Abstract
Novel benzothiazole Schiff bases L1 [1-((4,6-difluorobenzo[d]thiazol-2-ylimino)methyl) naphthalen-2-ol], L2 [3-((4,6-difluorobenzo[d]thiazol-2-ylimino) methyl)benzene-1,2-diol], L3 [2-((4,6-difluorobenzo[d]thiazol-2-ylimino)methyl)-5-methoxyphenol], L4 [2-((4,6-difluorobenzo[d]thiazol-2-ylimino)methyl)-4-chlorophenol] and their binary Cu(II) complexes were synthesized. The structures of all the compounds have been discussed on the basis of elemental analysis, FT-IR, NMR, UV-Visible, ESI-Mass, TGA, ESR, SEM, powder XRD and magnetic moments. Based on the analytical and spectral data a square planar geometry has been assigned to all complexes in which the Schiff bases act as monobasic bidentate ligands, coordinating through the azomethine nitrogen and phenolic oxygen atom. DNA binding ability of these complexes was studied on CT-DNA by using UV-Vis absorption, fluorescence and viscometry. DNA cleavage ability of the complexes was examined on pBR322 DNA by using gel electrophoresis method. All the DNA binding studies reveal that they are good intercalators. The bioefficacy of the ligands and their complexes was examined against the growth of bacteria and fungi in vitro to evaluate their antimicrobial potential. The screening data revealed that the complexes showed more antimicrobial activity than the corresponding free ligands.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA is a significant cellular receptor, many chemicals bring to bear their antitumor effects by binding to DNA and by this means change the replication of DNA and inhibit the growth of the cancer cells, which is the basis of preparing new and more efficient antitumor drugs [1–3]. During the last decade several transition metal complexes have been used as tools for understanding DNA structure, as agents for mediation of DNA cleavage or as chemotherapeutic agents. Benzothiazole nucleus possesses broad range of biological properties such as antitumour [4, 5], cytotoxic [6], antiglutamate/antiparkinson [7], inhibition of enzymes [8]. Moreover, Schiff bases derived from 2-aminobenzothiazole are of great importance due to their wide applications as antimicrobial, anti-inflammatory, anti-degenerative and anti-HIV agents [9–13]. There is an increasing prominence and evergreen curiosity in the field of metal containing drugs. Medicinal inorganic chemistry covering a wide applications of metals in therapeutics and diagnostics. The biological behaviour of Cu(II) complexes has been subjected to intense investigation for DNA binding and cleavage activities [14] for novel chemotherapeutics and highly sensitive diagnostic agents [15]. Metal complexes can bind to DNA via covalent or non-covalent type of binding (intercalating, groove or electrostatic). Copper has a long history of medicinal application, the anticancer and antiparasitic properties of Cu(II) complexes have increased the interest of scientists in their thorough studies [16, 17]. Synthesis, characterization, biological activity and DNA interaction of Cu(II) complexes of isoxazole Schiff bases were reported earlier from our laboratory [18, 19].
The aim of the present study is to synthesize the desired Cu(II) complexes from benzothiazole cored Schiff bases and to study the DNA binding and cleavage efficiency. Further, to investigate their effect on pathogenic strains of bacteria and fungi.
Experimental
Materials
All the chemicals and solvents used were purchased from Sigma-Aldrich, Merck and Hi media Ltd. All the solvents such as methanol, chloroform, petroleum ether, acetone and water were purified by standard procedures. The pBR322 DNA and CT-DNA were purchased from Genei Biolab, Bangalore, India and stored at 4 °C. All the buffer solutions used in the DNA interaction studies were prepared from doubly distilled water.
Instrumentation
The elemental analysis of Schiff bases and their Cu(II) complexes was carried out by using Perkin Elmer 240C (USA) elemental analyzer. 1H-NMR and 13C-NMR spectra of the Schiff bases were recorded on Bruker 400 MHz NMR instrument using TMS as internal reference. FT-IR spectra of the compounds were recorded in the range 4000–400 cm−1 on Perkin-Elmer Infrared model 337 using KBr pellets. Electronic spectra were recorded on Shimadzu UV-VIS 1601 spectrophotometer. ESI mass spectra of all compounds were recorded on VG AUTOSPEC mass spectrometer. Magnetic susceptibilities of Cu(II) complexes were measured on Gouy balance model 7550 using Hg[Co(SCN)4] as standard. Melting temperatures of the ligands and their complexes were determined on Polmon instrument (Model No. MP-96). Copper content of the complexes was estimated by atomic absorption spectroscopy after decomposing the complexes with concentrated HNO3 using GBC Avanta 1.0 AAS. The ESR spectra were recorded at liquid nitrogen temperature on JES-FA200 ESR spectrometer (JEOL-Japan). The thermo gravimetric analysis was carried out in a dynamic nitrogen atmosphere with a heating rate of 10 °C min−1 using a Shimadzu TGA-50H in the temperature range of 27–1000 °C. Viscosity measurements were performed using Ostwald viscometer (Vensil). Powder X-ray diffraction (XRD) analysis was recorded on X’pert Pro diffractometer. The morphology of ligands and complexes was measured by SEM (scanning electron microscopy, Zeiss evo18).
General Procedure for the Synthesis of Schiff Bases
Hot methanolic solution of 4,6-difluorobenzo[d]thiazol-2-amine (10 mmol) was magnetically stirred in a round bottom flask followed by drop wise addition of 2-hydroxy naphthalene-1-carbaldehyde / 2,3-dihydroxy benzaldehyde / 2-hydroxy-4-methoxy benzaldehyde / 5-chloro-2-hydroxy benzaldehyde (10 mmol). The reaction mixture was then refluxed with stirring at 60–80 °C for 2 h. After cooling, the yellow solid precipitate was collected by filtration. The resultant product was washed and recrystallized with methanol. The Schiff bases were dried in vacuum desiccators over anhydrous CaCl2. The purity of the ligands was checked by TLC.
Ligand L 1: Yield 78 %. M. P: 175 °C. Mol Wt: 340. Anal. Calc. (C18H10F2N2OS) (%): C, 63.52; H, 2.96; N, 8.23. Found: C, 63.58; H, 3.02; N, 8.26. IR (KBr) (cm−1): υ(OH) 3468; υ(CH=N) 1623; υ(C-O) 1219. UV (CHCl 3 ) λmax/nm(cm−1): 385 (25974); 261 (38314). 1 H NMR (400 MHz, CDCl 3 ): δ = 14.29 (s, 1H), 10.18 (s, 1H), 8.30 (d, J = 8.5 Hz, 1H), 7.95 (d, J = 9.0 Hz, 1H), 7.78 (d, J = 8.0 Hz, 1H), 7.65–7.60 (m, 1H), 7.45–7.41 (m, 1H), 7.37–7.34 (m, 1H), 7.12 (d, J = 9.0 Hz, 1H), 7.05–6.99 (m, 1H), Fig. S1. 13 C NMR (100 MHz, CDCl 3 ): δ = 176.1, 160.5, 159.8, 158.4, 157.9, 151.0, 139.4, 138.2, 136.8, 133.6, 129.1, 127.3, 125.0, 123.7, 117.3, 108.1, 104.2, 100.2. MS (ESI): m/z = 341 [M + H]+.
Ligand L 2: Yield 75 %. M. P: 168 °C. Mol Wt: 306. Anal. Calc. (C14H8F2N2O2S) (%): C, 58.26; H, 2.44; N, 7.50. Found: C, 59.02; H, 2.50; N, 7.53. IR (KBr) (cm−1): υ(OH) 3468; υ(CH=N) 1633; υ(C-O) 1247. UV (CHCl 3 ) λmax/nm(cm−1): 387 (25839); 322 (31055); 263 (38022). 1 H NMR (400 MHz, CDCl 3 ): δ = 10.23 (s, 1H), 7.74 (s, 2H), 7.51–7.42 (m, 1H), 7.17–7.09 (m, 3H), 6.84–6.79 (m, 1H). 13 C NMR (100 MHz, CDCl 3 ): δ = 192.8, 166.6, 157.2, 155.1, 152.0, 149.7, 146.1, 137.5, 133.3, 122.5, 121.3, 119.5, 103.7, 101.0, Fig. S2. MS (ESI): m/z = 307 [M + H]+.
Ligand L 3: Yield 76 %. M. P: 172 °C. Mol Wt: 320. Anal. Calc. (C15H10F2N2O2S) (%): C, 56.24; H, 3.15; N, 8.73. Found: C, 56.29; H, 3.01; N, 8.71. IR (KBr) (cm−1): υ(OH) 3317; υ(CH=N) 1614; υ(C-O) 1247. UV (CHCl 3 ) λmax/nm(cm−1): 383 (26109); 263 (38022). 1 H NMR (400 MHz, CDCl 3 ): δ = 9.28 (s, 1H), 7.86 (s, 1H), 7.84 (s, 1H), 7.51–7.41 (m, 3H), 7.15–7.09 (m, 2H), 3.86 (s, 3H). 13 C NMR (100 MHz, CDCl 3 ): δ = 170.6, 166.1, 162.9, 157.6, 154.9, 152.3, 149.5, 137.3, 133.6, 113.0, 108.1, 104.4, 102.4, 100.9, 55.6. MS (ESI): m/z = 321 [M + H]+.
Ligand L 4: Yield 70 %. M. P: 169 °C. Mol Wt: 324. Anal. Calc. (C14H7F2N2ClOS) (%): C, 51.78; H, 2.17; N, 8.63. Found: C, 51.59; H, 2.09; N, 8.81. IR (KBr) (cm−1): υ(OH) 3471; υ(CH=N) 1647; υ(C-O) 1257. UV (CHCl 3 ) λmax/nm(cm−1): 333 (30030); 264 (37878). 1 H NMR (400 MHz, CDCl 3 ): δ = 12.06 (s, 1H), 9.27 (s, 1H), 7.50 (d, J = 2.7 Hz, 1H), 7.45–7.41 (m, 1H), 7.39–7.36 (m, 1H), 7.06–7.01 (m, 2H), Fig. S3. 13 C NMR (100 MHz, CDCl 3 ): δ = 174.8, 161.9, 130.5, 159.1, 157.2, 139.8, 135.6, 132.4, 130.9, 125.7, 120.9, 118.3, 104.5, 102.2. MS (ESI): m/z = 325 [M + H]+.
Synthesis of Binary Cu(II) Complexes
The preparation of all Cu(II) complexes were described in Scheme 1.
The complexes [Cu(L1)2], [Cu(L2)2], [Cu(L3)2] & [Cu(L4)2] were prepared by using the following general procedure. Hot methanolic solution (10 mL) of copper acetate monohydrate (0.5 mmol) was added drop wise to a hot methanolic solution (10 mL) of the corresponding Schiff bases (1 mmol). After complete addition, the mixture was refluxed with stirring at 60–80 °C for 3 h. The solid precipitate was separated out upon cooling. The precipitate was filtered under suction and washed with methanol and petroleum ether, finally kept for drying in vacuum desiccators over anhydrous CaCl2.
[Cu(L 1 ) 2 ]: Yield 70 %. M. P: 240 °C. Mol Wt: 742. Anal. Calc. (C36H18CuF4N4O2S2) (%): C, 58.26; H, 2.44; N, 7.50; Cu, 8.56. Found: C, 58.22; H, 2.50; N, 7.55; Cu, 8.51. IR (KBr)(cm−1): υ(C=N) 1612, υ(C-O) 1238, υ(M-O) 589, υ(M-N) 429. ESR: g║ = 2.2778, g⊥ = 2.0594, G = 4.6767. UV-Vis (DMSO) λmax/nm(cm−1): 257 (38610), 339 (23498), 381 (26246), 525 (19047). μeff (BM):1.83. MS (ESI): 765 [M + Na]+.
[Cu(L 2 ) 2 ]: Yield 72 %. M. P: 235 °C. Mol Wt: 673. Anal. Calc. (C28H14CuF4N4O4S2) (%): C, 49.89; H, 2.09; N, 8.13; Cu, 9.43. Found: C, 49.95; H, 2.15; N, 8.19; Cu, 9.48. IR (KBr)(cm−1): υ(C=N) 1622, υ(C-O) 1257, υ(M-O) 587, υ(M-N) 441. ESR: g║ = 2.3792, g⊥ = 2.0700, G = 5.4171. UV-Vis (DMSO) λmax/nm(cm−1): 260 (38461), 342 (29239), 385 (25974), 520 (19230). μeff (BM):1.84. MS (ESI): 691 [M + NH4]+.
[Cu(L 3 ) 2 ]: Yield 69 %. M. P: 230 °C. Mol Wt: 701. Anal. Calc. (C30H18CuF4N4O4S2) (%): C, 52.37; H, 2.20; N, 8.73; Cu, 9.05. Found: C, 52.43; H, 2.26; N, 8.81; Cu, 9.10. IR (KBr)(cm−1): υ(C=N) 1609, υ(C-O) 1256, υ(M-O) 550, υ(M-N) 429. ESR: g║ = 2.3689, g⊥ = 2.0658, G = 5.6063. UV-Vis (DMSO) λmax/nm(cm−1): 259 (38610), 314 (31847), 411 (24330). μeff (BM):1.74. MS (ESI): 624 [M + Na]+.
[Cu(L 4 ) 2 ]: Yield 74 %. M. P: 247 °C. Mol Wt: 708. Anal. Calc. (C28H12CuF4N4O2S2Cl)(%): C, 47.30; H, 1.70; N, 7.88; Cu, 9.97. Found: C, 47.33; H, 1.66; N, 7.81; Cu, 9.91. IR (KBr)(cm−1): υ(C=N) 1635, υ(C-O) 1262, υ(M-O) 559, υ(M-N) 462. ESR: g║ = 2.3719, g⊥ = 2.0613, G = 6.0668. UV-Vis (DMSO) λmax/nm(cm−1): 259 (38610), 339 (29498), 383 (26109), 517 (19342). μeff (BM):1.77. MS (ESI): 726 [M + NH4]+.
DNA Binding Studies
UV–Vis Spectroscopic Studies
The UV-Visible absorption spectroscopic method is used to study the DNA binding nature of metal complexes. The absorption titrations were carried out in Tris–HCl buffer (5 mM Tris–HCl/50 mM NaCl, pH 7.2) at 25 °C by keeping the Cu(II) complex concentration constant (10 μM) and varying the concentration of the CT-DNA from 0 to 10 μM. The solution of CT-DNA in the buffer gave a ratio of UV absorbance of 1.8: 1.9 at 260 nm and 280 nm, which indicates the CT-DNA was sufficiently free of protein [20]. The CT-DNA concentration per nucleotide was measured by employing an extinction coefficient of 6600 M−1 cm−1 at 260 nm [21]. To eliminate the absorbance of CT-DNA itself, equal increments of CT-DNA were added to both the sample solution and the reference solution.
Fluorescence Study
By the fluorescence spectral method, the relative binding of the complexes to CT-DNA was studied with an EB-bound CT-DNA solution in Tris-HCl buffer (pH 7.2) at 25 °C were excited at 350 nm by varying concentrations of complexes from 0 to 60 μM. The emission intensity showed reduction upon addition of the complex. The effect of the metal complexes on the emission intensity of EB-bound CT-DNA was used to determine the binding properties of the metal complexes. The binding constants were calculated from the slopes of the lines in the plot of the fluorescence intensity versus complex concentration. The relative binding of the complexes to CT-DNA was determined by Stern–Volmer eq. I0/I = 1 + KSV r.
Where I0 and I are the fluorescence intensities in the absence and presence of complexes respectively, KSV is a linear Stern–Volmer constant, and r is the concentration of complex to that of DNA.
Viscosity Measurements
Ostwald capillary viscometer was used to study the viscosities of CT-DNA at temperature 30 ± 0.1 °C in a thermostatic water bath. CT- DNA concentration (100 μM) was kept constant and complex concentrations (0–100 μM) were varied in Tris–HCl buffer (pH 7.2) solution. The flow time of the solution was measured by a digital stop-watch. An average flow time was taken after three times measurement for the same complex solution to maintain the accuracy. A plot was made between (η/ηo)1/3 and the ratio of the concentration of the Cu(II) complexes to CT-DNA, where η is the viscosity of CT-DNA in the presence of the complex and ηo is the viscosity of CT-DNA alone. Viscosity values were calculated from the experimental flow time of CT-DNA containing solutions corrected from the flow time of buffer alone.
DNA Cleavage Experiments
DNA cleavage experiments were performed with pBR322 DNA at pH 7.2 in Tris–HCl buffer solution by agarose-gel electrophoresis method by varying concentration of Cu(II) complexes. Oxidative DNA cleavage was monitored by treating pBR322 DNA with H2O2 and photolytic DNA cleavage in presence of UV light. The loading dye 0.25 % bromophenol blue is added to the samples and made to a total volume of 16 μL. The samples were incubated at 37 °C for 2 h. The samples were loaded in the agarose gel and electrophoresis was carried out at 75 V for 1 h.
Biological Evaluation
The ligands and their Cu(II)complexes were screened for their in vitro anti bacterial activity against gram negative Escherichia coli (E. coli), Pseudomonas putida (P. putida) and Klebsiella pneumoniae (K. pneumoniae) and gram positive Bacillus subtilis (B. subtilis) and Staphylococcus aureus (S. aureus), Ampicillin was used as standard drug. The antifungal activity was screened against Aspergillus niger (A. niger) and Candida albicans (C. albicans) by using Ketoconazole as standard drug. Nutrient agar was used as the medium for antimicrobial activity [22]. The sterilized medium was inoculated with the suspension of the microorganism and then poured into a petridish. The test compounds (1 mg/mL in DMSO solvent) were impregnated on the paper disc and then placed on the solidified medium. The plates were kept for incubation at 37 °C for 24 h.
Results and Discussion
All Cu(II) complexes are coloured, stable at room temperature and non-hygroscopic and melt at high temperatures. All the complexes are soluble in DMF and DMSO but insoluble in water, alcohol and chloroform. The analytical data obtained for all complexes are in good agreement with the theoretical values and the metal to ligand ratio is 1:2.
Spectral Characterization
FT-IR Spectra
The IR spectra of free Schiff base ligands were compared with the spectra of their corresponding Cu(II) complexes to understand the coordinating mode of the Schiff base ligands to Cu(II) centre in the complexes. A representative IR spectra of L3 and [Cu(L3)2] are given in Figs. S4 and S5. The Schiff base ligands showed a strong band in the region 1614–1642 cm−1 which corresponds to the azomethine (C = N) group. Further, this band is shifted to a lower frequency region 1609–1635 cm−1 indicating co-ordination of azomethine nitrogen to Cu(II) ion [23–25]. A strong intense band observed at 1219–1257 cm−1 in Schiff bases due to phenolic C–O stretching is shifted to higher frequency by 1238–1262 cm−1 indicating Schiff bases are coordinated through the phenolic oxygen to the metal atom. A broad band appeared in the range of 3317-3471 cm−1 attributed to υOH in the Schiff bases, which is disappeared in complexes indicating deprotonation of the phenolic proton prior to coordination. The co-ordination of the azomethine nitrogen and phenolic oxygen are further supported by the appearance of two new non ligand bands at 520–589 and 429–462 cm−1 due to υM–O and υM–N respectively [26, 27]. The IR absorption frequencies are given in Table 1.
Electronic Spectra and Magnetic Moments
The magnetic moment values (μeff) of all Cu(II) complexes are in the range of 1.74–1.84 BM which reveals the presence of a single unpaired electron [28, 29]. The Schiff base ligands exhibited characteristic high energy absorption bands in the range of 260–263 nm and 314–344 nm which are attributed to intraligand π-π* transitions. Other low energy bands observed at 381–387 nm are due to n-π* transition of non bonding electrons of azomethine nitrogen of Schiff base ligands [30, 31]. The Cu(II) complexes show a broad band in the lower frequency region 411–525 nm corresponds to the d-d transition due to 2B1g → 2Eg transition. A representative electronic spectra of L1 and [Cu(L1)2] are given in Fig. S6. Based on electronic spectral data and magnetic moment values a square planar geometry is assigned to all Cu (II) complexes [32].
SEM
The surface morphology of ligands and their metal complexes was investigated by scanning electron micrograph analysis. All the compounds were analyzed by EDX analysis to find the elements present on surface. SEM photographs and EDX graphs of L1, L2, [Cu(L1)2], [Cu(L2)2] are given in Fig. 1 and Fig. S7. The SEM analysis of ligands L1, L2, L3 and L4 showed irregular cubes, rod shaped, irregular tube and elongated flake like structures respectively and the complexes [Cu(L1)2], [Cu(L2)2], [Cu(L3)2] & [Cu(L4)2] resemble twisted fibre, granular, grass like, irregular particle shaped structures respectively. The EDX analysis of ligands contains C, H, N, O, S, F and Cu(II) complexes contain C, H, N, O, S, F & Cu elements. The appearance of no other elemental peaks in the spectra assures the purity of the compounds.
Powder XRD
The powder XRD patterns of all the compounds were recorded at 2θ = 10-80o range. The powder XRD patterns of L1, L3, [Cu(L1)2], [Cu(L3)2] are shown in Fig. 2 and Fig. S8 , display sharp crystalline peaks for the ligands indicating their crystalline nature, whereas the complexes do not exhibit well defined sharp peaks possibly due to their amorphous nature. The crystallite sizes are calculated from XRD peak by using the Debye-Scherrer’s equation (D = 0.9 λ / βcosθ). The crystallite sizes calculated for ligands are 26 nm (L1), 17 nm (L2), 26 nm (L3), 14 nm (L4) and for complexes are 17 nm [Cu(L1)2], 23 nm [Cu(L2)2], 8 nm [Cu(L3)2], 17 nm [Cu(L4)2].
ESR Spectra
The liquid state X band ESR spectra of all the Cu(II) complexes were recorded at liquid nitrogen temperature (77 K). The ESR spectra of Cu(II) complexes provide the information about the extent of the delocalization of unpaired electron. The ESR spectra of [Cu(L1)2], [Cu(L2)2] are shown in Fig. S9 and Fig. 3. The g║, g⊥ and G values are given in Table 2. The trend observed g║ > g⊥ > ge(2.0023) is a typical of Cu(II) d9 system with axial symmetry and the unpaired electron is localized in dx 2 -y 2 orbital with 2B1g ground state of Cu(II) square planar complex [33, 34]. The g║ values are found to be less than 2.3 suggesting the covalent nature of metal-ligand bond. The G values are more than 4 indicating negligible exchange interaction between the Cu(II) centres [35].
ESI Mass Spectral Studies
The ligands and their Cu(II) complexes have been studied for their mass spectral analysis. The ESI mass spectra of ligands exhibit [M + H]+ peaks at m/z 341(L1), 307(L2), 321(L3) and 325(L4). The ESI mass spectra of complexes exhibit peaks at m/z 765 [M + Na]+ for [Cu(L1)2], 691 [M + NH4]+ for [Cu(L2)2], 624 [M + Na]+ for [Cu(L3)2] and 726 [M + NH4]+ for [Cu(L4)2]. The m/z fragments of complexes confirm the stoichiometry of the complexes as [Cu(L)2] type. The representative mass spectra of L1, L2, [Cu(L1)2] and [Cu(L2)2] are given in Figs. S10-S13.
Thermal Analysis
The thermal decomposition of Cu(II) complexes was studied using thermo gravimetric technique. The experiment was performed at temperature 27 °C to 1000 °C at heating rate of 10 °C min−1 under nitrogen atmosphere. The TGA curves of complexes [Cu(L1)2], [Cu(L2)2] are shown in Fig. S14. The TGA curves of the complexes show a two step decomposition process. All the complexes are stable up to 200 °C confirming that there is no coordinated water molecules present in the complex molecule. The first weight loss occurred in the range of 209–336 °C corresponding to the partial loss of ligand moiety. The second mass loss is appeared at 375–789 °C corresponding to the complete departure of the complexes and the loss of their ligand portion. The residue was the copper oxide (CuO).
DNA Binding Studies
UV-Vis Spectroscopic Studies
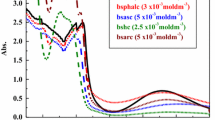
Electronic absorption spectroscopy is an effective technique to study the binding mode and extent of binding metal complexes with DNA [36, 37]. Absorption spectra of all the Cu(II) complexes were recorded in the absence and presence CT-DNA and shown in Fig. 4. The binding mode of the Cu(II) complexes to DNA through intercalation is characterized by the change in absorbance (hypochromism) and red shift in wavelength [38]. Hypochromism is observed due to the presence of aromatic chromophore which might facilitate the interaction of the complexes with the CT-DNA bases via non covalent π-π* interactions. When the complexes intercalate to the base pairs of CT-DNA, the π* orbital of the intercalated ligand in the complexes can couple with π orbital of the DNA base pairs, and then decreasing the π-π* transition energies. On the other hand, the coupling π* orbitals are partially filled by electrons, thus decreasing the transition probabilities [39, 40]. The absorption spectra of the Cu(II) complexes show a band at 407–379 nm. Upon incremental addition of CT-DNA, the band underwent hypochromism with a slight red shift. The Kb (intrinsic binding constant) is calculated from a plot of [DNA]/(ε a-ε f) Vs [DNA] using the equation: [DNA]/(ε a-ε f) = [DNA]/(ε b-ε f) +1/Kb(ε b-ε f), where [DNA] is the concentration of DNA in base pairs, εa is the apparent coefficient of Aobsd/[complex], εf and εb represent the extinction coefficients of the free and fully bound forms of the complex, respectively [41]. The binding constant Kb is calculated by the ratio of slope to the intercept. The binding constant Kb for [Cu(L1)2] is 8.5 ± 0.02 × 106 M−1,[Cu(L2)2] is 8.6 ± 0.02 × 105 M−1, [Cu(L3)2] is 2.86 ± 0.02 × 106 M−1 and [Cu(L4)2] is 1.91 ± 0.02 × 106 M−1. The Kb values are relatively less with the potential intercalators like ethidium bromide (EB) (Kb = 7 × 107 M−1) [42].
UV-Vis absorption spectra of complexes [Cu(L1)2], [Cu(L2)2], [Cu(L3)2] and [Cu(L4)2] in the absence (dashed line) and presence (solid line) of increasing amounts of CT-DNA in Tris HCl buffer (pH 7.2) at 25 °C. Conditions: [Complex] = 10 μM, [DNA] = 0–10 μM. Arrow (↓) shows the hypochromic and red shift upon increasing CT-DNA concentration. Inset: linear plot for the calculation of the intrinsic DNA binding constant, Kb
Fluorescence Study
Fluorescence experiments were performed to gain support for the binding mode of the complexes with CT-DNA. EB is one of the most sensitive fluorescent probes which can bind with DNA. There is an increase in the intensity of fluorescence spectra of EB in presence of CT-DNA because of intercalation between the DNA basepairs [43, 44]. The quenching of EB bound to DNA is calculated with the addition of increasing concentration of metal complexes. The Fig. 5 is showing the decrease in the intensity of EB with increase in the concentration of Cu(II) complexes. The reduction in the emission intensity is due to displacement of EB bound DNA with Cu(II) complexes [45]. The Ksv values calculated for all the complexes are 6.4 ± 0.02 × 104 M−1 [Cu(L1)2], 3.3 ± 0.02 × 104 M−1 [Cu(L2)2], 1.6 ± 0.02 × 104 M−1 [Cu(L3)2], 1.3 ± 0.02 × 104 M−1 [Cu(L4)2].
Fluorescence emission spectra of CT-DNA–EB system in the absence and presence of increasing concentration of complexes [Cu(L1)2], [Cu(L2)2], [Cu(L3)2] and [Cu(L4)2] in Tris HCl buffer (pH 7.2) at 25 °C. Conditions: [Complex] = 0–60 μM, [DNA] = 125 μM, [EB] = 12.5 μM. Arrow (↓) shows the emission intensity decreases upon increasing concentration of the complexes. Inset: I0/I versus r
Viscosity Measurements
The viscometric measurements were also carried out to further confirm the mode of interaction between the Cu(II) complexes and CT-DNA. A classical intercalation can cause an increase in the viscosity of DNA since it increases the separation between base pairs at intercalation site, hence an increase in overall length of the double helix [46, 47]. However a partial or non classical intercalation of the complex results in bending of the DNA helix, which decreases the effective length of DNA, with a concomitant decrease in its viscosity. The effects of all the Cu(II) complexes on the viscosity of DNA at 30 ± 0.1 °C are shown in Fig. 6. The relative specific viscosity of DNA is determined by increasing the concentration of the complex while the CT-DNA concentration was kept constant. By increasing the complex concentration to DNA the viscosity of DNA increases gradually which leads to an intercalative mode of binding [48]. These results revealed that the binding order of the complexes is found to be [Cu(L1)2] > [Cu(L2)2] > [Cu(L3)2] > [Cu(L4)2]. Among all Cu(II) complexes the increase in viscosity is more in the case of [Cu(L1)2]. This is due to lengthening of the DNA duplex by the extended planarity of bicyclic naphthalene moiety of [Cu(L1)2] which increases the intercalation compare to remaining complexes with monocyclic phenyl ring moiety [49].
DNA Cleavage Experiments
The DNA cleavage ability of Cu(II) complexes is investigated by agarose gel electrophoresis method with super coiled pBR322 DNA at different complex concentrations in the presence of H2O2 and UV light. The ability of the complexes in DNA cleavage is estimated by the conversion of DNA from Form I to Form II and Form III. During electrophoresis the fastest migration is detected in the supercoiled form (Form I). While the scission occurs on one strand the supercoiled form relaxes to generate nicked form (Form II), if both strands are cleaved linear form (Form III) appears which migrates in between Form I and Form II [50]. The concentration effect has been studied for the complexes by increasing the concentrations from 20 to 70 μM. As shown in Figs. 7 and 8 with increase in the complex concentration the circular supercoiled DNA converted to nicked DNA. Oxidative DNA cleavage of all the complexes were carried out in the presence of H2O2, there is no cleavage was observed in lane 1 and lane 2, while increasing the concentration of complexes (lane 3–8) form II is increased. The above results indicate that complexes can cleave the pBR322 DNA by oxidative cleavage via formation of hydroxyl radical. Photolytic DNA cleavage was performed in presence of UV irradiation at 345 nm, no DNA cleavage was observed in lane 1. From lane 2–8 DNA cleavage was observed by converting supercoiled DNA into nicked form, stating that cleavage capacity of the complexes is more at higher concentrations.
Oxidative cleavage of supercoiled pBR322 DNA(0.2 μg, 33.3 μM) at 37 °C in 5 mM Tris. HCl/5 mM NaCl buffer by the [Cu(L1)2] and [Cu(L2)2]. a Lane 1, DNA control; Lane 2, DNA + H2O2 (1 mM); Lane 3–8, DNA + H2O2 (1 mM) + [Cu(L1)2] (20, 30, 40, 50, 60 and 70 μM) respectively. b Lane 1, DNA control; Lane 2, DNA + H2O2 (1 mM); Lane 3–8, DNA + H2O2 (1 mM) + [Cu(L2)2] (20, 30, 40, 50, 60 and 70 μM) respectively
Photoactivated cleavage of supercoiled pBR322 DNA (0.2 μg, 33.3 μM) at 37 °C in 5 mM Tris. HCl/5 mM NaCl buffer by Cu(L1)2, Cu(L2)2 using UV irradiation of wavelength 345 nm. a Lane 1, DNA control; Lane 2–7, DNA + [Cu(L1)2] (20, 30, 40, 50, 60, 70 and 80 μM) respectively. b Lane 1, DNA control; Lane 2–7, DNA + [Cu(L2)2] (20, 30, 40, 50, 60, 70 and 80 μM) respectively
Biological Evaluation
The in vitro antimicrobial screening of all the ligands and their Cu(II) complexes were tested against the bacterial species E. coli, P. Putida, K. pneumoniae, B. subtilis and S. aureus and fungal species A. niger, and C. albicans by paper disc method. Ampicillin (antibacterial) and Ketoconazole (antifungal) were used as reference drugs. The antimicrobial activity stated that Cu(II) complexes show higher activity than corresponding free Schiff base ligands. The inhibition zone values of the compounds are summarized in Table 3. The inhibition zone values of all the Cu(II) complexes revealed that the complexes exhibit moderate to strong antimicrobial activity when compared with the standard drug (Fig. 9). The increase in antimicrobial activity of the complexes can be explained on the basis of Overtone’s concept [51] and Tweedy’s Chelation theory [52]. According to Overtone’s concept of cell permeability, the lipid membrane that surrounds the cell favors the passage of only the lipid soluble materials makes liposolubility as an important factor, which controls the antimicrobial activity. On chelation with metal ions the polarity of the metal ions reduced due to overlapping of ligand orbital and the partial sharing of its positive charge with the donor groups [53]. This process increases the lipophilicity of metal complexes [54].
Conclusion
Four novel Schiff bases and their binary Cu(II) complexes have been synthesized and characterized by analytical and spectral data. From the obtained data a square planar geometry has been assigned to all Cu(II) complexes. The UV-Vis absorption, fluorescence, and viscosity measurements revealed that the complexes bind with CT-DNA through an intercalation mode. From the DNA cleavage studies of Cu(II) complexes it is observed that the complexes effectively cleave supercoiled pBR322 DNA in the presence of H2O2 and also UV light. In vitro antimicrobial activity studies of Schiff base ligands and their Cu(II) complexes revealed that the activity increases upon coordination. Among all the complexes highest activity is observed for Cu(L1)2.
References
Song YM, Wu Q, Yang PJ, Luan NN, Wang LF, Liu YM (2006) DNA binding and cleavage activity of Ni(II) complex with all-trans retinoic acid. J Inorg Biochem 100:1685–1691
Tan C, Liu J, Chen L, Shi S, Ji L (2008) Synthesis, structural characteristics DNA binding properties and cytotoxicity studies of a series of Ru(III) complexes. J Inorg Biochem 102:1644–1653
Zuber G, Quada JC Jr, Hecht SM (1998) Sequence selective cleavage of a DNA Octanucleotide by chlorinated Bithiazoles and Bleomycins. J Am Chem Soc 120:9368–9369
Hall IH, Peaty NJ, Henry JR, Easmon J, Heinisch G, Purstinger G (1999) Investigations on the Mechanism of Action of the Novel Antitumor Agents 2-Benzothiazolyl, 2-Benzoxazolyl, and 2-Benzimidazolyl Hydrazones Derived from 2-Acetylpyridine. Arch Pharm 332:115–123
Hutchinson I, Bradshaw TD, Matthews CS, Stevens MFG, Westwell AD (2003) Antitumour benzothiazoles. Part 20:3′-Cyano and 3′-Alkynyl-substituted 2-(4′-Aminophenyl)benzothiazoles as new potent and selective analogues. Bioorg Med Chem Lett 13:471–474
D.M AE-A, Etaiw SEH, Ali EA (2013) Synthesis, spectroscopic, cytotoxic aspects and computational study of N-(pyridine-2-ylmethylene)benzo[d]thiazol-2-amine Schiff base and some of its transition metal complexes. J Mol Struct 1048:487–499
Benazzouz A, Boraud T, Dubedat P, Boireau A, Stutzmann JM, Gross C (1995) Riluzole prevents MPTP-induced parkinsonism in the rhesus monkey: a pilot study. Eur J Pharmacol 284:299–307
Arjunan V, Sakiladevi S, Rani T, Mythili CV, Mohan S (2012) FTIR, FT-Raman, FT-NMR, UV–visible and quantum chemical investigations of 2-amino-4 methylbenzothiazole. Spectrochim Acta A 88:220–231
Ma Y, Cao L, Kawabata T, Yoshino T, Yang B.B, Okada S (1998) Cupric nitrilotriacetate induces oxidative DNA damage and apoptosis in human leukemia HL-60 cells. Free Radic Biol Med 25:568–575.
Nalawade AM, Nalawade RA, Patange SM, Tase DR (2013) Thiazole Containing Schiffs Bases and Their Transition Metal Complexes. Int Eng Sci Inven 2:1–4
Li H, Li J, Chen H, Zhang Y, Huang D (2011) Synthesis and crystal structure of charge transfer complex (CTC) of 2-aminobenzothiazole with its Schiff Base. J Chem Crystallogr 41:1844–1849
Raman N, Selvan A, Sudharsan S (2011) Metallation of ethylenediamine based Schiff base with biologically active Cu(II), Ni(II) and Zn(II) ions: synthesis, spectroscopic characterization, electrochemical behaviour, DNA binding, photonuclease activity and in vitro antimicrobial efficacy. Spectrochim Acta A 79:873–883
Arjmand F, Mohani B, Parveen S (2006) New Dihydro OO'Bis(Salicylidene) 2,2' Aminobenzothiazolyl borate complexes: kinetic and Voltammetric studies of Dimethyltin copper complex with guanine, adenine, and calf thymus DNA. Bioinorg Chem Appl 1 http://dx.doi.org/10.1155/BCA/2006/32896.
Rajarajeswari C, Ganeshpandian M, Palaniandavar M, Riyasdeen A, Akbarsha MA (2014) Mixed ligand copper(II) complexes of 1,10-phenanthroline with tridentate phenolate/pyridyl/(benz)imidazolyl Schiff base ligands: covalent vs non-covalent DNA binding, DNA cleavage and cytotoxicity. J Inorg Biochem 140:255–268
Wu H, Yuan J, Huang X, Kou F, Liu B, Jia F, Wang K, Bai Y (2012) Two zinc(II) and cadmium(II) complexes based on the V-shaped ligand 2,6-bis(2 benzimidazolyl)pyridine: synthesis, crystal structure, DNA-binding properties and antioxidant activities. Inorg Chim Acta 390:12–21
Easmon J, Pürstinger G, Heinisch G, Roth T, Fiebig HH, Holzer W, Jäger W, Jenny M, Hofmann J (2001) Synthesis, cytotoxicity, and antitumor activity of copper(II) and iron(II) complexes of 4 N-Azabicyclo[3.2.2]nonane thiosemicarbazones derived from acyl Diazines. J Med Chem 44:2164–2171
Tahghighi A (2014) Importance of metal complexes for development of potential leishmanicidal agents. J Organomet Chem 770:51–60
Chityala V K, kumar K S, Ramesh M, Parthasarathy T, Shivaraj (2014) DNA Cleavage, Cytotoxic Activities, and Antimicrobial Studies of Ternary Copper(II) Complexes of Isoxazole Schiff Base and Heterocyclic Compounds. Hindawi Pub bioinochem and app doi:http://dx.doi.org/10.1155/2014/691260.
Kumar MP, Tejaswi S, Rambabu A, Kalalbandi VKA, Shivaraj (2015) Synthesis, crystal structure, DNA binding and cleavage studies of copper(II) complexes with isoxazole Schiff bases. Polyhedron 102:111–120
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J Mol Biol 3:208–218. doi:10.1016/s0022-2836(61)80047-8
Kumar CV, Asuncion EH (1993) DNA binding studies and site selective fluorescence sensitization of an anthryl probe. J Am Chem Soc 115:8547–8553. doi:10.1021/ja00072a004
Tandon VK, Yadav DB, Singh RV, Vaish M, Chaturvedi AK, Shukla PK (2005) Synthesis and biological evaluation of novel 1,4-naphthoquinone derivatives as antibacterial and antiviral agents. Bioorg Med Chem Lett 15:3463–3466. doi:10.1016/j.bmcl.2005.04.075
Bhunora S, Mugo J, Bhaw-Luximon A, Mapolie S, Van Wyk J, Darkwa J, Nordlander E (2011) The use of Cu and Zn salicylaldimine complexes as catalyst precursors in ring opening polymerization of lactides: ligand effects on polymer characteristics. Appl Organomet Chem 25:133–145
Youssef NS, El-Zahany E, El-Seidy AMA, Caselli A, Fantauzzi S, Cenini S (2009) Synthesis and characterisation of new Schiff base metal complexes and their use as catalysts for olefin cyclopropanation. Inorg Chim Acta 362:2006–2014
Percy GC, Thornton DA (1973) Infrared spectra of N-aryl salicylaldimine complexes substituted in both aryl rings. J Inorg Nucl Chem 35:2319–2327
Nakamoto K (1997) Infrared and Raman spectra of inorganic and coordination compounds, fifth edn. Wiley-Interscience, New York
Bellamy LJ (1980) The Infrared Spectra of Complex Molecules, second edn. Chapmann and Hall, London
Cotton FA, Wilkinson G (1972) Advanced Inorganic Chemistry, third edn. Interscience Publisher, New York
Lever ABP (1984) Inorganic Electronic Spectroscopy, second edn. Elsevier, Amsterdam
Zhou Y, Ye X, Xin F, Xin X (1999) Solid state self-assembly synthesis of cobalt(II), nickel(II), copper(II) and zinc(II) complexes with a bis-Schiff base. Transit Met Chem 24:118–120
Ramesh R, Maheswaran S (2003) Synthesis, spectra, dioxygen affinity and antifungal activity of Ru(III) Schiff base complexes. J Inorg Biochem 96:457–462
Unver H, Hayvali Z (2010) Synthesis, spectroscopic studies and structures of square-planar nickel(II) and copper(II) complexes derived from 2-{(Z)-[furan-2 ylmethyl]imino]methyl}-6-methoxyphenol. Spectrochim Acta A 75:782–788
Patel RN, Singh N, Shukla KK, Chauhan UK, Nicols -Gutierrez J, Castineiras A (2004) Magnetic, spectroscopic, structural and biological properties of mixed-ligand complexes of copper(II) with N,N,N ′,N″,N″-pentamethyldiethylenetriamine and polypyridine ligands. Inorg Chim Acta 357:2469–2476
Kivelson D, Neiman R (1961) ESR studies on the bonding in copper complexes. J Chem Phys 35:149–155
Hathaway BJ, Billing DE (1970) The electronic properties and stereochemistry of mono-nuclear complexes of the copper(II) ion. Coord Chem Rev 5:143–207
Barton JK, Danishefsky AT, Goldberg JM (1984) Tris(phenanthroline)ruthenium(II): stereoselectivity in binding to DNA. J Am Chem Soc 106:2172–2176
Tysoe SA, Morgan RJ, Baker AD, Strekas TC (1993) Spectroscopic investigation of differential binding modes of DELTA and LAMBDA.-Ru(bpy)2(ppz)2 with calf thymus DNA. J Phys Chem 97:1707–1711
Liu HK, Sadler PJ (2011) Metal complexes as DNA intercalators. Acc Chem Res 44:349–359
Pyle AM, Rehmann JP, Meshoyrer R, Kumar CV, Turro NJ, Barton JK (1989) Mixed-ligand complexes of ruthenium(II): factors governing binding to DNA. J Am Chem Soc 111:3051–3058
Li TR, Yang ZY, Wang BD, Qin DD (2008) Synthesis, characterization, antioxidant activity and DNA-binding studies of two rare earth(III) complexes with naringenin-2-hydroxy benzoyl hydrazone ligand. Eur J Med Chem 43:1688–1695
Sampath K, Sathiyaraj S, Jayabalakrishnan C (2013) DNA binding, DNA cleavage, antioxidant and cytotoxicity studies on ruthenium(II) complexes of benzaldehyde 4-methyl-3-thiosemicarbazones. Spectrochim Acta A 105:582–592
Waring MJ (1965) Complex formation between ethidium bromide and nucleic acids. J Mol Biol 13:269–282
Meyer-Almes FJ, Porschke D (1993) Mechanism of intercalation into the DNA double helix by ethidium. Biochemistry 32:4246–4253
Lepecq JB, Paoletti C (1967) A fluorescent complex between ethidium bromide and nucleic acids: physical-chemical characterization. J Mol Biol 27:87–106
Chen J, Wang X, Chao Y, Zhu JH, Zhu YG, Li YZ, Xu Q, Guo ZJ (2007) A Trinuclear copper(II) complex of 2,4,6-Tris(di-2-pyridylamine)-1,3,5-triazine shows prominent DNA cleavage activity. Inorg Chem 46:3306–3312. doi:10.1021/ic0614162
Tarui M, Doi M, Ishida T, Inoue M, Nakaike S, Kitamura K (1994) DNA-binding characterization of a novel anti-tumour benzo[a]phenazine derivative NC-182: spectroscopic and viscometric studies. Biochem J 304:271–279
Guo H, Lu J, Ruan Z, Zhang Y, Liu Y, Zang L, Jiang J, Huang J (2012) Synthesis, DNA-binding, cytotoxicity, and cleavage studies of unsymmetrical oxovanadium complexes. J Coord Chem 65:191–204
Satyanarayana S, Dabrowiak JC, Chaires JB (1993) Tris(phenanthroline)ruthenium(II) enantiomer interactions with DNA: mode and specificity of binding. Biochemistry 32:2573–2584
Zhen X S, Ye H B, Zhang L Q, Liu G J, Li H, Ji N L (1998) Synthesis characterization and effect of ligand planarity of [Ru(bipy)2 L)2+ on DNA binding affinity.
Sigman DS (1986) Nuclease activity of 1,10-phenanthroline-copper ion, nuclease activity of 1,10-phenanthroline-copper ion. Acc Chem Res 19:180–186
Belai S, Landreau A, Djebbar S, Benali-Baitich O, Bouet G, Bouchara JP (2008) Synthesis, characterization and antifungal activity of a series of manganese(II) and copper(II) complexes with ligands derived from reduced N,N′-O-phenylenebis(salicylideneimine). J Inorg Biochem 102:63–69
Tweedy BG (1964) Possible mechanism for reduction of elemental sulfur by Monilinia fructicola. Phytopathology 55:910–914
Sharma AK, Chandra S (2011) Complexation of nitrogen and Sulphur donor Schiff's base ligand to Cr(III) and Ni(II) metal ions: synthesis, spectroscopic and antipathogenic studies. Spectrochim Acta A 78:337–342
Chohan ZH (2004) Synthesis and biological properties of Cu(II) complexes with 1,1′-Disubstituted Ferrocenes. Synth React Inorg met Org Chem 34:833–846
Acknowledgments
We express our sincere thanks to the Head, Department of Chemistry for providing the necessary facilities. We are thankful to the Director, CFRD, Osmania University, Hyderabad, and the Director, IICT, Hyderabad, and the SAIF, IIT Bombay for providing spectral and analytical data. We are also thankful to DST-PURSE, DST-SERB and UGC-UPE (FAR) for providing financial assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tejaswi, S., Kumar, M.P., Rambabu, A. et al. Synthesis, Structural, DNA Binding and Cleavage Studies of Cu(II) Complexes Containing Benzothiazole Cored Schiff Bases. J Fluoresc 26, 2151–2163 (2016). https://doi.org/10.1007/s10895-016-1911-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10895-016-1911-3