Abstract
CircRNAs are a member of noncoding RNAs and have been verified to play an important regulatory role in cancers. In CRC, the regulatory mechanisms of various circRNAs have not been elucidated. The expression of circPACRGL and miR-330-3p was detected with qRT-PCR. The protein expression of CDK4, MMP-9, Bcl-2, Bax, cellular nucleic acid-binding protein (CNBP) and β-actin was measured with western blot. Cell proliferation was analyzed using MTT assay, colony formation assay, and EDU assay. Cell apoptosis was detected using flow cytometry. Cell migration and invasion were measured with wound healing and transwell invasion assay. Luciferase reporter assay and RIP assay was used to determine the relationship of among miR-330-3p, circPACRGL and CNBP in CRC cells. In this study, we found that circPACRGL and CNBP expressed high and miR-330-3p expressed low in CRC tissues and cells. Functional experiments showed that inhibition of circPACRGL reduced cell proliferation, migration and invasion in CRC. In addition, knockdown of circPACRGL contributed to cell apoptosis in CRC. Dual-luciferase report assay determined that circPACRGL was a miR-330-3p sponge molecular and CNBP was a target of miR-330-3p. Reversed experiments showed that the effects of sh-circPACRGL transfection on CRC cells were rescued by up-regulating CNBP expression. In this study, we for the first time found a novel regulatory network of circPACRGL in CRC. The results manifested that circPACRGL affected tumor growth by targeting miR-330-3p/CNBP axis in CRC, highlighting the potential of circPACRGL as a therapeutic target for colorectal cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer (CRC) is a common malignant tumor in the world, and its incidence in China continues to rise [1]. Owing to the widespread use of colorectal screening and surgery, the overall survival rate of CRC has significantly enhanced [2]. Because of the various causes of CRC, the difficulty in treatment is also increased [3]. Therefore, an in-depth understanding of the pathogenesis of CRC contributed to the development of targeted treatment, as well as effective biopharmaceutical therapy.
In the recent years, emerging evidence has determined that noncoding RNAs (ncRNAs) exert a vital regulatory function in tumors. Both circRNAs and microRNAs are ncRNAs and they are vital to the development and diagnosis of tumors [4,5,6,7,8]. CircRNAs are mainly localized in the cytoplasm without cap structure and polyA tail, and are circRNAs linked by exons of primary gene [9, 10]. CircRNA was notified to engage in the regulation of the occurrence and development of a variety of cancers, which has great research potential [11]. For example, circRNA UBAP2 promote cell progression in osteosarcoma through sponging miR-143 [12]. CircRNA-5692 was identified to inhibit hepatocellular carcinoma through modulating miR-328-5p/DAB2IP axis [13]. In CRC cancers, circRNA_CBL.11, circPACRGL and circRNA_0000392 have been verified to be related to cell proliferation, metastasis and invasion [14,15,16]. Some scholars have proposed the potential of using circRNA as a novel biomarker for the clinical diagnosis and treatment of cancer [8, 17, 18]. For example, Yang et al., reported that circRNA_PTK2 was positively correlated with tumor growth and metastasis, as well as poorer survival rates, then has been verified to be a novel therapeutic target for CRC metastasis [19]. Not only that, but miRNAs also have similar effects. The miRNA length is only 22-25nt, and it is an important regulator in regulating the post-transcriptional level of genes [20]. The most important mechanism of action of circRNAs is known to affect transcription silencing, translation and specific mRNA degradation by binding to miRNAs [21]. Therefore, circRNAs are important regulators in cancers. However, the functions of many important circRNAs in CRC are not clear, which is of great value for further study.
In this paper, we mainly studied the role of circPACRGL in CRC tumor development and experimentally verified its regulatory mechanism. CircPACRGL expression was elevated in CRC, and inhibition of circPACRGL could inhibit the development of CRC cells by regulating miR-330-3p/cellular nucleic acid-binding protein (CNBP) axis.
Materials and methods
Tissue collection, cell culture and transfection
CRC tissues and paired normal tissues were selected from 39 patients in Jiangxi University of Chinese Medicine. This test received the approval of the Ethics Committee of Jiangxi University of Chinese Medicine. Informed written consent were gained from patients. All patients did not undergo preoperation treatment. All samples were restored at -80℃ for subsequent analysis.
CRC cell lines (SW480, SW620, LOVO, HT29) and normal cell line (NCM460) procured from RiboBio Co. (Guangzhou, China) were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal bovine serum (FBS, Thermo, Waltham, MA, USA), 1% penicillin/streptomycin at 37 °C with 5% CO2.
Short hairpin RNA (shRNA) targeting circPACRGL (sh-circPACRGL) or CNBP (sh-CNBP), pcDNA-CNBP, miR-330-3p mimic, inhibitor, and their negative controls (sh-NC, pcDNA, mimic NC, and inhibitor NC) were procured from Genepharma (Shanghai, China). These oligos and plasmids were introduced into SW480 and LOVO cells through Lipofectamine 2000 (Invitrogen, CA, USA).
Quantitative real-time PCR
The total RNA from CRC tissues and cells was isolated via TRIzol reagent (Invitrogen) and quantified by spectrophotometry. TaqMan® MicroRNA real-time PCR assay reagent was exploited for the detection of miR-330-3p expression. For the expression of circPACRGL and CNBP, M-MLV Reverse Transcriptase (Invitrogen) and SYBR Green Real-time PCR kit (Invitrogen) were applied. GAPDH or U6 was employed as the normalized gene for circPACRGL, CNBP and miR-330-3p, respectively. The 2−ΔΔCt method was used to evaluate the data [22]. The primer sequences were represented in Table 1.
Western blot
Cells and tissues were isolated via RIPA buffer (ThermoFisher Scientific, 1 ml per 100 mm dish) and concentrations were estimated by BCA Protein Assay Kit (Beyotime, Shanghai, China). Western blot analyses were performed as previously described [23]. 20 µg proteins were added into the wells of the SDS–polyacrylamide gel electrophoresis (SDS-PAGE) gel. After running at 60 V for 10 min and developing the voltage to 120 V for 1 h, the bands were electro-transferred onto PVDF membranes (Millipore). Then, the membranes were incubated with antibodies against CDK4, MMP9, Bcl-2, Bax, CNBP, and β-actin (1:2000, Cell Signaling Technology, Danvers, MA, USA) at 4 °C overnight, followed by incubating with goat antimouse IgG (1:2000 dilution, Cell Signaling Technology) at 37 °C for 1 h. Finally, the blots were measured by PierceTM ECL western blotting substrate (ThermoFisher Scientific).
Cell proliferation
MTT assay kit (Sigma, St. Louis, MO, USA) was utilized to evaluate cell proliferation. 2 × 103 transfected cells were plated into the 96-well plates. After being cultured for 48 h, the cells were nurtured with 20 uL MTT solution (5 mg/mL) at 37 °C for 4 h. Subsequently, 200 µL of dimethyl sulfoxide (DMSO; Sigma) were added into each well for 30 min. Cell proliferation was detected at a wavelength of 490 nm using the Labsystems MultiSkan Ascent Type 354 microplate reader (Thermo Labsystems, Waltham, MA, USA).
Colony formation assay
The transfected cells (1000 cells/per well) were plated into six-well plates. 10 days later, colonies were settled with 10% formaldehyde for a half hour and spotted by 0.5% crystal violet (Sigma-Aldrich, San Francisco, CA, USA). The colonies were imaged with NIKON camera and counted.
Cell apoptosis
Flow cytometry was implemented to evaluate cell apoptosis. Transfected cells were trypsinized and washed 3 times with PBS, then spotted with FITC-Annexin V and PI (BD Biosciences, San Jose, CA) for 30 min. The apoptosis rate was estimated using flow cytometry (BD Biosciences).
Transwell
Cell invasion abilities were conducted via transwell chambers precoated with Matrigel (BD Biosciences). Briefly, transfected cells in the DMEM medium were added into the upper chamber, while the lower chamber was supplemented with DMEM medium including 10% FBS. 24 h later, the cells on the upper chamber were removed and spotted with 0.2% crystal violet for 30 min. Cell invasion was calculated using an inverted microscope.
Wound healing assay
In brief, about 4000 transfected cells were plated on to the six-well plates and maintained until 70–90% confluence. Then, a sterile pipette tip was utilized to make a “wound”, and the morphological images of the cells were recorded by a microscope at 0 h and 24 h.
Luciferase reporter assay
CircPACRGL wt, circPACRGL mut, CNBP 3’UTR wt and CNBP3’UTR mut sequences were cloned into pGL3 (Promega, Madison, WI, USA) to construct luciferase reporter vectors. These vectors and miR-330-3p were co-introduced into the cells by Lipofectamine 2000 (Invitrogen) for 48 h. Relative luciferase activity was estimated by Dual-Luciferase Reporter Assay System (Promega).
RIP assay
The Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit (Merck Life Science (Shanghai) Co., Ltd. Shanghai, China) was utilized to measure the ability to bind between miRNA and circRNA/mRNA. Briefly, transfected cells were fixed with formaldehyde and lysed in lysis buffer. Then, 100 μL of cell lysates in RIP buffer was hatched with magnetic beads integrated with the human anti-Argonaute2 (Ago2) antibody (Cell Signaling Technology) or negative control IgG (Cell Signaling Technology). After purified with proteinase K, the precipitation of RNA was examined by qPCR.
EdU assay
Briefly, transfected cells were cultured on a six-well plate overnight. The EdU assay was fulfilled using a Cell-Light EdU DNA Cell Proliferation Kit (RiboBio, Shanghai, PR, China) for testing cells as per manufacturer’s instructions.
Immunohistochemistry staining (IHC)
The tumor was settled in paraformaldehyde overnight and embedded in paraffin. Then the slices were incubated with CNBP, Ki-67, MMP9, and MMP2 antibodies (1:150). After incubating with the secondary streptavidin horseradish peroxidase conjunct antibody at room temperature, the signals were visualized in 3,3’-diaminobenzidine (ZLI9018, ZSGBBIO, China). All images were obtained by a light microscope.
Animal experiments
BABL/c female nude mice were procured from the Animal Center of Central South University. Animal experiments received the approval of the Ethical Committee for Animal Research of Jiangxi University of Chinese Medicine. The LOVO cells introduced with sh-circPACRGL and sh-NC were subcutaneously inoculated into mice. The tumor volume was calculated every 7 days for 30 days using the formula: 0.5 × length × wide2. Finally, the mice were euthanized and the tumors were weighed.
Statistical analysis
The data are presented as the mean ± standard deviation (SD) using the GraphPad Prism (GraphPad Software, San Diego, CA, USA). Student's t test was used for the comparison, and the correlation relationship among the expression of miR-330-5p, circPACRGL and CNBP was analyzed by Pearson's correlation analysis. P < 0.05 was considered statistically significant.
Results
CircPACRGL was highly expressed in CRC tissues and cells
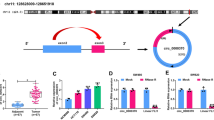
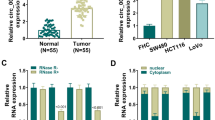
Here, 39 pairs of CRC tumor tissues and adjacent normal tissues, as well as CRC cell lines (SW480, SW620, LOVO and HT29) and normal cells (NCM460 cells) were used. As displayed in Fig. 1A and B, the analysis of qRT-PCR suggested that circPACRGL expression was remarkably enhanced in CRC tissues and cells as against with adjacent tissues and cells. We selected SW480 and LOVO cells for followed experiments. CircPACRGL is derived from the PACRGL gene (Fig. 1C), and harbors a loop structure in SW480 and LOVO cells (Fing 1D). Moreover, circPACRGL is more stable than linear PACRGL through actinomycin D treatment (Fig. 1E). These results suggested that circPACRGL played a role in CRC.
CircPACRGL was overexpressed in CRC tissues and cells. A The expression of circPACRGL was measured in tumor tissues and adjacent tissues in CRC. B The expression of circPACRGL was detected in CRC cells and normal cells. C The genomic loci of PACRGL and circPACRGL. D The expression of circPACRGL and liner PACRGL was analyzed after treatment with RNase R in SW380 and LOVO cells. E The expression of circPACRGL and liner PACRGL was analyzed after treatment with RNase R at 6, 12, 24 h in SW380 and LOVO cells
Knockdown of circPACRGL inhibited cell growth in CRC
To investigate the function of circPACRGL, sh-circPACRGL and sh-NC were introduced into SW480 and LOVO cells, and sh-circPACRGL sharply inhibited the expression of circPACRGL in SW480 and LOVO cells (Fig. 2A). The analysis of MTT assay and colony formation assay determined that inhibition of circPACRGL significantly reduced cell viability in SW480 and LOVO cells (Fig. 2B and C). Furthermore, EDU, transwell and wound healing assays determined that sh-circPACRGL transfection inhibited cell invasion and migration (Fig. 2D–F). Moreover, the analysis of flow cytometry showed that cell apoptosis was promoted by sh-circPACRGL transfection (Fig. 2G). As displayed in Fig. 2H, the protein levels of CDK4, MMP3, and Bcl-2 were obviously suppressed while the protein level of Bax was significantly elevated in the sh-circPACRGL group (Fig. 2H). Thus, the reduction of circPACRGL could decrease cell proliferation, invasion and migration in CRC.
Knockdown of circPACRGL inhibited cell growth in CRC. A and B qRT-PCR displayed that circPACRGL expression was inhibited in SW480 and LOVO cells with transfection of sh-circPACRGL. B, C and D MTT assay B, colony formation C and EDU assay D determined that cell proliferation was suppressed in SW480 and LOVO cells with transfection of sh-circPACRGL. E and F Transwell assay E and wound healing assay F were used to detect cell invasion and migration in SW480 and LOVO cells. G Flow cytometry suggested that cell apoptosis was increased in SW480 and LOVO cells with transfection of sh-circPACRGL. H Western blot indicated that the protein expression of CDK4, MMP3, and Bcl-2 was reduced while Bax protein expression was induced in SW480 and LOVO cells with transfection of sh-circPACRGL. *P < 0.05
CNBP was involved in the cellular progression of CRC
As shown in Fig. 3A-D, CNBP level was heightened in CRC tissues and cells, and strongly connected with circPACRGL, implying CNBP had roles in CRC cell progression. Knocked down CNBP could inhibit cell proliferation, migration and invasion significantly, while it induced cell apoptosis in SW480 and LOVO cells (Fig. 4A–G). Meanwhile, decreasing the expression of CNBP could reduce the protein expressions of CDK4, MMP9, and Bcl-2, and increase the protein expression of Bax in SW480 and LOVO cells (Fig. 4H). These effects were consistent with the effect of sh-circPACRGL on CRC cells. Therefore, it is reasonable to speculate that circPACRGL and CNBP may have some relationship in the regulation of CRC cell growth.
CNBP was involved in the cellular progression of CRC. A qRT-PCR displayed that circPACRGL expression was inhibited in SW480 and LOVO cells with transfection of sh-CNBP. B, C, and D MTT assay B, colony formation C and EDU assay D determined that cell proliferation was suppressed in SW480 and LOVO cells with transfection of sh-CNBP. E and F Transwell assay E and wound healing assay F were used to detect cell invasion and migration in sh-CNBP and sh-NC groups of SW480 and LOVO cells. G Flow cytometry suggested that cell apoptosis was increased in SW480 and LOVO cells with transfection of sh-CNBP. H Western blot indicated that the protein expression of CDK4, MMP3, and Bcl-2 was reduced while Bax protein expression was induced in SW480 and LOVO cells with transfection of sh-CNBP. *P < 0.05
Promotion of CNBP could reverse the suppressive effects of low circPACRGL on cell growth
To verify the relation of circPACRGL and CNBP in CRC, rescue experiments was applied and sh-circPACRGL was cotransfected with pcDNA-NC or pcDNA-CNBP into SW480 and LOVO cells. As displayed in Fig. 5A, the protein expression of CNBP was inhibited by sh-circPACRGL transfection, which was reversed by up-regulation of CNBP in SW480 and LOVO cells. More than that, reduction of circPACRGL repressed cell proliferation, invasion and migration but facilitated cell apoptosis, which was impaired by induction of CNBP in SW480 and LOVO cells (Fig. 5B - H). Besides, the influences of sh-circPACRGL on CDK4, MMP9, Bax, and Bcl-2 protein were overturned by pcDNA-CNBP (Fig. 5I). Thus, circPACRGL regulated cell growth by modulating CNBP expression in CRC.
Promotion of CNBP could reverse the suppressive effects of low circPACRGL on cell growth. A pcDNA-CNBP transfection induced CNBP expression in SW380 and LOVO cells by using western blot. B and C Western blot determined that the protein level of CNBP was repressed by sh-circPACRGL transfection, which increased by pcDNA-CNBP transfection in SW480 B and LOVO C cells. D and E colony formation assay D and EDU assay E showed that cell proliferation was inhibited by sh-circPACRGL transfection, which increased by pcDNA-CNBP transfection in SW480 and LOVO cells. F and G Transwell assay F and wound healing assay G showed that cell migration and invasion were reduced by sh-circPACRGL transfection, which was induced by pcDNA-CNBP transfection in SW480 and LOVO cells. H Flow cytometry showed that cell apoptosis was increased by circPACRGL transfection, which was inhibited by pcDNA-CNBP transfection in SW480 and LOVO cells. I Western blot indicated that the protein expression of CDK4, MMP3, and Bcl-2 was reduced while Bax protein expression was induced in SW480 and LOVO cells with transfection of sh-circPACRGL, which was reversed by pcDNA-CNBP transfection in SW480 and LOVO cells. *P < 0.05
CircPACRGL regulated CRC cell progression through miR-330-3p/CNBP axis
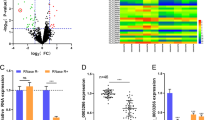
To further study the regulatory mechanism of circPACRGL in CRC, the prediction of the bioinformatics analysis showed that circPACRGL has binding sites with miR-330-3p (Fig. 6E) and CNBP was the target of miR-330-3p (Fig. 6F). In addition, miR-330-3p expression was significantly decreased in CRC tissues and cells (Fig. 6A and B). Besides, miR-330-3p was inversely associated with circPACRGL and CNBP in CRC tissues (Fig. 6C and D). Moreover, luciferase activity was markedly downregulated when the miR-330-3p was binding to circPACRGL wt or CNBP 3’UTR wt, but not circPACRGL mut or CNBP 3’UTR mut (Fig. 6G and I). In addition, RIP assay showed that anti-Ago2 antibody could decrease the levels of endogenous circPACRGL and miR-330-3p in SW480 and LOVP cells (Fig. 6H). Moreover, CircPACRGL protein expression was inhibited by down-regulation of circPACRGL, which was overturned by inhibition of miR-330-3p (Fig. 6J). Thus, the data determined that circPACRGL regulated CNBP through sponging miR-330-3p in CRC.
circPACRGL regulated CRC cell progression through miR-330-3p/CNBP axis. A and B The expression of miR-330-3p was measured in tumor tissues and cells. C Pearson's correlation analysis was used to determine the relationship between miR-330-3p and circPACRGL in CRC tissues. D Pearson's correlation analysis was used to determine the relationship between miR-330-3p and CNBP in CRC tissues. E The bioinformatics analysis predicted that miR-330-3p was a target miRNA of circPACRGL through binding sites. F The bioinformatics analysis predicted that CNBP was a target miRNA of miR-330-3p through binding sites. G and I Luciferase reporter assay determined that the luciferase activity was decreased when miR-330-3p binding to circPACRGL wt G and CNBP 3’UTR wt I, not circPACRGL mut and CNBP 3’UTR mut in SW480 and LOVO cells. H RIP assay determined that circPACRGL and miR-330-3p were enriched in SW480 and LOVO cells. J Western blot determined that the expression of CNBP was decreased by sh-circPACRGL transfection, which was reversed by miR-330-3p transfection in SW480 and LOVO cells. *P < 0.05
Knockdown of circPACRGL inhibited tumor progression in vivo
To further investigate whether circPACRGL affected tumor growth, we established mouse models by subcutaneous injection of sh-NC and sh-circPACRGL transfection, and measured tumor volume every seven days. The results showed that the tumor volume was significantly reduced after circPACRGL was knocked out (Fig. 7A). After 27 days of injection, the tumor was removed and weighed, and it was found that the silencing of circPACRGL resulted in tumor reduction (Fig. 7B). In addition, inhibition of circPACRGL can decrease the expression of circPACRGL and CNBP, and increase the level of miR-330-3p (Fig. 7C and D). IHC analysis showed that CNBP, ki-67, MMP9, and MMP2 were decreased in the sh-circPACRGL group as opposed to that in the sh-NC group, indicating that knockdown of circPACRGL could inhibit tumor proliferation and invasion in vivo (Fig. 7E). Therefore, circPACRGL silencing inhibited tumor growth in CRC by regulating miR-330-3p/CNBP axis.
Knockdown of circPACRGL inhibited tumor progression in vivo. A Photograph illustrated tumors from xenograft mice. And the growth curve of tumor volume was displayed. B Sh-circPACRGL resulted in a decline of tumor weight. C and D Knockdown of circPACRGL inhibited circPACRGL and CNBP expression, and induced miR-330-3p expression. E IHC analysis showed that CNBP, ki-67, MMP9, and MMP2 were decreased in the sh-circPACRGL group. *P < 0.05.
Discussion
Herein, circPACRGL expression was elevated in CRC tissues and cells, which was consistent with preceding research [14]. Moreover, circPACRGL silencing impeded colorectal cancer cell proliferation, migration and invasion, and promoted apoptosis in CRC cells.
It is well known that each circRNA does not play roles in cancer alone, which has the most important role to act as ceRNAs, which bind to miRNAs to regulate mRNA expression; thereby, affecting the translation of downstream proteins. Herein, we found that miR-330-3p is downregulated in colorectal cancer tissues and cells, and further confirmed that miR-330-3p is a miRNA of circPACRGL. MiR-330-3p was reported to play multiple functions in human diseases and was closely linked with cell progression and poor prognosis [24,25,26,27,28,29]. Thus, miR-330-3p is essential in the regulatory network of circPACRGL. Meanwhile, we further predicted and verified that CNBP is the target of miR-330-3p. Nora B Calcaterra et al. summarized that CNBP is a multifunctional nucleic acid chaperone involved in cell death and proliferation and may be related to transcriptional and/or translational level regulation [30]. More than that, CNBP was also related to tumorigenesis and development, which directly regulate tumor-promoting gene expression [31]. Moreover, we found that circPACRGL affected the expression of miR-330-3p and miR-330-5p regulated the expression of CNBP in CRC cells. Based on the above evidence and findings, we predicted that circPACRGL regulates cell growth and apoptosis by modulating the miR-330-3p/CNBP axis. Further reverse experiments showed that promotion of CNBP could reverse the oppressive effects of low circPACRGL on cell growth. Thus, circPACRGL promoted cell proliferation, migration and invasion but inhibited cell apoptosis in colorectal cancer via regulation of the miR-330-3p/CNBP axis. Whether the circPACRGL/miR-330-3p/CNBP axis has a role in other tumors is unclear, which could be explored in the future.
In this paper, circPACRGL functioned as a miR-330-3p sponge molecular to elevate CNBP expression, resulting in the promotion of CRC progression, which expanded the understanding of circRNA-mediated CRC progression and supported circPACRGL as a potential therapeutic target in CRC. The weakness of this study is the small clinical sample size. In the future, the function of circPACRGL needs to be validated and the signaling pathways it regulates and whether it regulates CRC growth through other ceRNA pathways need to be investigated.
Data Availability
Not Applicable.
References
Zhao P, Dai M, Chen W, Li N (2010) Cancer Trends in China. Jpn J Clin Oncol 40:281–285
Vega P, Valentin F, Cubiella J (2015) Colorectal cancer diagnosis: pitfalls and opportunities. World Journal of Gastrointestinal Oncology 7:422–433
Whittemore AS, Wuwilliams AH, Lee MM, Shu Z, Gallagher RP, Dengao J, Lun Z, Xianghui W, Kun C, Jung DL (1990) Diet, physical activity, and colorectal cancer among Chinese in North America and China. J Natl Cancer Inst 82:915–926
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S (2014) circRNA Biogenesis Competes with Pre-mRNA Splicing. Mol Cell 56:55–66
Ebbesen KK, Hansen TB, Kjems J (2017) Insights into circular RNA biology. RNA Biol 14:1035–1045. https://doi.org/10.1080/15476286.2016.1271524
Frankel LB, Lund AH (2012) MicroRNA regulation of autophagy. Carcinogenesis 33:2018–2025
Kristensen LS, Hansen TB, Venø MT, Kjems J (2018) Circular RNAs in cancer: opportunities and challenges in the field. Oncogene 37:555–565
Meng S, Zhou H, Feng Z, Xu Z, Tang Y, Li P, Wu M (2017) CircRNA: functions and properties of a novel potential biomarker for cancer. Mol Cancer 16:94. https://doi.org/10.1186/s12943-017-0663-2
Pervouchine DD (2019) Circular exonic RNAs: When RNA structure meets topology. Biochim Biophys Acta Gene Regul Mech 1862:194384. https://doi.org/10.1016/j.bbagrm.2019.05.002
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao L, Zhang Y, Wu YM, Dhanasekaran SM, Engelke CG, Cao X, Robinson DR, Nesvizhskii AI, Chinnaiyan AM (2019) The Landscape of Circular RNA in Cancer. Cell 176:869-881.e13. https://doi.org/10.1016/j.cell.2018.12.021
Yu T, Wang Y, Fan Y, Fang N, Wang T, Xu T, Shu Y (2019) CircRNAs in cancer metabolism: a review. J Hematol Oncol 12:90. https://doi.org/10.1186/s13045-019-0776-8
Zhang H, Wang G, Ding C, Liu P, Wang R, Ding W, Tong D, Wu D, Li C, Wei Q (2017) Increased circular RNA UBAP2 acts as a sponge of miR-143 to promote osteosarcoma progression. Oncotarget 8:61687–61697
Liu Z, Yu Y, Huang Z, Kong Y, Hu X, Xiao W, Quan J, Fan X (2019) CircRNA-5692 inhibits the progression of hepatocellular carcinoma by sponging miR-328-5p to enhance DAB2IP expression. Cell Death Dis 10:900. https://doi.org/10.1038/s41419-019-2089-9
Li H, Jin X, Liu B, Zhang P, Chen W, Li Q (2019) CircRNA CBL.11 suppresses cell proliferation by sponging miR-6778–5p in colorectal cancer. BMC Cancer 19:826. https://doi.org/10.1186/s12885-019-6017-2
Shang A, Gu C, Wang W, Wang X, Sun J, Zeng B, Chen C, Chang W, Ping Y, Ji P, Wu J, Quan W, Yao Y, Zhou Y, Sun Z, Li D (2020) Exosomal circPACRGL promotes progression of colorectal cancer via the miR-142-3p/miR-506-3p- TGF-β1 axis. Mol Cancer 19:117. https://doi.org/10.1186/s12943-020-01235-0
Xu H, Liu Y, Cheng P, Wang C, Liu Y, Zhou W, Xu Y, Ji G (2020) CircRNA_0000392 promotes colorectal cancer progression through the miR-193a-5p/PIK3R3/AKT axis. J Exp Clin Cancer Res 39:283. https://doi.org/10.1186/s13046-020-01799-1
Zhang HD, Jiang LH, Sun DW, Hou JC, Ji ZL (2017) CircRNA: a novel type of biomarker for cancer. Breast Cancer 25:1–7
Chen D, Chenyue Z, Jiamao L, Xinyu S, Haiyong W (2018) Screening differential circular RNA expression profiles reveal that hsa_circ_0128298 is a biomarker in the diagnosis and prognosis of hepatocellular carcinoma. Cancer Manag Res 10:1275–1283
Yang H, Li X, Meng Q, Sun H, Wu S, Hu W, Liu G, Li X, Yang Y, Chen R (2020) CircPTK2 (hsa_circ_0005273) as a novel therapeutic target for metastatic colorectal cancer. Mol Cancer 19:13. https://doi.org/10.1186/s12943-020-1139-3
Allantaz F, Cheng DT, Bergauer T, Ravindran P, Rossier MF, Ebeling M, Badi L, Reis B, Bitter H, D’Asaro M (2012) Expression profiling of human immune cell subsets identifies miRNA−mRNA regulatory relationships correlated with cell type specific expression. PLoS ONE 7:e29979
Feng Z, Ruyou X, Yingnan W, Xiaoying L, Shuang Z, Wenying H (2018) Comprehensive analysis of circRNA expression pattern and circRNA-miRNA-mRNA network in the pathogenesis of atherosclerosis in rabbits. Aging 10(9):2266–2283
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Yao S, Zheng P, Wu H, Song LM, Ying XF, Xing C, Li Y, Xiao ZQ, Zhou XN, Shen T, Chen L, Liu YH, Lai MD, Mei L, Gao TM, Li JM (2015) Erbin interacts with c-Cbl and promotes tumourigenesis and tumour growth in colorectal cancer by preventing c-Cbl-mediated ubiquitination and down-regulation of EGFR. J Pathol 236:65–77. https://doi.org/10.1002/path.4502
Chen T, Yang Z, Liu C, Wang L, Yang J, Chen L, Li W (2019) Circ_0078767 suppresses non-small-cell lung cancer by protecting RASSF1A expression via sponging miR-330-3p. Cell Prolif 52:e12548. https://doi.org/10.1111/cpr.12548
Huang Y, Sun H, Ma X, Zeng Y, Pan Y, Yu D, Liu Z, Xiang Y (2020) HLA-F-AS1/miR-330-3p/PFN1 axis promotes colorectal cancer progression. Life Sci 254:117180. https://doi.org/10.1016/j.lfs.2019.117180
Jin Z, Jia B, Tan L, Liu Y (2019) miR-330-3p suppresses liver cancer cell migration by targeting MAP2K1. Oncol Lett 18:314–320. https://doi.org/10.3892/ol.2019.10280
Shen L, Yi S, Huang L, Li S, Bai F, Lei S, Breitzig M, Czachor A, Sun H, Zheng Q, Wang F (2019) miR-330-3p promotes lung cancer cells invasion, migration, and metastasis by directly targeting hSOD2b. Biotechnol Appl Biochem 66:21–32. https://doi.org/10.1002/bab.1691
Wei CH, Wu G, Cai Q, Gao XC, Tong F, Zhou R, Zhang RG, Dong JH, Hu Y, Dong XR (2017) MicroRNA-330-3p promotes cell invasion and metastasis in non-small cell lung cancer through GRIA3 by activating MAPK/ERK signaling pathway. J Hematol Oncol 10:125. https://doi.org/10.1186/s13045-017-0493-0
Wang H, Chen SH, Kong P, Zhang LY, Zhang LL, Zhang NQ, Gu H (2018) Increased expression of miR-330-3p: a novel independent indicator of poor prognosis in human breast cancer. Eur Rev Med Pharmacol Sci 22:1726–1730. https://doi.org/10.26355/eurrev_201803_14587
Calcaterra NB, Armas P, Weiner AM, Borgognone M (2010) CNBP: a multifunctional nucleic acid chaperone involved in cell death and proliferation control. IUBMB Life 62:707–714. https://doi.org/10.1002/iub.379
Lee EH, Lee TA, Yoo HJ, Lee S, Park B (2019) CNBP controls tumor cell biology by regulating tumor‐promoting gene expression. Molecular Carcinogenesis.
Acknowledgements
None.
Funding
This work was supported by Funded project of “Double first-class” discipline (Traditional Chinese Medicine) construction in Jiangxi Province [NO.JXSYLXK-ZHY1054]; Science and Technology Research Project of Education Department of Jiangxi Province [No.201210, 200143]; Scientific planning and Planning Fund project of Jiangxi Health Commission [NO.20195637, 20165401]; Science and Technology Project of Jiangxi Administration of Traditional Chinese Medicine [NO.2019A106]; 2019 University-level Innovation and Entrepreneurship Training Plan for College Students of Jiangxi University of Chinese Medicine [NO.201910412045]; Csco-Hengrui Cancer Research Fund (Y-HR2017-128).
Author information
Authors and Affiliations
Contributions
All authors have been involved in the management of the patient and in the conception of the manuscript. The specific authors’ contribution is as follows. Haiyun Liu and Zhijian He: writing the drafting of the manuscript. Qianxia Lin, Wenchun Zhang and Xiaomin Wang: data collection. Yulong Ji, Yongqing Fang and Yuan Hu: analysis and interpretation. Xinrong Hu: software, reviewing and editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This study was permitted by the Ethics Committee of Jiangxi University of Chinese Medicine.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, H., Fang, Y., Hou, B. et al. CircPACRGL promoted cell proliferation, migration and invasion as well as inhibited cell apoptosis in colorectal cancer via regulation of the miR-330-3p/CNBP axis. Mol Cell Biochem 478, 1633–1644 (2023). https://doi.org/10.1007/s11010-022-04543-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-022-04543-9