Abstract
Nelsonia is a Mexican endemic genus of woodrat and includes only two uncommon species: N. neotomodon and N. goldmani. This genus is of great phylogenetic interest, but has been ignored in most taxonomic studies and revisions due to the scarcity of its representatives in museum collections. The phylogenetic position of this genus is poorly known, and its interspecific and intraspecific relationships are unclear. The aim of this study was to infer the phylogenetic position of Nelsonia within the Cricetidae family, to determine whether the two species are monophyletic groups, and to assess the intraspecific genetic variation of N. goldmani. We amplified two mitochondrial markers: cytochrome b and cytochrome oxidase subunit 1 from specimens collected recently and museum samples from the type locality in 1903 to 1981 sampling half the known individuals reported to date. Sequences were analyzed by Bayesian and maximum likelihood to generate a phylogenetic hypothesis. Divergence time was performed to infer the biogeographic history. The genus Nelsonia was monophyletic and sister group of the current Xenomys, Hodomys, and Neotoma diverging in the middle-late Miocene. Nelsonia goldmani was also monophyletic, diverged from N. neotomodon during the late Miocene, and was formed by four highly divergent lineages. Further evidence may support a rank of full species for each of the four clades. Our results along with fossil data suggest that likely the genus Nelsonia diverged from Repomys or Protorepomys in the region of Californian-Rocky Mountains of the United States of America, and posteriorly invaded the Western Sierra Madre and Transmexican Volcanic Belt in Mexico.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nelsonia Merriam, 1897, is a distinctive woodrat genus of phylogenetic interest. It is endemic to Mexico and includes only two species: the western diminutive woodrat (Nelsonia neotomodon Merriam, 1987) and the Goldman’s diminutive woodrat (Nelsonia goldmani Merriam, 1903). They are rare and poorly known species, considered under special protection by the Mexican government. These woodrats inhabit highlands on rocky cliffs, slopes, and ravines in humid and temperate environments of conifer and cloud forests (García-Mendoza and López-Gonzáles 2005; León-Tapia and Cervantes 2019a). They occur allopatrically, N. neotomodon inhabiting only the Western Sierra Madre (WSM), while N. goldmani is found throughout the Transmexican Volcanic Belt (TVB).
This genus is of great phylogenetic interest due to the taxonomic troubles since its discovery (Carleton 1980). This is because it has been ignored in most taxonomic revisions due to the scarcity of the biological material in museum collections (Bradley et al. 2007; Miller and Engstrom 2008; León-Tapia and Cervantes 2019a). Few hypotheses have been proposed about its phylogenetic position; some morphological characters suggested that Nelsonia met many of the requirements that might be expected in an ancestor of Neotoma (Hooper 1954) and later morphological systematics proposed that Nelsonia is closely related to Neotoma, Hodomys, and Xenomys with a hypothetical ancestral position to them (Carleton 1980). Another hypothesis pointed out that the ancestral position of Nelsonia to Neotoma could be dismissed because Nelsonia presents derived dental characters (May 1981). However, a recent study based on molar characters revealed that Nelsonia represents the last survivor of the mostly extinct tribe Galushamyina (Martin and Zakrzewski 2019). Although the external morphology of these four genera are not similar, other morphological characters such as the molar pattern show similarities (Hooper 1954; Carleton 1980). However, some primitive morphological characters, karyotypic data (Hooper and Musser 1964), and spermatozoa (León-Tapia and Cervantes 2019b) suggested phylogenetic affinity with the group of peromyscines (Bradley et al. 2007).
Because of this phylogenetic uncertainly, the genus Nelsonia has had different taxonomic placements over time (Miller and Engstrom 2008). Molecular studies suggested close phylogenetic relationships among Xenomys, Neotoma, and Hodomys, but no specimens of Nelsonia were included in such works due to the difficulty of obtaining samples (Edwards et al. 2001; Edwards and Bradley 2001, 2002; Reeder et al. 2006; Longhofer and Bradley 2006; Miller and Engstrom 2008). With the molecular markers ND3, ND4L, ND4, and the tRNA arginine, one sample of N. neotomodon was phylogenetically related to Neotoma (Engel et al. 1998), but no samples of Xenomys and Hodomys were included.
On the other hand, the two species of Nelsonia have differing taxonomic placements as well; N. goldmani was included as subspecies of N. neotomodon (Hooper 1954), but external coloration and principally the presence of an anterorbital zygomatic notch in N. goldmani (Engstrom et al. 1992) and differences in the baculum and phallus (León-Tapia and Cervantes 2019b) distinguished the two species morphologically. These morphological differences may suggest that two lineages are present in the genus, but no formal phylogenetic analysis has been performed.
Currently, N. neotomodon is a monotypic species, while N. goldmani have two subspecies: N. g. goldmani and N. g. cliftoni. This taxonomic status was proposed on the differences of coloration and skull variation in samples of N. goldmani (Engstrom et al. 1992). The TVB where N. goldmani inhabits is one of the most complex mountain ranges in the world because of its topography and the variability of altitudes, environments, and climates (Gómez-Tuena et al. 2005; Ferrusquía-Villafranca et al. 2010), resulting in an intricate distribution pattern and high number of exclusive species that suggest this region is an evolutionarily active area (Escalante et al. 2004, 2007; Mastretta-Yanes et al. 2015). Hence, the historic process of the TVB could have had an effect on the isolation of populations of N. goldmani and genetic structure could be detected.
Therefore, the aim of the present study was to perform a phylogenetic inference to test the position of the genus Nelsonia within the Cricetidae family. To assess if N. goldmani in the TVB is a monophyletic group. Finally, to determine the intraspecific genetic variation in N. goldmani and whether monophyletic groups are formed between the two subspecies.
Material and Methods
Taxonomic Sampling
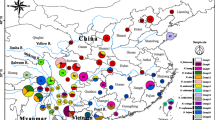
We assessed tissue samples of a total of 24 individuals of Nelsonia, two of N. neotomodon from museum collections and 22 of N. goldmani, five collected in a previous study (León-Tapia and Cervantes 2019a) and 17 skins from museums in the United States collected between 1903 to 1985, including specimens of the holotype series (Supplementary Material Table S1). These samples represent almost half the total number of individuals collected since the description of N. goldmani in 1903 (Fig. 1). Furthermore, we included 31 specimens of 13 cricetid species from the Colección Nacional de Mamíferos (CNMA) of Instituto de Biología, Universidad Nacional Autónoma de México. A total of 48 DNA sequences of cytocrome b (Cytb) and 19 DNA sequences of cytocrome oxidase subunit 1 (COI) of additional cricetid species were downloaded from GenBank and BOLD Systems in order to form a representative group of the subfamilies Sigmodontinae, Tylominae, and tribes of the Neotominae subfamily. All accession numbers of the samples sequenced in this study and downloaded sequences are available as Supplementary Material Table S1.
Localities of Nelsonia specimens included in this study with the five lineages found in the phylogenetic tree: N. netomodon as stars and N. goldmani as remaining geometric figures. The orange polygon represents the Transmexican Volcanic Belt and green polygon to Western Sierra Madre according to Morrone et al. (2017). Information of localities can be found in Supplementary Material Table S1
DNA Extraction, Amplification, and Sequencing
Genomic DNA from tissues and skins was extracted using the kits Quiagen Dneasy Tissue Kit® (Quiagen Inc., Valencia, CA, USA) and AxyPrep™ Multisource Genomic DNA Miniprep (Axygen Scientific Inc., Union City, CA, USA). For the museum samples, we obtained 1–3 mm2 of ventral skin that prior to DNA extraction were washed three times using 200 μl of STE solution (Hillis et al. 1996) and incubated at 56 °C each time.
We amplified the complete Cytb and COI mitochondrial genes by polymerase chain reaction (PCR; Saiki et al. 1988) using the thermal cycler parameters reported by Edwards and Bradley (2002) and Ivanova et al. (2007). We used two external and 14 internal primers reported in the literature for Cytb, and six external primers for COI (Irwin et al. 1991; Smith and Patton 1993; Smith and Patton 1999; Ivanova et al. 2007). In addition, two internal primer for Cytb and eight for COI were designed with the program OligoAnalyzer 3.1 (Integrated DNA Technologies, Inc., Coralville, IA, USA; Table 1) for the degraded DNA from skins. The DNA extraction and PCR reactions were performed independently by species and locality in order to avoid cross-contamination in skin samples as precautionary measures suggested in the literature (Martínková and Searle 2006; Sefc et al. 2007; Wandeler et al. 2007; Töpfer et al. 2011). The amplicons were purified using the kits QIAquick PCR purification (Quiagen Inc., Valencia, CA, USA) and AxyPrep™ PCR Clean-up (Axygen Scientific Inc., Union City, CA, USA), sequenced using the BigDye™ Terminator v1.1 Cycle Sequencing Kit, and analyzed with a 3100 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). All the resulting sequences were visually inspected and edited in BioEdit 7.0.9.0 (Hall 1999). In addition, some COI samples of the Cricetidae family were processed using the Ivanova et al. (2012) protocol at Guelph University, Canada as part of the project of bar-code of life in the CNMA (Supplementary Material Table S1).
Sequences were aligned with ClustalW algorithm in MEGAX (Kumar et al. 2018), and the final sequences deposited in the NCBI Genbank (accession numbers MK878906–MK878985; Supplementary Material Table S1). The Cytb and COI matrices were merged in Mesquite v3.04 (Maddison and Maddison 2015) to generate a final alignment concatenated.
Phylogenetic Reconstruction
The best fitting partitioning scheme (concatenated Cytb and COI matrices) and nucleotide substitution model were selected under the Akaike Information Criterion (AIC) in PartitionFinder Version 1.1 (Lanfear et al. 2012), and used for the phylogenetic inferences. Phylogenetic analyses were conducted with the Maximum Likelihood (ML) and Bayesian inference (BI) methods using sequences of Sigmodon mascotensis and Sigmodon hispidus as outgroup.
For ML inference we used RaxML v8 (Stamatakis 2014) with 1000 rapid bootstrap pseudoreplicates as nodal support. The BI was done in MrBayes v3.2.6 (Ronquist et al. 2012) using a random starting tree and unlinked parameters; the analysis was conducted with two independent runs and four Markov chains, 10 million generations, and sampling every 1000. Convergence and stationarity were assessed through values of the average standard deviation of split frequencies with Tracer 1.7.1 (Rambaut et al. 2018). The first 20% of generations were discarded as burnin and with the remaining trees a majority rule consensus tree with posterior probabilities was constructed.
Genetic Distances and Diversity
For each mitochondrial marker, the number of haplotypes, haplotype diversity, number of segregating sites, and nucleotide diversity were estimated with the package pegas v0.11 (Paradis 2010) in R software v3.3.2 (R Foundation for Statistical Computing, Vienna, Austria). The genetic mean distances between the principal clades were calculated under Kimura 2-parameter (K2p) substitution model using MEGAX (Kumar et al. 2018). Finally, in order to determine isolation by distance, we performed a Mantel Test estimating the euclidian geographic distances using the corrected coordinates (León-Tapia and Cervantes 2019a) with the K2p genetic distances using a Pearson correlation and 10,000 permutations to assess the significance of the observed value under the simulations with the package ade4 v1.7–13 (Chessel et al. 2004) in R software.
Divergence Time Estimation
We calculated divergence dates between taxa using BEAST v2.6.3 (Bouckaert et al. 2019) with the dataset partitioned and evolutionary models estimated in PartitionFinder. We used four calibration dates based on the available fossil record, rather than secondary calibrations in order to avoid a false distribution of age estimates (Schenk 2016). First, we used Copemys as the ancestor of all Neotominae after diverging from Sigmodontinae (Steppan et al. 2004), Repomys gustelyi as likely ancestor of genus Nelsonia (May 1981). The earliest fossil records were Copemys sp. 23.03–5.33 Myr (Williams et al. 2003) and Repomys gustelyi 10.3–1.8 Myr (Albright 1999). Two additional fossils for Reithrodontomys (1.8–6.89 Myr) and Neotoma (5.3–10.3 Myr) according to Steppan and Schenk (2017) were used. Two independent runs were carried out with 20 million generations, sampling every 2000. The relaxed clock log normal model was used with a Yule model diversification process. The convergence and stationarity within chains were checked in Tracer 1.7.1 (Rambaut et al. 2018). LogCombiner v2.6.3 was used to merge the two runs and discard 20% of trees as burnin, whereas the consensus tree was generated with TreeAnotator v2.6.3.
Data Availability
All data generated or analyzed during this study are included in this published article, supplementary information files, and are available from the corresponding author on reasonable request. All nucleotide sequences are available in Bold Systems DataBase (http://www.boldsystems.org/) and NCBI GenBank repository (https://www.ncbi.nlm.nih.gov/genbank) under the accession numbers included in the Supporting Information Table S1.
Results
Phylogenetic Reconstruction
The concatenated matrix of Cytb and COI consisted of 1801 base pairs (bp) for all cricetid species, amplifying the complete Cytb (1143 pb) for all specimens of Nelsonia, and 650 pb of the COI except for one sample of N. goldmani (USNM 125815). The best scheme was for Cytb and COI subsets and the best evolutionary model was GTR + I + Г for each marker (−Lnl = 23,057.48, AIC = 46,734.96148). The two phylogenetic analyses recovered similar topologies (Fig. 2) retrieving the clades with Sigmodon (subfamily Sigmodontinae), Nyctomys, Tylomys, and Ototylomys (subfamily Tylomyinae), and the subfamily Neotominae: Baiomys, Scotinomys (tribe Baiomyini), Reithrodontomys, Osgoodomys, Peromyscus (tribe Reithrodontomyini), Xenomys, Hodomys, Neotoma (tribe Neotomini), and Nelsonia.
Phylogenetic tree of Nelsonia obtained by Maximum Likelihood and Bayesian Inference based on cytochrome b and cytochrome oxidase subunit 1 mitochondrial markers. Values in branches show nodal support for each inference (bootstrap/posterior distribution). Within N. goldmani four major clades are shown
The genus Nelsonia was monophyletic with a high support values, a bootstrap support (BS) of 99% for ML analysis and a posterior distribution (PD) of 1.00 for BI. This clade was the sister group to the genera Xenomys, Hodomys, and Neotoma with 69% BS and 1.00 PD. On the other hand, N. neotomodon and N. goldmani were monophyletic with high nodal support.
Within N. goldmani, four clades were retrieved (Figs. 1 and 2): clade I with samples from the easternmost localities in Estado de México state with 92% BS and 1.00 PD; clade II with individuals from southern Pátzcuaro at the center of Michoacán state and nodal support of 100% BS and 1.00 PD; clade III with samples from western Michoacán state with low BS (46%) and high PD (1.00); and clade IV with specimens from the westernmost localities in southern Jalisco and northern Colima with high BS and PD values (88% and 1.00, respectively). Although low BS values (40–54%) determined the intraspecific relationships among these clades, the PD values were high (1.00). Specimens of clade I, II, and III correspond to the subspecies N. goldmani goldmani, while samples from clade IV correspond to the subspecies N. goldmani cliftoni.
Genetic Distances and Diversity
Within N. goldmani samples, we found 21 haplotypes for Cytb and 16 for COI; the nucleotide frequencies for Cytb were: A = 0.34, C = 0.28, G = 0.12, and T = 0.26, whereas for COI were: A = 0.29, C = 0.24, G = 0.17, and T = 0.30. The overall haplotype diversity was high for Cytb and COI, 0.995 and 0.976, respectively. The number of segregating sites were 259 and 114, the nucleotide diversity 0.0781 and 0.0624 for Cytb and COI, respectively. The haplotype diversity and nucleotide diversity were high for the four clades of N. goldmani (Table 2).
The K2p genetic distances showed that genus Nelsonia was divergent from the other clades (Table 3): the mean genetic distances within N. goldmani were 8.5% (0.26–15.73%) for Cytb and 6.9% (0.0–13.9%) for COI. On the other hand, the genetic distances among the four clades within N. goldmani were high in Cytb (7.2–9.8%) and COI (5.8–10.9%) principally between the farthest localities (Table 4). Finally, Mantel test showed that the correlation for the two mitochondrial genes was significant, for Cytb r = 0.259 (P = 0.008) and COI r = 0.644 (P < 0.001).
Divergence Time Estimation
The time divergence tree was similar to the ML and BI trees (Fig. 3). The genus Nelsonia diverged from the genera of Neotomini group by 11.13 Myr (7.64–14.88 Myr, 95% HDP) in the middle to late Miocene. The split between N. goldmani and N. neotomodon occurred 7.41 Myr (5.08–10.27 Myr, 95% HDP). The divergence times within the four lineages of N. goldmani occured from the late Miocene to early Pleistocene. The first cladogenetic event of clade I took place 5.07 Myr (3.16–7.21 Myr, 95% HDP), the divergence of clade II occurred 4.15 Myr (2.51–5.97 Myr, 95% HDP), and the third one between clade III and IV at 3.51 Myr (2.11–5.11 Myr, 95% HDP).
Divergence time reconstructed by Bayesian Inference in Beast using cytochrome b and cytochrome oxidase subunit 1 mitochondrial markers for the genus Nelsonia. Numbers indicate the mean divergence time in millions of years with the minimum and maximum value of the 95% highest posterior density in square brackets and showed as blue bars in the tree. The red nodes denote the fossil calibration points. Colors below the tree depict the geological time scale limits as reference. The roman numbers refer to the four lineages recovered within N. goldmani
Discussion
Phylogenetic Relationships of Nelsonia
The phylogenetic hypothesis resulting from this study was consistent with previous phylogenetic analyses and different molecular markers, for example Reeder et al. (2006), Miller and Engstrom (2008), and Steppan and Schenk (2017). These studies recovered the subfamilies Sigmodontinae, Tylomyinae, Neotominae, and the subclades within Neotominae that correspond to the Baiomini, Neotomini, and Reithrodontomyini tribes.
The monophyly of Nelsonia as sister group of the current species of Neotomini including Neotoma, Hodomys, and Xenomys was concordant with paleontological studies, in which the dental derived characters of Nelsonia are divergent from the Neotomini tribe (May 1981), and are useful for considering it as a highly derived member of the Galushamyini tribe (Martin and Zakrzewski 2019). Therefore, considering the dental and molecular characters, Nelsonia must be considered as the unique extant member of the tribe Galushamyini.
Genetic Relationships within Nelsonia
The monophyly with the high genetic distances between N. neotomodon and N. goldmani supports the morphological hypothesis generated with cranial characters that these forms represent two distinctive taxa. Engstrom et al. (1992) pointed out that the main discrimination of these groups was based on the presence/absence of the anteorbital zygomatic notch and Carleton (1980) considered its absence in N. neotomodon as a plesiomorphic character.
The monophyly with high haplotype diversity, nucleotide diversity, and genetic distances within N. goldmani was consistent with the four clades recovered, which showed a clear instraspecific genetic structure supported by significant isolation by distance with four distinctive lineages. Clade I with three specimens of two localities at the center of the TVB was the sister group of the remaining specimens; the next cladogenetic event was the south Pátzcuaro specimens (clade II), and the third cladogenetic split was between specimens from Tancítaro and Patambán (clade III) and those from the western portion of the geographic distribution (clade IV). The genetic distances among the clades (Table 4) showed high divergence values as the 6–13% reported in other rodent populations in the TVB, such as the pocket gophers Cratogeomys merriami, C. tylorhynus, and C. gymnurus (Demastes et al. 2002), and species of the rodent genera Baiomys, Neotoma, Reithrodontomys, and Sigmodon (Bell et al. 2001; Edwards and Bradley 2002; Amman and Bradley 2004; Carroll et al. 2005).
Our results revealed that the previously recognized subspecies based on morphological traits were congruent, to a certain extent, with the molecular intraspecific relationships. In the description of N. g. cliftoni, several cranial differences were reported with regard to N. g. goldmani, implying the existence of two subspecies (Genoways and Jones 1968). Nonetheless, Engstrom et al. (1992) demonstrated that such differences were due to the comparison of young specimens of N. g. goldmani with adult specimens of N. g. cliftoni, and the only meaningful cranial difference between them was the length of rostrum being greater in N. g. cliftoni. Another distinctive characteristic of N. g. cliftoni is the pelage coloration, which is paler dorsally and laterally than in N. g. goldmani, but it has been demonstrated in rodents and lagomorphs that this character can vary in response to specific environmental conditions (Demastes et al. 2002; Stoner et al. 2003; Hoekstra et al. 2005).
Our results recovered the clade IV with samples of N. g. cliftoni, but specimens of N. g. goldmani were grouped in three lineages. Based on the high genetic distances in N. goldmani (Table 4), the genetic species concept defined as “a group of genetically compatible interbreeding natural populations that is genetically isolated from other such groups” (Baker and Bradley 2006: 645) could explain more completely the taxonomy of the four clades detected in the analyses. Hence, based on this context, the four clades likely could represent four species rather than only one. However, more morphological or ecological evidence is necessary to clarify if they represent different species, as well as including new specimens, such as the recent individual discovered in the new locality in Morelos (González-Cózatl et al. 2016). We suggest that the two subspecies should be maintained until new evidence is presented: N. g. cliftoni (clade IV) with individuals from the western TVB, and N. g. goldmani (clade I, II, and III) with individuals from central TVB.
Biogeographic History of Nelsonia
Based on molar structures, May (1981) and recently Martin and Zakrzewski (2019) with phylogenetic analyses on molar characters, established that the extinct genus Repomys or Protorepomys could represent the ancestor lineages of Nelsonia. The main fossil record of R. gustelyi, R. maxumi, R. panacaensis, R. arizonensis, and P. mckayensis are distributed in the southwestern part of the USA (Martin and Zakrzewski 2019; Paleobiology Database https://paleobiodb.org) within the biogeographic region known as Californian-Rocky Mountains (Escalante et al. 2013). The mountain extensions in Mexico, such as the WSM, have worked as a historic corridor facilitating the dispersion process from north to south. Several mammalian species from the southwestern mountains of USA invaded the WSM that worked as dispersion route mainly in periods with higher moisture and suitable vegetation in highlands (Ferrusquía-Villafranca et al. 2010). Therefore, it is likely that that Nelsonia after diverging from Repomys or Protorepomys invaded the WSM in Mexico, but additional evidences are needed to confirm this hypothesis.
Based on morphological studies Engstrom et al. (1992) hypothesized that N. neotomodon and N. goldmani diverged by a vicariant process, result of a historical disruption of gene flow probably in Pliocene-Pleistocene or earlier. Our results showed that the divergence of the lineages from WSW (N. neotomodon) and TVB (N. g. cliftoni and N. g. goldmani) occurred during the late Miocene (Fig. 3) matching with a warmer and more humid climate than today (Micheels et al. 2007), which is characterized globally by increased aridity and seasonality and with the spread of grasslands and desert scrub in several areas of North America (Badgley et al. 2014). The increased temperatures from the Oligocene peaked at 17 Myr, posteriorly forming permanent ice sheets in Antarctica causing cooling in North America during the late Miocene-Pleistocene (Maguire and Stigall 2008). This climatic and changing vegetation likely had a strong influence for the cladogenetic event between lineages of WSM and TVB. Since the biogeographic isolation of the WSM and TVB approximately from 25 to 17 Myr ago (Ferrari et al. 1999; Gómez-Tuena et al. 2005), these regions could have contained the environmental suitability that Nelsonia required until the forest distribution was interrupted between the SWS and TVB by climate change. This hypothesis is supported by the facts that the ecological niche of N. goldmani is influenced by temperature climatic variables (León-Tapia and Cervantes 2019a) and N. neotomodon presents a clear increase pattern in the zygomatic plate breath of the specimens from north to south (Engstrom et al. 1992). This likely suggests that Nelsonia after divergence from Repomys or Protorepomys invaded the WSW from north to south and subsequently the TVB, restricting N. neotodomon in the WSM and remaining lineages at the TVB. Other rodent species diverged at similar time such as the genus Habromys, which originated in the Pliocene and diversified during the Pleistocene (León-Paniagua et al. 2007). However, it is necessary to include more samples of N. neotomodon, and to gather more evidence that links the dynamic history of the WSM and TVB with the speciation event between N. neotomodon and N. goldmani.
In addition to the isolation by distance explanation, for the observed genetic differentiation of the four lineages from TVB, the geologic history of the TVB could have had a strong impact on the diversification of the four lineages due to the formation of geographic barriers and new habitats (Ceballos et al. 2010; Ferrusquía-Villafranca et al. 2010). Besides, the dramatic climatic fluctuations registered during the late Miocene-Pleistocene could have modified the habitat limits of the lineages in several areas; this is supported by the cyclic altitudinal displacements, expansions, and contractions of the conifer forests (Martin and Harrell 1957; Van Devender 1990; McDonald 1993). This occurred particularly during the warm Pliocene period over 3 Myr (Contoux et al. 2012; Boer et al. 2017), in which the climate change resulted in a temperature increase, decrease of precipitation and a reduction of available soil moisture in North America that led to a subsequent reduction in the geographical coverage of forest-type vegetation replacing it with more open-type vegetation, such as grasslands and shrubland (Prescott et al. 2018).
These climatic changes could have isolated the four lineages in highlands during the interglacial periods as that recorded for conifer trees (Moreno-Letelier and Piñero 2009) and fragmentation in cloud mountain forests (Van Devender 1990). These factors have been related with the diversification processes in others montane species, such as rodents (Sullivan et al. 2000; Demastes et al. 2002; Amman and Bradley 2004; Hafner et al. 2005; León-Paniagua et al. 2007), plants (Rodríguez-Banderas et al. 2009; Ruiz-Sánchez and Specht 2013; Pérez-Crespo et al. 2017; Rodríguez-Gómez et al. 2017), insects (Anducho-Reyes et al. 2008; Arriaga-Jiménez et al. 2018), birds (McCormack et al. 2008; Zarza et al. 2016), and reptiles (Alvarado-Díaz and Campbell 2004; Zaldivar-Riverón et al. 2005; Bryson et al. 2011; Sunny et al. 2018). Thus, the complex geologic history, topographic, environmental heterogeneity, and climatic history of the TVB, undoubtedly influenced the geographic distribution of Nelsonia lineages. This biogeographic region has been scarcely explored for small mammals and the present study provides new information that highlights the need to carry out further research on endemic rodents from that region and their evolutionary history.
Conclusions
Genus Nelsonia is a monophyletic group and sister to the current three genera in the tribe Neotomini, which is concordant with molar phylogenetic analyses that support Nelsonia as the unique extant member of the Galushamyini tribe. Nelsonia diverged from the Neotomini tribe in the middle-late Miocene, and subsequently, N. neotomodon from WSW and the remaining lineages from TVB in the late Miocene. Four main lineages of N. goldmani from TVB had a high genetic differentiation, but more samples and new evidence are needed to confirm if the four clades are different species. The evolutionary history of these species is not fully understood; however, the western mountains in the United States and WSW played a significant role in the biogeographic history of this genus to finally settle down in the TVB, where the dramatic climatic changes isolated genetically the four evolutionary lineages recognized herein.
Data Availability
The data that support the findings of this study are openly available in GenBank; accession numbers, specimen information, and primers used are available as supplementary information.
References
Albright LB (1999) Magnetostratigraphy and biochronology of the San Timoteo Badlands, southern California, with implications for local Pliocene–Pleistocene tectonic and depositional patterns. Geol Soc Am Bull 111(9):1265–1293. https://doi.org/10.1130/0016-7606(1999)111<1265:MABOTS>2.3.CO;2
Alvarado-Díaz J, Campbell JA (2004) A new montane rattlesnake (Viperidae) from Michoacán Mexico. Herpetologica 60(2):281–286. https://doi.org/10.1655/03-40
Amman BR, Bradley RD (2004) Molecular evolution in Baiomys (Rodentia: Sigmodontinae) evidence for a genetic subdivision in B. musculus. J Mammal 85(1):162–166. https://doi.org/10.1644/1545-1542(2004)085<0162:MEIBRS>2.0.CO;2
Anducho-Reyes MA, Cognato AI, Hayes JL, Zuniga G (2008) Phylogeography of the bark beetle Dendroctonus mexicanus Hopkins (Coleoptera: Curculionidae: Scolytinae). Mol Phylogenet Evol 49(3):930–940. https://doi.org/10.1016/j.ympev.2008.09.005
Arriaga-Jiménez A, Rös M, Halffter G (2018) High variability of dung beetle diversity patterns at four mountains of the Trans-Mexican Volcanic Belt. PeerJ 6: e4468. https://doi.org/10.7717/peerj.4468
Badgley C, Smiley TM, Finarelli JA (2014) Great basin mammal diversity in relation to landscape history. J Mammal 95(6):1090–1106. https://doi.org/10.1644/13-mamm-s-088
Baker RJ, Bradley RD (2006) Speciation in mammals and the genetic species concept. J Mammal 87(4):643–662. https://doi.org/10.1644/06-MAMM-F-038R2.1
Bell DM, Hamilton MJ, Edwards CW, Wiggins LE, Martínez RM, Strauss RE, Bradley RD, Baker RJ (2001) Patterns of karyotypic megaevolution in Reithrodontomys: evidence from a cytochrome-b phylogenetic hypothesis. J Mammal 82(1):81–91. https://doi.org/10.1644/1545-1542(2001)082<0081:POKMIR>2.0.CO;2
Boer B, Haywood AM, Dolan AM, Hunter SJ, Prescott CL (2017) The transient response of ice volume to orbital forcing during the warm late Pliocene. Geophys Res Lett 44(20):10486–10494. https://doi.org/10.1002/2017GL073535
Bouckaert R, Vaughan TG, Barido-Sottani J, Duchêne S, Fourment M, Gavryushkina A, Heled J, Jones G, Kühnert D, De Maio N, Matschiner M, Mendes FK, Müller NF, Ogilvie HA, du Plessis L, Alex Popinga A, Andrew Rambaut A, David Rasmussen D, Igor Siveroni I, Marc A. Suchard, Wu CH, Xie D, Zhang C, Stadler T, Drummond AJ (2019) BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol 15(4): e1006650. https://doi.org/10.1371/journal.pcbi.1006650
Bradley RD, Durish ND, Rogers DS, Miller JR, Engstrom MD, Kilpatrick CW (2007) Toward a molecular phylogeny for Peromyscus: evidence from mitochondrial cytochrome-b sequences. J Mammal 88(5):1146–1159. https://doi.org/10.1644/06-MAMM-A-342R.1
Bryson RW, Murphy RW, Lathrop A, Lazcano-Villareal D (2011) Evolutionary drivers of phylogeographical diversity in the highlands of Mexico: a case study of the Crotalus triseriatus species group of montane rattlesnakes. J Biogeogr 38(4):697–710. https://doi.org/10.1111/j.1365-2699.2010.02431.x
Carleton MD (1980) Phylogenetic relationships in neotomine-peromyscine rodents (Muroidea) and a reappraisal of the dichotomy within New World Cricetinae. Occas Pap Mus Zool Univ Mich 157:1–146
Carroll DS, Peppers LL, Bradley RD (2005) Molecular systematics and phylogeography of the Sigmodon hispidus species group. In: Sánchez-Cordero V, Medellín RA (eds) Contribuciones Mastozoológicas en Homenaje a Bernardo Villa. CONABIO, Mexico, pp 87–100
Ceballos G, Arroyo-Cabrales J, Ponce E (2010) Effects of Pleistocene environmental changes on the distribution and community structure on the mammalian fauna of Mexico. Quat Res 73(3):464–473. https://doi.org/10.1016/j.yqres.2010.02.006
Chessel D, Dufour AB, Thioulouse J (2004) The ade4 package-I- One-table methods. R News 4:5–10
Contoux C, Ramstein G, Jost A (2012) Modelling the mid-Pliocene warm period climate with the IPSL coupled model and its atmospheric component LMDZ5A. Geosci Model Dev 5(3):903–917. https://doi.org/10.5194/gmd-5-903-2012
Demastes JW, Spradling TA, Hafner MS, Hafner DJ, Reed DL (2002) Systematics and phylogeography of pocket gophers in the genera Cratogeomys and Pappogeomys. Mol Phylogenet Evol 22(1):144–154. https://doi.org/10.1006/mpev.2001.1044
Edwards CW, Bradley RD (2001) Molecular phylogenetics of the Neotoma floridana species group. J Mammal 82(3):791–798. https://doi.org/10.1644/1545-1542(2001)082<0791:MPOTNF>2.0.CO;2
Edwards CW, Bradley RD (2002) Molecular systematics of the genus Neotoma. Mol Phylogenet Evol 25(3):489–500. https://doi.org/10.1016/S1055-7903(02)00294-4
Edwards CW, Fulhorst CF, Bradley RD (2001) Molecular phylogenetics of the Neotoma albigula species group: further evidence of a paraphyletic assemblage. J Mammal 82(2):267–279. https://doi.org/10.1644/1545-1542(2001)082<0267:MPOTNA>2.0.CO;2
Engel ST, Hogan KM, Taylor JF, Davis SK (1998) Molecular systematics and paleobiogeography of the South American rodents. Mol Biol Evol 15(1):35–49. https://doi.org/10.1093/oxfordjournals.molbev.a025845
Engstrom MD, Sánchez-Herrera O, Urbano-Vidales G (1992) Distribution, geographic variation, and systematic relationships within Nelsonia (Rodentia:Sigmodontinae). Proc Biol Soc Wash 105:867–881
Escalante T, Morrone JJ, Rodríguez-Tapia G (2013) Biogeographic regions of North American mammals based on endemism. Biol J Linnean Soc 110(3):485–499. https://doi.org/10.1111/bij.12142
Escalante T, Rodríguez G, Gámez N, León-Paniagua L, Barrera O, Sanchéz-Cordero V (2007) Biogeografía y conservación de los mamíferos. In: Luna I, Morrone JJ, Espinosa D (eds) Biodiversidad de la Faja Vólcanica Transmexicana. UNAM, D.F., pp 485–502
Escalante T, Rodríguez G, Morrone JJ (2004) The diversification of Nearctic mammals in the Mexican transition zone. Biol J Linnean Soc 83(3):327–339. https://doi.org/10.1111/j.1095-8312.2004.00386.x
Ferrari L, López-Martínez M, Aguirre-Díaz G, Carrasco-Núñez G (1999) Space-time patterns of Cenozoic arc volcanism in central Mexico: from the Sierra Madre Occidental to the Mexican Volcanic Belt. Geology 27(4):303–306. https://doi.org/10.1130/0091-7613(1999)027<0303:STPOCA>2.3.CO;2
Ferrusquía-Villafranca I, Arroyo-Cabrales J, Martínez-Hernández E, Gama-Castro J, Ruiz-González J, Polaco OJ, Johnson E (2010) Pleistocene mammals of México: a critical review of regional chronofaunas, climate change response and biogeographic provinciality. Quat Int 217(1–2):53–104. https://doi.org/10.1016/j.quaint.2009.11.036
García-Mendoza DF, López-Gonzáles GC (2005) Diminutive woodrat (Nelsonia neotomodon) in Chihuahua, Mexico. Southwest Nat 50(4):530–506. https://doi.org/10.1894/0038-4909(2005)050[0503:DWNNIC]2.0.CO;2
Genoways HH, Jones JK (1968) A new mouse of the genus Nelsonia from southern Jalisco, México. Proc Biol Soc Wash 81:97–100
Gómez-Tuena A, Orozco-Esquivel MT, Ferrari L (2005) Petrogénesis ígnea de la Faja Volcánica Transmexicana. Bol Soc Geol Mex 57(3):227–283. https://doi.org/10.18268/bsgm2005v57n3a2
González-Cózatl FX, Vallejo RM, Arellano E (2016) First record of Nelsonia goldmani in the state of Morelos, Mexico. Rev Mex Biodivers 87(2):545–547. https://doi.org/10.1016/j.rmb.2015.12.001
Hafner MS, Light JE, Hafner DJ, Brant SV, Spradling TA, Demastes JW (2005) Cryptic species in the Mexican pocket gopher, Cratogeomys merriami. J Mammal 86(6):1095–1108. https://doi.org/10.1644/05-MAMM-A-064R1.1
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hillis DM, Moritz G, Mable BK (1996) Molecular Systematics. Second ed. Sinauer, Sunderland, Massachusetts
Hoekstra HE, Krenz JG, Nachman MW (2005) Local adaptation in the rock pocket mouse (Chaetodipus intermedius): natural selection and phylogenetic history of pupolations. Heredity 94(2):217–228. https://doi.org/10.1038/sj.hdy.6800600
Hooper TE (1954) A synopsis of the cricetine rodent genus Nelsonia. Occas Pap Mus Zool Univ Mich 558:1–12
Hooper TE, Musser GG (1964) The glans penis in Neotropical cricetines (Family Muridae) with comments on classification of muroid rodents. Occas Pap Mus Zool Univ Mich 123:1–57
Irwin DM, Kocher TD, Wilson AC (1991) Evolution of the cytochrome b gene in mammals. J Mol Evol 32(2):37–55. https://doi.org/10.1007/BF02515385
Ivanova NV, Clare EL, Borisenko AV (2012) DNA Barcoding in mammals. In: Kress W, Erickson D (eds) DNA Barcodes. Methods in Molecular Biology (Methods and Protocols). Humana Press, Totowa, pp 153-182. https://doi.org/10.1007/978-1-61779-591-6_8
Ivanova NV, Zemlak TS, Hanner RS, Hebert PDN (2007) Universal primer cocktails for fish DNA barcoding. Mol Ecol Notes 7(4):544–548. https://doi.org/10.1111/j.1471-8286.2007.01748.x
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29(6):1695–1701. https://doi.org/10.1093/molbev/mss020
León-Paniagua L, Navarro-Sigüenza AG, Hernández-Baños BE, Morales JC (2007) Diversification of the arboreal mice of the genus Habromys (Rodentia: Cricetidae: Neotominae) in the Mesoamerican highlands. Mol Phylogenet Evol 42(3):653–664. https://doi.org/10.1016/j.ympev.2006.08.019
León-Tapia MA, Cervantes FA (2019a) Noteworthy records and ecological niche modeling of the rare and endangered Goldman’s diminutive woodrat Nelsonia goldmani (Rodentia: Cricetidae) endemic to central Mexican highlands. Mammalia 83(4):330–342. https://doi.org/10.1515/mammalia-2018-0023
León-Tapia MA, Cervantes FA (2019b) First contribution to the description of reproductive structures of Nelsonia goldmani (Rodentia: Cricetidae). Therya 10(2):155–160. https://doi.org/10.12933/therya-19-747 ISSN 2007-3364
Longhofer LK, Bradley RD (2006) Molecular systematics of the genus Neotoma based on DNA sequences from intrón 2 of the alcohol dehydrogenase gene. J Mammal 87(5):961–970. https://doi.org/10.1644/05-MAMM-A-355R1.1
Maddison WP, Maddison DR (2015) Mesquite: a modular system for evolutionary analysis. Version 3.04. http://mesquiteproject.org.
Maguire KC, Stigall AL (2008) Paleobiogeography of Miocene Equinae of North America: a phylogenetic biogeographic analysis of the relative roles of climate, vicariance, and dispersal. Palaeogeogr Palaeoclimatol Palaeoecol 267(3–4):175–184. https://doi.org/10.1016/j.palaeo.2008.06.014
Martin RA, Zakrzewski RJ (2019) On the ancestry of woodrats. J Mammal 100(5):1564–1582. https://doi.org/10.1093/jmammal/gyz105
Martin PS, Harrell BE (1957) The Pleistocene history of temperate biotas in Mexico and eastern United States. Ecology 38(3):468–480. https://doi.org/10.2307/1929892
Martínková N, Searle JB (2006) Amplification success rate of DNA from museum skin collections: a case study stoats from 18 museums. Mol Ecol Notes 6(4):1014–1017. https://doi.org/10.1111/j.1471-8286.2006.01482.x
Mastretta-Yanes A, Moreno-Letelier A, Piñero D, Jorgensen T, Emerson B (2015) Biodiversity in the Mexican highlands and the interaction of geology, geography and climate within the Trans-Mexican Volcanic Belt. J Biogeogr 42(9):1586–1600. https://doi.org/10.1111/jbi.12546
May SR (1981) Repomys (Mammalia: Rodentia gen. nov.) from the late Neogene of California and Nevada. J Vertebr Paleontol 1(2):219–230. https://doi.org/10.1080/02724634.1981.10011893
McCormack JE, Peterson AT, Bonaccorso E, Smith TB (2008) Speciation in the highlands of Mexico: genetic and phenotypic divergence in the Mexican jay (Aphelocoma ultramarina). Mol Ecol 17(10):2505–2521. https://doi.org/10.1111/j.1365-294X.2008.03776.x
McDonald JA (1993) Phytogeography and History of the Alpine-Subalpine Flora of Northeastern Mexico. Biological Diversity in Mexico: Origins and Distribution. Oxford University Press, New York
Micheels A, Bruch AA, Uhl D, Utescher T, Mosbrugger V (2007) A late Miocene climate model simulation with ECHAM4/ML and its quantitative validation with terrestrial proxy data. Palaeogeogr Palaeoclimatol Palaeoecol 253(1–2):251–270. https://doi.org/10.1016/j.palaeo.2007.03.042
Miller JR, Engstrom MD (2008) The relationships of major lineages within peromyscine rodents: a molecular phylogenetic hypothesis and systematic reappraisal, J Mammal 89(5):1279–1295. https://doi.org/10.1644/07-MAMM-A-195.1
Moreno-Letelier A, Piñero D (2009) Phylogeographic structure of Pinus strombiformis Engelm. across the Chihuahuan Desert filter-barrier. J Biogeogr 36(1):121–131. https://doi.org/10.1111/j.1365-2699.2008.02001.x
Morrone JJ, Escalante T, Rodríguez-Tapia G (2017) Mexican biogeographic provinces: map and shapefiles. Zootaxa 4277(2):277–279. doi: https://doi.org/10.11646/zootaxa.4277.2.8
Paradis E (2010) pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics 26(3):419–420. https://doi.org/10.1093/bioinformatics/btp696
Pérez-Crespo MJ, Ornelas JF, González-Rodríguez A, Ruiz-Sánchez E, Vásquez-Aguilar AA, Ramírez-Barahona S (2017) Phylogeography and population differentiation in the Psittacanthus calyculatus (Loranthaceae) mistletoe: a complex scenario of climate–volcanism interaction along the Trans-Mexican Volcanic Belt. J Biogeogr 44(11):2501–2514. https://doi.org/10.1111/jbi.13070
Prescott CL, Dolan AM, Haywood AM, Hunter SJ, Tindall JC (2018) Regional climate and vegetation response to orbital forcing within the mid-Pliocene warm period: a study using HadCM3. Global Planet Change 161:231–243. https://doi.org/10.1016/j.gloplacha.2017.12.015
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67(5):901-904. https://doi.org/10.1093/sysbio/syy032
Reeder SA, Darin SC, Edwards CW, Kilpatrick CW, Bradley RD (2006) Neotomine-peromyscine rodent systematics based on combined analyses of nuclear and mitochondrial DNA sequences. Mol Phylogenet Evol 40(1):251–258. https://doi.org/10.1016/j.ympev.2006.03.016
Rodríguez-Banderas A, Vargas-Mendoza CF, Buonamici A, Vendramin GG (2009) Genetic diversity and phylogeographic analysis of Pinus leiophylla: a post-glacial range expansion. J Biogeogr 36(9):1807–1820. https://doi.org/10.1111/j.1365-2699.2009.02104.x
Rodríguez-Gómez F, Oyama K, Ochoa-Orozco M, Mendoza-Cuenca L, Gaytán-Legaria R, González-Rodríguez A (2017) Phylogeography and climate-associated morphological variation in the endemic white oak Quercus deserticola (Fagaceae) along the Trans-Mexican Volcanic Belt. Botany 96(2):121–133. https://doi.org/10.1139/cjb-2017-0116
Ronquist F, Teslenko M, Van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542. https://doi.org/10.1093/sysbio/sys029
Ruiz-Sánchez E, Specht CD (2013) Influence of the geological history of the Trans-Mexican Volcanic Belt on the diversification of Nolina parviflora (Asparagaceae: Nolinoideae). J Biogeogr 40(7):1336–1347. https://doi.org/10.1111/jbi.12073
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Erlich, HA (1988) Primer- directed enzymatic amplification of DNA with a thermo-stable DNA polymerase. Science 239:487–491
Schenk JJ (2016) Consequences of secondary calibrations on divergence time estimates. PLoS ONE 11(1):e0148228. https://doi.org/10.1371/journal.pone.0148228
Sefc KM, Payne RB, Sorenson MD (2007) Single base errors in PCR products from avian museum specimens and their effect on estimates of historical genetic diversity. Conserv Genet 8(4):879–884. https://doi.org/10.1007/s10592-006-9240-8
Smith MF, Patton JL (1993) The diversification of South American rodents: evidence from mitochondrial sequence data for the Akodontine tribe. Biol J Linnean Soc 50(3):149–177. https://doi.org/10.1006/bijl.1993.1052
Smith MF, Patton JL (1999) Phylogenetic relationships and the radiation of Sigmodontine rodents in South America: evidence from cytochrome b. J Mammal Evol 6:89–128. https://doi.org/10.1023/A:1020668004578
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Steppan SJ, Adkins RM, Anderson J (2004) Phylogeny and divergence-date estimates of rapid radiations in muroid rodents based in multiple nuclear genes. Syst Biol 53(4):533–553. https://doi.org/10.1080/10635150490468701
Steppan SJ, Schenk JJ (2017) Muroid rodent phylogenetics: 900-species tree reveals increasing diversification rates. PLoS ONE 12(8):e0183070. https://doi.org/10.1371/journal.pone.0183070
Stoner CJ, Bininda-Emonds ORP, Caro T (2003) The adaptive significance of coloration in lagomorphs. Biol J Linnean Soc 79(2):309–328. https://doi.org/10.1046/j.1095-8312.2003.00190.x
Sullivan J, Arellano E, Rogers DS (2000) Comparative phylogeography of Mesoamerican highland rodents: concerted versus independent response to past climatic fluctuations. Am Nat 155:755–768. https://doi.org/10.1086/303362
Sunny A, Monroy-Vilchis O, Zarco-González MM (2018) Genetic diversity and structure of Crotalus triseriatus, a rattlesnake of central Mexico. J Genet 97(5):1119–1130. https://doi.org/10.1007/s12041-018-1004-y
Töpfer T, Gamaut A, Haring E (2011) Utility of arsenic-treated bird skins for DNA extraction. BMC Res Notes 4:197. https://doi.org/10.1186/1756-0500-4-197
Van Devender TR (1990) Late Quaternary Vegetation and Climate of the Chihuahuan Desert, United States and Mexico: The Last 40,000 Years of Biotic Change. University of Arizona Press, Tucson
Wandeler P, Hoeck PEA, Keller LF (2007) Back to the future: museum specimens in population genetics. Trends Ecol Evol 22(12):634–642. https://doi.org/10.1016/j.tree.2007.08.017
Williams M, Boardman GL, Shiebout JA, Kilbourne B, Nguyen H (2003) Miocene of Fort Polk, western Louisiana, 2002-2003. J Vertebr Paleontol 23:110
Zaldivar-Riverón A, Nieto-Montes de Oca A, Laclette JP (2005) Phylogeny and evolution of dorsal pattern in the Mexican endemic lizard genus Barisia (Anguidae: Gerrhonotinae). J Zool Syst Evol Res 43(3):243–257. https://doi.org/10.1111/j.1439-0469.2005.00308.x
Zarza E, Faircloth BC, Tsai WLE, Bryson RW Jr, Klicka J, Mccormack JE (2016) Hidden histories of gene flow in highland birds revealed with genomic markers. Mol Ecol 25(20):5144–5157. https://doi.org/10.1111/mec.13813
Acknowledgements
We thank Posgrado en Ciencias Biológicas of Universidad Nacional Autónoma de México (UNAM), Instituto de Biología for their support of this research. Partial funding was provided through a master scholarship from Consejo Nacional de Ciencia y Tecnología (CONACYT), Mexico to MALT (CVU 346631). Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica (grant IN215711-2, PAPIIT, UNAM) helped with the financing of laboratory materials and reagents, and Comisión Nacional para el Conocimiento y Uso de la Biodiversidad (CONABIO) helped with partial support through grant HB029. We thank Alex V. Borisenko and Natalia V. Ivanova for their help in processing samples in the BOLD SYSTEMS project (code FCMUN). Laura Margarita Márquez Valdelamar helped in sequencing samples at the Instituto de Biología. Curators and museums kindly approved and afforded the skin samples for the study: Celia López González (CIIDIR Durango), Candace McCaffery (Florida Museum of Natural History), Jack Dumbacher and Maureen Flannery (California Academy of Sciences), Alfred L. Gardner and Suzanne C. Peurach (National Museum of Natural History), Mark S. Hafner (Louisiana State University Museum of Natural Science), Jim Dines (Natural History Museum Los Angeles County), and Robert M. Timm (University of Kansas).
Funding
Partial funding was provided through a master scholarship from Consejo Nacional de Ciencia y Tecnología (CONACYT), Mexico to MALT (CVU 346631). Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica (grant IN215711–2, PAPIIT, UNAM) helped with the financing of laboratory materials and reagents, and Comisión Nacional para el Conocimiento y Uso de la Biodiversidad (CONABIO) helped with partial support through grant HB029.
Author information
Authors and Affiliations
Contributions
Both authors contributed to the study conception and design. Data collection and analysis were performed by MALT. The first draft of the manuscript was written by MALT, and both authors commented on previous versions of the manuscript. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest or competing interest.
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Code Availability
Not applicable.
Supplementary Information
ESM 1
(XLSX 18 kb)
Rights and permissions
About this article
Cite this article
León-Tapia, M.Á., Cervantes, F.A. Systematics and the Unexpected High Mitochondrial Genetic Divergence of Nelsonia goldmani (Rodentia: Cricetidae) from Mexican Highlands. J Mammal Evol 28, 939–951 (2021). https://doi.org/10.1007/s10914-020-09532-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10914-020-09532-7