Abstract
Quantitative structure–activity relationships (QSAR) and molecular docking studies have been performed on a series of 35 α-glucosidase inhibitory derivatives. The QSAR models have been developed by genetic algorithm-multiple linear regression (GA-MLR) and least squares-support vector machine (LS-SVM) methods to correlate the conformational descriptors to the inhibitory activity. The obtained models with 5 descriptors were validated and illustrated to be statistically significant. They had desirable prediction based on squared correlation coefficient (R2), cross-validated correlation coefficient (Q2), root-mean-squares error (RMSE) and Fisher (F) parameters (R2 = 0.951, Q2 = 0.931, RMSE = 0.121, and F = 114.629 for GA-MLR model, and R2 = 0.989, Q2 = 0.987, RMSE = 0.056 and F = 543.754 for LS-SVM model). The crucial descriptor named DELS was explored to have the highest correlation with the inhibitory activity and thus has been chosen to build a simple model. The QSAR model developed with this mono-descriptor showed appropriate results of the predicted model using LS-SVM method (R2 = 0.888, Q2 = 0.872, RMSE = 0.185 and F = 221.459). Also, molecular docking which focuses on the interaction between ligands and α-glucosidase in the protein active site considered different binding positions to find the best binding mode. It helped the QSAR study to propose more comprehensive details of the compounds structures and was used to design more active compounds. The most active designed compound had a high inhibitory activity of 9.22 that can be proposed for the treatment of diabetes type 2.
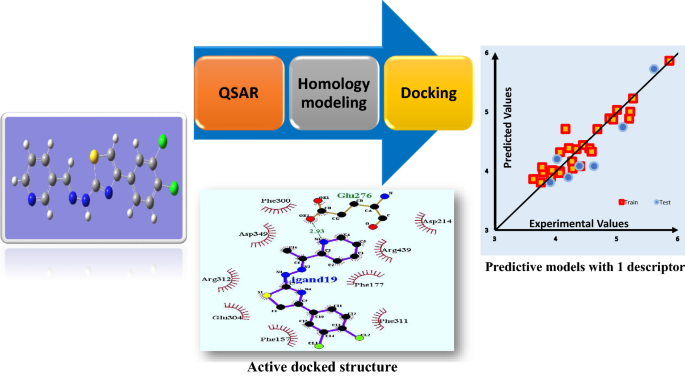
Graphic abstract
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Diabetes is known as the seventh reason for death worldwide [1], and about 438 million people will suffer from this disease by 2030 [2]. In the diabetes category, diabetes type 2 is the most common illness which includes about 80–90% of diabetic cases [3]. There are two important factors with a major influence on diabetic problems. The first one is the insulin hormone which is released by the pancreas and converts glucose to the required energy of the cells, and the second one is the enzyme α-glucosidase that breaks long-chain carbohydrates into small ones such as glucose and fructose. In the body of a diabetic patient, enough insulin is not released or it does not work properly, while α-glucosidase continues its activity. This leads to the aggregation of glucose in the blood (hyperglycemia) which can hurt different organs of the body, especially the nerves and blood vessels [4]. Therefore, inhibiting the catalytic activity of α-glucosidase is considered as a solution to control the amount of glucose in the blood particularly in individuals with diabetes mellitus type 2 [5, 6]. Some different glycosidic inhibitors of α-glucosidase have been used such as miglitol [7], voglibose and acarbose [8]; although these inhibitors are effective, they have some side effects such as flatulence, diarrhea and abdominal discomfort and have to be used in combination of other medications to increase efficiency [9]. Thus, great attention is paid to discover or design novel and efficient inhibitors. Among various admixtures, heterocyclic compounds are notable options. They can be used to synthesize and produce new drugs due to their synergy with most of the molecular targets [10]. Pyridine as a heterocyclic compound and primitive section of many natural compounds have received much attention to be used in a new generation of drugs [11].
Pharmacists believe that the chemical properties of any segment in a drug depend on its structure. So, structural knowledge is required to anticipate pharmaceutical function. In order to save time and investment in the process of designing effective medicines, more useful methods than trial and error are required and QSAR serves as a beneficial computer tool for this purpose. This method makes a rational relationship between the structure of compounds and their properties and finally predicts the biological activities of the compounds to be prepared. QSAR is a mathematical relationship between biological activity and chemical properties of compounds in form of Yi = Fi (X1, X2,…, Xn) where Yi is the dependent variable (IC50) and Xi refers to a molecular descriptor as the independent variable [12,13,14,15,16].
Molecular docking as a complementary tool for QSAR modeling is an advantageous method to calculate the descriptors that contain significant structural information of the compounds. The main role of this method is to seek different orientations of ligands in protein active sites as the receptor. In this way, molecular docking generates a series of complexes and predicts the best orientation for ligand binding [17].
A survey on recent works about the inhibition of α-glucosidase reveals that different compounds have been studied to inhibit its activity using a computer analytical tool as QSAR [1, 4, 7, 18, 19]. But there is not any investigation on arylated hydrazinyl thiazole derivatives that possess inhibition properties against this enzyme. Hence, in this study, the required effective concentration for causing 50% inhibition (IC50) of α-glucosidase for 35 arylated hydrazinyl thiazole-based pyridine derivatives has been predicted by the QSAR models. In this process, two different modeling methods, namely MLR and LS-SVM, were used to predict the inhibitory activity. On the other hand, molecular docking has been used to interpret the binding interactions of the compounds and calculate all available descriptors. These compounds were considered as ligands, and molecular docking describes the different binding positions of the ligands in the active sites of the target protein (α-glucosidase). This work was especially aimed to build the easiest model for the descriptor. This goal needs to follow the QSAR and molecular docking procedures. Then validation of the calculated models is required. After all of these efforts to find the most simple model, the statistical results of the predictive model will be compared with previous works in this scope. Moreover, some new compounds have been designed by combining QSAR and molecular docking results with improved inhibitory activities.
Materials and methods
Data set
The data set is obtained from the research of Ali et al. [11] shown in Table 1. It consists of 39 arylated hydrazinyl thiazole-based pyridine derivatives which were synthesized by two-level reaction patterns. These heterocyclic compounds consist of favorable α-glucosidase inhibitory activity. On the other hand, the new synthetic compounds have constructive likeliness as for example pyridine ring, thiazole ring and hydrazine moiety. The remarkable point is the existence of the same amidine moiety as in the antidiabetic agent “metformin.”
In this work, 4 compounds containing NO2 (ionic compounds) have been removed and the remained 35 derivatives were studied. The IC50 values varied in a range of 1.4–168 µM. They were converted to their equivalent pIC50 (− logIC50) values. Figure 1 and Table 1 exhibit the chemical structures and experimental inhibitory activity values of these compounds.
Geometry optimization of compounds
Three-dimensional (3D) structures of the compounds were pre-optimized based on minimum energy molecular geometries by the HyperChem package (Ver. 7.0) [20]. The RM1 optimization method was used for the initial optimization of compounds. The HyperChem output files were introduced to Gaussian software [21], and optimization of compounds was performed based on a semi-empirical (PM6) method with a frequency cycle to find the lowest energy level in every compound (the most stable state of a compound).
Molecular descriptors calculation and selection
QSAR modeling needs some favorite descriptors to describe the relationship between the chemical structure and activity of the molecules. There is various software with a different theoretical basis for this purpose. Here, Dragon software (Ver 7.0) [22] has been used to calculate descriptors. It contains about 4485 descriptors which are divided into several categories including topological and geometrical, ring descriptors, 2D autocorrelation, GETAWAY (GEometry, Topology and Atom-Weights AssemblY) descriptors, physical properties which include zero-, one-, two- and three-dimensional descriptors. In the first step, about 2100 descriptors were calculated for QSAR analysis. If two descriptors have a correlation coefficient higher than 0.9, one of them has to be excluded [23]. Also, all the duplicate and zero descriptors are useless descriptors that have to be removed. So, the number of remained descriptors is reduced to about 978 descriptors. To build the final QSAR model, these descriptors should be chosen proportional to the data set [24, 25].
Model construction and evaluation parameters
QSAR models were developed using the genetic algorithm (GA) technique. GA provided the final practical descriptors of the model, and GA-MLR as a linear method and LS-SVM as a nonlinear method were applied to construct the QSAR models. To evaluate the model, the data set was divided into two subsets: a test set and a train set. The model is built based on the train set, and its efficiency is analyzed based on its performance on the test set. The y-scrambling method was used to choose the test set. In this method, all compounds sorted with descending data and about 20% of the data was chosen as the test set (7–10 compounds).
Assessment of the model performance was performed via the leave-one-out (LOO) cross-validation method. This is the most popular method to evaluate a QSAR model. In this method, there is a sample set of n members. Each member would be set aside in turn, and the modeling would be applied to the other n − 1 remaining members. This process will be continued until all members are put aside once. Every time, the R2 parameter is evaluated and the values closer to unity lead to less error for activity prediction [26]. The applicability domain and some other important parameters such as RMSE and F have to be studied for a more thorough validation as an inevitable step of QSAR modeling.
As stated earlier, to verify a QSAR model, usually, the LOO cross-validation procedure is applied. The outcome represented by the cross-validated correlation coefficient (R2), which is calculated based on the below formula:
Here \( y_{i} \),\( \hat{y}_{i} \) and \( \bar{y} \) are the actual, estimated and averaged (over the entire data set) activities, respectively [25]. According to the literature, a good model should pass the following conditions [19, 27]:
Q2 coefficient of leave-one-out cross-validation, R2 squared correlation coefficient, k slope of the regression line through the origin, R 20 regression of the anticipated activities opposed to observed activities.
Other important statistical parameters that are required to have a perfect comparison between different models are defined as: S standard error of estimation, F Fischer ratio.
And RMSE values calculated as follows [27]:
where \( y_{i} \) the experimental value of the activity, \( y_{0} \) the predicted value of inhibitory activity using the model, \( n_{s} \) the number of molecules in the data set, lower values of “S” and “RMSE” together with a higher measure of “F” means that the model can forecast the biological activity with lower error, and it can reveal the high prediction potential of the QSAR models.
Applicability domain
The applicability domain is a theoretical space in which the predictions of QSAR are reliable. There are different approaches to determine the applicability domain, but here the most common method, i.e., the William plot is used. It involves the calculation of the standardized residuals versus leverage amounts. Calculation of the leverage (hi) for each compound and its threshold is defined in Eqs. (7) and (8), respectively. Compounds with leverage more than warning leverage (h*) usually had a great influence on the model. A point in the right side of h* with a residual more than 3 or less than − 3 is known as the over-fitted point.
In Eq. (7), xi is the descriptor vector of the query molecule and X is the k × n matrix containing the k descriptor values for the train molecules (n members).
In Eq. (8), k is the number of descriptors in the selected model, and n is defined as the number of objects in the train set [28].
Molecular docking
Molecular docking is an accurate approach to predict the binding affinity and orientation of ligands to the target molecules which is enzyme α-glucosidase in our study. Since the 3D structure of the protein was not available in the protein data bank, the homology modeling method is applied as an alternative solution. This method predicts the structure of an unknown protein based on the structure of similar proteins from the same family [29]. In this study, the homology modeling was used with the template 3A47 [30].
Then molecular docking was run by AutoDock 4.2 software [31]. For all docking parameters, standard values were used. A two-dimensional schematic representations of the docking results including binding sites and interactions of inhibitor with ligands were proposed using LIGPLOT [32].
QSAR and molecular docking could be applied for designing new inhibitors. According to the basic structures, i.e., arylated hydrazinyl thiazole-based pyridine scaffold, new inhibitors have been designed to reduce the inhibitory level. The results of QSAR and molecular docking of the main compounds were carefully investigated to detect the most effective basic structures. Then, the best structures were modified by replacing some of their branches with various useful components. So, some new basic structures were produced (about 126).
Results and discussion
MLR and LS-SVM models
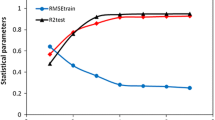
To find a statistically rational QSAR model, the number of independent variables has to be determined through a reliable approach so that in this study the final number of model descriptors was set by the “breaking point” method. This method is based on the sloping trend of statistical parameters versus the number of descriptors. Figure 2 shows that the slope of the breaking point diagram starts to drop off from the fifth descriptor. As far as the smallest suitable number of descriptors is concerned, the breaking point is the optimum number of descriptors [19], which is 5 in this case.
GA was used to select the most effective descriptors in a huge space of different features. The selected descriptors were then assessed to be incorporated in the final model. Consecutively, the final models were built on the 5 selected descriptors as presented in Table 2.
The linear function including the selected variables was obtained using GA-MLR method as below:
The equation indicates descriptors G1p and GATS4s have the highest coefficients in the model, and they have a direct relationship with pIC50. To better illustrate the influence of these variables, their correlation with each other and pIC50 was calculated and the results are collected in Table 3. It has been demonstrated that descriptor DELS provides the highest correlation with the inhibitory activity which makes it a crucial descriptor to build the model. So, a mono-descriptor model named simple model was made with descriptor DELS.
Evaluation of the models
GA-MLR model includes 5 final variables as the most influential descriptors. To assess the nonlinearity relation between the descriptors and pIC50 a reliable model was constructed based on 5 selected descriptors by use of the LS-SVM method. The results of this model were significant. So, it is a good solution to compare the predictive ability of the model through two different methods.
One of the evaluation methods is the comparison of statistical parameters related to QSAR models. In this case, parameters such as Q2, R2, RMSE, F and S were calculated for the MLR and LS-SVM models. These results for the GA-MLR model based on 5 descriptors and 10 test compounds in Table 4 represent a good prediction capacity. The model has a high multiple correlation coefficient (0.951) and a low prediction error. Figures 3 and 4 illustrate the calculated and experimental values of pIC50 for the train and test data set, respectively. The maximum prediction error was a 5.028% error which is acceptable.
The regression line indicates the comparison between predicted and experimental values in Fig. 5. Also, the residual graph of the MLR model with 5 final descriptors is shown in Fig. 6. As it is obvious, congestion of compounds either for train or for test set shows they are well distributed, and none of them has unaccepted distance from the fitted lines.
According to Fig. 7, in the applicability domain analysis, one point (compound 12) with a residual more than 3 in William plot was predicted with slightly higher error. These errors may be due to an error in experimental data. The other points all stayed in the determined applicability domain by William plot.
On the other hand, the statistical results of the LS-SVM model with 5 descriptors in Table 4 describe that the model can predict appropriately and it is more useful than its MLR model. Like the MLR model, in the LS-SVM model based on 5 descriptors, the residual graph in Fig. 8 demonstrates the proper distribution of the data set.
As it is shown in Fig. 9, in the William plot of this model all compounds stand in the applicability domain. As a result, both linear and nonlinear models have an acceptable predictive capacity for inhibitory activity calculation. The predicted values of pIC50 using these models are displayed in Table 1.
To ensure the stability of these models, they were validated with different test groups and nearly all of them represented good results. Table 5 indicates the average statistical values of ten new LS-SVM models. These results prove that the fitness of the model is not dependent on the selected test set as by varying the test and train set it still can predict satisfactorily. Therefore the models present favorable statistical results to be trusted as reliable predictive models.
Descriptors analysis to explore a simple model
The effectiveness of each descriptor in the QSAR model is investigated with sensitivity analysis. In this method, a descriptor is eliminated and the difference between RMSE values in this state and the base case (with all descriptors) is observed. A greater difference means that the descriptor had a more profound role in the model [27].
Figure 10 describes the calculated sensitivity test values to find the most effective descriptors in the model.
Different descriptors as independent variables in the linear equation come from several categories in Dragon descriptors and thus convey different structural information about the compounds. DELS is a topological descriptor with a positive sign in the MLR equation that discloses basic information about the size of molecules, degree of branching, flexibility and the overall shape topological indices which are 2D descriptors based on graph theory concepts [19]. Another essential descriptor, GATS4s, is the Geary autocorrelation of lag 4 weighted by I-state, containing information about the distribution of inherent state along with the topological structure [33]. The higher value of this descriptor leads to higher pIC50. H4m is H autocorrelation of lag 4/weighted by atomic masses which is a GETAWAY descriptor [34] whose lower values cause higher pIC50. Descriptor G1p is the 1st component symmetry directional WHIM index/weighted by atomic polarizabilities [35]. It has a positive sign in the MLR equation, and thus, the pIC50 value increases at higher values of this descriptor (the higher value of pIC50 indicates a lower value of inhibitory activity). The last descriptor D/Dtr05 is a ring descriptor [36] with a negative sign in the linear equation and a negative effect on pIC50 based on the sensitivity test.
As expected (according to Table 3) DELS descriptor had the main role among all descriptors. Results show that H4m, GATS4s, D/Dtr05, and G1p stand in the next places, respectively.
Table 3 illustrates that the DELS descriptor has a high correlation (0.796) with pIC50 which is verified by sensitivity analysis too. Therefore, it seems necessary to make a comparative study between the base model and the model constructed with this descriptor. To evaluate the simple QSAR model constructed by the use of the LS-SVM method, the statistical results were derived, and as reported in Table 6, they present a satisfactory accuracy.
According to this table, R2 = 0.888, Q2 = 0.872, RMSE = 0.185 and F = 221.459, which means the simple model can be a favorable model to predict pIC50 values of the compounds with a high degree of reliability. Besides, the regression line in Fig. 11 and residual diagram in Fig. 12 show the acceptable dispersion of compounds by the simple model.
Not only the selected model with a specific test group had good statistical results, but also other different test groups were studied and showed acceptable ability to predict the inhibitory activity. The final results of models based on the DELS descriptor with different test groups are summarized in Table 7.
As the final result, in this research, a simple model with only one descriptor (DELS) through the LS-SVM method was extracted to predict pIC50 values of α-glucosidase inhibitors with good statistical features. However, the other calculated models using GA-MLR and LS-SVM methods had better statistical results albeit with 5 variables and the nonlinear model had even better prediction capability. The best R2 value in previous studies is 0.872, and most of these studies had used just a single linear or nonlinear method to build their QSAR models. Hence, it seems necessary to compare different linear and nonlinear models to find the best model for pIC50 prediction. Table 8 presents a summarized survey on various works in this scope, and it can be observed that this work has better results in comparison with recent studies in this field. Therefore, the presented models can be useful to predict the inhibitory activity of these special α-glucosidase inhibitors.
Homology modeling
The baker’s yeast α-glucosidase was applied in the homology modeling approach. A suitable structural template was found for homology modeling in the Protein Data Bank (PDB) at the National Center for Biotechnology and Information (NCBI). The amino acid sequence of the α-glucosidase was inputted using BLAST and PSIBLAST algorithms and was retrieved with 72.51% identification to build the homology model that comprises 584 amino acid residues from the SWISS-PROT protein sequence data bank (http://www.expasy.org/sprot/; Accession No.). Figure 13 shows sequence alignment between yeast α-glucosidase and the template 3A47 taken from SWISS-MODEL site. The structure of the simulated protein was designed and is depicted in Fig. 14.
The Ramachandra servera was used to evaluate the accuracy of amino acid placements which was determined to be equal to 97% according to Fig. 15. In other words 97% of the amino acids have been located in allowable zones which indicates the high quality of the forecasted structure.
Molecular docking
Molecular docking was exerted on the compounds to calculate useful descriptors and considering different orientations of ligands in the α-glucosidase active site. All docking features were obtained by the use of AutoDock tools and binana [37]. Different models were established by these descriptors, but none of them had good statistical results as good as Dragon descriptors to apply the significant effect in QSAR modeling.
The different binding mode of ligands with protein was considered. The impressive interaction of the inhibitors with the diversified residues in the active site of the enzyme was gained.
Finding a rational relation between these compounds and their structures to understand how some compounds had the most activity depends on their structural properties, and often it is hard work. In this study, three of the most active compounds are shown in Fig. 16: compound 9, 19 and 28. The common residuals in these compounds comprise from Phe (177, 311, 157) and Arg (312, 439) groups. They had an effective role to improve pIC50 values. Also, it demonstrated the hydrophobic interactions between the enzyme and ligands, different atoms in the structures and their positions, the residuals, hydrogen binding and the other connections. The best binding position of protein with ligands in the active site receptor is useful to design and produce some new drugs.
Different descriptors of the calculated QSAR model already described physical and topological properties, geometry, ring structures and atom binding position have a significant effect on the inhibitory activity. Also, information from molecular docking processes can be used to understand the structure of the compounds with more details which helps QSAR explain compounds structurally and find the best compounds to produce medicine. So, according to the QSAR and docking findings, it is necessary to notice how atoms are gathered to construct the complexes.
Analysis of designed compounds
New inhibitors have been designed based on arylated hydrazinyl thiazole-based pyridine scaffold by QSAR and molecular docking approach. A study of the inhibitors reveals that halogen molecules (F and Cl atoms) and OH have a key role in increasing the inhibitory activity. The most active designed inhibitors are shown in Table S1 and Figure S1 (as supporting information in supplementary materials) with their structures and calculated pIC50 values using the presented MLR model based on 5 descriptors. All pIC50 values are better than the main inhibitors of the study. Of course, these values need to be verified experimentally after the synthesis of the designed compounds.
In the docking process, the correlation between free energy and pIC50 values was calculated for all designed compounds. Although the correlation has been improved in comparison with the main descriptors, it still does not have a significant value (− 0.226). Also, the interaction of molecules by different amino acids was investigated. The common residuals that have been repeated almost in all inhibitors are the Phe (157, 177, 158 and 311). Two structures with high activity had a hydrophobic interaction of His 239, Arg 312 and Asp 349. 2D representation of the most active new structures A1, A2 and B3 is shown in Fig. 17.
Conclusion
In the present study, two different approaches, namely GA-MLR and LS-SVM methods, were applied to establish linear and nonlinear QSAR models to predict the biological activity of a set of arylated hydrazinyl thiazole-based pyridine derivatives. Among various descriptors calculated, the 5 most potent descriptors were selected via GA to build the final QSAR model. DELS descriptor among the selected descriptors had the highest correlation (0.796) with pIC50. It was able to build a QSAR model solely with favorable prediction ability. In previous studies on α-glucosidase inhibition, the best-reported value for R2 was about 0.872, while in the present study with a QSAR model with 5 final descriptors the value of R2 is 0.989 in the nonlinear model and it is about 0.888 in the simple model (using descriptor DELS based on LS-SVM method). Thus the presented models even the simple model can forecast the inhibitory activity of the compounds with higher accuracy than the previous modeling studies. Also, branching information and the size of molecules that come from the DELS descriptor had been considered as the most effective subjects on inhibitory activities of the compounds. Three of the best predicted pIC50 values belong to compounds 9, 19 and 28 all have an aromatic ring connected to two branches of Cl atoms next to each other which reveals the fundamental role of halogen atoms in the inhibition of enzyme activities. Finally, the most active designed compounds (addressed as A1 in this study) had the best pIC50 value of 9.22 comparable to the basic data set.
References
Imran S, Taha M, Ismail NH, Kashif SM, Rahim F, Jamil W, Hariono M, Yusuf M, Wahab H (2015) Synthesis of novel flavone hydrazones: in-vitro evaluation of α-glucosidase inhibition, QSAR analysis and docking studies. Eur J Med Chem 105:156–170. https://doi.org/10.1016/j.ejmech.2015.10.017
Goldenberg RM (2011) Management of unmet needs in type 2 diabetes mellitus: the role of incretin agents. Can J Diabetes 35(5):518–527. https://doi.org/10.1016/S1499-2671(11)80008-0
Narender T, Madhur G, Jaiswal N, Agrawal M, Maurya CK, Rahuja N, Srivastava AK, Tamrakar AK (2013) Synthesis of novel triterpene and N-allylated/N-alkylated niacin hybrids as α-glucosidase inhibitors. Eur J Med Chem 63:162–169. https://doi.org/10.1016/j.ejmech.2013.01.053
Dinparast L, Valizadeh H, Bahadori MB, Soltani S, Asghari B, Rashidi MR (2016) Design, synthesis, α-glucosidase inhibitory activity, molecular docking and QSAR studies of benzimidazole derivatives. J Mol Struct 1114:84–94. https://doi.org/10.1016/j.molstruc.2016.02.005
Park H, Hwang KY, Kim YH, Oh KH, Lee JY, Kim K (2008) Discovery and biological evaluation of novel α-glucosidase inhibitors with in vivo antidiabetic effect. Bioorg Med Chem Lett 18(13):3711–3715. https://doi.org/10.1016/j.bmcl.2008.05.056
Park H, Hwang KY, Oh KH, Kim YH, Lee JY, Kim K (2008) Discovery of novel α-glucosidase inhibitors based on the virtual screening with the homology-modeled protein structure. Bioorg Med Chem 16(1):284–292. https://doi.org/10.1016/j.bmc.2007.09.036
Asadollahi-Baboli M, Dehnavi S (2018) Docking and QSAR analysis of tetracyclic oxindole derivatives as α-glucosidase inhibitors. Comput Biol Chem 76:283–292. https://doi.org/10.1016/j.compbiolchem.2018.07.019
Scott LJ, Spencer CM (2000) Miglitol: a review of its therapeutic potential in type 2 diabetes. Drugs 59(3):521–549. https://doi.org/10.2165/00003495-200059030-00012
Wang SL (2018) New novel α-glucosidase inhibitors produced by microbial conversion. Process Biochem 65:228–232. https://doi.org/10.1016/j.procbio.2017.11.016
Channar PA, Saeed A, Larik FA, Rashid S, Iqbal Q, Rozi M, Younis S, Mahar J (2017) Design and synthesis of 2, 6-di (substituted phenyl) thiazolo [3, 2-b]-1, 2, 4-triazoles as α-glucosidase and α-amylase inhibitors, co-relative pharmacokinetics and 3D QSAR and risk analysis. Biomed Pharmacother 94:499–513. https://doi.org/10.1016/j.biopha.2017.07.139
Ali F, Khan KM, Salar U, Taha M, Ismail NH, Wadood A, Riaz M, Perveen S (2017) Hydrazinyl arylthiazole based pyridine scaffolds: synthesis, structural characterization, in vitro α-glucosidase inhibitory activity, and in silico studies. Eur J Med Chem 138:255–272. https://doi.org/10.1016/j.ejmech.2017.06.041
Ghaslani D, Gorji ZE, Gorji AE, Riahi S (2017) Descriptive and predictive models for Henry’s law constant of CO2 in ionic liquids: a QSPR study. Chem Eng Res Des 120:15–25. https://doi.org/10.1016/j.cherd.2016.12.020
Hasanebrahimi G, Riahi S, Fini MF (2017) Exploring beneficial structural features of ionic surfactants for wettability alteration of carbonate rocks using QSPR modeling technique. J Mol Liq 240:196–208. https://doi.org/10.1016/j.molliq.2017.05.009
Mehraein I, Riahi S (2017) The QSPR models to predict the solubility of CO2 in ionic liquids based on least-squares support vector machines and genetic algorithm-multi linear regression. J Mol Liq 225:521–530. https://doi.org/10.1016/j.molliq.2016.10.133
Abbasi-Radmoghaddam Z, Riahi S, Gharaghani S, Mohammadi-Khanaposhtanai M (2020) Design of potential anti-tumor PARP-1 inhibitors by QSAR and molecular modeling studies. Mol Divers. https://doi.org/10.1007/s11030-020-10063-9
Rezaei B, Riahi S (2016) Prediction of CO2 loading of amines in carbon capture process using membrane contactors: a molecular modeling. J Nat Gas Sci Eng 33:388–396. https://doi.org/10.1016/j.jngse.2016.05.003
Liu Z, Liu Y, Zeng G, Shao B, Chen M, Li Z, Jiang Y, Liu Y, Zhang Y, Zhong H (2018) Application of molecular docking for the degradation of organic pollutants in the environmental remediation: a review. Chemosphere 203:139–150. https://doi.org/10.1016/j.chemosphere.2018.03.179
Liu Y, Ke Z, Cui J, Chen WH, Ma L, Wang B (2008) Synthesis, inhibitory activities, and QSAR study of xanthone derivatives as α-glucosidase inhibitors. Bioorg Med Chem 16(15):7185–7192. https://doi.org/10.1016/j.bmc.2008.06.043
Kraim K, Khatmi D, Saihi Y, Ferkous F, Brahimi M (2009) Quantitative structure activity relationship for the computational prediction of α-glucosidase inhibitory. Chemom Intell Lab Syst 97(2):118–126. https://doi.org/10.1016/j.chemolab.2009.03.006
Release H (2002) 7.5 for windows, molecular modeling system, Hypercube. Inc. http://www.hyper.com
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H (2009) Gaussian 09, Gaussian, Inc., Wallingford, CT, vol 32, pp 5648–5652
Todeschini R, Consonni V, Mauri A, Pavan M (2002) DRAGON software. Milano, Italy
Gagic Z, Nikolic K, Ivkovic B, Filipic S, Agbaba D (2016) QSAR studies and design of new analogs of vitamin E with enhanced antiproliferative activity on MCF-7 breast cancer cells. J Taiwan Inst Chem Eng 59:33–44. https://doi.org/10.1016/j.jtice.2015.07.019
Roy K, Kar S, Das RN (2015) Understanding the basics of QSAR for applications in pharmaceutical sciences and risk assessment. Academic Press, London
Golbraikh A, Tropsha A (2000) Predictive QSAR modeling based on diversity sampling of experimental datasets for the training and test set selection. Mol Divers 5(4):231–243. https://doi.org/10.1023/A:1021372108686
Hawkins DM, Basak SC, Mills D (2003) Assessing model fit by cross-validation. J Chem Inf Comput Sci 43(2):579–586. https://doi.org/10.1021/ci025626i
Gharaghani S, Khayamian T, Ebrahimi M (2013) Molecular dynamics simulation study and molecular docking descriptors in structure-based QSAR on acetylcholinesterase (AChE) inhibitors. SAR QSAR Environ Res 24(9):773–794. https://doi.org/10.1080/1062936X.2013.792877
Aouidate A, Ghaleb A, Ghamali M, Chtita S, Choukrad M, Sbai A, Bouachrine M, Lakhlifi T (2016) Combining DFT and QSAR studies for predicting psychotomimetic activity of substituted phenethylamines using statistical methods. J Taibah Univers Sci 10(6):787–796. https://doi.org/10.1016/j.jtusci.2016.07.001
Pitman MR, Menz RI (2006). Methods for protein homology modelling. In: Applied mycology and biotechnology, vol 6. Elsevier, pp 37–59
https://www.uniprot.org/uniprot/P53341.fasta. Accessed Sept 2018
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461. https://doi.org/10.1016/S1874-5334(06)80005-5
Laskowski RA, Swindells MB. (2011). LigPlot+: multiple ligand–protein interaction diagrams for drug discovery, pp 2778–2786. https://doi.org/10.1021/ci200227u
He J, Peng T, Yang X, Liu H (2018) Development of QSAR models for predicting the binding affinity of endocrine disrupting chemicals to eight fish estrogen receptor. Ecotoxicol Environ Saf 148:211–219. https://doi.org/10.1016/j.ecoenv.2017.10.023
Jouyban A, Shayanfar A, Ghafourian T, Acree WE Jr (2014) Solubility prediction of pharmaceuticals in dioxane + water mixtures at various temperatures: effects of different descriptors and feature selection methods. J Mol Liq 195:125–131. https://doi.org/10.1016/j.molliq.2014.02.012
Jukić M, Rastija V, Opačak-Bernardi T, Stolić I, Krstulović L, Bajić M, Glavaš-Obrovac L (2017) Antitumor activity of 3, 4-ethylenedioxythiophene derivatives and quantitative structure-activity relationship analysis. J Mol Struct 1133:66–73. https://doi.org/10.1016/j.molstruc.2016.11.074
Roy K, Das RN (2013) QSTR with extended topochemical atom (ETA) indices. 16. Development of predictive classification and regression models for toxicity of ionic liquids towards Daphnia magna. J Hazard Mater 254:166–178. https://doi.org/10.1016/j.jhazmat.2013.03.023
Durrant JD, McCammon JA (2011) BINANA: a novel algorithm for ligand-binding characterization. J Mol Gr Model 29(6):888–893. https://doi.org/10.1016/j.jmgm.2011.01.004
Acknowledgments
The authors would like to gratefully acknowledge the support from the Institute of Petroleum Engineering (IPE), University of Tehran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the supplementary material.
Rights and permissions
About this article
Cite this article
Izadpanah, E., Riahi, S., Abbasi-Radmoghaddam, Z. et al. A simple and robust model to predict the inhibitory activity of α-glucosidase inhibitors through combined QSAR modeling and molecular docking techniques. Mol Divers 25, 1811–1825 (2021). https://doi.org/10.1007/s11030-020-10164-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-020-10164-5