significantly decreased level of 1.9 % (Table 5). An intermediate level of linolenic acid was observed in the single homozygous mutant when compared to the double mutant and wild type, and the heterozygotes also have an intermediate linolenic acid content when compared to wild type and mutant within a single FAD3 locus (Table 5), indicating that both FAD3 genes function in a dosage-dependent manner. Statistical analysis using general linear models (GLM, SAS 9.3) showed a strong correlation between the linolenic acid content and both FAD3A and FAD3C genes with R 2 = 0.60 (P < 0.001). An KASP marker assay has been also developed for FAD2-1A from the PI 603452 source based on the single nucleotide deletion previously reported (Pham et al. 2011) and was designated as GSM029 (Table 2). The marker assay was robust and routinely used for genotyping in our breeding programs (data not shown).
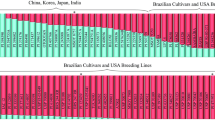
To validate the robustness of these SNP assays across different genetic backgrounds, we also tested these markers with a diverse panel of soybean germplasms consisting of 33 North American soybean ancestral lines and 87 PIs and soybean cultivars (Table S1). The result indicated that all the lines tested possess wild-type alleles for both GSM256 (FAD2-1B) and GSM257 (FAD2-1A), except for two original sources, 17D and PI 283327, and two experimental lines, S12-11641 and S12-11711, which are high oleic lines derived from the sources of 17D and PI 283327 (Table S1). As expected, the oleic acid level of these two lines is significantly higher than that of others. All 120 lines carry the wild-type allele of GSM029 (FAD2-1A) and have 16–27 % oleic acid content except for the original source PI 283327. Three lines: PI 506582, PI 417360, and Jogun have relatively higher oleic acid level (44.5, 50.3, and 40.7 %, respectively), but do not carry the target mutant alleles at either FAD2-1A or FAD2-1B loci. This is similar to what has been reported for a number of wild soybean lines with elevated oleic acid content by Pham et al. (2011). This suggests that these lines might carry other mutant alleles conferring high oleic content other than FAD2-1 alleles from the sources of 17D, PI 283327, and PI 603452. A follow-up study is necessary to confirm this hypothesis.
We also genotyped the panel of validation lines with low linolenic TaqMan assays, GSM329 (FAD3A), GSM254 (FAD3A), and GSM255 (FAD3C). The wild-type allele of GSM329 (FAD3A), present in all the lines except the original donor source C1640 and Benning low lin/low palm, agrees with their normal linolenic acid levels. Benning low lin/low palm is an NIL of Benning with the low linolenic and low palmitic acid traits introgressed from C1640 and N872122-4 (Burton et al. 1994). Similarly, all the lines genotyped possess wild-type alleles for both GSM254 (FAD3A) and GSM255 (FAD3C) except the soybean line TN10-5002 (Dr. Vince Pantalone, unpublished data), which was derived from the original donor, CX1512-44.
Two lines PI 506582 and PI 417360 have a relatively low level of linolenic acid, but carry wild-type alleles at both FAD3A and FAD3C variant SNPs queried. The low level of linolenic acid in these lines could be influenced by high oleic acid content, or the lines possess different alleles which control low linolenic acid. Two high oleic lines, S12-11641 and S12-11711 also have a relatively low level of linolenic acid, which was affected by high oleic contents in these lines. In summary, all these SNP allele calls were consistently corresponding to expected phenotypes, suggesting that these markers are functional and robust, and can be employed across different genetic backgrounds for marker-assisted selection.
The genetic basis of high oleic acid and low linolenic acid content in certain sources has been elucidated, and the functional SNP markers have been identified (Schlueter et al. 2007; Pham et al. 2010; Tang et al. 2005; Dierking and Bilyeu 2009; Bilyeu et al. 2005, 2006; Chappell and Bilyeu 2006). However, the existing SNP marker assays are not robust enough to be applied in soybean breeding programs with high-throughput settings. In this study, we have developed robust TaqMan or KASP functional marker assays for FAD2-1A (17D), FAD2-1A (PI 603452), and FAD2-1B (PI 283327) for the selection of high oleic acid content soybean from three sources, and FAD3A (C1640), FAD3A (CX1512-44), and FAD3C (CX1512-44) for the selection of low linolenic acid content soybean from two sources. TaqMan SNP marker assays are increasingly being applied in fungus, plant, and animal research because of their advantages over other assays (Kim et al. 2010; Giancola et al. 2006; Kulik 2011; Shen et al. 2009; Mori et al. 2008). They are high throughput and cost/time-efficient with low requirements of DNA concentration and quality (e.g., DNA extracted from soybean seeds), easy sample preparation, and simple PCR running protocol. The chemistry of TaqMan assays offers flexibility in designing markers with high allelic specificity, which makes the assays extremely accurate and robust. The clustering of data points from the assay makes the genotyping results straightforward and greatly benefits the SNP allele calling process. Moreover, it has been previously shown that the TaqMan assay is also powerful for detecting DNA deletions (Pham et al. 2014; Fedick et al. 2012).
Soybean seed oil composition, especially high oleic acid and low linolenic acid content, is of tremendous interest to consumers because of its nutritional importance and industrial applications. Breeding of high oleic acid and low linolenic acid content soybean cultivars becomes one of the most important breeding goals. The SNP TaqMan marker assays we developed for the selection of favorable oil composition from certain sources are robust in separating wild type, mutant, and heterozygote and provide more accurate results compared to other available marker assays. Therefore, the assays could be applied in soybean breeding programs to facilitate the marker-assisted selection for high oleic acid and low linolenic acid content traits with improved efficiency and precision. We have widely utilized these assays in our genotyping laboratories to facilitate the development of soybean lines with desired oil profiles from particular sources in an accurate and efficient manner.
References
Beuselinck PR, Sleper DA, Bilyeu KD (2006) An assessment of phenotype selection for linolenic acid using genetic markers. Crop Sci 46(2):747–750
Bilyeu KD, Palavalli L, Sleper DA, Beuselinck PR (2003) Three microsomal omega-3 fatty-acid desaturase genes contribute to soybean linolenic acid levels. Crop Sci 43:1833–1838
Bilyeu K, Palavalli L, Sleper D, Beuselinck P (2005) Mutations in soybean microsomal omega-3 fatty acid desaturase genes reduce linolenic acid concentration in soybean seeds. Crop Sci 45:1830–1836
Bilyeu K, Palavalli L, Sleper DA, Beuselinck P (2006) Molecular genetic resources for development of 1 % linolenic acid soybeans. Crop Sci 46:1913–1918
Bilyeu K, Gillman JD, LeRoy AR (2011) Novel FAD3 mutant allele combinations produce soybeans containing 1 % linolenic acid in the seed oil. Crop Sci 51:259–264
Burton JW, Wilson RF, Brim CA (1994) Registration of N79-2077-12 and N87-2122-4, two soybean germplasm lines with reduced palmitic acid in seed oil. Crop Sci 34:313
Chantarangsu S, Cressey T, Mahasirimongkol S, Tawon Y, Ngo-Giang-Huong N, Jourdain G, Lallemant M, Chantratita W (2007) Comparison of the TaqMan and LightCycler systems in evaluation of CYP2B6 516G > T polymorphism. Mol Cell Probe 21(5–6):408–411
Chappell AS, Bilyeu KD (2006) A GmFAD3A mutation in the low linolenic acid soybean mutant C1640. Plant Breed 125:535–536
Dierking EC, Bilyeu KD (2009) New sources of soybean seed meal and oil composition traits identified through TILLING. BMC Plant Biol 9:89. doi:10.1186/1471-2229-9-89
Fedick A, Su J, Treff NR (2012) Development of TaqMan allelic discrimination based genotyping of large DNA deletions. Genomics 99(3):127–131
Fehr WR (2007) Breeding for modified fatty acid composition in soybean. Crop Sci 47:S72–S87
Fehr W, Hammond E (2000) Reduced linolenic acid production in soybeans. US patent no 6 133 509 issued October 17, 2000
Fehr WR, Welke GA, Hammond EG, Duvick DN, Cianzio SR (1992) Inheritance of reduced linolenic acid content in soybean genotype-A16 and genotype-A17. Crop Sci 32:903–906
Gebauer SK, Psota TL, Kris-Etherton PM (2007) The diversity of health effects of individual trans fatty acid isomers. Lipids 42(9):787–799
Giancola S, McKhann HI, Berard A, Camilleri C, Durand S, Libeau P, Roux F, Reboud X, Gut IG, Brunel D (2006) Utilization of the three high-throughput SNP genotyping methods, the GOOD assay, Amplifluor and TaqMan, in diploid and polyploid plants. Theor Appl Genet 112(6):1115–1124
Gizlice Z, Carter TE, Burton JW (1994) Genetic base for North-American public soybean cultivars released between 1947 and 1988. Crop Sci 34:1143–1151
Houghton SG, Cockerill FR (2006) Real-time PCR: overview and applications. Surgery 139(1):1–5
Katan MB (1998) Health effects of trans fatty acids. Eur J Clin Invest 28(4):257–258
Keim P, Olson TC, Shoemaker RC (1988) A rapid protocol for isolating soybean DNA. Soybean Genet Newsl 15:150–152
Kim KS, Bellendir S, Hudson KA, Hill CB, Hartman GL, Hyten DL, Hudson ME, Diers BW (2010) Fine mapping the soybean aphid resistance gene Rag1 in soybean. Theor Appl Genet 120(5):1063–1071
Kulik T (2011) Development of TaqMan assays for 3AcDON, 15AcDON and NIV Fusarium genotypes based on Tri12 gene. Cereal Res Commun 39(2):200–214
Mori K, Imai K, Matsuda A, Ikeda Y, Naruse S, Hitora-Takeshita H, Nakano M, Taniguchi T, Omi N, Tashiro K, Kinoshita S (2008) LOXL1 genetic polymorphisms are associated with exfoliation glaucoma in the Japanese population. Mol Vis 14:1037–1040
Mozaffarian D, Aro A, Willett WC (2009) Health effects of trans-fatty acids: experimental and observational evidence. Eur J Clin Nutr 63:S5–S21
Pham AT, Lee JD, Shannon JG, Bilyeu KD (2010) Mutant alleles of FAD2-1A and FAD2-1B combine to produce soybeans with the high oleic acid seed oil trait. BMC Plant Biol 10:195. doi:10.1186/1471-2229-10-195
Pham AT, Lee JD, Shannon JG, Bilyeu KD (2011) A novel FAD2-1 A allele in a soybean plant introduction offers an alternate means to produce soybean seed oil with 85 % oleic acid content. Theor Appl Genet 123(5):793–802
Pham AT, Shannon JG, Bilyeu KD (2012) Combinations of mutant FAD2 and FAD3 genes to produce high oleic acid and low linolenic acid soybean oil. Theor Appl Genet 125(3):503–515
Pham AT, Bilyeu K, Chen PY, Boerma HR, Li ZL (2014) Characterization of the fan1 locus in soybean line A5 and development of molecular assays for high-throughput genotyping of FAD3 genes. Mol Breed 33(4):895–907
Rennie BD, Tanner JW (1991) New allele at the fan Locus in the soybean line A5. Crop Sci 31:297–301
Schlueter JA, Vaslenko-Sanders IF, Deshpande S, Yi J, Siegfried M, Roe BA, Schlueter SD, Scheffler BE, Shoemaker RC (2007) The FAD2 gene family of soybean: insights into the structural and functional divergence of a paleoplyploid genome. Crop Sci 47:S14–S26
Shen GQ, Abdullah KG, Wang QK (2009) The TaqMan method for SNP genotyping. Methods Mol Biol 578:293–306
Tang GQ, Novitzky WP, Griffin HC, Huber SC, Dewey RE (2005) Oleate desaturase enzymes of soybean: evidence of regulation through differential stability and phosphorylation. Plant J 44(3):433–446
Warner K, Fehr W (2008) Mid-oleic/ultra low linolenic acid soybean oil: a healthful new alternative to hydrogenated oil for frying. J Am Oil Chem Soc 85(10):945–951
Wilcox JR, Cavins JF (1985) Inheritance of low linolenic acid content of the seed oil of a mutant in Glycine max. Theor Appl Genet 71(1):74–78
Acknowledgments
We would like to acknowledge Dr. H. Roger Boerma at the University of Georgia for developing some of populations which were used in this research and Dale Wood and Steve Finnerty for their technical assistance in field/greenhouse planting. We also thank Colleen Wu and Tatyana Nienow for their assistance with sampling and genotyping. We appreciate the funding support from the United Soybean Board to this research.
Author contribution
ZS drafted the manuscript with edits from ZL, KB, and HN. ZS and NB involved in the data acquisition and marker validation and performed association analysis for all the markers in the study. AP designed all the TaqMan primers and probes. KB involved in the fatty acid profiling and initial SNP discovery. GS and KB contributed to the development of lines used to create populations and assay implementation. ZL provided oversight of the experiment and interpreted the results.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shi, Z., Bachleda, N., Pham, A.T. et al. High-throughput and functional SNP detection assays for oleic and linolenic acids in soybean. Mol Breeding 35, 176 (2015). https://doi.org/10.1007/s11032-015-0368-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0368-4