Abstract
α-Amylases are used in various biotechnological processes including the textile, paper, food, biofuels, detergents and pharmaceutical industries. In this study, a novel gene encoding α-amylase was cloned from marine bacterium Salinispora arenicola CNP193 and the protein was expressed in Escherichia coli. The α-amylase gene from S. arenicola CNP193 had a length of 1839 bp and encoded a α-amylase with an estimated molecular mass of 74 kDa. The optimum temperature and pH for the recombinant α-amylase was 50 °C and 7 respectively. Na+, K+ and Ca2+ increased the activity of the recombinant α-amylase whereas the enzyme was inhibited by Cu2+, Zn2+, Hg2+, Pb2+, Fe3+ and Mn2+. Thin layer chromatography results confirmed that monosaccharide, disaccharide and maltotriose are the hydrolysis products. The results of our study suggest that this enzyme has considerable potential in industrial applications.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Alpha-amylases (endo-1,4-α-d-glucan glucanohydrolase, EC3.2.1.1) hydrolyze the 1,4-α-glucosidic linkages within starch and related polysaccharides to produce different length oligosaccharides [1]. Amylases have biotechnological applications such as in starch liquefaction, textile, food and baking [2]. Amylases constitute about 30% of enzyme market [3]. Plants, animals and microorganisms produce different α-amylases. Cheaper cost, enhanced production and consistency makes microbial amylases very attractive for biotechnology industry [4]. Amylases have been reported from different microorganisms such as Bacillus, Aspergillus and Penicillium species [5]. However there is very less literature available for amylases from marine bacteria.
Microorganisms are well known source of enzymes [6, 7]. There is very less information available about the marine amylases and their industrial applications. The marine environment, particularly at the microbial level, has been relatively unexploited, but it has a significant potential as a source of industrial enzymes. Among marine microorganisms, studies of natural products and several enzymes have been reported from Salinispora strains. Salinispora is the first reported marine obligate actinomycete and have become model organisms for secondary metabolite discovery. However, Salinispora strains enzymes have not been studied much. Sequencing and recombinant techniques have led to the exploration of biotechnological potential of Salinispora strains Genome mining has enabled to explore novel biocatalysts from Salinispora strains. This provides an efficient way to the exploration of different enzymes of industrial significance [8].
There is an ongoing interest in the isolation of α-amylases with higher activity or different catalytic properties. Salinispora strains have potential to produce novel α-amylases with higher activity or high salt tolerance. Herein, the α-amylase coding gene from Salinispora arenicola CNP193 was mined, cloned and expressed in Escherichia coli, and the recombinant α-amylase was purified and characterized in this study.
2 Experimental
2.1 Strains and Culture Conditions
The S. arenicola CNP193 was kindly provided by Dr. Paul R Jensen, Center for Marine Biotechnology and Biomedicine, Scripps Institution of Oceanography, University of California, San Diego, USA. The genome of S. arenicola strain CNP193 was deposited in the Joint Genome Institute’s Integrated Microbial Genomes (IMG) database, http://img.jgi.doe.gov/cgi-bin/w/main.cgi (accession number: 2518285552). E. coli DH5α and E. coli BL21 (DE3) were purchased from Takara. The pGEM-T vector, pET-28a (+) and Ni–NTA His Tag Kit was purchased from Promega.
2.2 Cloning and Expression of the α-Amylase Gene in E. coli
All DNA manipulations were performed using standard protocols as described previously [9]. Taq DNA polymerase was used for gene amplification using primer pair i.e.
Forward primer, 5′- CCGGAATTCATGCCGGAGTACCCG- 3′,
Reverse primer, 5′- CCCAAGCTTCCGCCCCCGGTCACTGTT- 3′.
The purified PCR product was cloned into pGEM-T vector and sequenced. BLAST was used for the analysis of the protein sequence (https://blast.ncbi.nlm.nih.gov/Blast). Genedoc software package was used for multiple sequence alignment. Cleavage sites were predicted by the Signa lP 4.1 Server (http://www.cbs.dtu.dk/services/SignalP) was used for the prediction of cleavage sites.
The amplified PCR products from S. arenicola CNP193 were digested with EcoR I and Hind III and inserted into pET28a (+) expression vector digested with the same enzymes to produce plasmid pET28a (+)-Saa1-Amy. The plasmids were then transformed into E. coli BL21 (DE3). The recombinant strains were grown in 100 mL of LB broth supplemented with 100 μg/mL ampicillin at 37 °C under agitation (180 rpm). When cell density (A600) reached 0.8, the culture was induced with 0.02 mM IPTG at 20 °C for 24 h. Cells were harvested by centrifugation at 8000×g for 10 min at 4 °C. The cell pellet was re-suspended in of 25 mM Tris–HCl buffer (pH 7.0) followed by sonication at 4 °C using a sonifier (7 min, 10 s ON/25 s OFF, 50% amplitude). The crude extract was then centrifuged at 12,000×g for 20 min at 4 °C to separate the cell debris. The final supernatant was filtered with a 0.22 μm sterile syringe filter and stored at 4 °C until used.
2.3 Purification of the Recombinant Enzyme
The final supernatant was first purified by DEAE-sepherose ion exchange chromatography described previously [10]. The DEAE-Sepharose column is prewashed with 5 bed volumes of 20 mM Tris–HCl, pH 8.5, plus 70 mM NaCl before loading the sample described above. The column is eluted with a linear gradient from 0.1 to 0.5 M NaCl at a flow rate of 2 mL/min. Fractions were assayed for α-amylase activity. Fractions containing the peak of α-amylase activity were pooled, desalted and loaded to the Ni–NTA resin column (18 × 100 mm) that had been equilibrated with start buffer (20 mM sodium phosphate pH 6.5, 500 mM NaCl, and 5 mM imidazole), and eluted with a step-wise elution with the indicated concentrations of imidazole (10, 50, 80, 100, 200 and 500 mM) in 50 mM Tris–HCl buffer (pH 8.0) and 500 mM NaCl. Fractions containing α-amylase activity were pooled, desalted, and concentrated. The recombinant α-amylase was named as Saa1-Amy. Protein concentration was determined using Bradford protein assay kit (Bio-Rad, Hercules, CA) with bovine serum albumin as a standard. 12% sodium dodecyI sulfate (SDS) polyacrylamide was used for the estimation of molecular mass of the recombinant enzyme.
2.4 α-Amylase Activity Assay
The α-amylase enzyme activity was measured as described earlier [11]. The assay was performed by adding 0.15 mL of starch solution (1%, w/v) as a substrate to the appropriately diluted enzyme sample containing 0.1 mL of enzyme in 50 mM Tris–HCl buffer (pH 7). The reaction mixture was incubated for 15 min at 50 °C, at which temperatures the assayed enzymes were most active. The α-amylase activity was confirmed by measuring the amount of reducing sugars released during starch hydrolysis using the dinitrosalicylic acid method. After incubation at 50 °C for 15 min, the reaction was stopped by adding 2 mL of 3-5-dinitrosalicylic acid reagent. One unit of α-amylase activity was defined as the amount of enzyme causing release of reducing sugars equivalent to 1 μM of glucose from starch per min under the assay condition.
2.5 Characterization of Recombinant α-Amylase
Effect of different temperatures (15–60 °C) with 50 mM sodium phosphate buffer (pH 7) were studied for recombinant α-amylase. Saa1-Amy was preincubated at 25 °C, 35 °C, 45 °C, 55 °C and 65 °C for the thermal stability studies. pH range of 4–9 with 50 mM Britton Robinson buffer at 50 °C was performed to find the optimal pH for recombinant amylase activity. The effects of different metal ions on enzyme activity were determined by incubating the Saa1-Amy with different metal ions (1 mM) and soluble starch (1%, w/v) in a 0.1 M sodium phosphate buffer (pH 7) at 50 °C for 1 h, respectively.
2.6 Identification of Hydrolysis Products of the Recombinant α-Amylase Saa1-Amy
Thin-layer chromatography (TLC) was used for identification of hydrolysis products of the Saa1-Amy. Hydrolysis product was produced by incubating the Saa1-Amy containing soluble starch (1%, w/v) in a 0.1 M sodium phosphate buffer (pH 7.0) at 50 °C for 3 h, 6 h, 9 h, 12 h, separately. Glucose, sucrose and raffinose were dissolved in water (1%, w/v) and they were used as standard sample. The reaction samples (approx. 5μL) were spotted on a TLC plate and developed with a solvent system of n-butanol/acetic acid/water [2:1:1(v:v:v)]. The developed TLC plate was dried in natural condition and soaked into 10% (v/v) sulfuric acid in ethanol followed by baking at 115 °C for 10 min.
3 Results
3.1 Cloning and Expression of α-Amylase Gene in pET-28a (+)
An 1839-bp putative α-amylase-encoding gene, named Saa1, was amplified from the genomic DNA of S. aronicola CNP193. GenBank accession number of 1839-bp α-amylase-encoding gene, named Saa1 from S. arenicola CNP193 is MK403732. Saa1 gene was heterologously expressed in E. coli BL21 (DE3) cells. The protein was overexpressed after induction by 0.5 mM IPTG. The recombinant Saa1-Amy was purified using DEAE-Sepharose Fast Flow in first step. It is further purified in second step using Ni–NTA His Tag Kit. Total protein and enzyme activity as well as the yield of purified recombinant α-amylase from E. coli BL21 (DE3) is shown in Table 1. The yield of purified recombinant α-amylase was 22.8 mg.
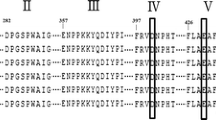
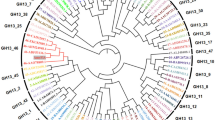
BLASTp of the translated protein sequence showed it had a 74% identity with a glucoamylase from Micromonospora sp. CNB394 and 59% identity with an amylase from Streptomyces sp. CNT302. The alignment of the amino acid sequence is shown in Fig. 1. The molecular weight of alpha-amylase being studied was determined by SDS-PAGE. Figure 2 shows a protein band with a molecular mass of about 74 kDa corresponding to the recombinant Saa1-Amy.
SDS-PAGE analysis of the α-amylase expression and purification. SDS-PAGE gel concentration was 12% and was stained by Coomassie R250. Lane M. marker (Genview, USA); Lane 1 and Lane 2, The pET28 a/Saa1 after induction in E. coli BL21; Lane 3, The recombinant α-amylase purified by DEAE-Sepharose ion exchange chromatography; Lane 4, The recombinant α-amylase purified by Ni–NTA chromatography
3.2 Recombinant α-Amylase Enzymatic Properties
Effect of temperature, stability, pH, and metal ions on the catalytic activity of Saa1-Amy was evaluated to characterize this enzyme. The optimal temperature for the recombinant α-amylase was 50 °C (Fig. 3), and it maintains more than 70% activity between 40 and 55 °C.
Thermostability results of recombinant α-amylase were shown in Fig. 4. The Saa1-Amy retained more than 85% of its activity at 50 and 60 °C at 60 min, whereas it lost 25% and 43% after heating at the same temperature for 90 min. The enzyme only showed less than 20% activity when heated at 65 °C for 150 min. Saa1-Amy is a mesophilic α-amylase and it shows a remarkable stability at medium temperature.
Table 2 shows the effects of metal ions on the catalytic activity of SAA1. Ca2+, Na+ And K+ significantly increased the α-amylase activity, whereas the other metal ions, such as, Hg+ and Pb2+, inhibited the enzyme activity.
The optimal pH of recombinant α-amylase was 7. The enzyme retained about 60% of its activity at pH 5, and the enzyme still have more than 60% of its activity at pH 9 (Fig. 5). Most of the enzymes give maximum activity at optimum around pH 7.
3.3 Hydrolysis Product Produced by Recombinant α-Amylase
The end products of starch hydrolysis by Saa1Amy were analyzed with thin layer chromatography. Saa1-Amy efficiently hydrolyzed starch to glucose, maltose, maltotriose and tetrasaccharide (Fig. 6). In the whole process of hydrolysis, we can find all the four kinds of hydrolysis products. This result implies that Saa1Amy randomly cleaves the 1,4-α-glucosidic linkages between adjacent glucose units in the linear amylose of starch.
The hydrolysis products of Saa1Amy on starch. Lane 1. Marker (glucose, saccharose, raffinose and maltotetraose); Lane 2, the hydrolysis products after incubating in 50 °C for 30 min; Lane 3, the hydrolysis products after incubating in 50 °C for 50 min; Lane 4, the hydrolysis products after incubating in 50 °C for 90 min; Lane 5, the hydrolysis products after incubating in 50 °C for 120 min; Lane 6, the hydrolysis products after incubating in 50 °C for 150 min; Lane 7, the hydrolysis products after incubating in 50 °C for 3 h; Lane 8, the hydrolysis products after incubating in 50 °C for 6 h; Lane 9, the hydrolysis products after incubating in 50 °C for 12 h
4 Discussion
Previous studies have shown that the α-amylases have been classified into more than 135 families based on amino sequence similarity [12]. Saa1 belongs to the GHF15 (glucoside hydrolase family 15). Multiple sequence alignment revealed that Saa1 shares the same conserved block marked in the shaded boxes. An effective way to increase the yield of α-amylase is by the recombinant expression [13]. Escherichia coli is the most commonly used expression host. Genetic background of E. coli is well understood and many tools are available for its genetic manipulation. Simple media is required for quick growth of E. coli. Heterologous proteins have been expressed and purified in E. coli [14].
The recombinant enzyme Saa1-Amy described in this study is the first α-amylase from marine S. aronicola CNP193 and expressed in E. coli. S. aronicola CNP193 amylase is an addition to the α-amylases that were previously obtained from other marine microorganisms such as A. haloplanctis [15], Nocardiopsis sp. [16], Pseudoalteromonas sp. [17], Zunongwangia profunda [18], A. agilis [19] and Bacillus sp. dsh191 [2].
The optimal temperature and pH for the recombinant α-amylase in this study was 50 °C and 7.0 respectively. Previous studies have reported optimum temperature of 50 °C for recombinant α-amylase expressed in E.coli [20]. In previous studies recombinant α-amylase expressed in E. coli has optimum pH 6.0. Recombinant α-amylase maximal activity was achieved at pH 6.0 [13]. (Recombinant α-amylase from marine bacterium Bacillus sp. dsh19-1 expressed in E. coli has maximal activity at pH 6.0 [2].
Ca2+, Na+ And K+ significantly increased the α-amylase activity, whereas the other metal ions, such as, Hg+ and Pb2+, inhibited the enzyme activity. The effects of metal ions on the catalytic activity of SAA1 may prove that the amylase is a metalloenzyme. Amylases are stimulated by Ca2+ [21]. Most of the known α-amylases are metalloenzymes, requiring Ca2+ [22,23,24]. This increase in the activity of recombinant α-amylase by metal ions is in accordance with the results reported previously [2].
In conclusion, a novel α-amylase (Saa1-Amy) has been discovered by genome mining of S. arenicola CNP193 and then successfully expressed. Based on the finding as discussed in this paper, it is concluded that the recombinant strain is one of the most promising mesophilic α-amylase production with large potential in industry and there is a great commercial value in further development of this recombinant enzyme.
References
MacGregor EA, Janecek S, Svensson B (2001) Relationship of sequence and structure to specificity in the alpha-amylase family of enzymes. Biochim Biophys Acta 1546(1):1–20
Dou S, Chi N, Zhou X et al (2018) Molecular cloning, expression, and biochemical characterization of a novel cold-active α-amylase from Bacillus sp. dsh19-1. Extremophiles. https://doi.org/10.1007/s00792-018-1034-7
Nikitkova AE, Haase EM, Scannapieco FA (2013) Taking the starch out of oral biofilm formation: molecular basis and functional significance of salivary alpha-amylase binding to oral streptococci. Appl Environ Microbiol 79(2):416–423
Xiao Z, Wu M, Grosse S et al (2014) Genome mining for new alpha-amylase and glucoamylase encoding sequences and high level expression of a glucoamylase from Talaromyces stipitatus for potential raw starch hydrolysis. Appl Biochem Biotechnol 172(1):73–86
Anamaria CS, María CR, Barbara AA et al (2019) Heterologous expression and biochemical characterization of a novel cold active α-amylase from the Antarctic bacteria Pseudoalteromonas sp. 2–3. Protein Expr Purif 155:78–85
Ahmed S, Mustafa G, Arshad M et al (2017) Fungal Biomass protein production from Trichoderma harzianum using rice polishing. Biomed Res Int. https://doi.org/10.1155/2017/6232793
Ahmed S, Riaz S, Jamil A (2009) Molecular cloning of fungal xylanases: an overview. Appl Microbiol Biotechnol 84(1):19–35
Ziemert N, Lechner A, Wietz M et al (2014) Diversity and evolution of secondary metabolism in the marine actinomycete genus Salinispora. Proc Natl Acad Sci USA 111:E1130–E1139
Maldonado LA, Fenical W, Jensen PR et al (2005) Salinispora arenicola gen. nov., sp. nov. and Salinispora tropica sp. nov., obligate marine actinomycetes belonging to the family Micromonosporaceae. Int J Syst Evol Microbiol 55:1759–1766
Mamo G, Hatti-Kaul R, Mattiasson KB (2006) A thermostable alkaline active endo-β-1-4-xylanase from Bacillus halodurans S7: purification and characterization. Enz Microb Technol 39(7):1492–1498
Xiao Z, Storms R, Tsang A (2006) Quantitative starch–iodine method for measuring alpha-amylase and glucoamylase activities. Anal Biochem 351(1):146–148
Kuriki T, Imanaka T (1999) The concept of the alpha-amylase family: structural similarity and common catalytic mechanism. J Biosci Bioeng 87(5):557–565
Zhao F, Song Q, Wang B et al (2019) Secretion of the recombination α-amylase in Escherichia coli and purification by the gram-positive enhancer matrix (GEM) particles. Int J Biol Macromol 123:91–96
Zafar M, Ahmed S, Khan MIM, Jamil A (2014) Recombinant expression and characterization of a novel endoglucanase from Bacillus subtilis in Escherichia coli. Mol Biol Rep 41(5):3295–3302
Feller G, Lonhienne T, Deroanne C et al (1992) Purification, characterization, and nucleotide sequence of the thermolabile alpha-amylase from the Antarctic psychrotroph Alteromonas haloplanctis A23. J Biol Chem 267(8):5217–5221
Zhang JW, Zeng RY (2008) Purification and characterization of a cold adapted α-amylase produced by Nocardiopsis, sp. 7326 isolated from Prydz Bay, Antarctic. Mar Biotechnol 10(1):75–82
Lu M, Wang S, Fang Y et al (2010) Cloning, expression, purification, and characterization of cold-adapted α-amylase from Pseudoalteromonas arctica GS230. Protein J 29(8):591–597
Qin Y, Huang Z, Liu Z (2014) A novel cold-active and salt-tolerant α-amylase from marine bacterium Zunongwangia profunda: molecular cloning, heterologous expression and biochemical characterization. Extremophiles 18(2):271–281
Kim SM, Park H, Choi JI (2017) Cloning and characterization of cold-adapted α-amylase from Antarctic Arthrobacter agilis. Appl Biochem Biotechnol 181(3):1048–1059
Li Z, Wu J, Zhang B et al (2015) AmyM, a novel maltohexaose-forming α-amylase from Corallococcus sp. strain EGB. Appl Environ Microbiol 81(6):1977–1987
Vieille C, Zeikus GJ (2001) Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol Mol Biol Rev 65(1):1–43
Gupta R, Gigras P, Mohapatra H et al (2003) Microbial α-amylases: biotechnological perspective. Process Biochem 38:1599–1616
Ghorbel RE, Maktouf S, Massoud EB et al (2009) New thermostable amylase from Bacillus cohnii US147 with a broad pH applicability. Appl Biochem Biotechnol 157(1):50–60
Chen J, Chen X, Dai J et al (2015) Cloning, enhanced expression and characterization of an α-amylase gene from a wild strain in B. subtilis WB800. Int J Biol Macromol 80:200–207
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 31772016), Jiangsu Province Marine Science and Technology Innovation Project (HY2018-10), the Priority Academic Program Development of Jiangsu Higher Education Institutions, the Scientific Research Foundation for the Returned Overseas Chinese Scholars, Six talent peaks project in Jiangsu Provincev (2016-SWYY-195), and Project “333” of Jiangsu Province.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, S., Ahmed, S. & Fang, Y. Cloning, Expression and Characterization of a Novel α-Amylase from Salinispora arenicola CNP193. Protein J 38, 716–722 (2019). https://doi.org/10.1007/s10930-019-09870-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-019-09870-3