Abstract
Metachromatic leukodystrophy (MLD) is a rare hereditary neurodegenerative disease caused by deficiency of the lysosomal enzyme arylsulfatase A (ARSA). This study described the clinical and molecular characteristics of 24 Chinese children with MLD and investigated functional characterization of five novel ARSA variants. A retrospective analysis was performed in 24 patients diagnosed with MLD at Guangzhou Women and Children’s Medical Center in South China. Five novel mutations were further characterized by transient expression studies. We recruited 17 late-infantile, 3 early-juvenile, 4 late-juvenile MLD patients. In late-infantile patients, motor developmental delay and gait disturbance were the most frequent symptoms at onset. In juvenile patients, cognitive regression and gait disturbance were the most frequent chief complaints. Overall, 25 different ARSA mutations were identified with 5 novel mutations.The most frequent alleles were p.W320* and p.G449Rfs. The mutation p.W320*, p.Q155=, p.P91L, p.G156D, p.H208Mfs*46 and p.G449Rfs may link to late-infantile type. The novel missense mutations were predicted damaging in silico. The bioinformatic structural analysis of the novel missense mutations showed that these amino acid replacements would cause severe impairment of protein structure and function. In vitro functional analysis of the six mutants, showing a low ARSA enzyme activity, clearly demonstrated their pathogenic nature. The mutation p.D413N linked to R alleles. In western blotting analysis of the ARSA protein, the examined mutations retained reduced amounts of ARSA protein compared to the wild type. This study expands the spectrum of genotype of MLD. It helps to the future studies of genotype-phenotype correlations to estimate prognosis and develop new therapeutic approach.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Metachromatic leukodystrophy (MLD, OMIM 250,100) is a rare inherited neurodegenerative disease. The estimated incidence of MLD is ranged from 1/40,000 to 1/160, 000 worldwide, however, higher incidence has been observed in some isolated populations (van Rappard et al. 2015; Shaimardanova et al. 2020). It is mainly caused by the deficient activity of arylsulfatase A (ARSA) that is encoded by arylsulfatase A gene (ARSA, OMIM 607,574) (van Rappard et al. 2015; Shaimardanova et al. 2020; Amr et al. 2021). ARSA is necessary for the hydrolysis of sulfatides and its deficiency leads to superabundant accumulation of sulfatides in lysosomal storage deposits of microglia, oligodendrocytes, Schwann cells, some neurons and macrophages in the central and peripheral nervous systems. As a result, demyelination occurs gradually, leading to progressive motor and cognitive deterioration as clinical characteristics. It is hypothesized that the differences in the residual enzyme activity of ARSA cause a great diversity of onset age and the severity of the disease. Generally, the lower the residual ARSA enzyme activity is, the earlier and the more severe the disease manifests. But the definite relationship has not been fully illuminated (Liaw et al. 2015; Kehrer et al. 2021). Based on the age of the first symptom onset, MLD is clinically divided into three subtypes: late-infantile type (6∼24 months), juvenile type (∼ 16 years, further classified into early-juvenile type if onset age younger than 6 years and late-juvenile type if onset age younger than 16 years) and adult type (> 16 years), in which the late-infantile type accounts for 50∼60% and presents with rapid progression and the most severity (Kehrer et al. 2021; Groeschel et al. 2016; Fumagalli et al. 2021).
The ARSA gene contains 8 exons spread about 3.2 kb on chromosome 22q13.33 and encodes a polypeptide of 509 amino acids. To date, more than 200 different ARSA gene mutations have been detected (Human Gene Mutation Database at the Institute of Medical Genetics in Cardiff: ARSA Gene: http://www.hgmd.cf.ac.uk). Among these mutations, approximately three-fourths of them are missense/nonsense mutations; other mutations include splicing, small deletions, small insertions, small indels, gross deletions and gross insertions of the ARSA gene. The ARSA mutations have been divided into two functional types: 0 alleles, resulting in extremely low enzymatic activity, and R alleles, resulting in detectable residual enzymatic activity (Fumagalli et al. 2021).
In this study, we described the clinical and molecular characteristics of 24 Chinese patients with MLD including five novel mutations and investigated functional characterization of six ARSA variants.
Materials and methods
Participants
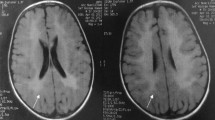
This is a longitudinal and retrospective study, which was conducted at Guangzhou Women and Children’s Medical Center (the National Children’s Medical Center for South Central Region). The diagnostic criteria of MLD in our cases were as follows: (1) suspected manifestations such as gait disturbance, developmental delays, motor weakness, muscle rigidity in the lower extremities, seizures, spastic paralysis, gradual loss of speech, intellectual disability, cognitive symptoms, impaired swallowing and so on; (2) diffuse, high-intensity signal in periventricular and subcortical white matter in brain magnetic resonance imaging; (3) low ARSA activity in leucocytes and/or (4) pathogenic mutations in ARSA gene (Beerepoot et al. 2020; Karaağaoğlu et al. 2020). Pre-symptomatic diagnosis was made with a definite family history and verified gene mutations (Chen et al. 2018). Finally, 24 children with MLD, who visited our hospital between July 2010 and April 2021, were enrolled in this study. They came from 21 unrelated families, for P1 and P2, P19 and P20, and P23 and P24 were siblings, respectively. None of them were descendants of consanguineous marriages.
Demographical data, clinical presentations, family history and significant examinations were collected from medical records. Informed consent was obtained from all patients’ parents. The study was approved by the Institutional Review Board of Guangzhou Women and Children’s Medical Center.
Measurement of ARSA activity
Leukocytes harvested from peripheral blood were suspended in filtered water and subjected to three rounds of sonication at 4℃. After a 17 h incubation period at 0℃, the reaction was stopped with NaOH and the absorption was measured at 515 nm. The activity of ARSA was quantified as nmol/mg.17 h in leukocytes (normal range 218 ~ 550 nmol/mg.17 h), by applying a synthetic substrate (p-nitrocatechol sulfate) (Strobel et al. 2020).
Molecular analysis of ARSA gene
Genomic DNA was extracted from peripheral blood leukocytes of all patients using a standard procedure. The primer sequences are listed in Table S1. All the exon sequences and intron-exon boundaries of the ARSA gene were amplified by polymerase chain reaction using corresponding primers, then the products were sequenced using an ABI 3730xl DNA Analyzer (Life Technologies Co., CA, USA). The Chromas software (Technelysium Pty Ltd, South Brisbane, Australia) was applied to read the sequencing chromatograms and align them with the reference sequence of ARSA gene (NM_000487.5). The novel mutations in the study were determined by comparing the Human Gene Mutation Database (HGMD) and the National Center for Biotechnology Information (NCBI) database. Genetic variants were searched in the Single Nucleotide Polymorphism Database (dbSNP) and the 1000 Genomes Project. Intronic variants were analyzed with GenSCAN (http://genes.mit.edu/GENSCAN.html) to determine whether the consensus sequence of any splice site was altered. Novel mutations were verified by direct sequencing of the PCR products in 100 unrelated healthy controls and comparing the Exome Aggregation Consortium (ExAC) database along with the NHLBI exome variant database.
Protein function prediction of novel mutations
To predict the effect of amino acid substitutions, we performed in silico analysis using the SIFT/PROVEAN (http://sift.jcvi.org) and Polyphen-2 (http://genetics.bwh.harvard.edu/pph2) web software. SIFT Score ranges from 0 to 1. The amino acid substitution is predicted as damaging if the score is ≤ 0.05, and tolerated if the score is > 0.05 (J. Craig Venter Institute, USA). The variant is predicted to be deleterious if the PROVEAN score is ≤ − 2.5), and neutral if the score is > − 2.5 (J. Craig Venter Institute, USA). Polyphen-2 prediction outcome can be one of “probably damaging”, “possibly damaging”, or “benign”.
Protein visualization and structural analysis
The effect of the missense mutations on the overall structure of the protein and on its activity was also investigated using the crystal structure of human ARSA protein (PDBID 1N2L). Virtual models of ARSA mutations and bioinformatics analyses were performed with the PyMOL (TM) Molecular Graphics System (Version 4.6.0).
Plasmid construct
The wild-type ARSA cDNA (GenBank: NC_000022.11) was linked to the enhanced green fluorescent protein (EGFP) reporter gene and cloned into the pcDNA3.1 plasmid (Invitrogen, Carlsbad, California). Six missense mutations of ARSA gene, including c.101G > A (p.G34D), c.221 C > G (p.S74C), c.239 C > T (p.T80I), c.272 C > T (p.P91L), c.581 C > G (p.P194R), c.1237G > A (p.D413N) were introduced in the wild-type cDNA fused to EGFP by a Quikchange Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA).
Cell culture and transient transfection
293FT cells were grown in Dulbecco’s modified Eagle’s medium (Gibco BRL, Grand Island, NJ, USA) supplemented with 10% fetal calf serum (Gibco BRL, Grand Island, NJ, USA), 100 U/mL penicillin and 100 U/mL streptomycin (Hyclone, Logan, UT, USA) at 37 °C in a humidified atmosphere enriched with 5% CO2. 293FT cells were transfected by pcDNA3.1-EGFP vector with wild-type ARSA cDNA and six mutants using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. After 72 h transfection, cells were harvested for ARSA enzyme assays and Western blotting. All transfections were performed in duplicate in three individual experiments.
ARSA activity assay
The harvested 293FT cells were sonicated in ddH2O and then protein concentrations were determined using Micro BCA Protein Assay Kit (Pierce, Rockford, IL, USA). The enzyme activity assay was performed.
Western blotting
The transfected cells were lysed by RIPA Lysis Buffer (Beyotime, Beijing, China) containing 1% PMSF (Beyotime, Beijing, China). Protein (30 µg) was loaded for electrophoresis on 10% SDS-PAGE gels and transferred to PVDF membranes. The membranes were then blocked with 5% non-fat milk in TBS, containing 0.1% Tween 20 for 1.5 h at room temperature, and then incubated with the respective primary antibody ARSA and GAPDH (Multi Sciences, Hangzhou, China) at 4 °C overnight. Finally, membranes were incubated with the HRP secondary antibody (Multi Sciences, Hangzhou, China) for 2 h at room temperature. The signals were detected using the ECL western blotting detection reagent (Pierce, Rockford, IL, USA).
Statistical analysis
The difference between the wild-type enzyme activities and the individual mutant enzyme activities was analyzed by GraphPad Prism 5 (GraphPad Software Inc., CA, USA) using a Student’s t-test. P < 0.05 was considered as statistically significant.
Results
General demographic characteristics and clinical manifestations
Twenty-four pediatric patients (11 boys and 13 girls) with diagnosed MLD from 21 unrelated families were finally enrolled in this study, eight of whom (33%) had positive family history. All of them were born after normal pregnancy and delivery, however, symptoms gradually occurred with time going. The median age of disease onset was 16.5 (15.0, 31.0) months and the age of diagnosis was 29.0 (22.8, 56.0) months, which was much later than the disease onset time (P = 0.001). Seventeen of them (71%) were categorized as late infantile type, and the remaining (29%) were of juvenile type (3 early-juvenile type and 4 late-juvenile type).
At disease onset, motor developmental delay and gait disturbance were the most frequent symptoms in the patients with late infantile type, followed by motor regression and muscle weakness. In the juvenile patients cognitive regression and gait disturbance were the most frequent chief complaints, then seizure. Half of all the participants (50%) were found to be hypertonic when they visited our hospital for the first time, and nearly one quarter were characterized with knee hyperreflexia (n = 7, 29%) or positive Babinski sign (n = 7, 29%), respectively.
Biochemical findings
The ARSA enzyme activity analyzed in peripheral blood leukocytes from the 24 patients ranged from 9.4 to 99.5 nmol/17 h/mg. The enzyme activity was reduced in all the patients. There was no significant difference in the ARSA enzyme activity between late-infantile and juvenile type.
Genetic findings
Sequencing analysis of ARSA gene identified 25 different mutations in 24 patients belong to 21 families (Table 1). Of the 25 mutations, 20 (80%) have been previously reported and 5 (20%) have never been described. Five novel mutations were detected: four missense mutations (p.G34D, p.T80I, p.P91L, p.D413N)) and one small deletion mutation (p.L422Dfs*3).
A total of 25 different mutations included 16 (64%) missense, 2 (8%) nonsense, 2 (8%) splicing, 3 (12%) small deletion, 1 (4%) small insertion and 1 (4%) synonymous mutation. Thus, the most prevalent ARSA variants were missense mutations, including 4 novel amino acid substitutions. The three most prevalent mutations were p.W320* (14.6%,7/48), p.G449Rfs (10.4%,5/48), and p.P91L (8.3%, 4/48). When combined, these three ARSA mutations represented 33.3% of the total ARSA alleles. Generally, the nonsense mutations leaded to the appearance of a premature stop codon, disturbed the synthesis of enzyme protein and influenced the function of the enzyme. Most of the nonsense mutations occurred in late-infantile type of MLD and showed more severe phenotype than the missense mutations. Meanwhile, in late-juvenile type of MLD, all the ARSA mutations were classified as missense mutations. While mutations were found in all exons except exon 6 and 7, they were concentrated in exon 2 (27.1%), exon5 (22.9%) and exon 8 (20.1%). None of these mutations were detected in 100 alleles of unrelated healthy controls.
In late-infantile type of MLD, we identified 19 mutations of the ARSA gene in the 17 patients: 10 (52.6%) missense, 3 (15.8%) small deletion, 2 (10.5%) nonsense, 2 (10.5%) splicing, 1 (5.3) small insertion and 1 (5.3%) synonymous mutations. In the early-juvenile type of MLD, the ARSA mutations were classified as follows among the 3 patients: 4 (80%) missense and 1 (25%) small insertion mutations. In late-juvenile type of MLD, we identified 6 mutations of ARSA gene in the 4 patients and all the mutations were missense mutations. Homozygosity for the mutations p.P91L, p.G156D, c.621_622delC, c.1344dupC were associated with late-infantile type of MLD.
Protein function prediction of novel mutations
The novel missense mutations (p.G34D, p.T80I, p.P91L and p.D413N) in the ARSA gene were predicted damaging by the SIFT/PROVEAN and Polyphen-2 web software.
Bioinformatic structural analysis
To investigate the possible consequences of the missense mutations at protein level, we visualized the crystal structure of human ARSA (Fig. 1).
-
p.G34D: The structural analysis of the mutation predicted that the replacement of a hydrophobic neutral glycine (G) with the acidic aspartic acid (D), occurring in the proximity of the residues D29 and D30 known to be involved in assisting the substrate binding as a key catalytic residue in the active site.
-
p.S74C: The S74 residue is adjacent to the active site residues in coordination of the calcium ion in the active sites of ARSA. Experimental data have shown that addition of divalent metal ions stimulates the activity of ARSA. Substitution from the serine (S) to the cystine (C) at this position may cause loss of full enzyme activity.
-
p.T80I: The T80 residue is adjacent to the active site residues and make structural interactions with the catalytic core. Substitution from the hydrophilic threonine(T) to the hydrophobic isoleucine(I) may influence the function of catalysis.
-
p.P91L: The P91 made the structure interaction with the residue locating in the catalytic core, This change of the proline (P) to the leucine (L) likely resulted in protein misfolding or catalytic inactivation.
-
p.P194R: The P194 located at the periphery of the molecule which may to be one of N-glycosylation sites. Substitution from the hydrophobic proline (P) to the basic arginine (R) may influence the activity of enzyme.
-
p.D413N: The D413 was involved in regulation of dimer-octamer association. This change of the acidic aspartic acid (D) to the neutral asparagine (N) may influence the protein folding and stability.
Characterization of the novel sequence variants
To determine the impact of mutants on protein function, we characterized six missense mutations: p.G34D, p.S74C, p.T80I, p.P91L, p.P194R and p.D413N. Although the mutations p.S74C and p.P194R were no longer novel since they were reported during the preparation of the present study (Wu et al. 2021), was also included in the analysis. Six mutant vectors were constructed by site-direct mutagenesis in 293FT cells and the ARSA activity in extracts of 293FT cells was detected. Using the same conditions, the wild-type, untransfected and 6 mutant proteins were analyzed for comparative purposes. The ARSA activity in extracts of 293FT cells transfected with the wild-type ARSA was 1950.14 ± 56.12 nmol/17 h/mg and that of untransfected cells was 135.74 ± 15.36 nmol/17 h/mg. The six mutants had very low ARSA activity and none of them showed significant activity above the background (Table 2; Fig. 2).
ARSA protein was analyzed by western blotting (Fig. 3). As shown in Fig. 2, the six mutations retained reduced amounts of ARSA protein compared to the wild type, suggesting a decrease in the total amount of ARSA protein in these mutants.
Discussion
MLD is named after the histopathological discovery of metachromatic particles (sulfatides) in the central and peripheral nervous systems and clinical symptoms occur gradually because of progressive accumulation of undegraded sulfatides in such organs (Gieselmann and Krägeloh-Mann 2010). Consequently, disease onset of MLD tend to be unclear and is easily overlooked by caregivers (Fumagalli et al. 2021; Wu et al. 2021; Harrington et al. 2019). As demonstrated in this study and other reports in the literature (Fumagalli et al. 2021; Beerepoot et al. 2020), the time of disease diagnosis is much later than that of disease onset, which, to some degree, reflects the lacking recognition of caregivers and inadequate knowledge on this disease of clinicians. Motor dysfunction is found to be the most frequent first symptom in this population, which is greatly in agreement with previous findings (Kehrer et al. 2021; Karaağaoğlu et al. 2020; Chen et al. 2018), strengthens our understanding of this disease and raises caregivers’ and physician’ vigilance of motion abnormality. Cognitive impairment is only observed in the participants with juvenile MLD, which is scarcely reported in patients with late-infantile type. Maybe the late-infantile patients are characterized with great development in gross motor and it is not easy for caregivers to recognize cognitive impairment.
In this study we reported the clinical and molecular characterization of a group of Chinese patients with MLD. The group included 17 patients with late-infantile type, 3 patients with early-juvenile type and 4 patients with late-juvenile type. In such a pediatric population, motor dysfunction was the predominant first symptom and positive neurological signs were found in large part of patients at their first visit. Compared with the patients with juvenile MLD, much more late infantile patients had hypertonia. The decrease of ARSA enzyme activity is the characteristic of MLD (Strobel et al. 2020; Doherty et al. 2019; Fluharty et al. 1983). There was no significant difference in assayed enzyme activity between late-infantile and juvenile type. In the 17 patients with late-infantile type, missense mutations were most frequently observed (10/19, 52.6%). However, late-infantile type of MLD was strongly associated with mutations including small deletion, nonsense, and splicing mutations. In this study, 64% of patients have private mutations and novel mutations were detected of a frequency of 20%.
The 25 mutations detected in the 24 patients reflected that most of the mutations were private mutations. However, the mutation p.W320* could be a hotspot mutation as seven patients showed this mutation. It results in a premature termination codon, predicted to cause a truncated or absent ARSA protein. It was first reported by Chen et al. in a Chinese patient of late-infantile type with homozygous form (Chen et al. 2018). It was also reported by Liaw et al. in two Chinese patients of late-infantile type with homozygous or compound heterozygous form (Liaw et al. 2015). The six patients with the mutation p.W320* in our study presented with late-infantile type.The mutation p.W320* may be link to late infantile type of MLD. The nonsense mutation p.W320* was found to be the most common one in our population, which was different from that of other populations. For example, c.465 + 1G > A was confirmed as the most frequent variant in a German population and c.931G > A was the most frequent one in a Turkish population (Liaw et al. 2015; Karaağaoğlu et al. 2020). Even in the mainland of China, the mutations c.459 + 1G > A and p.Pro426Leu were discovered to be the most common one in a medical center in North China (Chen et al. 2018).
The correlation between the phenotype and the genotype of MLD is heterogeneous (Santhanakumaran et al. 2022). In this study, we identified 17 cases of late-infantile type of MLD including 10 (52.6%) missense, 3 (15.8%) small deletion, 2 (10.5%) nonsense, 2 (10.5%) splicing, 1 (5.3) small insertion and 1 (5.3%) synonymous mutations. Among these mutations, we found an unexpected synonymous mutation p.Q155=. This synonymous mutation was first descibed by Chen et al. in two Chinese patients of late infantile type with compound heterozygous form (Chen et al. 2018). The 465 locus was the last base in the second exon of ARSA (Chen et al. 2018). The mutation was located between the second exon and second intron. Based on the RNA analysis, it was found that after the change of the 465 locus, RNA indeed produced a splicing mutation, which presented as a splicing jump of the 371–465 loci in the second exon (involved 95 bases) (Chen et al. 2018). Homozygosity for the mutations p.P91L, p.G156D, p.H208Mfs*46 and p.G449Rfs showed in the patients with the late-infantile type. It implied that the mutations p.P91L, p.G156D, p.H208Mfs*46 and p.G449Rfs may be link to the late- infantile type of MLD.
Three cases of early-juvenile type of MLD were identified 4 (80%) missense and 1 (20%) small insertion mutations. Four cases of late-juvenile were identified 6 (100%) missense mutations. The mutation p.R86W was comsidered to be an R mutation with detectable residual ARSA enzyme activity. Cesani et al. expressed mutated ARSA alleles in HeLa cells using lentiviral vectors including the mutation p.R86W (Cesani et al. 2016). The mutation p.R86W showed a residual activity and proved to associated with R alleles.
Our research further expanded the spectrum with 5 novel variants: four missense mutations (p.G34D, p.T80I, p.P91L, p.D413N) and one small deletion mutation (p.L422Dfs*3). These four novel missense mutations were predicted to be damaging as per the in-silico analysis using the web software. The crystal structure of ARSA and the substrate also indicated that the novel mutation may influence the activity of enzyme (Lukatela et al. 1998; von Bülow et al. 2001).
To verify the pathological impact of six ARSA variants on the ARSA activity, we carried out in vitro expression experiments among the mutations p.G34D, p.S74C, p.T80I, p.P91L, p.P194R and p.D413N. Meanwhile, the mutation p.S74C and p.P194R, although no longer novel since it was reported during the preparation of our study was also included in the analysis (Wu et al. 2021). Transfection of 293FT cells with wild-type ARSA cDNA resulted in an 14-fold increase of the enzyme activity compared to the activity of untransfected 293FT cells. The in vitro residual activity of the six mutants was significantly lower than the activity of the normal protein. Among the mutations, the mutation p.D413N showed a residual activity of 42.8% of the wildtype enzyme on average. In this study the mutation p.D413N was found in two siblings with late-juvenile type. Thus we suggested that the mutation p.D413N linked to R alleles. This mutation displayed about 50% of WT cells ARSA activity, the pathogenic character of this mutation should be proven in another way such as urinary sulfatide level measurement. ARSA protein was analyzed by western blotting. The six mutations retained reduced amounts of ARSA protein compared to the wild type, suggesting a decrease in the total amount of ARSA protein in these mutants.
The treatment options of MLD patients are limited, and the disease progression is always rapid and alarming. Based on the pathogenesis of MLD, new therapeutic approaches like hematopoietic stem cell transplantation, substrate reduction therapy, gene therapy and enzyme replacement therapy have been used with mixed results (Batzios and Zafeiriou 2012; Patil and Maegawa 2013; Kurtzberg 2022; Cabanillas et al. 2022; Fumagalli et al. 2022; Babcock et al. 2021). The phenomenon was observed in this study that early onset subtypes indicated worse outcomes after the treatment of hematopoietic stem cell transplantation, but no statistical conclusion was achieved maybe for relatively small sample size. On the whole, during the follow-up period, patients treated by transplantation did not show more promising outcomes than those treated by symptomatic therapies, which corroborates part of previous studies (Chen et al. 2016; Martin et al. 2013; Cable et al. 2011; Smith et al. 2010). Existed motor dysfunction at the time of transplantation and transplantation-related toxic effects may be the plausible reasons for distressing outcomes after transplantation treatment (Beschle et al. 2020).
In conclusion, this study enriches our understandings of MLD. In accordance with previous findings, motor dysfunction is proved as the predominant first symptom. The most frequent mutation of ARSA gene in this population is p.Trp320* and p.G449Rfs. The pathogenic spectrum is expanded by adding 5 novel variants. The mutation p.D413N was functional characterized as R alleles.
Data availability
The original contributions presented in the study are included in the article; further inquiries can be directed to the corresponding author.
References
Amr K, Fateen E, Mansour L et al (2021) Clinical, biochemical, and molecular characterization of metachromatic Leukodystrophy among Egyptian Pediatric patients: expansion of the ARSA Mutational Spectrum. J Mol Neurosci 71(5):1112–1130. https://doi.org/10.1007/s12031-020-01734-1
Babcock MC, Mikulka CR, Wang B et al (2021) Substrate reduction therapy for Krabbe disease and metachromatic leukodystrophy using a novel ceramide galactosyltransferase inhibitor. Sci Rep 11(1):14486. https://doi.org/10.1038/s41598-021-93601-1
Batzios SP, Zafeiriou DI (2012) Developing treatment options for metachromatic leukodystrophy. Mol Genet Metab 105(1):56–63. 10.1016/ j.ymgme.2011.10.002
Beerepoot S, van Dooren SJM, Salomons GS et al (2020) Metachromatic leukodystrophy genotypes in the Netherlands reveal novel pathogenic ARSA variants in non-caucasian patients. Neurogenetics 21(4):289–299. 10.1007/ s10048-020-00621-6
Beschle J, Döring M, Kehrer C et al (2020) Early clinical course after hematopoietic stem cell transplantation in children with juvenile metachromatic leukodystrophy. Mol Cell Pediatr 7(1):12. https://doi.org/10.1186/s40348-020-00103-7
Cabanillas Stanchi KM, Böhringer J, Strölin M et al (2022) Hematopoietic stem cell transplantation with mesenchymal stromal cells in children with Metachromatic Leukodystrophy. Stem Cells Dev 31(7–8):163–175. https://doi.org/10.1089/scd.2021.0352
Cable C, Finkel RS, Lehky TJ et al (2011) Unrelated umbilical cord blood transplant for juvenile metachromatic leukodystrophy: a 5-year follow-up in three affected siblings. Mol Genet Metab 102(2):207–209. https://doi.org/10.1016/j.ymgme.2010.10.002
Cesani M, Lorioli L, Grossi S et al (2016) Mutation update of ARSA and PSAP genes causing metachromatic leukodystrophy. Hum Mutat 37(1):16–27. https://doi.org/10.1002/humu.22919
Chen X, Gill D, Shaw P et al (2016) Outcome of early juvenile onset metachromatic leukodystrophy after unrelated cord blood transplantation: a Case Series and Review of the literature. J Child Neurol 31(3):338–344. https://doi.org/10.1177/0883073815595078
Chen L, Yan H, Cao B et al (2018) Identification of Novel ARSA mutations in Chinese patients with Metachromatic Leukodystrophy. Int J Genomics 2018:2361068. https://doi.org/10.1155/2018/2361068
Doherty K, Frazier SB, Clark M et al (2019) A closer look at ARSA activity in a patient with metachromatic leukodystrophy. Mol Genet Metab Rep 19:100460. https://doi.org/10.1016/j.ymgmr.2019.100460
Fluharty AL, Meek WE, Kihara H et al (1983) Pseudo arylsulfatase A deficiency: evidence for a structurally altered enzyme. Biochem Biophys Res Commun 112(1):191–197. https://doi.org/10.1016/0006-291x(83)91815-6
Fumagalli F, Zambon AA, Rancoita PMV et al (2021) Metachromatic leukodystrophy: a single-center longitudinal study of 45 patients. J Inherit Metab Dis 44(5):1151–1164. https://doi.org/10.1002/jimd.12388
Fumagalli F, Calbi V, Natali Sora MG et al (2022) Lentiviral haematopoietic stem-cell gene therapy for early-onset metachromatic leukodystrophy: long-term results from a non-randomised, open-label, phase 1/2 trial and expanded access. Lancet 399(10322):372–383. https://doi.org/10.1016/S0140-6736(21)02017-1
Gieselmann V, Krägeloh-Mann I (2010) Metachromatic leukodystrophy–an update. Neuropediatrics 2010 41(1):1–6. https://doi.org/10.1055/s-0030-1253412
Groeschel S, Kühl JS, Bley AE et al (2016) Long-term outcome of allogeneic hematopoietic stem cell transplantation in patients with juvenile metachromatic leukodystrophy compared with nontransplanted control patients. JAMA Neurol 73(9):1133–40. https://doi.org/10.1001/jamaneurol.2016.2067
Harrington M, Whalley D, Twiss J et al (2019) Insights into the natural history of metachromatic leukodystrophy from interviews with caregivers. Orphanet J Rare Dis 14(1):89. https://doi.org/10.1186/s13023-019-1060-2
Karaağaoğlu E, Topçu M, Akarsu NA et al (2020) Comprehensive clinical, biochemical, radiological and genetic analysis of 28 Turkish cases with suspected metachromatic leukodystrophy and their relatives. Mol Genet Metab Rep 25:100688. https://doi.org/10.1016/j.ymgmr.2020.100688
Kehrer C, Elgün S, Raabe C et al (2021) Association of age at onset and first symptoms with disease progression in patients with metachromatic leukodystrophy. Neurology 96(2):e255–e266. https://doi.org/10.1212/WNL.0000000000011047
Kurtzberg J (2022) Gene therapy offers new hope for children with metachromatic leukodystrophy. Lancet 399(10322):338–339. https://doi.org/10.1016/S0140-6736(22)00057-5
Liaw HR, Lee HF, Chi CS et al (2015) Late infantile metachromatic leukodystrophy: clinical manifestations of five Taiwanese patients and genetic features in Asia. Orphanet J Rare Dis 10:144. https://doi.org/10.1186/s13023-015-0363-1
Lukatela G, Krauss N, Theis K et al (1998) Crystal structure of human arylsulfatase A: the aldehyde function and the metal ion at the active site suggest a novel mechanism for sulfate ester hydrolysis. Biochemistry 37(11):3654–3664. https://doi.org/10.1021/bi9714924
Martin HR, Poe MD, Provenzale JM et al (2013) Neurodevelopmental outcomes of umbilical cord blood transplantation in metachromatic leukodystrophy. Biol Blood Marrow Transpl 19(4):616–624. https://doi.org/10.1016/j.bbmt.2013.01.010
Patil SA, Maegawa GH (2013) Developing therapeutic approaches for metachromatic leukodystrophy. Drug Des Devel Ther 7:729–745. https://doi.org/10.2147/DDDT.S15467
Santhanakumaran V, Groeschel S, Harzer K et al (2022) Predicting clinical phenotypes of metachromatic leukodystrophy based on the arylsulfatase A activity and the ARSA genotype?-Chances and challenges. Mol Genet Metab 137(3):273–282. https://doi.org/10.1016/j.ymgme.2022.09.009
Shaimardanova AA, Chulpanova DS, Solovyeva VV et al (2020) Metachromatic leukodystrophy: diagnosis, modeling, and treatment approaches. Front Med (Lausanne) 7:576221. https://doi.org/10.3389/fmed.2020.576221
Smith NJ, Marcus RE, Sahakian BJ et al (2010) Haematopoietic stem cell transplantation does not retard disease progression in the psycho-cognitive variant of late-onset metachromatic leukodystrophy. J Inherit Metab Dis 33(Suppl 3):S471–475. https://doi.org/10.1007/s10545-010-9240-1
Strobel S, Hesse N, Santhanakumaran V et al (2020) Optimization of enzyme essays to enhance reliability of activity measurements in leukocyte lysates for the diagnosis of Metachromatic Leukodystrophy and Gangliosidose. Cells 9(12):2553. https://doi.org/10.3390/cells9122553
van Rappard DF, Boelens JJ, Wolf NI (2015) Metachromatic leukodystrophy: Disease spectrum and approaches for treatment. Best Pract Res Clin Endocrinol Metab 29(2):261–273. https://doi.org/10.1016/j.beem.2014.10.001
von Bülow R, Schmidt B, Dierks T et al (2001) Crystal structure of an enzyme-substrate complex provides insight into the interaction between human arylsulfatase A and its substrates during catalysis. J Mol Biol 305(2):269–277. https://doi.org/10.1006/jmbi.2000.4297
Wu S, Hou M, Zhang Y et al (2021) Chinese cases of metachromatic leukodystrophy with the novel missense mutations in ARSA gene. J Mol Neurosci 71(2):245–251. https://doi.org/10.1007/s12031-020-01643-3
Acknowledgements
We would like to thank all professionals from Guangzhou Women and Children’s Medical Center involved in this study.
Funding
This work was supported by Guangzhou Municipal Science and Technology Project (No.202102080028), Natural Science Foundation of Guangdong Province (No.2022A1515012524), and the Internal Funds from Guangzhou Institute of Pediatrics, Guangzhou Women and Children’s Medical Center (NKE-PRE-2019-003) and the Youth Pilot Project of Guangzhou Institute of Pediatrics, Guangzhou Women and Children’s Medical Center (YIP-2016-010).
Author information
Authors and Affiliations
Contributions
Taoling Li & Yonglan Huang: These authors share first authorship and contributed equally to this work in investigation, Writing - Original Draft; Investigation, Methodology, Validation, Writing – review & editing. Chunyan Tao, Xi Yin, Xueying Su & Cuili Liang: Formal analysis, Methodology, Data curation. Minyan Jiang, Yanna Cai, Yunting Lin & Chunhua Zeng: Investigation, Data curation. Xiaoyuan Zhao: Formal analysis. Li Liu & Wen Zhang: These authors share corresponding authorship and contributed equally to this work in Conceptualization, Project administration, Supervision, Writing–review & editing.
Corresponding authors
Ethics declarations
The article is original and has not been published previously. All the stated authors approve its submission. The authors declare that there are no relevant financial or nonfinancial interests to disclose.
Ethics approval
The study was approved by the Institutional Review Board of Guangzhou Women and Children’s Medical Center (2015 − 112).
Consent to participate
Written informed consent was obtained from the parents of all patients.
Consent to publish
Not applicable.
Competing interests
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
ESM 1
(244 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, T., Huang, Y., Tao, C. et al. Biochemical and molecular analysis of pediatric patients with metachromatic leukodystrophy in South China: functional characterization of five novel ARSA variants. Metab Brain Dis 39, 753–762 (2024). https://doi.org/10.1007/s11011-024-01348-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-024-01348-1