Abstract
The study of rare diseases has long been an area of challenge for medical researchers, with agonizingly slow movement towards improved understanding of pathophysiology and treatments compared with more common illnesses. The push towards evidence-based medicine (EBM), which prioritizes certain types of evidence over others, poses a particular issue when mapped onto rare diseases, which may not be feasibly investigated using the methodologies endorsed by EBM, due to a number of constraints. While other trial designs have been suggested to overcome these limitations (with varying success), perhaps the most recent and enthusiastically adopted is the application of artificial intelligence to rare disease data. This paper critically examines the pitfalls of EBM (and its trial design offshoots) as it pertains to rare diseases, exploring the current landscape of AI as a potential solution to these challenges. This discussion is also taken a step further, providing philosophical commentary on the weaknesses and dangers of AI algorithms applied to rare disease research. While not proposing a singular solution, this article does provide a thoughtful reminder that no ‘one-size-fits-all’ approach exists in the complex world of rare diseases. We must balance cautious optimism with critical evaluation of new research paradigms and technology, while at the same time not neglecting the ever-important aspect of patient values and preferences, which may be challenging to incorporate into computer-driven models.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rare diseases have—and continue to—pose a significant challenge for not only the patients who have to live with these conditions, but also the clinicians and researchers attempting to treat and understand them. Despite low individual disease rates, rare diseases collectively impact an estimated 3.5–5.9% of the world’s population (approximately 263–446 million people) (Chung et al. 2022; Nguengang Wakap et al. 2020). In the United States, the Orphan Drug Act defines a rare disease or condition as one impacting less than 200,000 people in the country (a prevalence of < 64 per 100,000 people), while in Europe, this number sits at less than 1 in 2000 people in the general population (Behera et al. 2007; Brasil et al. 2019; Genetic and Rare Diseases Information Center 2023; Hampton 2006; Sernadela et al. 2017). In one-quarter of patients, receiving a diagnosis takes 5–30 years (while others are never diagnosed), typically after having seen countless care providers, undergoing numerous tests, and enduring years with a nameless set of symptoms that few seem to understand (Visibelli et al. 2023). Unfortunately, given the scarcity of treatments for many rare diseases, a patient’s challenges may not be over even once correctly diagnosed. Given that approximately 80% of rare diseases are thought to be genetic in origin, the past several decades’ explosion in genetic research and knowledge about the human genome has led to new insights—as well as questions—about the nature of various rare diseases. To add to the complexity and importance of better understanding such diseases, many present early in life (often from birth), and have a severe, sometimes fatal course. As such, the logistical and ethical challenges of studying diseases in paediatric populations has added additional considerations to rare disease research efforts. This is, of course, on top of the already obvious challenge of working with small population sizes.
In the broader field of medicine, questions surrounding how best to approach clinical research and care are nothing new. In a response to what was seen as biased clinical judgement unsupported by solid scientific backing arose the ‘evidence-based medicine’ (EBM) movement, which provided a set of guidelines by which to critically assess evidence and apply it to clinical practice (Evidence-Based Medicine Working Group 1992; Masic et al. 2008; Rosenberg and Sackett 1996). First proposed in 1981, through a series of articles in the Canadian Medical Association Journal (CMAJ), this approach—touted by its supporters as a ‘paradigm shift’ for practitioners and medical learners alike—described not only how to critically assess evidence, but also put forth a controversial evidence ‘hierarchy,’ placing certain methodologies (e.g. animal studies, case reports) towards the bottom (lowest confidence), and others (e.g. randomized controlled trials (RCTs), meta-analyses) at the top (highest confidence) (Bolignano and Pisano 2016; Burns et al. 2011; Evidence-Based Medicine Working Group 1992; Lester and O’Reilly 2015; Sur and Dahm 2011). EBM, it was claimed, would enable clinicians to critically examine evidence, as well as use the highest quality evidence from ‘unbiased’ research in making decisions for their patients. Unfortunately, this argument is not without its faults, many of which have been strongly voiced by those who have been critical of EBM (e.g. Goldenberg 2006; Kulkarni 2005; Tonelli 1998). Here, I will not provide a full discussion of the strengths and weaknesses of EBM, which have been explored at length in other papers. Rather, I aim to explore EBM’s pitfalls specifically in the context of rare diseases, particularly debates surrounding the generalizability of evidence, and challenges in applying the EBM hierarchy to small, complex patient populations. This will lay the foundation for understanding enthusiasm about artificial intelligence (AI) as a potential solution to the ‘rare disease problem.’
I am not the first to open this debate. Indeed, many have realized the impossibility of applying certain methodologies, like the RCT, to rare disease populations. As a result, numerous ‘solutions’ have been put forth in an attempt to overcome irresolvable limitations for those studying rare diseases. Given the importance of these new approaches to the progression of rare disease research, these alternate methodologies will also be briefly summarized within this discussion (see Table 1). However, the place I wish for us to focus our attention on is implementation of AI into this realm. There is no denying the benefits of new technologies applied in research and clinical care, such as machine learning (ML) algorithms that enable data analysis and pattern recognition far exceeding human capabilities. In the world of rare diseases, AI has been proposed as a revolution that circumvents not only the challenges of EBM-favoured methodologies, but even surpasses the constraints of study designs developed to overcome these challenges.
This paper’s novelty lies in how the conversation is further extended, drawing attention to the issues that exist in harnessing the exciting (and indeed, powerful) opportunities provided by artificial intelligence and applying it to patient data. In much the same way that EBM was touted as a ‘revolution’ in clinical medicine decades ago, there is increasingly a sense that AI technologies will ‘revolutionize’ medicine once again. Given the unique considerations for populations of patients with rare diseases, it is important to take pause and reflect on the logistic and ethical implications of such a ‘solution’ in the long and aggravating battle to make headway for individuals living with poorly understood, rare diseases.
Evidence-based medicine, generalizability, and the ‘rare disease problem’
In the early 1990s, EBM fully emerged, heralded as a “paradigm shift” that would allow clinicians to employ critical assessment of scientific evidence from the medical literature to answer important questions in their field of practice, thereby doing away with ‘biased’ professional judgement (Guyatt et al. 1992). Under the framework of EBM, former ways of making clinical decisions were considered ‘unsystematic,’ citing professional experience and understanding of the basic mechanisms of disease “necessary, but not sufficient guides for clinical practice” (Evidence-Based Medicine Working Group 1992). As described above, a critical component of the EBM paradigm is its evidence hierarchy, which aims to help clinicians employ evidence gathered from the most unbiased, rigorous methodologies. In this hierarchy, RCTs and meta-analyses reside at the top, while evidence from sources such as case reports and animal studies are considered far less favourable and more prone to bias (Fig. 1). Unsurprisingly, EBM has not gone without critiques (e.g. Charlton and Miles 1998; Goldenberg 2006; Kulkarni 2005; Tonelli 1998). A very important question, for the purpose of this conversation specifically as it pertains to rare diseases, is how results from methodologies such as RCTs may be applied to individual patients (Kulkarni 2005).
I believe it is important to acknowledge, as others have, that incorporating ‘evidence’ into medicine is not an all-or-nothing approach. As has been voiced, clinical decisions have always been ‘evidence-based,’ long before the EBM movement came into existence (Kulkarni 2005; Tonelli 1998). The key distinction between prior research traditions in clinical medicine and the EBM movement is, as Kulkarni rightly points out, “a fundamental difference in their philosophical assumptions about what things in clinical medicine are able to be studied (ontology) and how clinical medicine can and should be studied (epistemology and methodology).” As part of this heated debate is the all-important question about the nature of reality, which the EBM framework (with its focus on “systematic, unbiased observation”) purports to better study than other modes of critical inquiry (Kulkarni 2005). Of course, problems exist in this way of thinking, namely the idea that there is a single approach (indeed, any approach) that will enable us to uncover a completely impartial view about the nature of reality. As Maya Goldenberg points out, “our observations are ‘coloured’ by our background beliefs and assumptions (and therefore can never be, even under the most ideal circumstances or controlled experimental settings, the unmitigated perception of the nature of things” (Feyerabend 1978; Goldenberg 2006; Hanson 1958; Kuhn 1970, 1996). Recognizing this philosophical truth creates major issues for EBM and the hierarchy of evidence it claims will lead to medical ‘truths’ and overall improved patient outcomes. Critically, it implies that RCTs, meta-analyses, and other evidence high on the pyramid may not be as sound as once considered (Bolignano and Pisano 2016). And if this is the case, it begs the question: for populations where these methodologies are not even a possibility, such as rare diseases, what better approaches can be employed?
To add to this conversation, consideration from a philosophical perspective has gone further in exploring the crux of EBM itself—the nature of ‘evidence.’ As noted by Jeremy Howick, ‘evidence’ of effectiveness within clinical medicine is not as simple as favourable results from an RCT (or other method high on the EBM hierarchy) (Howick 2011). Instead, one must go deeper to consider the complexity of a concept like ‘evidence’ itself, and the ways this complexity intertwines further with bodies, minds, and natural human variation (Anjum et al. 2020). For instance, evidence that holds clinical utility must also be relevant to individual patients—something that cannot be determined empirically, no matter how rigorous the test. Yes, a new medication may be helpful in a patient managing their weight, for example—but this alone does not satisfy a ‘patient-relevant’ outcome. For this, one must further explore the notion of quality of life/what a patient considers to be a ‘better’ life for themselves—a philosophical debate that is beyond the scope of this paper, but of utmost importance to note within this discussion nonetheless. Beyond patient quality of life, one must also consider benefit-harm analyses when considering evidence. In the overwhelming majority of cases, evidence is presented in terms of statistical significance or effect sizes—while neglecting to explore the many possible side-effects that may come with its use (Howick 2011). Moreover, the relevance of these side effects to an individual patient is impossible to factor into analyses—for one patient, chronic constipation may be a minor annoyance; for another, it may significantly impact their overall wellbeing. An additional word should be said about the notion of ‘best available options’ in medicine, and the way this is often missed when looking solely at the evidence produced in support of a particular treatment, procedure, or other intervention impacting health. For any condition, different options will be available—for some conditions, this is very broad, for others, the number of options may be quite narrow. One option that always exists (but is regularly overlooked) is simply to do nothing at all. While a physician considering data, for instance, of a new pharmaceutical drug, may see strong ‘evidence’ from a trial—in that the drug checks all the boxes that EBM demands, aspects of the larger patient picture may be missed (Worrall, 2022). Particularly for patients with rare diseases, where creative, forward-thinking approaches may be required to determine what the best available option is—for their unique case at both the biological and personal level—EBM is not only about solid evidence, but solid evidence applied to unique cases that are as complex as the human condition itself.
As Simon Day points out in their discussion about EBM and rare diseases, other forms of evidence are critical in piecing together clinical pictures for patients with rare diseases, as quite obviously, “any data are better than none and good and reliable quality of data are better than poor quality and unreliable data” (Day, 2017). Using the common analogy for accuracy and precision—targets on a dartboard—Day makes the important point that in research (particularly research into diseases for which little is known), we may not even know where the “target” being aimed for is. Perfectly randomized trials do little to help if we have no idea what we are aiming at—whether that target be diagnoses, prognosis, treatment, or any other variable of interest (Figs. 2, 3). The key in the case of rare disease, made up of small populations of patients with complex illness, is to stop trying to solve the problem with tools unfit for the job (e.g. RCTs). Instead, we must turn to other forms of evidence, which prioritize quality of evidence over quantity or unnecessary fixation on fitting into the EBM hierarchy.
Reality of data collection. Given that the target is unknown (e.g. in developing a new treatment for a rare disease), we do not know whether the data is on target or not. We also do not know the relative precision—whether bullets are closely packed in relation to the actual size of the target.
This brings us to the important point of generalizability—which is an essential breaking point for those attempting to transplant rare disease research into the confines of EBM. When initially conceived, EBM was seen as a “way to close the gulf between good clinical research and clinical practice” (Rosenberg and Donald 1995; Tonelli 1998). And yet, one must reconcile themselves with the fact that this ‘gap’ between what is observed in research and our individual patient can never be fully resolved, as it “represents [both] an intrinsic, philosophical gap,” and well as an ethical gap (Malterud 1995; Tonelli 1998). The individual patient in front of us is not equivalent to patients documented in research, particularly in the context of methods like RCTs and meta-analyses where participant data may be pooled and difficult (if not impossible) to find reports of individual patient characteristics or measurements. A similar argument exists in rare disease care as that explored by Nancy Cartwright in the context of health policy. If we agree that a philosophical gap exists between the individual patient in front of us and outcomes reported in clinical literature (regardless of how ‘rigorous’ the method claims to be), then we can also agree that even transplanting a similar study protocol from one context (e.g. one patient population) to the next (a small sample of rare disease patients) can never yield precisely the same results (Cartwright 2013). It is very enticing to want to believe that just because a finding is observed somewhere (“there”), that it can be considered to apply widely, and therefore provide the same expectation in some new setting (“here”). Of course, many—even the layperson—would recognize that just because something works in one context, does not mean it can be applied to another. What might function as a fantastic beach umbrella here on Earth would lead to a sorry end for both umbrella and astronaut if employed as sun protection on Venus. Yet, this is precisely what the world of EBM has tried to force upon the study of challenging patient populations, such as those with rare diseases—and such thinking has influenced (as we shall soon see) the ‘solutions’ available to overcome our inability to run studies such as RCTs in these contexts. These ideas have been further explored in the work of Jonathan Fuller, who notes both the myth and fallacy of ‘simple extrapolation’—extrapolating the findings of trials to clinical practice through EBM (Fuller 2021). Within this argument, Fuller notes that a myth exists whereby statistics are wrongly seen as transposable metrics that can be applied from the context of their origin (studies that are ‘solidly constructed’ under EBM criteria) to general patient populations – what has been coined as the ‘myth of the golden risk ratio’ (Fuller 2021; Reiczigel et al. 2017). Furthermore, simple extrapolation carries a crucial fallacy rooted in ignorance, blindly concluding that effect sizes can be transplanted from highly controlled EBM contexts to other patient populations (or even, as is important for our discussion, individual patients), simply because contrary evidence is not provided. It should also be noted that even in cases where a clinician finds an RCT studied in a population similar to the patient in front of them, issues with external validity continue to remain underreported in such trials, and meta-analyses may even be worse for concealing biases (Borgerson 2009). Moreover, failing to incorporate unique patient values and preferences into care creates serious ethical dilemmas—and these factors are impossible to capture and translate from large, quantitative studies into the clinic.
New methodologies, none perfect: fitting rare diseases into EBM
While the key takeaway of this paper lies in the proceeding discussion of AI’s implementation into rare disease research and care, it is important that other methodologies are documented in brief. These approaches have been discussed at length by others, and I encourage those interested to explore each in more detail than what is provided here, which is by no means complete. Rather, this summary will serve as our bridge from a strict EBM paradigm into the newest paradigm of artificial intelligence and its offshoot, machine learning.
As demonstrated in Table 1 below, numerous different methodologies have been proposed to apply EBM principles to smaller patient populations, even down to the level of the individual patient. As with any study methodology, each has their own advantages and disadvantages and crucially, no single approach has been developed as a perfect fit for EBM research applied to rare diseases. For instance, ‘N-of-1’ trials are one such approach for studying individual patients, where (as the name suggests), a single person is included in the trial (Lillie et al. 2011). Here, the principles of crossover RCTs are applied to individual subjects, thus allowing the participant to act as their own control, and with the benefit that preferred treatments can be determined at the individual level (Abrahamyan et al. 2016; Tudur Smith et al. 2014). Unfortunately, as outlined below, N-of-1 trials, while an innovative solution, do suffer the same disadvantages as crossover trials, and meta-analyses based on N-of-1 trials suffer limitations in generalizing findings owing to unique, individual patient variability/characteristics.
Other methodologies face challenges due to the wide variation that exists in diseases themselves, with some approaches suitable for one condition, but perhaps impractical for another. As an example, randomized withdrawal designs/randomized discontinuation designs (whereby subjects receive an experimental treatment for a specified time, after which they are randomly assigned to continue the treatment or be switched to a placebo) have the potential to increase study efficacy because fewer patients are exposed to the placebo, but are limited to predictable, chronic/slowly progressing diseases (Tudur Smith et al. 2014). Conversely, other methods may be limited by the nature of the treatments themselves. For instance, factorial designs, which may be excellent for comparing multiple interventions and uncovering interactions between groups, cannot incorporate treatments that must be administered separately (which would exclude a necessary combination of interventions in the factorial design) (see Table 1). Altogether, these challenges (and more) highlighted in the table below point to the difficulties of applying EBM to small populations of rare disease patient cohorts. Even amongst these novel, rigorous techniques for uncovering clinical ‘evidence,’ no perfect fit for rare disease research emerges, encouraging us to consider the next section of this article, where the novelty of artificial intelligence in this context is explored.
Before moving on, this discussion would not be complete without considering some of the statistical and endpoint challenges that fed into the design of these new methodologies (and still plague many). Given the small sample sizes that are naturally part of studying rare diseases, it can be difficult—if not impossible—to recruit cohorts large enough to reach the standard 80% statistical power typically sought after by those looking for ‘high-quality’ research (as well as those funding this research). As Abrahamayan et al. (2016) point out in their discussion of alternative designs for clinical trials in rare diseases, nothing is inherently biased about small study sizes, but in cases where researchers do find significant P-values (usually set at less than 0.05, another topic of debate), the observed difference will be greater than the true value. In the [more likely] case where insignificant results are obtained, there is a high likelihood that the results may not even have the opportunity to be published in the first place (Abrahamayan et al. 2016). Moreover, many rare diseases arise from genetic causes, of which there may be unique genetic sub-groups that have different responses to the variable(s) under study (e.g. different responses to treatment; different safety profiles). Testing in multiple sub-groups (potentially important to gain a true picture of the study effect) may run into issues of even smaller, more diluted sample sizes. In some cases, researchers may decide to reserve sub-group testing for a final analysis stage to investigate different treatment effects across genetic populations. This too creates issues, where testing in multiple subgroups increases the risk of Type I error, as well as potentially increasing Type II error (particularly if the study was not designed to have sufficient power for treatment-subgroup interactions) (Abrahamayan et al. 2016; Korn 2013). A final consideration during analyses of rare disease populations lies in the challenge of working with small populations, which may have non-normal distributions. With many standard statistical analyses relying on assumptions of normality, researchers may run into issues with meeting necessary assumptions, as well as finding faulty results from analysis that appeared sound on the surface (Abrahamayan et al. 2016; Ludbrook 1995). When working with other illnesses that naturally have larger populations to draw participant samples from, the central limit theorem tends to minimize such concerns surrounding normality (Kwak and Kim 2017).
Beyond statistical challenges and the methodologies themselves, defining endpoints in rare disease research has been cited as yet another critical issue (Abrahamayan et al. 2016; Brown et al. 2018). It is agreed that primary endpoints of clinical trials should have well-defined and reliable measurements, which are also “clinically meaningful and relevant to the patient, readily measurable and sensitive to intervention” (Aronson 2005; Fleming and Powers 2012). Given that many rare diseases have poorly understood etiologies and disease progression, responses to treatments, and incurability (among other considerations), surrogate endpoints that are easy to measure are often employed (Abrahamayan et al. 2016). For instance, as discussed by Brown et al. (2018) in the context of rare oncological tumor research, overall survival (OS) is used as a “robust and realistic indicator of efficacy,” but may not be ideal or entirely realistic as an endpoint for rare tumor trials. For instance, in malignancies that have long survival times, using OS as a primary endpoint may lead to long studies that are not feasible, particularly given the number of patients needed to provide sufficient power to the study. In many rare diseases, surrogate outcomes that come from measures such as biomarkers are cheaper, easier, and faster to measure, as well as being more accepted by patients and clinicians alike (Abrahamayan et al. 2016; Aronson 2005). However, it should be noted that using biomarkers (e.g. lab results) as a primary endpoint may not lead to the most accurate or clinically relevant conclusions (particularly in cases where disease biology is poorly understood). Of course, using solely biological measurements as trial endpoints also brings in the all-important issue of neglecting patient values and preferences in the discussion. Other less easily definable—but no less important—measures, such as quality of life, pain levels, and so forth—may be better endpoints, but this brings in debates surrounding how best to produce ‘unbiased’ measures that can be generalized across patient populations.
Unfortunately, solutions to these challenges are yet to exist (and may never exist). As Bolignano et al. (2014) point out in the setting of rare renal diseases, perhaps the best option is agree to disagree, and focus more specifically on the quality of evidence, rather than quantity. Of course, meeting these goals does not happen in isolation, and the importance of collaboration across health centres and research groups cannot be understated. These collaborations are key, as we shall see, in the context of artificial intelligence and machine learning, our next and final point of discussion.
Machine learning in medicine: panacea to the ‘rare disease problem,’ or an even greater challenge?
As is evident from the preceding discussion, the proposed solutions to the ‘rare disease problem’ continue to have their challenges, and still strive to make themselves fit within an EBM methodological framework. Along with many other areas of medicine, a recent movement towards utilizing AI has entered the world of rare disease research. As most readers will be aware, artificial intelligence is not a new concept, though one that has grown in popularity over the past decade as technological advancements have radically altered what AI can accomplish. One of AI’s most promising elements is its ability to bring together and analyze data from numerous sources, including imaging, multi-omics, laboratory results/biomarkers, electronic health records, and so forth (Brasil et al. 2019). Two subsets of AI that are important to our discussion of medicine and rare diseases are machine learning (ML) and deep learning (DL). Machine learning, which constructs algorithms based on training datasets, can be used to produce outputs that can assist with diagnosis, prognosis, and treatment (Kufel et al. 2023). Deep learning takes this a step further, employing even more complex and abstract models. One of the most important benefits of ML and DL, as it applies to medicine, is its ability to find patterns within data (often enormous datasets) that would be challenging, if not impossible, for humans to recognize (Visibelli et al. 2023).
The parallels between the EBM and AI movements are striking, both seen as paradigm shifts for the clinical and research communities. In each case, EBM and ML models attempt to support clinical decision making, but vary in their epistemological methods (Scott et al. 2021). While EBM uses empirical research to drive inferences, ML focuses on using data mining methods to find patterns and associations in datasets. As Scott et al. (2021) point out in their article exploring the complimentary approaches of EBM and ML, “[o]bservational data and ML are useful when prospective research studies, especially RCTs, are not feasible because of ethical concerns, logistical barriers, limited timespans, cost, or inability to recruit patients and/or clinicians.” They go on to propose that ML could offer a new means of supporting clinical decisions, which is more closely tailored to an individual patient than the information derived from an RCT would be. In short, EBM is based on hypothesis-driven discoveries. ML, on the other hand, is data-driven.
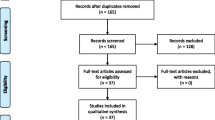
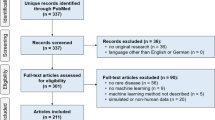
Currently, ML is being applied in numerous areas of rare disease research, primarily diagnosis and to some extent, prognosis and treatment discovery. As an example, for patients with genetic changes, AI algorithms have made great strides in helping us predict the significance of these variants (for instance, whether a specific variant is likely to be pathogenic and contributing to a patient’s phenotype, or simply an incidental finding) (Brasil et al. 2019). Phenotype and biochemical-driven diagnoses are another area where AI is increasingly being explored for rare disease, where computerized recognition of characteristic physical features present in imaging or derived from lab results, may point towards new ways of diagnosis and providing prognosis to patients (Hallowell et al. 2019; Visibelli et al. 2023). Although still in its infancy, using AI for research on treatments for rare diseases is also increasing, such as in models that can simulate therapeutic options to help guide more individualized treatments. According to a recent review by Visibelli et al. (2023), the most commonly applied algorithms are SVM (Support Vector Machine), RF (Random Forest) and ANN (Artificial Neural Networks), which can handle the complex, high-dimensional data that rare disease research demands. Most commonly, images were the sources of data input, which has implications for which rare diseases have the opportunity to be most rigorously studied.As an example, in a scoping review conducted by Schaefer et al. (2020), 211 studies from 32 countries investigating 74 rare diseases were identified. Of these studies, there was an overrepresentation of disease groups that have imaging data, such as neurologic diseases which often have CT, MRI, and other such scans to use for ML pattern recognition. This review also found that most studies of ML for rare diseases focused on diagnosis (40.8%) or prognosis (38.4%), with only a small proportion of studies where ML was applied specifically to improve treatment (4.7%) (Schaefer et al. 2020).
In further conversations about clinical evaluation of ML in medicine, the application of RCTs specifically for the purpose of assessing these models has been suggested (and simultaneously, brought into question). While the medical community calls for more RCTs to explore the reliability and validity of AI approaches in healthcare, assessing ML using EBM-based methodologies does not come without limitations. Whether studying treatments/interventions themselves, or new AI models used to study them, philosophers of science have been careful to point out the challenges of RCTs in the new age of AI (Genin and Grote 2021). These include threats to internal validity (such as level of physician experience/willingness to change decisions based on AI feedback, and other ‘physician effects’) and external validity (e.g. ‘novelty effects,’ where physicians involved in a study are not acclimated to AI technologies as they would be after routine use in the real-world). Many of the same EBM-based suggestions for improving studies, such as randomization and blinding, have been proposed to improve ECTs in medical AI (Genin and Grote 2021). Importantly, the common thread amongst most researchers of rare diseases—including those employing AI algorithms to better understand these conditions—is the need for international collaboration, with initiatives and networks that bring together both data and expertise to a common place.
Despite obvious—and well-deserved—excitement about the many possibilities that AI brings to the world of rare diseases, it is also critical that careful consideration is taken into the limits of AI, logistically as well as ethically. Here, I outline these issues in chronological order (see Table 2), starting at the model development stage. Early in the process of developing an ML or DL model, one of the most obvious issues is that of overfitting and by extension, lack of generalizability. While methods to develop ML models may vary, utilizing training sets to develop models that can then be applied to real-world data is a common feature. Fit your model too closely to the training dataset, and its outputs may not be able to extend to future contexts (Freiesleben and Grote 2023). There is a constant trade-off between fitting the algorithm to the data currently at hand, and having it perform accurately when presented with a new patient. In many areas of medicine, ML holds immense promise—for instance, when presented with thousands of imaging results to discriminate between healthy patients and those with a common disease. However, can the same be said for rare diseases, or is applying ML frameworks only opening an even greater black box and generalizability issue than EBM methodologies such as RCTs?
The answer, I contend, is not so simple—naturally owing to the incredibly varied nature of rare diseases. Let us imagine, for instance, a rare genetic disease resulting from a very specific mutation. We will call this disease X (Fig. 4). Let us also imagine, for argument’s sake, that disease X has full penetrance (every case of the mutation manifests with disease), and has the same, clearly observable, well-defined phenotype for every patient. In such a case, ML algorithms may (and I say ‘may’ with caution) be a wonderful solution to issues such as small sample sizes, where even a tiny patient cohort might provide rich insights into the underlying biology and potential treatments for the disease. ML in this context could allow us to make the most of data collected from few patients, which could then be extended to the care of future patients with the same mutation leading to disease X.
On the other hand (which is far more likely to be the case) let us also consider some disease Y, again a rare genetic condition resulting from single nucleotide mutations. However, perhaps disease Y presents with a variable phenotype, where some features are common to all patients (thus, giving them the ‘disease Y’ diagnosis), while other features are only observed in a select few. Let us further imagine that the phenotype depends on the precise mutation a patient carries. Rather than disease X, which is caused by a single mutation in a single gene, disease Y may be caused by multiple different mutations within a single gene. Or, even more challenging—a similar phenotype may be caused by different mutations in different genes. Quite quickly, we can see that even if ML can provide deep insights into the patients under current study, it is difficult to apply these findings to new patients with disease Y. Doing so may lead to erroneous, if not also dangerously misleading, conclusions. Even in the seemingly ‘clear-cut’ case of disease X, with incredibly small sample sizes of patients for certain rare conditions, it is impossible to know whether the currently reported cases represent the entirety of this patient population—past, present, and future. Just because I pick ten red balls out of a bag does not mean no other colours exist. We would need to empty out the entire bag to be sure (and never add any new balls). Increasing certainty can be found in additional reports of cases that either support or refute our current understanding of disease—the kind of case reports that EBM places at the bottom of its evidence pyramid. Yet, enough reports together may paint the nuanced clinical picture that quantitative methodologies will never fully be able to. Only once we have increased confidence in exactly what we are studying can we fully apply and rely on new, exciting possibilities like AI to the world of rare diseases (Fig. 4).
Hypothetical diseases ‘X’ and ‘Y’. A. Disease ‘X,’ which results from the same genetic change in every patient, and has the same outwardly observable disease characteristics. B. Disease ‘Y,’ which may result from multiple different mutations within the same gene, or different mutations in different genes. This disease may also have variation in disease presentation across patients
An additional consideration at the model development stage, with important implications for generalizability, is the need to include clinical experts in the feature selection process (the process by which variables are selected to include in ML algorithms). While data scientists can bring the expertise required to build complex models, decisions surrounding which inputs are clinically relevant to patients with specific diseases are essential. Otherwise, we run the risk of producing models that may spuriously predict a particular outcome, but with little connection to measures that are actually related to the disease pathology in question. For instance, a clinician may know that certain laboratory measurements are highly indicative of a disease or disease outcome, while other measurements hold no relevance to the patient at hand. Blood haemoglobin levels or liver function tests may be a critical biomarker for one condition, and not another. Just because data is available for certain features (input variables), does not mean it should be randomly integrated into models. Clinical experts need to work alongside computer scientists to ensure that models are being built with variables that have true clinical relevance and value to the patient in front of them.
At the stage of applying AI models to actual patient cohorts, the external validity of these algorithms must be considered (Scott et al. 2021; Visibelli et al. 2023). Without rigorous means of validating models in clinical settings, we have no way of knowing how effective these models are at delivering outputs that are accurate and meaningful to patients and their healthcare providers. When thinking about individuals with rare diseases, this becomes even more important, as described above in cases of overfitting to training datasets. The international community has yet to outline clear and specific guidelines to assess the external validity of ML models (Visibelli et al. 2023; Youssef et al. 2023). As with RCTs, which lack clear guidelines for assessing and reporting external validity—so too do these new AI-driven models. On top of external validity, ensuring that results from an algorithm are reproducible will be integral, just as EBM places a high degree of importance on the reproducibility of studies, and value of systematic reviews and meta-analyses (Beam et al. 2020; Scott et al. 2021). Reproducibility of results will be an important checkpoint in the external validation process to ensure that results are not simply due to chance, perhaps overfit to training data and unable to produce the same results when mapped onto real patients (especially those with complex, rare diseases).
At the stage of implementing ML and DL algorithms in clinical spaces, the question of interpretability becomes critical. Techniques like RCTs are opaque enough as it is to the healthcare providers who may be trying to make sense of them (Wadden 2021). When it comes to the ‘black box’ of computer algorithms, it will be even more essential that transparency is maintained, allowing clinicians to scrutinize these models, understanding what goes on ‘underneath the hood,’ and how output decisions are arrived at Zhang and Zhang 2023). This includes, as occurred with EBM, implementing new additions to the medical curriculum for learners and practicing physicians alike—so that care providers can critically assess the AI tools they are using. Otherwise, we create the danger of entering an era of ‘automated medicine,’ with clinicians blindly believing computer-generated outputs to be absolute truth. While EBM initially aimed to drive away what it saw as ‘biased’ clinical judgements, we must take care that the pendulum does not swing too far in the opposite direction, where perhaps even more biased algorithms take over, and future clinicians are left without the skillset and confidence to make individualized decisions based on professional experience, personal knowledge, and other available evidence (Genin and Grote 2021).
As a final point, ethical issues do not end with AI, but rather, only intensify. Hallowell et al. (2019) brings up a number of these considerations in their paper exploring big data phenotyping in rare diseases, such as the safeguards that must be put in place to handle patient data (especially if, as has been suggested, this data is pooled and used by numerous research groups and organizations, both academic and private) (Larson et al. 2020; Murdoch 2021; Safdar et al. 2020). Moreover, it is clear that large datasets of patient information hold great economic value, in that they can be used for means such as developing new diagnostic technologies and in the discovery of novel treatments (Martinho et al. 2021). How do we prevent patients from being taken advantage of—their health information turned into a commodity, perhaps with no benefit provided to them? What consent processes must take place to ethically create the larger datasets so desperately needed? How can we ensure patients do not feel like they are being turned into a nameless research subject—especially in the context of rare diseases, where patient numbers can be so few, and researchers can be so desperate for data? Along with these ethical issues, AI research must contend with many of the same questions the genetic research community did when introducing new genetic testing methodologies into its practice, such as how to deal with incidental findings.
A last, and very important limitation of tools based on artificial intelligence is the additional complexity when patient values and preferences are considered, which are much more challenging to incorporate into a computerized model than, for instance, objective imaging data. In many ways, AI has stepped into the place that EBM originally did, holding great promise for a ‘clean,’ logical way to go about making clinical decisions. Just as one of the major critiques of EBM was its inability to clearly address personal preferences and experiences of the individual patient, so too does this challenge plague an automated approach to medicine. Now, unlike decades ago when EBM emerged, the issue is even more pressing—can clinical decisions be trusted, if humans are no longer critically evaluating evidence themselves, but rather, relying on a computer to do this evaluation for them? In all cases, we must remember that patient wishes should feed into the ultimate output: the final, mutually-agreed upon decision from care providers and patients. The individual patient, above all, must exist at the finish line of decision-making, regardless of the approach (e.g. EBM, AI, etc.) used to arrive there.
Conclusion
Despite having small patient numbers at the individual level, rare diseases represent an important—and difficult—area of medical investigation. Unfit for the ‘tools’ (e.g. RCTs) highly advocated for in the EBM paradigm, clinicians and researchers alike are left to grapple with how best to study these conditions, and by extension, provide the best care possible to their patients. Despite numerous modified methodologies having been proposed to make rare disease research fit within the scope of EBM, these trial designs come with their own disadvantages, and none are perfect in overcoming every limitation. As with other fields of medicine, artificial intelligence is gaining increasing attention for a potential solution to these issues. However, given the complex, opaque nature of computer algorithms and their outputs, important considerations must take place before thoughtlessly bringing new technology into clinical practice. This article outlined logistic and philosophical factors that must be addressed to ensure safe, accurate, and reliable use of machine learning in the world of rare disease research and care. While artificial intelligence is a powerful tool, it is one that can easily be misapplied. It is important that in the early stages of its integration into healthcare decisions, consistent checks and balances be put in place to ensure the best for our patients—and to ensure that those with rare diseases (whose data makes advancement in the field possible)—also benefit from medical progress. Will AI hold the key for improving rare disease research and care, or only complicate matters further? Likely both, though only time will tell. Until then, clinicians will be left to grapple with the ongoing challenge that infrequent, poorly understood diseases present—and patients with rare diseases, left to grapple with continued questions that far too often, go unanswered.
Data availability
No data was collected in preparing this manuscript. All information may be found by referencing the cited sources listed within the manuscript.
References
Abrahamyan, L., B.M. Feldman, G. Tomlinson, M.E. Faughnan, S.R. Johnson, I.R. Diamond, and S. Gupta. 2016. Alternative designs for clinical trials in rare diseases. American Journal of Medical Genetics. Part C, Seminars in Medical Genetics 172 (4): 313–331. https://doi.org/10.1002/ajmg.c.31533.
Anjum, R.L., S. Copeland, and E. Rocca. 2020. Medical scientists and philosophers worldwide appeal to EBM to expand the notion of ‘evidence.’ BMJ Evidence-Based Medicine 25 (1): 6–8. https://doi.org/10.1136/bmjebm-2018-111092.
Aronson, J.K. 2005. Biomarkers and surrogate endpoints. British Journal of Clinical Pharmacology 59 (5): 491–494. https://doi.org/10.1111/j.1365-2125.2005.02435.x.
Bartlett, R.H., D.W. Roloff, R.G. Cornell, A.F. Andrews, P.W. Dillon, and J.B. Zwischenberger. 1985. Extracorporeal circulation in neonatal respiratory failure: a prospective randomized study. Pediatrics 76 (4): 479–487.
Beam, A.L., A.K. Manrai, and M. Ghassemi. 2020. Challenges to the reproducibility of machine learning models in health care. JAMA 323 (4): 305–306. https://doi.org/10.1001/jama.2019.20866.
Behera, M., A. Kumar, H.P. Soares, L. Sokol, and B. Djulbegovic. 2007. Evidence-based medicine for rare diseases: implications for data interpretation and clinical trial design. Cancer Control: Journal of the Moffitt Cancer Center 14 (2): 160–166. https://doi.org/10.1177/107327480701400209.
Bolignano, D., E.V. Nagler, W. Van Biesen, and C. Zoccali. 2014. Providing guidance in the dark: rare renal diseases and the challenge to improve the quality of evidence. Nephrology, Dialysis, Transplantation: Official Publication of the European Dialysis and Transplant Association—European Renal Association 29 (9): 1628–1632. https://doi.org/10.1093/ndt/gft344.
Bolignano, D., and A. Pisano. 2016. Good-quality research in rare diseases: trials and tribulations. Pediatric Nephrology (berlin, Germany) 31 (11): 2017–2023. https://doi.org/10.1007/s00467-016-3323-7.
Böllert, F.G., J.Y. Paton, T.G. Marshall, J. Calvert, A.P. Greening, and J.A. Innes. 1999. Recombinant DNase in cystic fibrosis: a protocol for targeted introduction through n-of-1 trials. Scottish cystic fibrosis group. The European Respiratory Journal 13 (1): 107–113. https://doi.org/10.1183/09031936.99.13105399.
Borgerson, K. 2009. Valuing evidence: bias and the evidence hierarchy of evidence-based medicine. Perspectives in Biology and Medicine 52 (2): 218–233. https://doi.org/10.1353/pbm.0.0086.
Brasil, S., C. Pascoal, R. Francisco, V. Dos Reis Ferreira, P.A. Videira, and A.G. Valadão. 2019. Artificial intelligence (AI) in rare diseases: is the future brighter? Genes 10 (12): 978. https://doi.org/10.3390/genes10120978.
Brown, C.H., T.R. Ten Have, B. Jo, G. Dagne, P.A. Wyman, B. Muthén, and R.D. Gibbons. 2009. Adaptive designs for randomized trials in public health. Annual Review of Public Health 30: 1–25. https://doi.org/10.1146/annurev.publhealth.031308.100223.
Brown, J., R.W. Naumann, W.E. Brady, R.L. Coleman, K.N. Moore, and D.M. Gershenson. 2018. Clinical trial methodology in rare gynecologic tumor research: strategies for success. Gynecologic Oncology 149 (3): 605–611. https://doi.org/10.1016/j.ygyno.2018.04.008.
Burns, P.B., R.J. Rohrich, and K.C. Chung. 2011. The levels of evidence and their role in evidence-based medicine. Plastic and Reconstructive Surgery 128 (1): 305–310. https://doi.org/10.1097/PRS.0b013e318219c171.
Cartwright, N. 2013. Knowing what we are talking about: Why evidence doesn’t always travel. Evidence & Policy 9 (1): 97–112. https://doi.org/10.1332/174426413X662581.
Charlton, B.G., and A. Miles. 1998. The rise and fall of EBM. QJM: Monthly Journal of the Association of Physicians 91 (5): 371–374. https://doi.org/10.1093/qjmed/91.5.371.
Chow, S.C., and M. Chang. 2008. Adaptive design methods in clinical trials—a review. Orphanet Journal of Rare Diseases 3: 11. https://doi.org/10.1186/1750-1172-3-11.
Chung, C.C.Y., Hong Kong Genome Project, A.T.W. Chu, and B.H.Y. Chung. 2022. Rare disease emerging as a global public health priority. Frontiers in Public Health 10: 1028545. https://doi.org/10.3389/fpubh.2022.1028545.
Day, S. 2010. Evidence-based medicine and rare diseases. Advances in Experimental Medicine and Biology 686: 41–53. https://doi.org/10.1007/978-90-481-9485-8_3.
Djulbegovic, B., and G.H. Guyatt. 2017. Progress in evidence-based medicine: a quarter century on. Lancet (london, England) 390 (10092): 415–423. https://doi.org/10.1016/S0140-6736(16)31592-6.
Evidence-Based Medicine Working Group. 1992. Evidence-based medicine. A new approach to teaching the practice of medicine. JAMA 268 (17): 2420–2425. https://doi.org/10.1001/jama.1992.03490170092032.
Feldman, B., E. Wang, A. Willan, and J.P. Szalai. 2001. The randomized placebo-phase design for clinical trials. Journal of Clinical Epidemiology 54 (6): 550–557. https://doi.org/10.1016/s0895-4356(00)00357-7.
Feyerabend, P. 1978. Against method. London: Verso.
Fleming, T.R., and J.H. Powers. 2012. Biomarkers and surrogate endpoints in clinical trials. Statistics in Medicine 31 (25): 2973–2984. https://doi.org/10.1002/sim.5403.
Freiesleben, T., and T. Grote. 2023. Beyond generalization: a theory of robustness in machine learning. Synthese (dordrecht) 202 (4): 109. https://doi.org/10.1007/s11229-023-04334-9.
Fuller, J. 2021. The myth and fallacy of simple extrapolation in medicine. Synthese (dordrecht) 198 (4): 2919–2939. https://doi.org/10.1007/s11229-019-02255-0.
Gallin, J.I., D.W. Alling, H.L. Malech, R. Wesley, D. Koziol, B. Marciano, E.M. Eisenstein, M.L. Turner, E.S. DeCarlo, J.M. Starling, and S.M. Holland. 2003. Itraconazole to prevent fungal infections in chronic granulomatous disease. The New England Journal of Medicine 348 (24): 2416–2422. https://doi.org/10.1056/NEJMoa021931.
Genetic and Rare Diseases Information Center. (2023). What is a Rare Disease? National Center for Advancing Translational Sciences. https://rarediseases.info.nih.gov/about
Genin, K., and T. Grote. 2021. Randomized controlled trials in medical AI: a methodological critique. Philosophy of Medicine 2: 1–15. https://doi.org/10.5195/POM.2021.27.
Goldenberg, M.J. 2006. On evidence and evidence-based medicine: lessons from the philosophy of science. Social Science & Medicine (1982) 62 (11): 2621–2632. https://doi.org/10.1016/j.socscimed.2005.11.031.
Guimaraes, P., and Y. Palesch. 2007. Power and sample size simulations for randomized play-the-winner rules. Contemporary Clinical Trials 28 (4): 487–499. https://doi.org/10.1016/j.cct.2007.01.006.
Guyatt, G., J. Cairns, D. Churchill, et al. 1992. Evidence-based medicine: a new approach to teaching the practice of medicine. JAMA 268 (17): 2420–2425. https://doi.org/10.1001/jama.1992.03490170092032.
Guyatt, G., D. Sackett, D.W. Taylor, J. Chong, R. Roberts, and S. Pugsley. 1986. Determining optimal therapy–randomized trials in individual patients. The New England Journal of Medicine 314 (14): 889–892. https://doi.org/10.1056/NEJM198604033141406.
Guyatt, G.H., A. Heyting, R. Jaeschke, J. Keller, J.D. Adachi, and R.S. Roberts. 1990. N of 1 randomized trials for investigating new drugs. Controlled Clinical Trials 11 (2): 88–100. https://doi.org/10.1016/0197-2456(90)90003-k.
Hallowell, N., M. Parker, and C. Nellåker. 2019. Big data phenotyping in rare diseases: some ethical issues. Genetics in Medicine: Official Journal of the American College of Medical Genetics 21 (2): 272–274. https://doi.org/10.1038/s41436-018-0067-8.
Hampton, T. 2006. Rare disease research gets boost. JAMA 295 (24): 2836–2838. https://doi.org/10.1001/jama.295.24.2836.
Hanson, N.R. 1958. Patterns of discovery. Cambridge: Cambridge University Press.
Heymach, J.V., B.E. Johnson, D. Prager, E. Csada, J. Roubec, M. Pesek, I. Spásová, C.P. Belani, I. Bodrogi, S. Gadgeel, S.J. Kennedy, J. Hou, and R.S. Herbst. 2007. Randomized, placebo-controlled phase II study of vandetanib plus docetaxel in previously treated non-small cell lung cancer. Journal of Clinical Oncology: Official Journal of the American Society of Clinical Oncology 25 (27): 4270–4277. https://doi.org/10.1200/JCO.2006.10.5122.
Howick, Jeremy. 2011. The philosophy of evidence-based medicine. Wiley-Blackwell: BMJ Books.
Huntington Study Group. 2001. A randomized, placebo-controlled trial of coenzyme Q10 and remacemide in Huntington’s disease. Neurology 57 (3): 397–404. https://doi.org/10.1212/wnl.57.3.397.
Ilowite, N.T., K. Prather, Y. Lokhnygina, L.E. Schanberg, M. Elder, D. Milojevic, J.W. Verbsky, S.J. Spalding, Y. Kimura, L.F. Imundo, M.G. Punaro, D.D. Sherry, S.E. Tarvin, L.S. Zemel, J.D. Birmingham, B.S. Gottlieb, M.L. Miller, K. O’Neil, N.M. Ruth, C.A. Wallace, and C.I. Sandborg. 2014. Randomized, double-blind placebo-controlled trial of the efficacy and safety of rilonacept in the treatment of systemic juvenile idiopathic arthritis. Arthritis & Rheumatology (hoboken, N.J.) 66 (9): 2570–2579. https://doi.org/10.1002/art.38699.
Johnson, S.R., B.M. Feldman, J.E. Pope, and G.A. Tomlinson. 2009. Shifting our thinking about uncommon disease trials: the case of methotrexate in scleroderma. The Journal of Rheumatology 36 (2): 323–329. https://doi.org/10.3899/jrheum.071169.
Katz, N. 2009. Enriched enrollment randomized withdrawal trial designs of analgesics: focus on methodology. The Clinical Journal of Pain 25 (9): 797–807. https://doi.org/10.1097/AJP.0b013e3181b12dec.
Korn, E.L., L.M. McShane, and B. Freidlin. 2013. Statistical challenges in the evaluation of treatments for small patient populations. Science Translational Medicine 5 (178): 178ar3. https://doi.org/10.1126/scitranslmed.3004018.
Kufel, J., K. Bargieł-Łączek, S. Kocot, M. Koźlik, W. Bartnikowska, M. Janik, Ł Czogalik, P. Dudek, M. Magiera, A. Lis, I. Paszkiewicz, Z. Nawrat, M. Cebula, and K. Gruszczyńska. 2023. What is machine learning, artificial neural networks and deep learning?-Examples of practical applications in medicine. Diagnostics (basel, Switzerland) 13 (15): 2582. https://doi.org/10.3390/diagnostics13152582.
Kuhn, T. 1970. Reflections on my critics. In Criticisms and the growth of knowledge, ed. I. Lakatos and A. Margraves, 231–278. Cambridge: Cambridge University Press.
Kuhn, T. 1996. The structure of scientific revolutions, 3rd ed. Chicago: University of Chicago Press.
Kulkarni, A.V. 2005. The challenges of evidence-based medicine: a philosophical perspective. Medicine, Health Care, and Philosophy 8 (2): 255–260. https://doi.org/10.1007/s11019-004-7345-8.
Kwak, S.G., and J.H. Kim. 2017. Central limit theorem: The cornerstone of modern statistics. Korean Journal of Anesthesiology 70 (2): 144–156. https://doi.org/10.4097/kjae.2017.70.2.144.
Larson, D.B., D.C. Magnus, M.P. Lungren, N.H. Shah, and C.P. Langlotz. 2020. Ethics of using and sharing clinical imaging data for artificial intelligence: a proposed framework. Radiology 295 (3): 675–682. https://doi.org/10.1148/radiol.2020192536.
Lester, J.N., and M. O’Reilly. 2015. Is evidence-based practice a threat to the progress of the qualitative community? arguments from the bottom of the pyramid. Qualitative Inquiry 21 (7): 628–632. https://doi.org/10.1177/1077800414563808.
Lillie, E.O., B. Patay, J. Diamant, B. Issell, E.J. Topol, and N.J. Schork. 2011. The n-of-1 clinical trial: the ultimate strategy for individualizing medicine? Personalized Medicine 8 (2): 161–173. https://doi.org/10.2217/pme.11.7.
Lovell, D.J., E.H. Giannini, A. Reiff, G.D. Cawkwell, E.D. Silverman, J.J. Nocton, L.D. Stein, A. Gedalia, N.T. Ilowite, C.A. Wallace, J. Whitmore, and B.K. Finck. 2000. Etanercept in children with polyarticular juvenile rheumatoid arthritis. Pediatric rheumatology collaborative study group. The New England Journal of Medicine 342 (11): 763–769. https://doi.org/10.1056/NEJM200003163421103.
Liu, P.Y., S. Dahlberg, and J. Crowley. 1993. Selection designs for pilot studies based on survival. Biometrics 49 (2): 391–398.
Ludbrook, J. 1995. Issues in biomedical statistics: comparing means under normal distribution theory. The Australian and New Zealand Journal of Surgery 65 (4): 267–272. https://doi.org/10.1111/j.1445-2197.1995.tb00626.x.
Malterud, K. 1995. The legitimacy of clinical knowledge: towards a medical epistemology embracing the art of medicine. Theoretical Medicine 16 (2): 183–198. https://doi.org/10.1007/BF00998544.
Martinho, A., M. Kroesen, and C. Chorus. 2021. A healthy debate: exploring the views of medical doctors on the ethics of artificial intelligence. Artificial Intelligence in Medicine 121: 102190–102190. https://doi.org/10.1016/j.artmed.2021.102190.
Masic, I., M. Miokovic, and B. Muhamedagic. 2008. Evidence based medicine—new approaches and challenges. Acta Informatica Medica: AIM: Journal of the Society for Medical Informatics of Bosnia & Herzegovina : Casopis Drustva Za Medicinsku Informatiku BiH 16 (4): 219–225. https://doi.org/10.5455/aim.2008.16.219-225.
Mok, E., G. Letellier, J.M. Cuisset, A. Denjean, F. Gottrand, C. Alberti, and R. Hankard. 2009. Lack of functional benefit with glutamine versus placebo in Duchenne muscular dystrophy: a randomized crossover trial. PLoS ONE 4 (5): e5448. https://doi.org/10.1371/journal.pone.0005448.
Montgomery, A.A., T.J. Peters, and P. Little. 2003. Design, analysis and presentation of factorial randomised controlled trials. BMC Medical Research Methodology 3: 26. https://doi.org/10.1186/1471-2288-3-26.
Murdoch, B. 2021. Privacy and artificial intelligence: challenges for protecting health information in a new era. BMC Medical Ethics 22 (1): 1–122. https://doi.org/10.1186/s12910-021-00687-3.
Nguengang Wakap, S., D.M. Lambert, A. Olry, C. Rodwell, C. Gueydan, V. Lanneau, D. Murphy, Y. Le Cam, and A. Rath. 2020. Estimating cumulative point prevalence of rare diseases: analysis of the orphanet database. European Journal of Human Genetics: EJHG 28 (2): 165–173. https://doi.org/10.1038/s41431-019-0508-0.
Oddis, C.V., A.M. Reed, R. Aggarwal, L.G. Rider, D.P. Ascherman, M.C. Levesque, R.J. Barohn, B.M. Feldman, M.O. Harris-Love, D.C. Koontz, N. Fertig, S.S. Kelley, S.L. Pryber, F.W. Miller, H.E. Rockette, RIM Study Group. 2013. Rituximab in the treatment of refractory adult and juvenile dermatomyositis and adult polymyositis: a randomized, placebo-phase trial. Arthritis and Rheumatism 65 (2): 314–324. https://doi.org/10.1002/art.37754.
Reiczigel, J., J. Singer, and Z.S. Lang. 2017. Exact inference for the risk ratio with an imperfect diagnostic test. Epidemiology and Infection 145 (1): 187–193. https://doi.org/10.1017/S0950268816002028.
Rosenberg, W., and A. Donald. 1995. Evidence based medicine: an approach to clinical problem-solving. BMJ (clinical Research Ed.) 310 (6987): 1122–1126. https://doi.org/10.1136/bmj.310.6987.1122.
Rosenberg, W.M., and D.L. Sackett. 1996. On the need for evidence-based medicine. Theriogenology 51 (3): 212–217.
Ruperto, N., D.J. Lovell, P. Quartier, E. Paz, N. Rubio-Pérez, C.A. Silva, C. Abud-Mendoza, R. Burgos-Vargas, V. Gerloni, J.A. Melo-Gomes, C. Saad-Magalhães, F. Sztajnbok, C. Goldenstein-Schainberg, M. Scheinberg, I.C. Penades, M. Fischbach, J. Orozco, P.J. Hashkes, C. Hom, L. Jung, Pediatric Rheumatology Collaborative Study Group. 2008. Abatacept in children with juvenile idiopathic arthritis: a randomised, double-blind, placebo-controlled withdrawal trial. Lancet (london, England) 372 (9636): 383–391. https://doi.org/10.1016/S0140-6736(08)60998-8.
Safdar, N.M., J.D. Banja, and C.C. Meltzer. 2020. Ethical considerations in artificial intelligence. European Journal of Radiology 122: 108768–108768. https://doi.org/10.1016/j.ejrad.2019.108768.
Schaefer, J., M. Lehne, J. Schepers, F. Prasser, and S. Thun. 2020. The use of machine learning in rare diseases: a scoping review. Orphanet Journal of Rare Diseases 15 (1): 1–145. https://doi.org/10.1186/s13023-020-01424-6.
Scott, I., D. Cook, and E. Coiera. 2021. Evidence-based medicine and machine learning: a partnership with a common purpose. BMJ Evidence-Based Medicine 26 (6): 290–294. https://doi.org/10.1136/bmjebm-2020-111379.
Sernadela, P., L. González-Castro, C. Carta, E. van der Horst, P. Lopes, R. Kaliyaperumal, M. Thompson, R. Thompson, N. Queralt-Rosinach, E. Lopez, L. Wood, A. Robertson, C. Lamanna, M. Gilling, M. Orth, R. Merino-Martinez, M. Posada, D. Taruscio, H. Lochmüller, P. Robinson, and J.L. Oliveira. 2017. Linked registries: connecting rare diseases patient registries through a semantic web layer. BioMed Research International 2017: 8327980. https://doi.org/10.1155/2017/8327980.
Sibbald, B., and C. Roberts. 1998. Understanding Controlled Trials. Crossover Trials. BMJ (clinical Research Ed.) 316 (7146): 1719. https://doi.org/10.1136/bmj.316.7146.1719.
Stallard, N., and S. Todd. 2003. Sequential designs for phase III clinical trials incorporating treatment selection. Statistics in Medicine 22 (5): 689–703. https://doi.org/10.1002/sim.1362.
Sur, R.L., and P. Dahm. 2011. History of evidence-based medicine. Indian Journal of Urology: IJU: Journal of the Urological Society of India 27 (4): 487–489. https://doi.org/10.4103/0970-1591.91438.
Tan, S.B., K.B. Dear, P. Bruzzi, and D. Machin. 2003. Strategy for randomised clinical trials in rare cancers. BMJ (clinical Research Ed.) 327 (7405): 47–49. https://doi.org/10.1136/bmj.327.7405.47.
Tonelli, M.R. 1998. The philosophical limits of evidence-based medicine. Academic Medicine: Journal of the Association of American Medical Colleges 73 (12): 1234–1240. https://doi.org/10.1097/00001888-199812000-00011.
Tudur Smith, C., P.R. Williamson, and M.W. Beresford. 2014. Methodology of clinical trials for rare diseases. Best Practice & Research. Clinical Rheumatology 28 (2): 247–262. https://doi.org/10.1016/j.berh.2014.03.004.
van der Lee, J.H., J. Wesseling, M.W. Tanck, and M. Offringa. 2008. Efficient ways exist to obtain the optimal sample size in clinical trials in rare diseases. Journal of Clinical Epidemiology 61 (4): 324–330. https://doi.org/10.1016/j.jclinepi.2007.07.008.
Visibelli, A., B. Roncaglia, O. Spiga, and A. Santucci. 2023. The impact of artificial intelligence in the odyssey of rare diseases. Biomedicines 11 (3): 887. https://doi.org/10.3390/biomedicines11030887.
Wadden, J.J. 2021. Defining the undefinable: the black box problem in healthcare artificial intelligence. Journal of Medical Ethics 48 (10): 764–768. https://doi.org/10.1136/medethics-2021-107529.
Ware, J.H. 1989. Investigating therapies of potentially great benefit: ECMO. Statistical Science 4 (4): 298–306.
Whelan, D.B., K. Dainty, and J. Chahal. 2012. Efficient designs: factorial randomized trials. The Journal of Bone and Joint Surgery. American 94 (Suppl 1): 34–38. https://doi.org/10.2106/JBJS.L.00243.
Worrall, J. 2002. What evidence in evidence-based medicine? Philosophy of Science 69 (S3): S316–S330. https://doi.org/10.1086/341855.
Youssef, A., M. Pencina, A. Thakur, T. Zhu, D. Clifton, and N.H. Shah. 2023. External validation of AI models in health should be replaced with recurring local validation. Nature Medicine 29 (11): 2686–2687. https://doi.org/10.1038/s41591-023-02540-z.
Zhang, J., and Z.-M. Zhang. 2023. Ethics and governance of trustworthy medical artificial intelligence. BMC Medical Informatics and Decision Making 23 (1): 7–7. https://doi.org/10.1186/s12911-023-02103-9.
Acknowledgements
The author would like to thank Alexandra Calzavara and Dr. Prateek Lala for their input on this manuscript’s topic and suggestions during the research and writing process. The author would also like to acknowledge the use of the design software, Canva, for development of this manuscript’s figures.
Funding
No funding was received to assist with the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Olivia Rennie conceptualized the topic of this manuscript, and completed all research involved in reviewing published literature in this area. Olivia Rennie wrote the manuscript and manually created all tables and figures.
Corresponding author
Ethics declarations
Competing interests
The author has no competing interests to declare that are relevant to the content of this article.
Ethical approval
This scientific contribution provides a review of published literature and expert input/experience in the field. No ethical approval was required for conducting the necessary research in preparing this manuscript.
Informed consent
No human or non-human subjects were involved in research for this manuscript. No consent was required in preparing this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rennie, O. Navigating the uncommon: challenges in applying evidence-based medicine to rare diseases and the prospects of artificial intelligence solutions. Med Health Care and Philos 27, 269–284 (2024). https://doi.org/10.1007/s11019-024-10206-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11019-024-10206-x