Abstract
This study aimed to characterize the expression status and potentially mechanistic involvement of SNHG7 in pituitary adenoma. Relative expression of SNHG7 and miR-449a was analyzed by real-time PCR. Cell viability was measured with Cell Counting Kit-8 (CCK-8). Cell apoptosis was determined by PI/Annexin V double staining followed by flow cytometry analysis. Cell invasion and migration were analyzed by wound healing and transwell assays, respectively. The regulatory action of miR-449a on SNHG7 was interrogated by luciferase reporter assay. We also investigated the pro-tumor activity of SNHG7 with the MMQ xenograft tumor mouse model. We identified the aberrant up-regulation of SNHG7 in pituitary adenoma both in vivo and in vitro, which associated with poor survival outcome. siRNA-mediated SNHG7-knockdown decreased cell viability, increased apoptosis and compromised migration and invasion. We further predicted and validated that SNHG7 negatively regulated miR-449a via sponging. Concurrent inhibition of miR-449a restored cell viability, apoptosis, migration and invasion influenced by SNHG7-deficiency. Most importantly, we demonstrated that SNHG7-silencing delayed xenograft tumor progression, which was accompanied with increased miR-449a and decreased Ki67 intensity. Our study highlighted the essential oncogenic properties of the SNHG7/miR-449a axis in pituitary adenoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pituitary adenoma is a common human malignancy derived from the pituitary gland, affecting about 16% of the general population (Ezzat et al. 2004). However, the clinically evident pituitary adenomas that trigger medical intervention are relatively rare and affecting about 0.1% of the population. Based on their biological functioning, pituitary adenomas are roughly categorized into three classes including benign adenoma, invasion adenoma and carcinoma, and most adenomas are benign where only 35% have invasion potential and 0.1% as malignant carcinomas.
Multiple risk factors have been documented to be associated with the morbidity and mortality of pituitary adenoma. Among which, the multiple endocrine neoplasia type 1 (MEN1) is a relatively rare hereditary genetic aberrance, occurring in 1 out of 30,000 individuals (Newey and Thakker 2011). In addition, the Carney complex (NAME and LAMB syndrome) predisposes the development of growth hormone-producing pituitary tumors (McCarthy et al. 1986). Clinical diagnosis of this disease is frequently base on constellation of related symptoms and pituitary tuberculoma, which is confirmed by hormone level test and radiographic imaging of the pituitary such as CT scan and MRI (Lin et al. 2018). Treatment options for pituitary adenomas depend on the type and size of tumor. Prolactinomas are usually administrated with quinagolide or cabergoline with the aim to decrease tumor size and alleviate symptoms, and sometimes in combination with radiation therapy, proton therapy or surgery for large tumor mass (Chanson et al. 2015). Despite of advances in both diagnosis and therapeutics of pituitary adenomas, the insightful understanding especially at the molecular level is still critically important for better clinical management.
Long non-coding RNAs (lncRNAs) are class of oligonucleotide with the average length of more than 200 nt and without evident protein-coding potential (Kapranov et al. 2007). Diverse physiological functions of lncRNAs are increasingly acknowledged, including gene transcriptional regulation, post-transcriptional modulation, epigenetic regulation and scaffold complexation (Ransohoff et al. 2018). Assembled evidences have uncovered the fundamental involvements of lncRNAs in human cancers as well. In the context of pituitary adenomas, for instance, the lncRNA CLRN1-AS1 was found to serve as a tumor suppressor by inhibiting autophagy via de-activating the Wnt/β-catenin signaling pathway (Wang et al. 2019). LncRNA H19 inhibited mTORC1 by disrupting 4E-BP1/Raptor interaction in pituitary tumors (Wu et al. 2018).
On the other hand, small nucleolar RNA host gene 7 (SNHG7) has been extensively investigated in a number of tumors with established oncogenic role through distinct signaling pathways. For example, Chen et al. showed that SNHG7-knockdown suppressed cell migration and proliferation in bladder cancer via activation of the Wnt/β-catenin pathway (Chen et al. 2019). Zhang et al. reported that SNHG7 overexpression exacerbated cell invasion and migration by regulating the miR-34a/Snail/EMT (epithelial-mesenchymal transition) pathway in gastric cancer (Zhang et al. 2020). In breast cancer, Sun et al. suggested that SNHG7 played critical roles in malignant behaviors and was involved in EMT initiation and Notch-1 signaling via interacting with miR-34a (Sun et al. 2019). The study performed by Qi et al. uncovered the oncogenic role of SNHG7 in prostate cancer via modulation the miR-503/cyclin D1 axis, which consequently contributed to cell proliferation and cell cycle progression (Qi et al. 2018). Han et al. provided evidences showing that SNHG7-silencing partially suppressed EMT process in prostate cancer by modulating miR-324-3p/WNT2B signaling (Han et al. 2019). However, the expression pattern and potential mechanistic involvement of SNHG7 in pituitary adenomas are still elusive.
Here we set out to clarify this issue both in vitro and in vivo. And we further elucidated the molecular events underlying the oncogenic activities of SNHG7 in this disease. Our results highlighted the importance of SNHG7/miR-449a in pituitary adenoma.
Materials and methods
Tissue samples
A total of 30 pituitary tumor tissues with paired adjacent normal tissues were collected at Hebei General Hospital from Dec. 2014 to Nov. 2015. The written informed consent was received from the participants. Tissue samples were flash frozen in liquid nitrogen and stored at −80 °C until use. The protocol was approved by the Institutional Ethics Committee of Hebei General Hospital.
Cell culture
Rat pituitary, GH1, RC-4B/C, GH3 and MMQ cell lines were obtained from the American Type Culture Collection (ATCC, VA, USA). Cells were routinely cultured in F-12 K medium (Invitrogen, CA, USA) supplemented with 12.5% FBS (Gibco, MA, USA) and 1% antibiotics in CO2 incubator (5%) at 37 °C. Cell identities were validated by short tandem repeat profiling and mycoplasma contamination was tested by regular PCR. Transfection was conducted with Lipofectamine 2000 (Invitrogen, MA, USA) with ~70% efficiency as tested using a GFP-positive control.
Real-time PCR
Total RNAs from both cells and tissues were extracted using the TRIzol Reagent (Invitrogen, MA, USA) following the manufacturer’s protocol. cDNA was synthesized using the TIANScript II cDNA kit (Tiangen, Beijing, China). Real-time PCR was performed on the ABI-7900 PCR System (Applied Biosystems, CA, USA) with the SYBR Green Master Mix kit (Invitrogen, CA, USA). The fold-changes were calculated by the 2-ΔΔCt method. The primer sequences were listed as below:
-
SNHG7 Forward: 5′-AGGCTGAAGTTACAGGTC-3′,
-
SNHG7 Reverse: 5′-TTGGCTCCCAGTGTCTTA-3′,
-
U6 Forward: 5′-CCAGTGCAGGGTCCGAGGT-3′,
-
U6 Reverse: 5′-CCAGTGCAGGGTCCGAGGT-3′,
-
miR-449a Forward: 5′-CGGGGTACCGTTTCAGTGGAGGTGTCT-3′,
-
miR-449a Reverse: 5′-CCGCTCGAGCCTGTAGCCAAGAACTGC-3′,
-
β-actin Forward: 5′-CGTGACATTAAGGAGAAGCTG-3′,
-
β-actin Forward: 5′-CTAGAAGCATTTGCGGTGGAC-3′.
Cell counting Kit-8 (CCK-8) assay
Cell viability was determined by CCK-8 (Beyotime, China) according to the provider’s instructions. Briefly, the indicated cells were seeded in the 96-well culture plate (5 × 103 cells/well) and subjected to consecutive culture. The absorbance at 450 nm was measured in triplicate at indicated time on a microplate reader (Bio-Tek, VT, USA).
Cell apoptosis analysis
The indicated cells were prepared into single-cell suspension by trypsinization. The cell apoptosis was determined by Annexin V-FITC/PI Apoptosis Detection kit (Kaiji Biotechnology, Nanjing, China) following the manufacturer’s protocol. Flow cytometry was performed on CytoFlex (Beckman Coulter, CA, USA), where viable cells are negative to both probes, apoptotic cells are Annexin positive, necrotic cells are PI positive and Annexin negative.
Wound healing assay
Cells were plated into 6-well plates and a scratch wound was created by sterile pipette tips. The detached cells were washed off with PBS and cells were continuously cultured in serum-free medium. Gap closure was regularly monitored and captured under a light microscope.
Transwell assay
Cell invasion was evaluated using the transwell chamber (Corning, NY, USA). The indicated cells were prepared into single cell solution and poured into the upper inserts which was precoated with 0.1% Matrigel (BD BioSciences, NJ, USA). The lower compartment was supplied with complete culture medium. After 12 h of incubation, the invaded cells were fixed and stained with crystal violet solution. The images were taken under an inverted microscope, and cells were counted from five independent areas.
Dual-luciferase assay
The luciferase assay was performed to investigate the regulatory potential of miR-449a on SNHG7. Full-length SNHG7 transcript was fused to luciferase reporter vector pGL4 (Promega, WI, USA). Co-transfection with miR-449a was achieved by Lipofectamine 2000 (Invitrogen, MA, USA). Luciferase activity was determined using Bright-Glo Luciferase Assay System (Promega, WI, USA) on the Multi-Mode microplate reader (Bio-Tek, VT, USA).
Immunohistochemistry assay (IHC)
The xenograft tissues were first fixed in 10% formaldehyde and embedded in paraffin. The blocks were sectioned into 5 μm thickness with a microtome (Leica, Wetzlar, Germany). The section was probed with primary anti-Ki67 antibody (Santa Cruz Biotechnology, 1:500, TX, USA) at 4 °C overnight. The HRP-conjugated secondary antibody was applied next for 2 h at room temperature, and developed using the DAB substrate (Sigma-Aldrich, MO, USA).
Xenograft mice model
The animal study was approved by the Institutional Animal Care and Use Committee of Hebei General Hospital. MMQ cells (1× 106) with stable SNHG7-knockdown or control shRNA were subcutaneously injected into the lower flank of nude mice (Vital River, Beijing, China). Tumor volumes were measured with a digital caliper and estimated with the formula: 0.5 × length×width2.
Statistics analysis
Statistical analysis was performed with SPSS 23.0 (SPSS, IL, USA) and processed with GraphPad PRISM 7 (GraphPad, CA, USA). Every experiment was repeated at least three times as biological replicates. The intra-group comparison was analyzed with Student’s unpaired t-test. Correlation was examined using Pearson’s χ2 analysis. P value was set as <0.05 to be statistical difference.
Results
SNHG7 expressed higher in pituitary adenoma tissues and cell lines
We first set out to determine the relative expression of SNHG7 in pituitary adenoma both in vivo and in vitro. Quantitation of transcript levels by real-time PCR showed significant up-regulation of SNHG7 in pituitary adenoma tissue samples in comparison with adjacent normal tissue controls (Fig. 1a). This was further consolidated in cell culture, and remarkably higher SNHG7 was noticed in GH1, RC-4B/C, GH3 and MMQ cells in comparison with rat pituitary cells (Fig. 1b). More importantly, the high abundance of SNHG7 was associated with evidently unfavorable prognosis in comparison with low-SNHG7 group in pituitary adenomas patients (Fig. 1c). Expression levels of SNHG7 in all samples were ordered and the median expression level was used to differentiate between high/low expression of SNHG7. Therefore, we characterized the aberrant over-expression of SNHG7 in pituitary adenoma, which indicated potential oncogenic properties in this disease.
SNHG7 expressed higher in pituitary adenomas tissues and cell lines. a RT-qPCR was performed to determine the relative expressions of SNHG7 in pituitary adenomas tissues and adjacent normal tissues. b RT-qPCR was performed to estimate the relative expressions of SNHG7 in pituitary adenomas cell lines and normal cells. c Survival curves was analyzed by Kaplan–Meier survival analysis, p = 0.0342 by log-rank test. The results are displayed as the mean ± SD, **P < 0.01
SNHG7 knockdown suppressed pituitary adenoma cell proliferation, migration and invasion
Next, we sought to investigate the potential oncogenic contributions of SNHG7 in pituitary adenoma cells. To this end, we first established SNHG7-knockdown cell lines derived from both GH3 and MMQ cells with specific siRNA. The knockdown efficiency was evaluated by real-time PCR, and approximately 50% of reduction was achieved in both cell lines as shown in Fig. 2a. We noticed that cell viability measured up to 4 days was greatly compromised by SNHG3-deficiency (Fig. 2b). The PI/Annexin V double staining results demonstrated notable induction of cell apoptosis by SNHG7-knockdown (Fig. 2c). We further assessed the possible impacts of SNHG7 on metastasis-related behaviors including migration and invasion in pituitary adenoma cells. As shown in Fig. 2d, the gap closure was delayed by SNHG7-silencing in both GH3 and MMQ cells, which suggested suppressed migration elicited by down-regulation of SNHG7. Consistently, the invasive capacity as measured by transwell assay displayed significant reduction in both SNHG7-depleted GH3 and MMQ cells (Fig. 2e). For convenient comparison, we provided the statistical results alongside the representative images acquired from both wound healing and transwell assays. Therefore, our data supported the oncogenic potential of SNHG7 in pituitary adenoma, and SNHG7-knockdown significantly suppressed cell proliferation, migration and invasion in pituitary adenoma in vitro.
SNHG7 knockdown suppressed pituitary adenomas cell proliferation, migration and invasion. a Knockdown efficiency of si-SNHG7 was confirmed by qPCR. After transfection of si-SNHG7, b the cell viability was detected by CCK-8 assay; c apoptosis was detected by cytometry using the Annexin V-FITC/PI Apoptosis Detection kit (viable cells are negative to both probes; apoptotic cells are Annexin positive; necrotic cells are PI positive and Annexin negative); d cell migration was measured by wound healing assay; e cell invasion was estimated by transwell assay. The results are displayed as the mean ± SD, **P < 0.01
SNHG7 sponged miR-449a
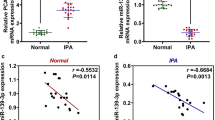
Next, we decided to elucidate the molecular mechanism underlying the oncogenic property of SNHG in pituitary adenoma. Here we employed the online algorithm Starbase 2.0 (starbase.sysu.edu.cn/starbase2/index.php) to predict the target microRNAs (miRNAs, miRs) of SNHG7. There were a total of 25 miRNAs predicted as potential targets of SNHG7. We were interested in miR-449a for the following three reasons: 1) we verified the predicted top 20 miRNAs by luciferase reporter activity assay, and found 7 miRNAs including miR-449a were significantly regulated by SNHG7; 2) miR-449a has been identified as a tumor suppressor in a series of cancer types, such as endometrial cancer, breast cancer and gastric cancer; 3) there existed reports showing that the SNHG7/miR-449a axis was involved in thyroid cancer. Based on the above reasons, we focused our limited research resource into miR-449a, while leaving the other candidates for future studies. Alignment between miR-449a and putative binding site of SNHG7 was shown in Fig. 3a. The regulatory effect of miR-449a on SNHG7 was interrogated with luciferase reporter assay. Exogenous miR-449a tremendously inhibited SNHG-7-fused luciferase activities in both GH3 and MMQ cells (Fig. 3b). In line with this observation, siRNA-mediated specific knockdown of SNHG7 markedly induced up-regulation of endogenous miR-449a (Fig. 3c). Conversely, ectopic introduction of miR-449a mimics inhibited expression of SNHG7 in both GH3 and MMQ cells (Fig. 3d). Contrary to the up-regulated SNHG7, we observed marked down-regulation of miR-449a in pituitary adenoma tissues in comparison with adjacent normal tissues (Fig. 3e). Our data uncovered the sponging action of SNHG7 on miR-449a and suggested the mutually negative regulation between SNHG7 and miR-449a in pituitary adenoma.
SNHG7 sponged miR-449a. a Schematic presentation of the wildtype and mutant SNHG7 binding sites with miR-449a by Starbase 2.0. b Luciferase activity was performed with SNHG7-wt or SNHG7-mut reporter co-transfection with miR-449a mimics. c miR-449a expression was measured when si-SNHG7 was transfected. d SNHG7 expression was measured when miR-449a mimics were transfected. e Expression patterns of miR-449a in pituitary adenomas tissues and adjacent normal tissues. wt, wild type; mut, mutant. The results are displayed as the mean ± SD, **P < 0.01
MiR-449a mediated the regulation in pituitary adenoma cell proliferation and metastasis induced by SNHG7
Despite of regulation of miR-449a by SNHG7, the potential involvement and contribution of miR-449a in the oncogenic activity of SNHG7 in pituitary adenoma were still to be defined. Here we specifically inhibited miR-449a in the context of SNHG7-knockdown via employment of miR-449a specific inhibitor. The inhibitory effects were first evaluated by real-time PCR, which demonstrated that more than 50% of reduction was achieved in both GH3 and MMQ cells (Fig. 4a). Concurrent treatment with miR-449a inhibitor greatly restored the compromised cell viability by SNHG7-knockdown in both GH3 and MMQ cells (Fig. 4b). Similarly, cell apoptosis induced by SNHG7 depletion in both GH3 and MMQ cells was partially relieved by miR449a inhibition (Fig. 4c). Furthermore, both cell migration (Fig. 4d) and invasion (Fig. 4e) were significantly accelerated by miR-449a inhibition in SNHG7-deficient GH3 and MMQ cells. For convenient comparison, we provided the statistical results acquired from cell apoptosis, wound healing and transwell assays in Fig. 4f. Our results suggested that miR-449a predominantly mediated oncogenic signaling of SNHG7 in pituitary adenoma.
miR-449a mediated the regulation in pituitary adenomas cell proliferation and metastasis induced by SNHG7. a Knockdown efficiency of miR-449a inhibitor was confirmed by qPCR. After transfection of si-SNHG7 with miR-449a or miR-NC, b the cell viability was detected by CCK-8 assay; c apoptosis was detected by cytometry using the Annexin V-FITC/PI Apoptosis Detection kit (viable cells are negative to both probes; apoptotic cells are Annexin positive; necrotic cells are PI positive and Annexin negative); d cell migration was measured by wound healing assay; e cell invasion was estimated by transwell assay; f Quantitative analysis were presented. The results are displayed as the mean ± SD, **P < 0.01 vs. si-NC + miR-NC group; ##P < 0.01 vs. si-SNHG7 + miR-NC group
SNHG7 knockdown inhibited tumorigenesis in vivo
All previous evidences in support of oncogenic activity of SNHG7 were acquired from in vitro cell culture, conclusion based on which was potentially compromised by artifacts involved in experiment procedure. To consolidate our observations, we further investigated the impact of SNHG7-deficiency on tumor progression in vivo by employing a xenograft mouse model. Growth of SNHG7-silenced MMQ cell-derived xenograft tumor was significantly delayed in comparison with control tumor (Fig. 5a). Xenograft tumors resected from SNHG7-deficient group were much smaller than the ones from the control group (Fig. 5b). We confirmed the up-regulation of miR-449a in SNHG7-deficient MMQ xenograft tumors at the endpoint, and around 3-fold up-regulation of endogenous miR-449a was noticed (Fig. 5c). In line with the tumor-suppressive effect of SNHG7-deficiency, our IHC result demonstrated tremendous reduction of cell proliferative marker Ki67 in SNHG7-depleted xenograft tumors (Fig. 5d). In summary, we provided both in vitro and in vivo evidences supporting the oncogenic activity of SNHG7 in pituitary adenoma.
SNHG7 knockdown inhibited tumor genesis in vivo. MMQ cells expressing sh-SNHG7 or sh-NC stably were injected into mice, a tumor size was analyzed weekly. b Tumor weight was measured 5 weeks. c miR-449a expression was detected. (D) Ki67 expression was determined by ICH. The results are displayed as the mean ± SD, **P < 0.01
Discussion
Despite of accumulative evidences pointing to its oncogenic properties in diverse human cancers, the expression pattern and mechanistic involvement of SNHG7 in pituitary adenoma were still largely unknown currently. Here we analyzed the abundance of SNHG7 transcript in pituitary adenoma both in vivo and in vitro. We characterized the aberrant over-expression of SNHG7, which was closely associated with unfavorable clinical prognosis and hinted oncogenic its activity. We further experimentally demonstrated that siRNA-mediated SNHG7-knockdown significantly inhibited cell viability, induced cell apoptosis and compromised cell migrative and invasive capacities in two different pituitary adenoma cell lines. We further took advantage of an online bioinformatic tool to discover the potential target of SNHG7 and identified miR-449a as the top candidate. We uncovered the sponging effect of SNHG7 on miR-449a. Both endogenous SNHG7 and luciferase reporter activity were inhibited by ectopic miR-449a, and conversely, siRNA-mediated SNHG7-depletion led to increased miR-449a. Notably, co-administration with miR-449a specific inhibitor greatly restored cell viability, migration and invasion, which were suppressed by SNHG7-depletion in pituitary adenomas cells, while attenuated cell apoptosis. These individual effects collectively contributed to the anti-tumor activity of miR-449a inhibitor. Most importantly, we provided in vivo evidences in support of the oncogene role of SNHG7 with MMQ xenograft tumor mouse model.
SNHG7-depletion significantly inhibited tumor progression with accompanied persistent up-regulation of endogenous miR-449a and decreased cell proliferative index. Our data firstly unraveled the oncogenic property of SNHG7 in pituitary adenoma through modulation of cell viability, apoptosis, migration and invasion. We proposed the mechanistic involvement of miR-449a in this scenario and highlighted the anti-tumor activity of miR-449a inhibitor in this disease. Our data supported further investigations into the therapeutic values of miR-449a antagonists in pituitary adenoma both in vitro and in clinic. In this regard, the elucidation of the SNHG7/miR-499a signaling in pituitary adenoma expanded both of the current diagnostic and therapeutic avenues.
So far, accumulative studies pointed to the anti-tumor properties of miR-449a in a number of human cancers. For instance, Huang et al. proposed miR-499 and its target Flot2 as prognostic biomarkers for glioma (Huang et al. 2019). Li et al. showed that forced overexpression of miR-449 suppressed papillary thyroid carcinoma cell proliferation through targeting the RET-β-catenin signaling axis (Li et al. 2016). Jang et al. identified the low abundance and its causal linkage of miR-449 in gynecologic clear cell carcinoma (Jang et al. 2014). In gastric cancer, the study performed by Bou et al. demonstrated that miR-449 compromised cell proliferation and was down-regulated (Bou Kheir et al. 2011). Zhang et al. suggested that miR-499-silencing stimulated cell migration and invasion in breast cancer via targeting TPD52 (Zhang et al. 2017). This phenomenon was consolidated by the investigation conducted by Jiang et al., which uncovered the involvement of CREPT-mediated Wnt/β-catenin inhibition (Jiang et al. 2019). In hepatocellular cell lines, Zhang et al. reported that miR-449 exerted proliferation inhibitory roles via blocking the lipid metabolic pathway related to SIRT1 (Zhang et al. 2014), while Buurman et al. indicated that histone deacetylases stimulated hepatocellular growth factor pathway through negative suppression of miR-449 (Buurman et al. 2012). The novel finding presented by Qu et al. exhibited that lncARSR was selectively packaged in exosome and transferred to render sunitinib resistance to renal cancer as a competing endogenous RNA (Qu et al. 2016). In colon cancer, Fang et al. uncovered that miR-449 suppressed cell proliferation of stem cells via negatively regulating CCND1 and E2F3 expression (Fang et al. 2013). In agreement with the well-established tumor suppressor role, here we provided evidences in support of the consistent function of miR-449 in pituitary adenoma, and proposed inhibition of miR-449a as a promising therapeutic intervention for this disease.
Noteworthily, here we only predicted and experimentally validated that miR-449a exert anti-tumor function at the downstream of aberrantly up-regulated SNHG7 in pituitary adenoma, whereas the target gene of miR-449a in this scenario was out of the range of our current investigation. However, identification of potential target genes which were closely regulated by miR-449a in pituitary adenoma was fundamentally essential to a complete understanding of the SNHG7/miR-449a signaling. This would definitely be our priority in the following study and achievable with both bioinformatics prediction and experimental validation. We also would not exclude any previously recognized miR-449a target such as c-Met in lung cancer (Luo et al. 2013), CDK6 in gastric cancer (Li et al. 2014) and TPD52 in breast cancer (Zhang et al. 2017).
In summary, here we for the first time uncovered the oncogenic potential of SNHG7 in pituitary adenoma. The aberrant overexpression of SNHG7 promoted cell proliferation, suppressed apoptosis and stimulated migrative and invasive cell behaviors. In addition to mechanistic elucidation, we exemplified the therapeutic application of miR-449a inhibitor in cell culture, which suggested further investigations in vivo and in clinic.
References
Bou Kheir T, Futoma-Kazmierczak E, Jacobsen A, Krogh A, Bardram L, Hother C, Grønbæk K, Federspiel B, Lund AH, Friis-Hansen L (2011) miR-449 inhibits cell proliferation and is down-regulated in gastric cancer. Mol Cancer 10:29. https://doi.org/10.1186/1476-4598-10-29
Buurman R, Gürlevik E, Schäffer V, Eilers M, Sandbothe M, Kreipe H, Wilkens L, Schlegelberger B, Kühnel F, Skawran B (2012) Histone deacetylases activate hepatocyte growth factor signaling by repressing microRNA-449 in hepatocellular carcinoma cells. Gastroenterology 143:811–820 e815. https://doi.org/10.1053/j.gastro.2012.05.033
Chanson P, Raverot G, Castinetti F, Cortet-Rudelli C, Galland F, Salenave S, French Endocrinology Society non-functioning pituitary adenoma w-g (2015) Management of clinically non-functioning pituitary adenoma. Ann Endocrinol (Paris) 76:239–247. https://doi.org/10.1016/j.ando.2015.04.002
Chen Y, Peng Y, Xu Z, Ge B, Xiang X, Zhang T, Gao L, Shi H, Wang C, Huang J (2019) Knockdown of lncRNA SNHG7 inhibited cell proliferation and migration in bladder cancer through activating Wnt/beta-catenin pathway. Pathol Res Pract 215:302–307. https://doi.org/10.1016/j.prp.2018.11.015
Ezzat S, Asa SL, Couldwell WT, Barr CE, Dodge WE, Vance ML, IE MC (2004) The prevalence of pituitary adenomas: a systematic review. Cancer 101:613–619. https://doi.org/10.1002/cncr.20412
Fang Y, Gu X, Li Z, Xiang J, Chen Z (2013) miR-449b inhibits the proliferation of SW1116 colon cancer stem cells through downregulation of CCND1 and E2F3 expression. Oncol Rep 30:399–406. https://doi.org/10.3892/or.2013.2465
Han Y, Hu H, Zhou J (2019) Knockdown of LncRNA SNHG7 inhibited epithelial-mesenchymal transition in prostate cancer though miR-324-3p/WNT2B axis in vitro. Pathol Res Pract 215:152537. https://doi.org/10.1016/j.prp.2019.152537
Huang S, Zheng S, Huang S, Cheng H, Lin Y, Wen Y, Lin W (2019) Flot2 targeted by miR-449 acts as a prognostic biomarker in glioma. Artif Cells Nanomed Biotechnol 47:250–255. https://doi.org/10.1080/21691401.2018.1549062
Jang SG, Yoo CW, Park SY, Kang S, Kim HK (2014) Low expression of miR-449 in gynecologic clear cell carcinoma. Int J Gynecol Cancer 24:1558–1563. https://doi.org/10.1097/IGC.0000000000000267
Jiang J, Yang X, He X, Ma W, Wang J, Zhou Q, Li M, Yu S (2019) MicroRNA-449b-5p suppresses the growth and invasion of breast cancer cells via inhibiting CREPT-mediated Wnt/beta-catenin signaling. Chem Biol Interact 302:74–82. https://doi.org/10.1016/j.cbi.2019.02.004
Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, Bell I, Cheung E, Drenkow J, Dumais E, Patel S, Helt G, Ganesh M, Ghosh S, Piccolboni A, Sementchenko V, Tammana H, Gingeras TR (2007) RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316:1484–1488. https://doi.org/10.1126/science.1138341
Li LP, Wu WJ, Sun DY, Xie ZY, Ma YC, Zhao YG (2014) miR-449a and CDK6 in gastric carcinoma. Oncol Lett 8:1533–1538. https://doi.org/10.3892/ol.2014.2370
Li Z, Huang X, Xu J, Su Q, Zhao J, Ma J (2016) miR-449 overexpression inhibits papillary thyroid carcinoma cell growth by targeting RET kinase-beta-catenin signaling pathway. Int J Oncol 49:1629–1637. https://doi.org/10.3892/ijo.2016.3659
Lin P, Liu X, Wang S, Li X, Song Y, Li L, Cai S, Wang X, Chen J (2018) Diagnosing pituitary adenoma in unstained sections based on multiphoton microscopy. Pituitary 21:362–370. https://doi.org/10.1007/s11102-018-0882-6
Luo W, Huang B, Li Z, Li H, Sun L, Zhang Q, Qiu X, Wang E (2013) MicroRNA-449a is downregulated in non-small cell lung cancer and inhibits migration and invasion by targeting c-Met. PLoS One 8:e64759. https://doi.org/10.1371/journal.pone.0064759
McCarthy PM, Piehler JM, Schaff HV, Pluth JR, Orszulak TA, Vidaillet HJ Jr, Carney JA (1986) The significance of multiple, recurrent, and "complex" cardiac myxomas. J Thorac Cardiovasc Surg 91:389–396
Newey PJ, Thakker RV (2011) Role of multiple endocrine neoplasia type 1 mutational analysis in clinical practice. Endocr Pract 17(Suppl 3):8–17. https://doi.org/10.4158/EP10379.RA
Qi H, Wen B, Wu Q, Cheng W, Lou J, Wei J, Huang J, Yao X, Weng G (2018) Long noncoding RNA SNHG7 accelerates prostate cancer proliferation and cycle progression through cyclin D1 by sponging miR-503. Biomed Pharmacother 102:326–332. https://doi.org/10.1016/j.biopha.2018.03.011
Qu L, Ding J, Chen C, Wu ZJ, Liu B, Gao Y, Chen W, Liu F, Sun W, Li XF, Wang X, Wang Y, Xu ZY, Gao L, Yang Q, Xu B, Li YM, Fang ZY, Xu ZP, Bao Y, Wu DS, Miao X, Sun HY, Sun YH, Wang HY, Wang LH (2016) Exosome-transmitted lncARSR promotes sunitinib resistance in renal cancer by acting as a competing endogenous RNA. Cancer Cell 29:653–668. https://doi.org/10.1016/j.ccell.2016.03.004
Ransohoff JD, Wei Y, Khavari PA (2018) The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol 19:143–157. https://doi.org/10.1038/nrm.2017.104
Sun X, Huang T, Liu Z, Sun M, Luo S (2019) LncRNA SNHG7 contributes to tumorigenesis and progression in breast cancer by interacting with miR-34a through EMT initiation and the Notch-1 pathway. Eur J Pharmacol 856:172407. https://doi.org/10.1016/j.ejphar.2019.172407
Wang C, Tan C, Wen Y, Zhang D, Li G, Chang L, Su J, Wang X (2019) FOXP1-induced lncRNA CLRN1-AS1 acts as a tumor suppressor in pituitary prolactinoma by repressing the autophagy via inactivating Wnt/beta-catenin signaling pathway. Cell Death Dis 10:499. https://doi.org/10.1038/s41419-019-1694-y
Wu ZR, Yan L, Liu YT, Cao L, Guo YH, Zhang Y, Yao H, Cai L, Shang HB, Rui WW, Yang G, Zhang XB, Tang H, Wang Y, Huang JY, Wei YX, Zhao WG, Su B, Wu ZB (2018) Inhibition of mTORC1 by lncRNA H19 via disrupting 4E-BP1/raptor interaction in pituitary tumours. Nat Commun 9:4624. https://doi.org/10.1038/s41467-018-06853-3
Zhang H, Feng Z, Huang R, Xia Z, Xiang G, Zhang J (2014) MicroRNA-449 suppresses proliferation of hepatoma cell lines through blockade lipid metabolic pathway related to SIRT1. Int J Oncol 45:2143–2152. https://doi.org/10.3892/ijo.2014.2596
Zhang Z, Wang J, Gao R, Yang X, Zhang Y, Li J, Zhang J, Zhao X, Xi C, Lu X (2017) Downregulation of microRNA-449 promotes migration and invasion of breast cancer cells by targeting tumor protein D52 (TPD52). Oncol Res 25:753–761. https://doi.org/10.3727/096504016X14772342320617
Zhang Y, Yuan Y, Zhang Y, Cheng L, Zhou X, Chen K (2020) SNHG7 accelerates cell migration and invasion through regulating miR-34a-Snail-EMT axis in gastric cancer. Cell Cycle 19:142–152. https://doi.org/10.1080/15384101.2019.1699753
Author information
Authors and Affiliations
Contributions
All authors participated in the design, interpretation of the studies and analysis of the data and review of the manuscript; Xiongfei Yue, Ce Dong, Zhanying Ye, Lin Zhu, Xiaoyang Zhang, Xiaoyan Wang, Feng Mo, Zheng Li, Baogen Pan conducted the experiments, Baogen Pan wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
None.
Ethical approval
The animal study was approved by the Institutional Animal Care and Use Committee of Hebei General Hospital.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yue, X., Dong, C., Ye, Z. et al. LncRNA SNHG7 sponges miR-449a to promote pituitary adenomas progression. Metab Brain Dis 36, 123–132 (2021). https://doi.org/10.1007/s11011-020-00611-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-020-00611-5