Abstract
Context
Species are influenced by factors operating at multiple scales, but multi-scale species distribution and abundance models are rarely used. Though multi-scale species distribution models outperform single-scale models, when compared through model selection, multi- and single-scale models built with computer learning algorithms have not been compared.
Objectives
We compared the performance of models using a simple and accessible, multi-scale, machine learning, species distribution and abundance modeling framework to pseudo-optimized and unoptimized single-scale models.
Methods
We characterized environmental variables at four spatial scales and used boosted regression trees to build multi-scale and single-scale distribution and abundance models for 28 bird species. For each species and across species, we compared the performance of multi-scale models to pseudo-optimized and lowest-performing unoptimized single-scale models.
Results
Multi-scale distribution models consistently performed as well or better than pseudo-optimized single-scale models and significantly better than unoptimized single-scale models. Abundance model performance showed a similar, but less pronounced pattern. Mixed-effects models, that controlled for species, provided strong evidence that multi-scale models performed better than unoptimized single-scale models. Although mean improvement in model performance across species appeared minor, for individual species, arbitrary selection of scale could result in discrepancies of up to fourteen percent for area of suitable habitat and population estimates.
Conclusions
Scale selection should be explicitly addressed in distribution and abundance modeling. The multi-scale species distribution and abundance modeling framework presented here provides a concise and accessible alternative to standard pseudo-scale optimization while addressing the scale-dependent response of species to their environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Predictions of species’ distributions and abundances are valuable in addressing current and future conservation challenges. Species distribution models (SDMs) that tie species occurrence throughout a landscape to the environment are now common practice (e.g. Fournier et al. 2017; Dalgarno et al. 2017; Evangelista et al. 2018; Reino et al. 2018). High-resolution land-use and land-cover data that can inform SDMs are freely available (United States Geological Survey 2011). For many species, the high resolution of current land cover data may be too small to capture species-environment relationships (e.g. species respond to larger scales than the 30 m grid cells). To investigate such relationships, researchers must recharacterize environmental data to more biologically appropriate scales.
The appropriate scale at which to characterize environmental covariates is species- and process-dependent (Wiens 1989; Wiens and Milne 1989; Levin 1992; Mayor et al. 2009; McGarigal et al. 2016). Within a single species and ecological process (e.g. occurrence or abundance), the results from one scale may not translate to another (Mayor et al. 2009). To characterize the comprehensive influence of an environmental covariate, multiple scales must be examined.
Scale optimization is the specific consideration and evaluation of the spatial scale at which a species most strongly responds to an environmental covariate. In its simplest form, ‘pseudo-scale optimization’ is performed by comparing the performance of models (e.g. AUC or AIC) with environmental variables characterized at different scales (McGarigal et al. 2016). More commonly, scale optimization is conducted in multiple steps (Boscolo and Metzger 2009; Pennington and Blair 2011; Timm et al. 2016; Stevens and Conway 2020). First, each environmental covariate is modeled at different spatial scales and model performance is compared. The scale of each covariate’s top performing model is considered that covariate’s optimal scale. These optimized covariates can then be combined into multi-scale models, which can increase model predictive performance (McGarigal et al. 2016; Timm et al. 2016). Though cumbersome, scale optimized multi-scale models outperform pseudo-scale optimized models (Stevens and Conway 2019). This scale optimized multi-scale framework can be adapted to SDMs (e.g. Stevens and Conway 2019; Stevens and Conway 2020). More sophisticated models that allow for a change in the strength of environmental covariate influence have been recently developed as well (Miguet et al. 2017; Moll et al. 2020).
SDMs run with machine learning algorithms, such as boosted regression trees (BRTs), seem to work under a completely different paradigm. First, model selection is rarely a component. Second, though multi-scale SDMs are used (e.g. Shirley et al. 2013; Halstead et al. 2019; Hallman and Robinson 2020), the multi-scale nature of the models is rarely discussed explicitly. Third, since BRTs are robust to overfitting, many environmental covariates characterized at multiple scales are included in the same model. BRTs can also be used to model abundance (Wilsey et al. 2016), with zero-inflated BRTs more appropriately dealing with the many zeroes associated with avian count data (Johnston et al. 2015). To our knowledge, no one has explicitly compared multi-scale and single-scale SDMs or abundance models run with the BRT algorithm.
Though multi-scale models are clearly important (Baladrón et al. 2016; Benítez-López et al. 2017), studies frequently select a scale without direct examination of the scale-dependence of a species’ response (McGarigal et al. 2016). We use a landscape-scale study of birds in western Oregon to explore the performance of multi-scale SDMs and abundance models in a boosted regression tree (BRT) framework. We assess the range of improvement in model performance from single- to multi-scale models by explicitly comparing our unoptimized (worst performing) single-scale models and our pseudo-optimized (best performing) single-scale models, to multi-scale models. Further, we assess the strength of the species-specificity of this improvement. Most importantly, we provide an easy to use method of multi-scale SDM and abundance modeling that we hope increases the accessibility of multi-scale models.
Methods
Count data, location, and study species
We studied 28 species of passerine that were specifically selected to encompass a variety of habitat preference and prevalence, factors that are known to affect species distribution model performance (Hallman and Robinson 2020). Point counts were conducted from 2011 to 2013 at 2231 locations across Benton and Polk Counties, Oregon. Benton and Polk counties fall within Oregon’s Coast Range and Willamette Valley ecoregions (Thorson et al. 2003). Land cover within these counties consists primarily of wet coniferous forests in the coastal mountain range and a mix of agricultural and urbanized lands in the Willamette Valley. Coast Range forests are a mix of federal and private lands and include the full range of forest ages from recently harvested to old-growth.
All avian point counts were conducted by (author) WDR. Roadside counts were spaced every 0.8 km along accessible roads. Off-road counts were conducted along 200 m grids in two National Wildlife Refuges. Roadside counts were primarily along narrow, lightly traveled rural logging and farming roads. Each count lasted 5 min and was visited once during the study. All birds seen or heard at all distances from observer were included. Field methods to account for imperfect detection were implemented, but for this study we chose to use raw unadjusted counts to make modeling methods applicable to the majority of freely available avian occurrence and abundance data (Sullivan et al. 2009; Hudson et al. 2017).
Environmental data
We compiled 25 freely available rasters of environmental variables to use in our models (Table 1). We downloaded 30-m resolution Oregon land cover data from United States Geological Survey’s National Gap Analysis Project (United State Geological Survey 2011) and selected 13 land cover classifications to use as model variables. We downloaded 30-m resolution data on forest structure and composition from Landscape Ecology, Modeling, Mapping and Analysis’s gradient nearest neighbor structure maps (Landscape Ecology, Modeling, Mapping, and Analysis 2014). We selected eight variables describing forest structure and composition to include in our models. We downloaded a 10-m resolution Digital Elevation Model from the Oregon Spatial Data Library (Oregon Spatial Data Library 2017), rescaled it to a matching 30-m resolution, and used it to calculate slope and compound topographic index (Yang et al. 2005), an index of topographic soil wetness. We used elevation, slope, and compound topographic index as the topographic variables in our models. We also downloaded a shapefile of Oregon’s waterways from the Oregon Spatial Data Library (Oregon Spatial Data Library 2016) and calculated a 30-m resolution distance to water as a final environmental variable in our models.
We characterized our environmental variables at four distinct scales (Boscolo and Metzger 2009; Shirley et al. 2013). We used focal statistics in ArcMap to calculate mean values for structural and topographic variables at 165 m, 315 m, 615 m, and 1215 m radii from cell centers. For land cover variables, we used focal statistics to calculate percent land cover at each of the same four scales. These radii were selected to characterize the environmental variables at scales ranging from the local point count scale to small landscapes. They are large enough to include multiple territories of all species and are within a range of scales previously shown to be important to territorial songbirds (Boscolo and Metzger 2009; Baladrón et al. 2016). Furthermore, smaller scales would not be adequately represented by the unadjusted unlimited-distance counts used here as many detected birds would fall outside of the area in which habitat was characterized.
SDMs and abundance model framework
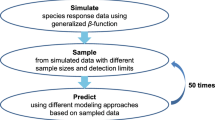
We used BRTs for occurrence and abundance modeling (Fig. 1; Elith et al. 2008). Although there is no single best SDM method (Qiao et al. 2015), BRTs are often among the top performing SDM algorithms (Guisan et al. 2007; Elith and Graham 2009; García-Callejas and Araújo 2016). For each model run, we withheld a random 20 percent of the data for model evaluation. We used the remaining 80 percent for model training. When comparing model performance more generally, it is better to use a checkerboard approach to partition data into training and evaluation sets (Wilsey et al. 2016). In this study, however, because our objective is to compare models from the same dataset, all data are subject to the same spatial structure, and checkerboarding is unnecessary. To evaluate the stochasticity of splitting the data into training and evaluation data sets, as well as the inherent stochasticity of boosted regression trees, we ran 20 iterations of each model (Barker et al. 2014; Wilsey et al. 2016). We optimized learning rates of boosted regression trees to allow best models of a minimum of 1000 trees. We held tree complexity and bag fraction constant at 3 and 0.75, respectively (Elith et al. 2008; Johnston et al. 2015). We used AUC (area under the curve) to evaluate SDM performance. Although AUC may be an inappropriate metric of true SDM performance (Lobo et al. 2008), it is a valid measure of SDM performance within a single species and study extent (Edrén et al. 2010).
We ran abundance models using a zero-inflation framework (Fig. 1; Johnston et al. 2015). Hurdle models such as these require data to pass an initial “hurdle” before proceeding with the model (Kulhanek et al. 2011; Oppel et al. 2012). Specifically, we (1) ran SDMs as above, (2) used species prevalence as a threshold to convert continuous habitat suitability to binomial suitable and unsuitable habitat (Liu et al. 2005) (3) restricted our data to suitable habitat, then (4) ran Poisson BRTs with the same parameters as the above SDMs to model abundance within suitable habitat. Due to outliers and non-normal distributions of data, we evaluated abundance models with Kendall’s rank correlation. True abundance at any site is an unknown latent variable (Welsh et al. 2013). Without accounting for imperfect detection, we estimated an index of abundance (i.e. relative abundance; Wilsey et al. 2016). For brevity, however, we use the term abundance throughout.
Pseudo-scale optimization
For each species, we conducted pseudo-scale optimization by running single-scale SDMs at each of the five environmental scales (McGarigal et al. 2016). The single-scale SDM with the greatest mean AUC was designated the pseudo-optimized single-scale SDM. The single-scale SDM with the lowest mean AUC was designated the unoptimized single-scale SDM. We developed multi-scale models by running SDMs that included all environmental variables at all scales. For abundance models, we conducted pseudo-scale optimization as described above using Kendall’s rank correlation as the metric of model performance. As above, the models with the greatest and lowest mean Kendall’s rank correlations were designated the pseudo-optimized and unoptimized single-scale abundance models, respectively. We developed multi-scale abundance models by running Poisson BRTs that included all environmental variables at all scales.
Multi-scale and single-scale model comparison
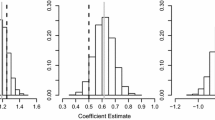
We used mixed-effects models to compare multi-scale, unoptimized single-scale, and pseudo-optimized single-scale model performance. We assigned multi-scale, unoptimized single-scale, and pseudo-optimized single-scale to each SDM or abundance model output as a categorical variable. We treated species as a random effect and used AUC and Kendall’s rank correlation as the dependent variable for SDMs and abundance models, respectively. Random effects account for species-specific differences in the predictability of occurrence and abundance, which is not of interest in this study. Significantly positive coefficients indicate higher performance for that subset of models. For mixed-effects models with AUC as the dependent variable, we used beta distributions, but results did not differ, so we report only normal mixed-effects models here.
To further clarify the differences between multi-scale SDMs, pseudo-optimized single-scale SDMs, and unoptimized single-scale SDMs, we calculated mean AUC for each species and the percent difference from the species mean AUC for each model. The species mean AUC was defined as the mean AUC from all models of a single species. By calculating percent difference from the mean performance of each species, species-specific effects are mitigated and contrasts between the performance of multi-scale models and single-scale models across species are more evident. We used an identical approach to examine differences in Kendall’s rank correlation from abundance models. Finally, we used t-tests to compare AUCs from multi-scale SDMs to unoptimized and pseudo-optimized single-scale SDMs for each species individually. Similarly, we used t-tests to compare Kendall’s rank correlations from multi-scale abundance models to unoptimized and pseudo-optimized single-scale abundance models for each species individually.
Multicollinearity and overfitting
Because multi-scale models include each covariate at each scale, the number of covariates can be high and multicollinearity and overfitting must be considered. Multicollinearity is not problematic for predictive models if the collinearity structure is retained in the test and training datasets (Dormann et al. 2012). Though BRTs are robust to overfitting, due to the high number of variables in multi-scale models, we felt that it was essential to explicitly test for overfitting. To do this, we ran models with the full environmental covariates and two reduced covariates sets: (1) only the most influential covariates and (2) only three topographic covariates (Shirley et al. 2013). To select the most influential covariates for each species, we used the relative influence of each covariate on occurrence or abundance (Elith et al. 2008; Illan et al. 2014). Due to the stochasticity inherent in BRT and data splitting, relative covariate influence changed from iteration to iteration. To address this, we counted the number of times each covariate was within the top five influential covariates in the 20 iterations of the full model. We did this for occurrence and abundance separately at each of the four scales. Any covariate that was within the top five influential covariates in at least five of the 20 models was added to the influential environmental covariate models. The number of influential environmental covariates ranged from 6 to 11 and 7 to 11 in SDMs and abundance models, respectively.
To check for overfitting in SDMs, we ran a mixed-effects model on the AUCs of multi-scale SDMs built on the three separate covariate sets: influential, topographic, and full. Species was treated as a random effect to control for species-specific differences. To check for overfitting in the abundance models, we ran a similar mixed-effects model on the Kandall’s rank correlation for multi-scale models built on the three covariates sets with species as a random effect. In both mixed-effects models, higher model performance in the influential or topographic covariate sets, which have far fewer variables, would indicate overfitting in the full covariate models.
Results
Species distribution models
The mixed-effects model comparing the AUC of multi-scale, unoptimized, and pseudo-optimized single-scale models showed a strong negative effect of unoptimized single-scale models across species (Table 2). Contrastingly, there was only a minor difference in multi-scale and pseudo-optimized single-scale models. At the individual species level, multi-scale SDMs performed significantly better than unoptimized single-scale SDMs in 89% (25/28) of our study species (Fig. 2, Online Resource 1). Multi-scale SDMs performed significantly better than pseudo-optimized single-scale SDMs in only 21% (6/28) of our study species. Multi-scale SDMs had higher mean AUC than pseudo-optimized single-scale models in 93% (26/28) of our study species, and never performed significantly worse than single-scale models.
Mean model performance (AUC) and 95% confidence intervals for the unoptimized single-scale, pseudo-optimized single-scale, and multi-scale SDMs for 28 passerine species. Significantly higher performance of multi-scale models than unoptimized and pseudo-optimized single-scale models indicated by * and **, respectively
Mean multi-scale AUCs ranged from 0.70 to 0.94 with an overall mean AUC of 0.81 across species (Figs. 2 & 3a, Online Resource 2). Sixty-eight percent (19/28) of our study species had mean multi-scale AUCs above 0.80, and 14% (4/28) had mean multi-scale AUCs above 0.90. Both overall SDM performance and the improvement in AUC between single-scale and multi-scale models was species-specific. The mean improvement in AUC between pseudo-optimized single-scale and multi-scale SDMs was 0.008 and the maximum improvement for any species was 0.034; improvements of one and four percent, respectively. The mean improvement in AUC between unoptimized single-scale and multi-scale SDMs was 0.031, with a maximum improvement of 0.112; improvements of 4 and 14%, respectively. We found no evidence of overfitting in any of our SDMs or abundance models (Online Resource 3).
Boxplots of a mean percent difference in model performance (AUC) from overall species mean in unoptimized single-scale, pseudo-optimized single-scale, and multi-scale SDMs for 28 species of passerine and b mean percent difference in model performance (Kendall’s Predictive Correlation) from overall species mean in unoptimized single-scale, pseudo-optimized single-scale, and multi-scale abundance models for 28 species of passerine
Although we found a clear scale-dependence of species’ response to the environment across species, the strength of that dependence was strongly species-specific (Online Resource 2). Among six species of warbler, two and three pseudo-optimized single-scales were identified for SDMs and abundance models, respectively. Likewise, three species of wren had three pseudo-optimized single-scales for both SDMs and abundance models. We found a negative relationship between scale and the number of species for which that scale was best. For example, in our SDMs there were eleven species that had a pseudo-optimized single-scale of 165 m, while only two had a pseudo-optimized single-scale of 1215 m.
Abundance models
The mixed-effects model comparing the Kendall’s rank correlations of multi-scale, unoptimized single-scale, and pseudo-optimized single-scale models showed a negative effect of unoptimized single-scale models across species (Table 2). There was nearly no difference in multi-scale and pseudo-optimized single-scale models. Multi-scale abundance models performed significantly better than unoptimized single scale abundance models in 39% (11/28) of our study species (Fig. 4, Online Resource 3). Multi-scale abundance models performed significantly better than pseudo-optimized single-scale abundance models in only one species but never performed significantly worse than single-scale models. The pseudo-optimized single-scale for abundance was species-specific and differed from the pseudo-optimized single-scale for SDMs in 50% (14/28) of our study species (Online Resource 2).
Mean model performance (Kendall’s predictive correlation) and 95% confidence intervals for the unoptimized single-scale, pseudo-optimized single-scale, and multi-scale abundance models for 28 passerine species. Significantly higher performance of multi-scale models than unoptimized and pseudo-optimized single-scale models indicated by * and **, respectively
Mean multi-scale Kendall’s rank correlations ranged from 0.25 to 0.60 with an overall mean of 0.42 across species (Figs. 4 and 3b, Online Resource 2). Fifty-seven percent (16/28) of our study species had mean multi-scale Kendall’s rank correlation above 0.40 and 29% (8/28) had mean multi-scale Kendall’s rank correlation above 0.5. Both overall abundance model performance and the improvement in Kendall’s rank correlation between single-scale and multi-scale models was species-specific. There was no mean improvement in performance between pseudo-optimized single-scale and multi-scale abundance models across species. The maximum improvement in Kendall’s rank correlation between pseudo-optimized single-scale and multi-scale abundance models was 0.037, or nine percent. The mean improvement in Kendall’s rank correlation between unoptimized single-scale and multi-scale SDMs was 0.022, with a maximum improvement of 0.050; five and 13 percent, respectively.
Discussion
The need for explicit consideration of scale
Our mixed-effects models showed that across species multi-scale models performed as well or better than pseudo-optimized single-scale models in both distribution and abundance modeling. The significantly poorer performance of unoptimized single-scale distribution and abundance models indicates that choosing an arbitrary environmental scale risks creating less predictive models. The consideration of scale through pseudo-optimized or multi-scale models will not unilaterally increase model performance. The arbitrary selection of a single environmental scale could result in a model with similar performance. The advantage to pseudo-optimized and multi-scale models is therefore in removing the guesswork and ensuring the use of environmental scales that result in higher performing models.
Results from the investigation of individual species strengthen our findings that (1) multi-scale distribution models perform as well or better than pseudo-optimized single-scale models and (2) the explicit consideration of environmental scale through multi-scale or pseudo-optimized single-scale modeling ameliorates the risks of arbitrary scale selection. Though at the individual species level multi-scale models performed significantly better than pseudo-optimized single-scale models in only 21% of the study species, multi-scale distribution models outperformed pseudo-optimized single-scale models in all but two species. If we were to increase sample sizes by running more iterations of each model, the number of significantly higher performing multi-scale models would no doubt increase. This indicates that multi-scale models do perform better than pseudo-optimized single-scale models. The risk of arbitrary scale selection is highlighted at the individual species level by the significantly lower performance of unoptimized single-scale models in nearly 90% of the study species. Results from abundance models appear to corroborate these findings.
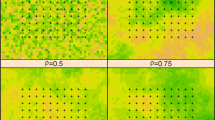
The degree of improvement between the modeling frameworks is species-specific and can be ecologically significant. Although multi-scale distribution models generally outperformed pseudo-optimized models, the mean improvement was relatively minor. The improvement between unoptimized single-scale models and multi-scale models, however, was much higher, especially for specific species. Brown Creeper, for example, had an improvement of over 0.1 AUC from unoptimized to multi-scale distribution models. Researchers interested in identifying suitable habitat and estimating populations often follow a similar process to what we used here: (1) run a SDM to estimate habitat suitability, (2) use a threshold such as prevalence to convert continuous suitability to binomial suitable and unsuitable habitat, and (3) run an abundance or density model to get population estimates within suitable habitat. When we use unoptimized single-scale models to estimate suitable habitat, abundance, and population size for Brown Creeper (converted abundances to densities naively assuming 200 m area surveyed), we estimate 13 percent more suitable habitat and a 14 percent smaller population size in our study area than when we use multi-scale models (Fig. 5). For common species such as Brown Creeper, these differences have little conservation impact, but similar discrepancies in species of conservation concern could result is poorly informed management decisions.
Advantages of our multi-scale approach
First, this multi-scale approach is a simple and concise method of explicitly accounting for environmental scale in distribution and abundance models. BRTs, as seen in this study, are robust to overfitting. This allows researchers to include environmental covariates characterized at many scales in a single multi-scale model. Pseudo-scale-optimization techniques generally include making a model for each environmental scale, then comparing model performance or AIC. More comprehensive approaches include making separate models for each environmental covariate at each environmental scale, to optimize each covariate individually. Theoretically, this allows optimization to account for a species responding to different environmental forces at different scales. In practice, this results in a large number of complex models, or few environmental covariates considered. Such complexity may be the reason that scale-optimization remains uncommon (McGarigal et al. 2016). Even studies that specifically address scale may fail to explicitly investigate the appropriateness of the scales they use (e.g. Fournier et al. 2017). The multi-scale approach employed here allows species to respond to difference environmental covariates at different scales all within a single model.
Second, by including all environmental covariates characterized at each scale in the same model, we allow for the relative influence of each to adjust in relation to other included covariates (Table 3). As described above, other multi-scale model frameworks optimize the scale of each covariate separately before combining the optimized covariates together in a single multi-scale model. These covariates, however, may have different optimal scales in single-covariate models than multi-scale models that include other similarly optimized covariates. Covariates that explain overlapping variance at one scale may explain more residual variance at other scales. Studies that include such comprehensive approaches may therefore use final models that fail to incorporate the optimal scale for each variable. This issue is irrelevant with the multi-scale modeling method presented here.
Considerations when choosing scales
No single scale is appropriate for all organisms (Wiens 1989; Levin 1992; Nadeau et al. 2017). The negative relationship between scale and the number of species for which that scale was best is likely a product of species selection. Most of our study species have small territory sizes that fall within the smaller radii used. SDMs of species with larger territories, such as raptors, would likely require topographic and land cover variables to be characterized at larger scales than considered here. Scale must be considered for each species independently.
We based the scales used in our study on similar songbird distribution modeling efforts (e.g. Boscolo and Metzger 2009; Shirley et al. 2013; Baladrón et al. 2016). We chose to use unlimited distance counts because we wanted our methods to be applicable to the eBird database. Given that birds were frequently detected over 100 m from the observer, we chose not to include smaller environmental scales that might be relevant to single territory sizes. This framework, however, could be used to build multi-level models with scales that range from microhabitat structures to landscapes (Meyer and Thuiller 2006). The inclusion of vastly different scales might further improve model performance and with this framework, the inclusion of poorly predictive scales should have little effect on overall model performance.
The pseudo-optimized single-scale of any ecological process is also influenced by the environmental covariates examined. There is a hierarchical structure of driving environmental forces in species’ distributions (Pearson and Dawson 2003). Climatic factors drive species’ distributions at the largest scales. Topographic then land cover factors are the dominant environmental forces driving distributions as scale decreases. Local factors such as habitat structure and composition drive distributions at more local scales. Biotic interactions may be important, especially at smaller scales where competitive exclusion may cause unoccupied sites in otherwise suitable habitat (Pearson and Dawson 2003; Araújo and Luoto 2007; Wiens et al. 2009). Recently, authors have argued that biotic interactions can and should be included at all scales (Wisz et al. 2013; Blois et al. 2013), but such interactions can be complex and difficult to model. Since environmental factors influence biotic interactions, environmental variables can be good predictors of distributions that are driven by biotic interactions (Godsoe et al. 2016). The pseudo-optimized single-scale is therefore dependent on the environmental covariates included in models. If only climatic factors were included, pseudo-optimized single-scales would increase. When using spectral bands to characterize the environment for SDMs, Shirley et al. (2013) found a most-influential-scale of nearly three times the largest scale used in our study. Spectral bands likely represent a mix of environmental forces that fall higher on the hierarchy than those considered here.
Caveat on modeling abundance
The less pronounced pattern in abundance model performance between single- and multi-scale models may be due to the inherent complexity in modeling abundance. Raw point counts can be poor indicators of abundance at the point count scale (Toms et al. 2006). At small scales, stochastic events may be highly influential on processes such as abundance and abundance is a larger scale process than occurrence (Wiens et al. 1989). Furthermore, behavioral interactions that were not considered here, may play a larger role at smaller spatial scales (Wiens et al. 2009). Studies that model abundance at larger spatial grains tend to have higher predictive performance (i.e. Illan et al. 2014; Barker et al. 2014).
Imperfect detection also adds to the noise in abundance modeling (Nichols et al. 2009). Estimation of imperfect detection can be used to create offsets to model “true” abundance in place of the raw observed abundance (Solymos et al. 2013). In doing so, “true” abundance is treated as an unverifiable latent variable (Welsh et al. 2013). Inclusion of detection probability may provide a useful extension to our modeling framework in the future. For a more complete discussion of the complexity in modeling abundance please refer to the Online Resource 4.
References
Araújo MB, Luoto M (2007) The importance of biotic interactions for modelling species distributions under climate change. Global Ecol Biogeogr 16:743–753
Baladrón AV, Isacch JP, Cavalli M, Bó MS (2016) Habitat selection by burrowing owls Athene cunicularia in the pampas of Argentina: a multiple-scale assessment. Acta Ornithol 51:137–150
Barker N, Cumming S, Darveau M (2014) Models to predict the distribution and abundance of breeding ducks in Canada. Avian Conserv and Ecol. https://doi.org/10.5751/ACE-00699-090207
Benítez-López A, Viñuela J, Mougeot F, García JT (2017) A multi-scale approach for identifying conservation needs of two threatened sympatric steppe birds. Biodivers Conserv 26:63–83
Blois JL, Zarnetske PL, Fitzpatrick MC, Finnegan S (2013) Climate change and the past, present, and future of biotic interactions. Science 341:499–504
Boscolo D, Metzger JP (2009) Is bird incidence in Atlantic forest fragments influenced by landscape patterns at multiple scales? Landsc Ecol 24:907–918
Dalgarno S, Mersey J, Gedalof Z, Lemon M (2017) Species-environment associations and predicted distribution of Black Oystercatcher breeding pairs in Haida Gwaii, British Columbia. Canada Avian Conserv Ecol. https://doi.org/10.5751/ACE-01094-120209
Dormann CF, Elith J, Bacher S, Buchmann C, Carl G, Carré G, Marquéz JRG, Gruber B, Lafourcade B, Leitão PJ, Münkemüller T (2012) Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography 36:27–46
Edrén SMC, Wisz MS, Teilmann J, Dietz R, Söderkvist J (2010) Modelling spatial patterns in harbour porpoise satellite telemetry data using maximum entropy. Ecography 33:698–708
Elith J, Graham CH (2009) Do they? How do they? WHY do they differ? On finding reasons for differing performances of species distribution models. Ecography 32:66–77
Elith J, Leathwick JR, Hastie T (2008) A working guide to boosted regression trees. J Anim Ecol 77:802–813
Evangelista PH, Mohamed AM, Hussein IA, Saied AH, Mohammed AH, Young NE (2018) Integrating indigenous local knowledge and species distribution modeling to detect wildlife in Somaliland. Ecosphere. https://doi.org/10.1002/ecs2.2134
Fournier A, Barbet-Massin M, Rome Q, Courchamp F (2017) Predicting species distribution combining multi-scale drivers. Glob Ecol Conserv 12:215–226. https://doi.org/10.1016/j.gecco.2017.11.002
García-Callejas D, Araújo MB (2016) The effects of model and data complexity on predictions from species distributions models. Ecol Model 326:4–12
Godsoe W, Franklin J, Blanchet FG (2016) Effects of biotic interactions on modeled species’ distribution can be masked by environmental gradients. Ecol Evol 7:654–664
Guisan A, Graham CH, Elith J, Huettmann F (2007) Sensitivity of predictive species distribution models to change in grain size. Divers Distrib 13:332–340
Hallman TA, Robinson WD (2020) Deciphering ecology from statistical artefacts: competing influence of sample size, prevalence and habitat specialization on species distribution models and how small evaluation datasets can inflate metrics of performance. Divers Distrib. https://doi.org/10.1111/ddi.13030
Halstead KE, Alexander JD, Hadley AS, Stephens JL, Yang Z, Betts MG (2019) Using a species-centered approach to predict bird community responses to habitat fragmentation. Landsc Ecol 34:1919–1935
Hudson M-AR, Francis CM, Campbell KJ, Downes CM, Smith AC, Pardieck KL (2017) The role of the North American breeding bird survey in conservation. Condor 119:526–545
Illan JG, Thomas CD, Jones JA, Wong WK, Shirley SM, Betts, MG (2014) Precipitation and winter temperature predict long-term range-scale abundance changes in Western North American birds. Glob Change Biol 20:3351–3364
Johnston A, Fink D, Reynolds MD, Hochachka WM, Sullivan BL, Bruns NE, Hallstein E, Merrifield MS, Matsumoto S, Kelling S (2015) Abundance models improve spatial and temporal prioritization of conservation resources. Ecol Appl 25:1749–1756
Kulhanek SA, Leung B, Ricciardi A (2011) Using ecological niche models to predict the abundance and impact of invasive species: application to the common carp. Ecol Appl 21:203–213
Landscape Ecology, Modeling, Mapping, and Analysis (2014) GNN Structure Maps. https://lemma.forestry.oregonstate.edu/data/structure-maps. Accessed 6 Sep 2016
Levin SA (1992) The problem of pattern and scale in ecology: the Robert H. MacArthur award lecture. Ecology 73:1943–1967
Liu C, Berry PM, Dawson TP, Pearson RG (2005) Selecting thresholds of occurrence in the prediction of species distributions. Ecography 28:385–393
Lobo JM, Jiménez-Valverde A, Real R (2008) AUC: a misleading measure of the performance of predictive distribution models. Global Ecol Biogeogr 17:145–151
Mayor SJ, Schneider DC, Schaefer JA, Mahoney SP (2009) Habitat selection at multiple scales. Ecoscience 16:238–247
McGarigal K, Wan HY, Zeller KA, Timm BC, Cushman SA (2016) Multi-scale habitat selection modeling: a review and outlook. Landsc Ecol 31:1161–1175
Meyer CB, Thuiller W (2006) Accuracy of resource selection functions across spatial scales. Divers Distrib 12:288–297
Miguet P, Fahrig L, Lavigne C (2017) How to quantify a distance-dependent landscape effect on a biological response. Methods Ecol Evol 8:1717–1724
Moll RJ, Cepek JD, Lorch PD, Dennis PM, Robison T, Montgomery RA (2020) At what spatial scale(s) do mammals respond to urbanization? Ecography. https://doi.org/10.1111/ecog.04762
Nadeau CP, Urban MC, Bridle JR (2017) Coarse climate change projections for species living in a fine-scaled world. Glob Change Biol 23:12–24
Nichols JD, Thomas L, Conn PB (2009) Inferences about landbird abundance from count data: recent advances and future directions. Model Demogr Process Mark Popul. https://doi.org/10.1007/978-0-387-78151-8_9
Oppel S, Meirinho A, Ramirez I, Gardner B, O'Connell AF, Miller PI, Louzao M (2012) Comparison of five modelling techniques to predict the spatial distribution and abundance of seabirds. Biol Conserv 156:94–104
Oregon Spatial Data Library (2017) Oregon 10m digital elevation model (DEM). https://spatialdata.oregonexplorer.info/geoportal/details;id=7a82c1be50504f56a9d49d13c7b4d9aa. Accessed 30 Nov 2015
Oregon Spatial Data Library (2016) Oregon rivers. https://spatialdata.oregonexplorer.info/geoportal/details;id=01606665b1034dc6877fbad58bb9879a. Accessed 28 Jun 2016
Pearson RG, Dawson TP (2003) Predicting the impacts of climate change on the distribution of species: are bioclimate envelope models useful? Global Ecol Biogeogr 12:361–371
Pennington DN, Blair RB (2011) Habitat selection of breeding riparian birds in an urban environment: untangling the relative importance of biophysical elements and spatial scale. Divers Distrib 17:506–518
Qiao H, Soberón J, Peterson AT (2015) No silver bullets in correlative ecological niche modelling: insights from testing among many potential algorithms for niche estimation. Methods Ecol Evol 6:1126–1136
Reino L, Triviño M, Beja P, Araújo MB, Figueira R, Segurado P (2018) Modelling landscape constraints on farmland bird species range shifts under climate change. Sci Total Environ 625:1596–1605
Shirley SM, Yang Z, Hutchinson RA, Alexander JD, McGarigal K, Betts MG (2013) Species distribution modelling for the people: unclassified landsat TM imagery predicts bird occurrence at fine resolutions. Divers Distrib 19:855–866
Sólymos P, Matsuoka SM, Bayne EM, Lele SR, Fontaine P, Cumming SG, Stralberg D, Schmiegelow FKA, Song SJ (2013) Calibrating indices of avian density from non-standardized survey data: making the most of a messy situation. Methods Ecol Evol 4:1047–1058
Stevens BS, Conway CJ (2019) Predicting species distributions: unifying model selection and scale optimization for multi-scale occupancy models. Ecosphere 10:e02748
Stevens BS, Conway CJ (2020) Predictive multi-scale occupancy models at range-wide extents: Effects of habitat and human disturbance on distributions of wetland birds. Divers Distrib 26:34–48
Sullivan BL, Wood CL, Iliff MJ, Bonney RE, Fink D, Kelling S (2009) eBird: A citizen-based bird observation network in the biological sciences. Biol Conserv 142:2282–2292
Thorson TD, Bryce SA, Lammers DA, Woods AJ, Omernik JM, Kagan J, Pater DE, Comstock JA (2003) Ecoregions of Oregon
Timm BC, McGarigal K, Cushman SA, Ganey JL (2016) Multi-scale Mexican spotted owl (Strix occidentalis lucida) nest/roost habitat selection in Arizona and a comparison with single-scale modeling results. Landsc Ecol 31:1209–1225
Toms JD, Schmiegelow FKA, Hannon SJ, Villard M-A (2006) Are point counts of boreal songbirds reliable proxies for more intensive abundance estimators? Auk 123:438
United States Geological Survey (2011) National gap analysis project. https://gapanalysis.usgs.gov/gaplandcover/data/download/. Accessed 7 Dec 2015
Welsh AH, Lindenmayer DB, Donnelly CF (2013) Fitting and Interpreting Occupancy Models. PLoS ONE 8:e52015
Wiens JA (1989) Spatial scaling in ecology. Funct Ecol 3:385–397
Wiens JA, Milne BT (1989) Scaling of ‘landscapes’ in landscape ecology, or, landscape ecology from a beetle’s perspective. Landsc Ecol 3:87–96
Wiens JA, Stralberg D, Jongsomjit D, Howell CA, Snyder MA (2009) Niches, models, and climate change: assessing the assumptions and uncertainties. P Natl Acad Sci 106:19729–19736
Wilsey CB, Jensen CM, Miller N (2016) Quantifying avian relative abundance and ecosystem service value to identify conservation opportunities in the Midwestern US. Avian Conserv Ecol. https://doi.org/10.5751/ACE-00902-110207
Wisz MS, Pottier J, Kissling WD, Pellissier L, Lenoir J, Damgaard CF, Dormann CF, Forchhammer MC, Grytnes JA, Guisan A, Heikkinen RK, Høye TT, Kühn I, Luoto M, Maiorano L, Nilsson MC, Normand S, Öckinger E, Schmidt NM, Termansen M, Timmermann A, Wardle DA, Aastrup P, Svenning JS (2013) The role of biotic interactions in shaping distributions and realised assemblages of species: implications for species distribution modelling. Biol Rev 88:15–30
Yang X, Chapman GA, Young MA, Gray JM (2005) Using compound topographic index to delineate soil landscape facets from digital elevation models for comprehensive coastal assessment. MODSIM 2005 International Congress on Modelling and Simulation. Modelling and Simulation Society of Australia and New Zealand, Canberra, pp 1511–1517
Acknowledgements
We appreciate assistance with surveys from R. Moore, database work and maintenance from R. DeMoss, and data entry from the Robinson Lab. The Robinson graduate lab provided helpful comments and discussion. The work was supported by the generous endowment of the Bob and Phyllis Mace Watchable Wildlife Professorship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hallman, T.A., Robinson, W.D. Comparing multi- and single-scale species distribution and abundance models built with the boosted regression tree algorithm. Landscape Ecol 35, 1161–1174 (2020). https://doi.org/10.1007/s10980-020-01007-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-020-01007-7