Abstract
Due to its widespread use and water solubility, poly(vinyl alcohol) (PVA) has the potential to find its way into various water or soil ecosystems. Despite the fact that many bacterial species with the capacity of utilizing PVA have been found and described, the influences of some environmental factors on their capabilities to biodegrade PVA have not been adequately studied. Therefore, study was made of the effects of two environmental factors on PVA degradation exhibited by two Sphingomonas strains. Both strains originated from common wastewater treatment plants, and proved to be considerably sensitive to increased inorganic salt concentrations; in brief, 13.3 mmol/l either of phosphate or chloride ions significantly delayed the degradation process or inhibited it entirely. In contrast to such halosensitivity, both strains were able to rapidly utilize PVA under suitable conditions, even when low inoculum sizes were applied. Initial cell densities, ranging from 100 to 107 cells/ml, were used in two series of degradation trials and PVA degradation occurred in all cases; merely delays extending over several days in the degradation process were noted when inoculum sizes of 100–103 cells/ml were applied.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Poly(vinyl alcohol) (PVA) is a synthetic polymer prepared by a hydrolysis of poly(vinylacetate). Its water solubility and widespread use by the textile and chemical industries and in glazed paper production mean it has the potential to enter different water environments, from which it could be removed by microbial degradation. Many bacterial genera responsible for PVA degradation have previously been reported, hence the biodegradation of PVA materials and bacterial biochemical pathways of PVA decomposition have already been reviewed [1, 2].
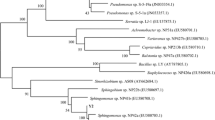
Bacteria able to degrade PVA include representatives of the genera Pseudomonas [3,4,5,6,7,8], Alcaligenes [9], Bacillus [10], Brevibacillus [11, 12], Sphingomonas [13,14,15], Achromobacter [16], Microbacterium and Paenibacillus [17], Sphingopyxis [18, 19], Streptomyces [20], Novosphingobium [21] and Geobacillus [12], although some of these require certain substances, often pyrroloquinoline quinone (PQQ), for successful PVA utilization.
Numbering amongst the most important degraders are gram-negative bacteria of the sphingomonad group, a specific cluster of Alpha-proteobacteria, comprising the genera Sphingomonas, Sphingopyxis, Sphingobium, Novosphingobium and Sphingosinicella [22, 23]. Although an overarching concept concerning several genera has not been widely accepted [24], it is impossible to ignore that some such described examples of these genera rank amongst the most active PVA degraders. Firstly, Tokiwa et al. [13] isolated Sphingomonas sp. TJ-7, which was capable of PVA biodegradation; the strain utilized approximately 60% of organic carbon over 5 days at 30 °C. Later, Kim et al. [14] obtained Sphingomonas sp. SA3 that was capable of degrading 85% of 5 g/l PVA during 5 days, in co-culture with SA2 bacterium with the ability to supply PQQ. Furthermore, Sphingomonas sp. was reported to be a member of an efficient mixed culture that degraded 1 g/l PVA over 5 days, during which Sphingomonas sp. increased in relative abundance at the initial stage of degradation [25]. In 2006, Yamatsu et al. [18] documented isolation of a new, effective, PQQ-independent strain, Sphingopyxis sp. PVA3. Therein, the bacterium utilized 90% of PVA after 6 days of cultivation (at an initial concentration of 1 g/l). The same genus was revealed to be the correct one for identifying the strain 113P3, as isolated by Hatanaka et al. [26] and primarily identified as Pseudomonas sp. Moreover, Hu et al. [19] reclassified the strain as Sphingopyxis sp. 113P3, and showed that the strain was able to consume more than 90% of PVA (5 g/l) in 4 days. In the most recent work, Rong et al. [21] isolated Novosphingobium sp. P7, which had the capacity to almost totally degrade 8 g/l PVA during 1 week of incubation in the presence of methionine as a growth factor.
In addition to possessing the possibility to utilize PVA, Sphingomonads (sensu latu) are known to degrade other xenobiotic polymers, such as polyethylene glycol or polyaspartate [27, 28], as well as various xenobiotic, low-molecular weight compounds [29]. The specific abilities to degrade polymers shown by Sphingomonads have undergone detailed study [30,31,32], therein pertaining to Sphingopyxis sp. 113P3; the authors proved that PVA is incorporated directly through the outer cell membrane to the periplasmic space, wherein it is subjected to the activities of PVA dehydrogenase and oxidized-PVA hydrolase. Despite amassing such great knowledge on the nature of PVA utilization by Sphingomonads, some environmental aspects concerning their PVA degradation, such as the effect of inorganic salt concentration or the minimum cell count required to initiate the degradation process have yet to be investigated. However, these aspects may be crucial to PVA biodegradation, both in the wastewater treatment process and in different types of surface water, within which degraders could be relatively scarce and salt concentrations might significantly vary. With this in mind, herein it was decided to study these two effects on PVA biodegradation with two Sphingomonas strains isolated from different wastewater treatment plants.
Materials and Methods
Biological Samples (Materials)
Two samples of activated sludge were used. One of them originated from a wastewater treatment plant (herein after referred to as “WWTP”) in Otrokovice, treating both municipal and industrial wastewater, while the other was obtained from a WWTP in Zlín, which largely treats municipal wastewater.
Chemicals
The PVA-MOWIOL 5-88—was purchased from Clariant (Germany). The PQQ was sourced from Fluka, while the ordinary chemicals were obtained from local suppliers.
Media Composition
The basal mineral medium (BMM, in g per litre if not otherwise stated): KH2PO4 0.18; Na2HPO4·12H2O 1.92; MgSO4·7H2O 0.1; Fe(NH4)2(SO4)2·6H2O 0.03; CaCl2·2H2O 0.01; NH4Cl 0.3; and NaCl 0.5, trace element solution 2 ml [33].
The PVA agars (in g/l of BMM if not otherwise stated): agar 19 and PVA 0.5; some portions were supplemented with PQQ 20 μg/l.
Compositions of mineral media (MM) with different phosphate concentrations constituted several modifications of BMM, containing 3.33, 6.67, 10, 13.3, 16.7 and 20 mmol/l of phosphate ions, which were prepared and designated MM3, MM7, MM10, MM13, MM17 and MM20, respectively. The MM7 medium was identical to the BMM.
Compositions of mineral media with different NaCl concentrations comprised five modifications of MM3 medium, containing 0, 3.33, 6.67, 13.3 and 26.7 mmol/l NaCl, which were prepared and designated NaCl 0, NaCl 3, NaCl 7, NaCl 13 and NaCl 27, respectively.
Compositions of mineral media with different KCl concentrations encompassed five modifications of MM3 medium, containing 0, 3.33, 6.67, 13.3 and 26.7 mmol/l KCl, which were prepared and designated KCl 0, KCl 3, KCl 7, KCl 13 and KCl 27, respectively.
Isolation of PVA-Degrading Bacteria
Both samples of activated sludge were acclimated in BMMs containing 200 mg/l PVA for 12 days at 25 °C. The suspensions obtained were then diluted by the serial method, and several dilutions were spread on the surface of two types of PVA agar, with and without the addition of PQQ. Mineral agars without PVA were also used in parallel. After incubation for 10 days at 25 °C in darkness, distinct colonies growing on the PVA agars with PQQ were picked up, purified and preserved at −80 °C in glycerol.
Identification of Bacterial Strains
This was carried out by the State Veterinary Institute in Prague on a commercial base. In brief, the DNA of pure strains were extracted by the Easy-DNA™ kit (Invitrogen) and MicroSeq Full Gene 16S rDNA PCR kit and MicroSeq Full Gene 16S rDNA Sequencing kit (Applied Biosystems) were used in accordance with the directions specified by the manufacturers. The sequences acquired for 16S rDNA were compared using BLAST at the NCBI website.
PVA Degradation Tests
All biodegradation tests were conducted in 500 ml glass bottles containing 100 ml of sterile BMM, or certain types of MM amended with PVA 0.5 g/l and PQQ 20 µg/l. When testing the effects of salts, the bottles were inoculated with 100 µl of bacterial suspension corresponding to the 1st degree of the McFarland scale (approx. 3 × 108 cells/ml). In tests of inoculum size, the suspensions were microscopically checked (Olympus CX41 featuring phase contrast), and then diluted to the required density to achieve initial cell concentrations of 100–107 cells/ml in degradation tests. Dilution of the suspensions involved utilizing the appropriate mineral media with PVA and PQQ.
All the test bottles were incubated at 25 °C in darkness using a laboratory shaker. The growth exhibited by cells was monitored by optical density measurement at 600 nm (OD600) on a double beam spectrophotometer (Unicam UV 500, ThermoSpectronic). Determining the concentration of PVA in culture supernatant entailed use of an iodometry micro-method, occurring once the cells had been removed by centrifugation (10,000×g, 10 min, 4 °C). In addition, the concentration of dissolved organic carbon (DOC) in the supernatant was determined via an automatic analyser (Shimadzu 5000). Every experiment was carried out in triplicate.
PVA Assay
Microplates were used to determine PVA concentration [15, 34]. Following centrifugation, 20 µl of a given sample, 42 µl boric acid solution (40 g/l) and 10 µl I2/KI solution (12.7 g I2 and 40 g KI per litre) were pipetted into a microplate well. Absorbance was measured at 660 nm by a microplate spectrophotometer (TECAN Sunrise, USA). The PVA concentrations present were calculated via a calibration curve.
Bacterial Growth on Tryptone or PVA
Growth tests were carried out in microplates. Portions of 190 µl of the desired types of mineral media supplemented with either 3 g/l tryptone or 1 g/l PVA were pipetted into every microplate well (each type of MM in eight wells). These wells were inoculated with 10 µl bacterial suspension, as prepared in the appropriate type of MM, and the plates were incubated at 25 °C. Blank wells inoculated with sterile MM and blank wells without substrate were produced in parallel. The growth of strains was monitored by OD600 measurement by a microplate spectrophotometer.
Results
Isolation and Identification of Bacterial Strains
Two strains of degrading bacteria were obtained from acclimated sludge suspensions using PVA agars amended with PQQ. Meanwhile, very slight growth of bacteria was recorded on the PVA agars without PQQ, whereas hardly any colony formation was found on the mineral agars. Both the isolates obtained were discerned as yellow-pigmented, non-fermenting rods, able to grow at temperatures ranging from 10 to 30 °C. For colony formation they required at least 7 days of incubation. Besides exhibiting growth on PVA agar with PQQ, they were able to grow on mineral agar amended with tryptone (1–3 g/l), during 5–6 days at 30 °C. Preliminary tests for PVA degradation showed that the strains possessed negligible potential for utilizing PVA without PQQ (data not shown). The strain designated OT2 was isolated from the WWTP in Otrokovice, treating mixed wastewater, while strain JK2 originated from the municipal WWTP in Zlín. Said bacterial strains were identified by 16S rDNA sequence analysis. Gene sequencing (1412 bp) revealed that the strains were members of the genus Sphingomonas and closely related to S. koreensis (Acc. No. NR 024998.1, similarity 98%). The sequences of the both strains were identical, so much so that the sequence of Sphingomonas sp. JK2 was deposited in the GenBank under the Acc. No. HQ727996.
Effect of Salt Concentrations on PVA Degradation
Initially, investigation was made into the effect of phosphate ion concentration. Phosphate salts are generally used as buffering agents in laboratory experiments, and they are almost ubiquitous at various concentrations in different waste and surface waters. The basal concentration of phosphate ions was set at 6.67 mmol/l, while a lower one and several elevated concentrations (of up to 20 mmol/l) were tested. The ratio of basic to dibasic phosphates remained constant in all tests, ensuring the same initial pH of the media. The courses of PVA degradation in both strains are given in Figs. 1 and 2.
The findings obtained showed that PVA degradation was very similar in both strains, but not the precisely the same. Besides the data given in Figs. 1 and 2, appropriate cell growth and decrease in DOC concentrations were found in all cases pertaining to decline in PVA concentration (data not shown). As indicated, the phosphate ion concentration of 6.67 mmol/l was seen as optimal for polymer utilization in both strains. Slightly different results were found in the case of 3.33 and 10.0 mmol/l; while under such conditions the process appeared moderately slowed down in the OT2 strain, nearly optimal degradation was recorded in the JK2 strain. For both strains, applying 13.3 mmol/l caused a significant delay in PVA decomposition, whereas the two highest concentrations of phosphate ions inhibited it completely. In order to find out if heightened phosphate concentrations can merely inhibit PVA degradation or inhibit common cell multiplication, growth tests on tryptone under different phosphate ion concentrations were prepared and performed. The results of these are given in Figs. 3 and 4.
Despite the crucial importance of phosphates as buffering agents and as a phosphorus source for bacterial metabolism, under the condition of 3 g/l of tryptone as the growth substrate, even just 10.0 mmol/l of phosphate ions managed to substantially diminish the rate of growth of the Sphingomonad strains. So as to test if this impact was caused by the phosphates themselves or by increased osmotic pressure, new tests were prepared and research carried out into bacterial growth on PVA under elevated concentrations of chloride. The tests were performed in mineral media containing the lowest concentration of phosphates (3.33 mmol/l), amended with PVA, PQQ and chloride at several concentrations. Both sodium and potassium salts were applied to distinguish the potential effect of the cation type, the results obtained being given in Figs. 5 and 6.
Clear evidence was found for a negative effect of elevated concentrations of chloride ions in both strains; adding 13.3 and 26.6 mmol/l totally inhibited PVA utilization for both the sodium and potassium salts (13.3 mmol represents 0.78 g/l NaCl or 0.99 g/l KCl). Meanwhile, only slight differences were discerned in the properties of the strains and the effect of the cation type when supplementing with lesser quantities of chloride. Concentrations of sodium chloride at 3.33 and 6.67 mmol/l subtly promoted the growth of the JK2 strain, whilst potassium chloride inhibited it slightly (see Fig. 5). The OT2 bacterium turned out to be very sensitive to any rise in salinity (see Fig. 6), as adding either chloride actually slowed down the utilization of PVA; it was noted that the potassium salt demonstrated a more conspicuous impact than the sodium one.
Effect of Inoculum Size on PVA Degradation
In order to determine the minimal inoculum size required to the biodegradation process, two lots of experiments were conducted in media that had been optimized as previously in the study. The act of obtaining initial cell concentrations from 100 to 107 cells/ml involved inoculating differently diluted portions of bacterial suspension into degradation tests. The findings on PVA degradation and cell growth are given in Figs. 7 and 8.
Nearly the same results were found in both series of tests, and in all cases successful PVA degradation was observed, even when the tiniest amounts of inoculum were applied. Under such conditions, merely relatively short delays were recorded, approximately 5 days in length, in the degradative process, which contrasted with the highest amounts of inoculum. Almost negligible variation in the extent of PVA degradation was seen for the OT2 strain when minimal and maximal inoculum levels were used (See Fig. 8).
Discussion
Effect of Inorganic Salts on PVA Biodegradation
The sensitivity of PVA-degrading Sphingomonads to low concentrations of inorganic salts has not been previously described. Herein, the initial series of tests in basal mineral medium containing 0.5 g/l NaCl revealed that just phosphate ion concentrations of 3.33–10.0 mmol/l ensured suitable conditions for both given strains to degrade PVA. Similarly, when mineral media with a concentration of phosphate ions at 3.33 mmol/l was amended with 13.3 mmol/l chloride, PVA utilization was fully inhibited. This chloride concentration corresponds to 0.78 g/l NaCl or 0.99 g/l KCl, and such concentrations are tolerable by many bacterial species. However, a phenomenon of sensitivity by some Sphingomonads to low NaCl concentrations is known to exist, as described by several authors. For example, Han et al. [35] isolated Sphingomonas oligoaromativorans from forest soil, observing it to grow optimally at 0.1 g/l NaCl, and other authors documented Sphingomonad strains growing optimally in the absence of NaCl [36,37,38,39]. However, certain Sphingomonas isolates are able to grow under circumstances of wide-ranging NaCl concentrations, as described by Huo et al. [40], who isolated Sphingomonas rubra growing in 0–80 g/l NaCl. In all cases, if halosensitive Sphingomonads turn out to be key microbes for PVA degradation either in waste or surface waters, inorganic salt concentrations may be crucial and a deciding factor in determining polymer degradation or recalcitrance to the same. Furthermore, halosensitivity must be taken into account when isolating PVA-degrading Sphingomonas from different sources. Herein, 0.5 g/l NaCl was successfully used within isolating agars, while Tokiwa [13] and Kim [14] applied 0.01 and 0.1 g/l, respectively. Therefore, utilizing several isolating media with various degrees of inorganic salt concentration might be an important factor for successfully isolating such PVA-degrading Sphingomonads.
Effect of Inoculum Size
The actual amount of bacterial degraders in contaminated matrices is of great importance to initiating the biodegradation process and to the rate of removal of the pollutant. Several studies have investigated this topic, arriving at the suggestion that a definite conclusion does not exist. Some research has revealed the absolute relevance of inoculum size to the process. For example, Ramadan et al. [41] found that Pseudomonas cepacia was able to mineralize p-nitrophenol in lake water when the water was inoculated with 3.3 × 104–3.6 × 105 cells per ml, whereas no mineralization was found for limited inoculation of water at 3.3 × 102 cells/ml. Similarly, Miethling and Karlson [42] described significant mineralization of pentachlorophenol in soil after inoculating it with 108 cells of Sphingomonas chlorophenolica RA2 per g; the inoculum size of 106 cells/g resulted in strongly reduced mineralization rates and 104 cells/g which did not significantly differ from the control.
In contrast, findings by Ayed et al. [43] and Chen et al. [44] showed that inoculum size exhibited a rather moderate effect on the biodegradation of Malachite Green by Sphingomonas paucimobilis and phenanthrene by Sphingomonas sp., respectively. Ayed et al. arrived at this conclusion after conducting tests with inoculum at sizes ranging 0.001–1.0 OD, whereas Chen et al. found that the effect of inoculum size (103–106 MPN/g) was less significant than the effect of salinity.
Herein, the authors tested the effect of initial cell count (100–107 per ml) on the biodegradation process, and, to their knowledge, it constitutes the first work to study such a broad range of inoculum sizes. From this point of view, achieving successful PVA biodegradation with minimal levels of inoculum might constitute a promising finding, suggesting that the process could proceed in different water environments even with limited bacterial communities. Certainly, this result would also be determined by low PVA toxicity. It is worthy of note that there are no signs of a toxic effect exhibited by the polymer against bacteria at the concentrations applied herein or in research conducted by other authors. It could be surmised that this apparent lack of PVA toxicity represents a crucial factor in biodegradation of the same by minimally sized inoculums, in comparison with the results of other biodegradation processes accomplished by Sphingomonads as cited above, in which the use of inoculums of limited size led to a significantly diminished extent of degradation.
Conclusion
Two PVA-utilizing Sphingomonad strains, isolated from regular wastewater treatment plants, revealed noticeable sensitivity to slightly increased concentrations of common inorganic salts. This phenomenon may influence polymer degradation during the wastewater treatment process if such concentrations of phosphate or chloride ions are present. Nevertheless, under suitable conditions the bacteria are able to utilize PVA in the presence of inoculums of very small size, corresponding to several cells per millilitre, hence might potentially ensure polymer degradation in a broader spectrum of surface waters.
References
Chiellini E, Corti A, D’Antone S, Solaro R (2003) Prog Polym Sci 28:963
Kawai F, Hu X (2009) Appl Microbiol Biotechnol 84:227
Suzuki T, Ichihara Y, Yamada M, Tonomura K (1973) Agric Biol Chem Tokyo 37:747
Watanabe Y, Morita M, Hamada N, Tsujisaka Y (1975) Agric Biol Chem Tokyo 39:2447
Sakazawa C, Shimao M, Taniguchi Y, Kato N (1981) Appl Environ Microbiol 41:261
Hashimoto S, Fujita. M (1985) J Ferment Technol 63:471
Fukae R, Fujii T, Takeo M, Yamamoto T, Sato T, Maeda Y, Sangen O (1994) Polym J 26:1381
Bharathiraja B, Jayamuthunagai J, Jayakumar M, Kirubakaran MA, Vivek P, Chandran M, Kumar RP (2013) Asian J Chem 25:8663
Matsumura S, Shimura Y, Terayama K, Kiyohara T (1994) Biotechnol Lett 16:1205
Mori T, Sakimoto M, Kagi T, Sakai T (1996) Biosci Biotechnol Biochem 60:330
Lim JG, Park DH (2001) J Microbiol Biotechnol 11:928
Kim MN, Yoon MG (2010) Polym Degrad Stab 95:89
Tokiwa Y, Kawabata G, Jarerat A (2001) Biotechnol Lett 23:1937
Kim BC, Sohn CK, Lim SK, Lee JW, Park W (2003) J Ind Microbiol Biotechnol 30:70
Vaclavkova T, Ruzicka J, Julinova M, Vicha R, Koutny M (2007) Appl Microbiol Biotechnol 76:911
Lee JA, Kim MN (2003) Polym Degrad Stab 81:303
Choi K, Park C, Kim S, Lyoo W, Sang HL, Lee J (2004) J Microbiol Biotechnol 14:1009
Yamatsu A, Matsumi R, Atomi H, Imanaka T (2006) Appl Microbiol Biotechnol 72:804
Hu X, Mamoto R, Fujioka Y, Tani A, Kimbara K, Kawai F (2008) Appl Microbiol Biotechnol 78:685
Zhang Y, Li Y, Shen W, Liu D, Chen J (2006) World J Microbiol Biotechnol 22:625
Rong D, Usui K, Morohoshi T, Kato N, Zhou MH, Ikeda T (2009) J Environ Biotechnol 9:131
Takeuchi M, Hamana K, Hiraishi A (2001) Int J Syst Evol Microbiol 51:1405
Maruyama T, Park HD, Ozawa K, Tanada Y, Sumino T, Hiraishi K, Kato K (2006) Int J Syst Evol Microbiol 56:85
Yabuuchi E, Kosako Y, Fujiwara N, Nata T, Matsunaga I, Ogura H, Kobayashi K (2002) Int J Syst Evol Microbiol 52:1485
Chen J, Zhang Y, Du GC, Hua ZZ, Zhu Y (2007) Enzyme Microb Technol 40:1686
Hatanaka T, Kawahara T, Asahi N, Tsuji M (1995) Biosci Biotechnol Biochem 59:1229
Takeuchi M, Kawai F, Shimada Y, Yokota A (1993) Syst Appl Microbiol 16:227
Kawai F (1999) J Ind Microbiol Biotechnol 23:400
Stolz A (2009) Appl Microbiol Biotechnol 81:793
Klomklang W, Tani A, Kimbara K, Mamoto R, Ueda T, Shimao M, Kawai F (2005) Microbiology 151:1255
Mamoto R, Nagai R, Tachibana S, Yasuda M, Tani A, Kimbara K, Kawai F (2006) Microbiology 152:1941
Kawai F, Kitajima S, Oda K, Higasa T, Charoenpanich J, Hu X, Mamoto R (2013) Arch Microbiol 195:131
Muchova M, Ruzicka J, Julinova M, Dolezalova M, Houser J, Koutny M, Bunkova L (2006) Water Sci Technol 60:965
Marusincova H, Husarova L, Ruzicka J, Ingr M, Navratil V, Bunkova L, Koutny M (2013) Int Biodeterior Biodegrad 84:21
Han SI, Lee JC, Ohta H, Whang KS (2014) Int J Syst Evol Microbiol 64:1679
Choi TE, Liu QM, Yang JE, Sun S, Kim SY, Yi TH, Im WT (2010) J Microbiol 48:760
Son HM, Kook M, Tran HT, Kim KY, Park SY, Kim JH, Yi TH (2014) Antonie Van Leeuwenhoek 105:791
Ahn JH, Kim BC, Kim JS, Lee GH, Song J, Kwon SW, Weon HY (2015) J Microbiol 53:673
Wei S, Wang T, Liu H, Zhang C, Guo J, Wang Q, Zhang Z (2015) Int J Syst Evol Microbiol 65:4644
Huo YY, Xu XW, Liu SP, Cui HL, Li X, Wu M (2011) Int J Syst Evol Microbiol 61:1028
Ramadan MA, El-Tayeb O, Alexander M (1990) Appl Environ Microbiol 56:1392
Miethling R, Karlson U (1996) Appl Environ Microbiol 62:4361
Ayed L, Chaieb K, Cheref A, Bakhrouf A (2009) World J Microbiol Biotechnol 25:705
Chen J, Wong MH, Wong YS, Tam NF (2008) Mar Pollut Bull 57:695
Funding
This study was funded by IGA/FT/2016/012 project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Měrková, M., Julinová, M., Houser, J. et al. An Effect of Salt Concentration and Inoculum Size on Poly(Vinyl Alcohol) Utilization by Two Sphingomonas Strains. J Polym Environ 26, 2227–2233 (2018). https://doi.org/10.1007/s10924-017-1122-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10924-017-1122-2