Abstract
Phylogenetic trees illustrate the evolutionary history of genes and species. Although genes evolve along with the species they belong to, a species tree and gene tree are often not identical. The reasons for this are the evolutionary events at the gene level, like duplication or transfer. These differences are handled by phylogenetic reconciliation, which formally is a mapping between a gene tree nodes and a species tree nodes and branches. We investigate models of reconciliation with gene transfers replacing existing genes, which is a biologically important event, but has never been included in the reconciliation models. The problem is close to the dated version of the classical subtree prune and regraft (SPR) distance problem, where a pruned subtree has to be regrafted only on a branch closer to the root. We prove that the reconciliation problem including transfer with replacement is NP-hard, and that, if speciations and transfers with replacement are the only allowed evolutionary events, it is fixed-parameter tractable with respect to the reconciliation’s weight. We prove that the results extend to the dated SPR problem.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Duplications and transfers are events in evolution of genes, and one of the major reasons for discordance between species and gene trees. These differences are modeled by phylogenetic reconciliation (see for example Szöllsi et al. 2015).
The main evolutionary event investigated in this paper is gene transfer (Fig. 1). It involves two, possibly ancient, species existing at the same moment. The species that provides a transferred gene is called a donor species, and the species that receives the gene is called a recipient species.
The gene transfer. a A horizontal gene transfer between species existing at the same moment. Species A is a donor species, while species B is a recipient species, and it receives a new gene copy. b A gene exits the observed phylogeny, through a speciation or transfer, then it returns through a horizontal transfer. c The event from b can be represented with a diagonal transfer. d A transferred gene (\(x_2\)) replaces already present gene (l). The replaced gene l is lost. This event is called the replacement transfer or transfer with replacement, and is represented by a transfer and gene loss. Formally, \(\delta _T(e)=l\), where \(e=(x_1,x_2)\) is a transfer
As phylogenetic analysis never includes the totality of living species, in particular ancient species which can be extinct or not sampled (see transfer from the dead in Szöllsi et al. 2013), the donor species is not assumed to belong to the species phylogeny, but is related to it through one of its ancestors. By extension, this ancestor is considered as the donor, which yields a diagonal transfer or transfer to the future (Fig. 1b–d).
The recipient species either receives a new gene copy, or replaces an existing one (Fig. 1d). The latter event is called replacement transfer or transfer with replacement. In Choi et al. (2012) replacement transfer is called replacing horizontal gene transfer (HGT). They found that the replacing HGT and the additive HGT affect gene functions differently in Streptococcus. In Rice and Palmer (2006) HGT in plastid genomes is studied and the evidence of transfer with replacement is found. HGT with replacement also occurred in the evolution of eukaryotes (Keeling and Palmer 2008). HGT with replacement is used in several phylogenetic studies (Abby et al. 2012; Beiko and Hamilton 2006; Suchard 2005).
In this article we explore the algorithmic aspects of transfers with replacements when time constraints are imposed on transfers (i.e. they should not be directed to the past).
1.1 A review of previous results
There are two usual ways of detecting transfers by comparing species trees and gene trees. The one is reconciliation, and the other is computation of SPR (subtree prune and regraft) scenarios. Our work lies at the intersection of the two. Indeed, reconciliations can consider time constraints but so far have not included replacing transfers. On the other hand, SPR scenarios are a good model for replacing transfers, but never considered time constraints.
Time constraints result in—fully or partially—dated species trees. For a dated species tree, finding a reconciliation of minimum cost, in model with gene transfers, is usually polynomial time (Merkle et al. 2010; Doyon et al. 2010; Bansal et al. 2012). With an undated species tree, and partial time constraints it is usually NP-hard (Hallett and Lagergren 2001; Tofigh et al. 2011; Bansal et al. 2012), and can be fixed parameter tractable (Hallett and Lagergren 2001; Tofigh et al. 2011), or inapproximable (Dasgupta et al. 2006). If a constraint on time consistency of reconciliation scenarios is relaxed, the problem becomes polynomial (Tofigh et al. 2011; Bansal et al. 2012).
Some results go beyond finding one optimal solution. In Bansal et al. (2013) an algorithm that uniformly samples the space of optimal solutions is given, and it runs in polynomial time (per sample). In Chan et al. (2015) the space of all solutions is explored, while the space of optimal solutions is explored in Scornavacca et al. (2013). Probabilistic models allow sampling of solutions in larger spaces, according to likelihood distributions (Szöllsi et al. 2013).
Until now, only reconciliations with transfers without replacements are investigated. However, note that duplications with replacement, i.e. conversions, were recently introduced by Hasić and Tannier (2019). For a more detailed review on reconciliations see Szöllsi et al. (2015), Nakhleh (2012), and Doyon et al. (2011).
Transfers that replace existing genes are in close relation to the classical tree rearrangement operation subtree prune and regraft (SPR). For the definition of SPR refer to Song (2006) (for the rooted trees) and Allen and Steel (2001) (for the unrooted trees). This operation has never been integrated in the reconciliation models but is used to detect transfers when it is the only allowed evolutionary event at the gene scale. In consequence, these studies are limited to datasets where genes appear in at most one copy per species.
Computing SPR distance between two rooted binary phylogenetic trees on the same label set is NP-hard (Hein et al. 1996; Allen and Steel 2001; Bordewich and Semple 2005). The problem is also NP-hard for the unrooted binary trees (Hickey et al. 2008). If we take the SPR distance as a parameter, then both rooted (Bordewich and Semple 2005) and unrooted (Whidden and Matsen 2018) versions are fixed-parameter tractable (FPT).
There is an approximation algorithm of ratio 3 (Hein et al. 1996), as well as an ILP algorithm for calculating the exact rooted SPR distance (Wu 2009), which can also be determined by reducing it to conjunctive normal form (Bonet and John 2009) and using the existing SAT solvers.
Rooted SPR distance is equivalent to the number of trees in the maximum agreement forest (MAF) (Bordewich and Semple 2005), while in the unrooted version SPR distance is greater than or equal to the size of the MAF (Allen and Steel 2001).
A divide and conquer approach with MAF is used for computing the exact SPR distance in Linz and Semple (2011). A 2.5-approximation algorithm for the MAF problem on two rooted binary phylogenetic trees is presented in Shi et al. (2016). In Chen et al. (2015) an FPT algorithm for the rooted SPR, with complexity \(O(2.344^ k n)\) is presented, which is an improvement compared to \(O(2.42^ k n)\) (Whidden et al. 2010). The rooted SPR is investigated also for non binary trees (Whidden et al. 2016), and MAF for multiple trees (Chen et al. 2016). For a more complete review see Shi et al. (2013), and Whidden et al. (2016).
The SPR distance on a dated species tree, where only contemporaneous or transfers to the future are allowed, is mentioned in Song (2006), where it is investigated how many dated trees are one SPR operation away from a given dated tree. The complexity of the dated SPR distance computation is left open, and has an answer as a consequence of our results here.
1.2 The contribution of this paper
In this paper, we analyze the algorithmic complexity of finding a minimum reconciliation with replacing transfers in the presence of a dated species tree. If the speciations and replacement transfers are the only evolutionary events in a reconciliation, then finding a minimum dated SPR scenario is an equivalent problem.
We define a model of reconciliation with gene transfer followed by gene replacement, i.e. a transferred gene replaces a gene that is already present in the recipient species (Fig. 1d). We call this event transfer with replacement, and it is represented by a transfer and a loss (the gene that is replaced). We prove that finding a minimum reconciliation that includes transfer with replacement is NP-hard. If speciation and transfer with replacement are the only allowed events, then the problem is fixed parameter tractable with respect to a minimum reconciliation weight, and it is easily reducible to the dated SPR problem. Therefore the dated SPR is also NP-hard and FPT. Note that the hardness of dated SPR is not easily deduced from the hardness of the general SPR, because all the known proofs make an extensive usage of the possibility of time inconsistent SPRs.
We prove NP-hardness by a reduction from Max 2-Sat. Gadgets for variables and clauses are constructed, and used to assemble a reconciliation that we call a proper reconciliation. Hence the gadgets are subreconciliations of the proper reconciliation. Next, we state an obvious claim that relates an optimal Max 2-Sat solution with a minimum proper reconciliation. Then we prove that any minimum reconciliation can be transformed (in polynomial time) into a proper reconciliation of the same weight (therefore minimum).
In order to prove parametrized tractability, we introduce the normalized reconciliation. Intuitively, this reconciliation can be obtained from any reconciliation by raising nodes of gene tree G as much as possible. We achieve this by mapping the nodes of G to the species tree nodes and edges closer to the root, without affecting transfers, hence keeping the weight of a reconciliation. Then we give a branch and bound algorithm that returns a minimum reconciliation that is also normalized. Thanks to the normalization, we can have at most three cases in the branching algorithm, and every branching produces at least one transfer, so the depth of any branching procedure is at most k, i.e. \(3^k n\) is an approximate complexity of the algorithm.
2 Definitions
Phylogenetic reconciliation is a relation between a gene tree and a species tree. The gene trees, and species trees are used to explain the evolutions of genes and species. Because of the gene-level evolutionary events, a gene tree G and a species tree S are often not identical. This difference can be used to detect the evolutionary events that caused it. In order to emphasize the difference, we place a gene tree inside a species tree. Formally, it is done by a mapping \(\rho :V(G)\rightarrow V(S)\cup E(S)\), that places gene tree nodes inside the species tree. According to the placement of a gene tree node (and some other conditions), we infer the type of evolutionary events. For example, if \(\rho (x)\in V(S)\), where \(x\in V(G)\), then x is a speciation. If, on the other hand, \(\rho (x)\in E(S)\), then \(x'\) might represent transfer, duplication or some other event, which is determined by additional conditions. Also, the mapping \(\rho \) determines where inside S an evolutionary event took place.
More details, and formal definitions are given in the remaining of this section.
A phylogenetic tree here is a rooted tree T such that the root vertex, denoted by root(T), has degree 1 or 2. If \(deg(root(T))=1\), then the incident edge is called the root edge. If \(deg(root(T)))=2\), then there is no root edge. We denote the set of all leaves of the tree T by L(T). Trees are considered binary, meaning that the nodes have at most two children. We say that they are fully binary when all internal nodes have exactly two children.
Let \(x,y\in V(T)\cup E(T)\). If x is an ancestor of y in T, i.e. if x is in the path from y to root(T), then we write \(y\le _T x\) or \(y\le x\), defining a partial order on the nodes and edges. If \(y\not \le x\) and \(x\not \le y\), then we say that x and y are incomparable in T.
Let \(x\in V(T)\cup E(T)\). By \({{\textsc {p}}}^{{\textsc {v}}}_T(x)={{\textsc {p}}}^{{\textsc {v}}}(x)\) we define the minimal node such that \(x<{{\textsc {p}}}^{{\textsc {v}}}(x)\), and by \({{\textsc {p}}}^{{\textsc {e}}}_T(x)={{\textsc {p}}}^{{\textsc {e}}}(x)\) the minimal edge of T such that \(x<{{\textsc {p}}}^{{\textsc {e}}}(x)\). We call \({{\textsc {p}}}^{{\textsc {v}}}(x)\) the node parent, and \({{\textsc {p}}}^{{\textsc {e}}}(x)\) the edge parent. Let \({{\textsc {p}}}_T(x)={{\textsc {p}}}(x)=min\{{{\textsc {p}}}^{{\textsc {v}}}_T(x),{{\textsc {p}}}^{{\textsc {e}}}_T(x)\}\). Note that if \(x\in V(T)\), then \({{\textsc {p}}}(x)\in E(T)\), and if \(x\in E(T)\), then \({{\textsc {p}}}(x)\in V(T)\).
Let \(x\in V(T)\cup E(T)\), and F is a subforest of T. By \({{\textsc {p}}}^{{\textsc {v}}}_F(x)\) and \({{\textsc {p}}}^{{\textsc {e}}}_F(x)\) we define the minimal node and the minimal edge from F such that \(x<{{\textsc {p}}}^{{\textsc {v}}}_F(x)\) and \(x<{{\textsc {p}}}^{{\textsc {e}}}_F(x)\), if they exist. Otherwise, \({{\textsc {p}}}^{{\textsc {v}}}_F(x)\) or \({{\textsc {p}}}^{{\textsc {e}}}_F(x)\) is not defined.
If x is a node/edge in T, then T(x) is the maximal rooted subtree with root node/edge x. If \(A\subseteq L(T)\), then T(A) is the subtree of T with a root vertex of degree 2, and \(L(T(A))=A\).
Let \(x\in V(T)\), \(deg(x)=2\), \(y,z\in V(T)\) are the neighbours of x. If we remove x from T, and connect y and z, then we say that node x is suppressed.
Let \(f:A\rightarrow B\), \(g:C\rightarrow B\), \(C\subseteq A\), and \(g(x)=f(x)\) for all \(x\in C\). Then we write \(g=f/C\).
The next definition extends the partial order on the set \(V(T)\cup E(T)\) to the total order by introducing the date function. Intuitively, to every node and edge from T a date (i.e. a point in the past) is assigned. This derives from the fact that phylogenetic trees and reconciliations represent evolutionary events that happened at some point in the past. Figure 2 depicts a date function for trees S and \(S'\).
Definition 1
(Date function. Dated tree) Let T be a rooted tree and \(\tau :V(T)\cup E(T)\rightarrow [0,+\infty )\) such that \(\tau (L(T))=\{0\}\), \(x_1,x_2\in V(T)\cup E(T)\), \(x_1<x_2\)\(\implies \tau (x_1)<\tau (x_2)\). Function \(\tau =\tau _T\) is a date function on the tree T, and T is a dated tree.
Note that the edges of T are assigned a date. Although it might seem more natural to assign an interval to an edge, here is more convenient to assign a point (i.e. a date).
By a species treeS we mean a dated, fully binary tree with function \(\tau _S=\tau \), and \(\tau (s_1)\ne \tau (s_2)\), for all \(s_1,s_2\in V(S)\backslash L(S)\), such that \(s_1\ne s_2\).
Subdividing an edge means that a vertex is added to the edge. Formally, edge \(e=(x,y)\) is subdivided if a node z is added to the graph along with edges (x, z), (z, y), and the edge e is removed.
Definition 2
(Subdivision of a species tree) Let \(S'\) be a tree obtained from S by subdividing some edges, and \(\forall e=({{\textsc {p}}}^{{\textsc {v}}}(s), s)\in E(S)\), and \(\forall s_1\in V(S)\) for which \(\tau (s)<\tau (s_1)<\tau ({{\textsc {p}}}^{{\textsc {v}}}(s))\), \(\exists s'\in V(S')\backslash V(S)\), \(\tau (s_1)= \tau (s')\), and \(s<s'<{{\textsc {p}}}^{{\textsc {v}}}(s)\). Tree \(S'\) with these properties, and with the minimum number of nodes is called the subdivision of the species tree S.
Tree \(S'\) denotes the subdivision of a tree S. To nodes from \(S'\) even dates are assigned, while the edges are assigned odd dates. The dates are integers from 0 to 2n, where n is the number of the extant species. If \(e\in E(S')\), then \(v_e\) is the maximum node from S such that \(v_e < e\). Here, \(\tau (v_e)=2\), and \(\tau (e)=5\)
Note that the node \(s'\) from Definition 2 is obtained by subdividing some edge, and \(deg(s')=2\). Subdivision of a species tree is unique (Fig. 2). Also, \(L(S')=L(S)\) and \(root(S')=root(S).\) If \(e\in E(S')\), then \(v_e\) denotes the maximum element from the set \(\{x\in V(S)\mid x<e\}\). We assume that \(\tau (V(S')\cup E(S'))=\{0,1,\ldots ,2n\}\), where n is the number of the extant species in S, \(\tau (L(S))=\{0\}\), \(\tau (root(S))=2n\). Therefore if \(x\in V(S')\cup E(S')\), then \(\tau ({{\textsc {p}}}(x))=\tau (x)+1\).
We now define a gene tree species tree reconciliation. A gene tree G is a fully binary tree, which comes with a mapping \(\phi :L(G) \rightarrow L(S)\) that indicates the species in which genes are found in the data.
In the reconciliations that include evolutionary events where one gene replaces another (like conversions or replacement transfers) parts of a gene tree G that are not reconstructed can be important. Therefore, if a gene tree G is given, in order to find a reconciliation, we need to find a gene tree \(G'\) that contains G. We call \(G'\) an extension of G.
Definition 3
(Extension) Let T be a tree, and \(deg(root(T))=1\). A tree \(T'\) is said to be an extension of a tree T if \(root(T')=root(T)\), \(deg(root(T'))=1\), T can be obtained from \(T'\) by pruning some subtrees and suppressing nodes of degree 2.
Figure 3, among other things, depicts G and \(G'\). In Fig. 3b we have \(L(G)=\{l_1,\ldots ,l_6\}\), \(V(G)=\{x_1,\ldots ,x_6\}\cup L(G)\), \(L(G')=L(G) \cup \{l_7,l_8,l_9\}\), \(V(G')=V(G)\cup \{x_7,x_8,x_9,l_7,l_8,l_9\}\). We always have \(root(G')=root(G)\), \(V(G)\subseteq V(G')\), and \(L(G)\subseteq L(G')\).
Note \({{\textsc {p}}}^{{\textsc {v}}}_{G}(l_2)=x_3\), \({{\textsc {p}}}^{{\textsc {v}}}_{G'}(l_2)=x_8\), \({{\textsc {p}}}^{{\textsc {e}}}_{G}(l_2)=(x_3,l_2)\), and \({{\textsc {p}}}^{{\textsc {e}}}_{G'}(l_2)=(x_8,l_2)\). Let \(e=(x_8,l_2)\), then \({{\textsc {p}}}^{{\textsc {v}}}_{G}(e)=x_3\), \({{\textsc {p}}}^{{\textsc {v}}}_{G'}(e)=x_8\), \({{\textsc {p}}}^{{\textsc {e}}}_{G}(e)=(x_2,x_3)\), and \({{\textsc {p}}}^{{\textsc {e}}}_{G'}(e)=(x_7,x_8)\).
A fundamental notion, in a reconciliation between a species tree and a gene tree, is the mapping of nodes of G the edges and vertices of S, denoted by \(\rho \).
Definition 4
(Semireconciliation) Let \(G'\) be an extension of a gene tree G, S be a species tree, and \(S'\) be a subdivision of S. Next, let \(\tau \) be a date function on \(S'\), \(\phi :L(G)\rightarrow L(S)\), and \(\rho :V(G')\rightarrow V(S)\cup E(S')\) such that \(\rho /L(G) = \phi \) and \(\rho (root(G'))=\rho (root(G))=root(S)\). If \(x,y\in V(G')\), \(x<y\), and \(\rho (x)\) and \(\rho (y)\) are comparable in S, then \(\rho (x)\le \rho (y)\). We call a semireconciliation the 6-tuple \({\mathfrak {R}}=(G,G',S,\phi ,\rho , \tau )\).
We say that a node \(x\in V(G')\) is positioned or placed in \(s\in V(S)\cup E(S')\), if \(\rho (x)=s\).
In Fig. 3 we have \(L(S)=\{L_1,\ldots ,L_5\}\), \(V(S)=\{X_1,\ldots ,X_5\}\cup L(S)\), \(E(S)=\{E_1,\ldots ,E_9\}\). For \(S'\) (Fig. 3d) we have: \(L(S')=L(S)\), \(V(S')\) contains V(S) and some other non-labeled nodes, \(E(S')=\{E_1,E_2,E_{3,1},\)\(E_{3,2}, E_4, E_5,E_6,\)\(E_{7,1},E_{7,2}, E_{8,1},E_{8,2}, E_{9,1},E_{9,2},E_{9,3},E_{9,4}\}\).
Regarding mapping \(\rho \), we have: \(\rho (x_1)=X_1\), \(\rho (x_2)=X_2\), \(\rho (x_3)=E_2\), \(\rho (x_{10})=E_{9,4}\), \(\rho (x_{11})=E_5\), \(\rho (l_{10})=E_{7,2}\) (but it could be \(\rho (l_{10})=E_{7,1}\) as well).
Mapping \(\rho \), together with the model’s set of evolutionary events, determines which evolutionary event corresponds to nodes of \(G'\). We will give more details after each definition.
Note that the nodes of \(G'\) are not mapped into \(V(S')\backslash V(S)\). If \(\rho (x)=e'\in E(S')\) and \(e'\) is a part of \(e\in E(S)\), then we will write \(\rho (x)\in e\).
The next definition introduces subtrees of \(G'\) that are not in G.
Definition 5
(Lost subtree) A maximal subtree T of \(G'\) such that \(V(T)\cap V(G)=\emptyset \) is called a lost subtree. An edge from a lost subtree we call a lost edge.
Figure 3b contains three lost subtrees (dashed lines). Every lost subtree has exactly one edge. Figure 3c also has three lost subtrees, but one of them has three edges (the subtree with vertices \(x_7,x_{14},l_{10},l_{11}\)).
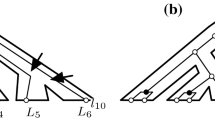
Examples of reconciliations. A species tree is S, gene tree is G, the subdivision of S is \(S'\), and an extension of the gene tree is \(G'\). The gene tree G in b is different from the gene tree G in (c). a A species tree S. \(V(S)=\{X_1,\ldots ,X_5,L_1,\ldots ,L_5\}\), \(root(S)=X_1\), \(root_E(S)=E_1\), \(E(S)=\{E_1,\ldots ,E_9\}\), \(L(S)=\{L_1,\ldots ,L_5\}\). b An LCA reconciliation. \(V(G)=\{x_1,\ldots ,x_6,l_1,\ldots ,l_6\}\), \(V(G')=V(G)\cup \{x_7,x_8,x_9,l_7,l_8,l_9\}\), \(L(G)=\{l_1,\ldots ,l_6\}\), \(L(G')=L(G)\cup \{l_7,l_8,l_9\}\). Duplications are \(x_3,x_6\). Speciations in G are \(x_2,x_4,x_5\), while speciations in \(G'\) that are not in G are \(x_7,x_8,x_9\). Losses are \(l_7,l_8,l_9\). We have three lost edges/subtrees (dashed lines). Node placement: \(\rho (x_1)=X_1\), \(\rho (x_2)=X_2\), \(\rho (x_3)=E_2\), \(\rho (l_9)=E_5\), etc. c The edge \((x_{12},x_{13})\) is a transfer. Since the loss \(l_9\) is paired with \(x_{11}\), \((x_{10},x_{11})\) is a transfer with replacement, i.e. (gene) \(x_{11}\) replaces \(l_9\). Node \(x_6\) is a conversion (and is paired with \(l_{11}\)). We have three lost subtrees. One lost subtree (with vertices \(x_7,x_{14},l_{10},l_{11}\)) has three edges. We have two non-free losses (\(l_8\) and \(l_{10}\)), and two free losses (\(l_9\) and \(l_{13}\)). d Tree \(S'\) is the subdivision of S. Edge \(E_9\) is subdivided into four parts, while each of the edges \(E_3,E_7,E_8\) is subdivided into two parts. The rest of the edges are not subdivided
A semireconciliation is a reconciliation without established gene evolutionary events. The next definitions introduce these events. We define evolutionary events only for genes. Every internal node of S is a speciation, which is an event where one species splits into two.
Gene speciation occurs when a species is split into two, and each species receives a copy of the gene. In Fig. 3b speciations are \(x_2,x_4,x_5,x_7,x_8,x_9\).
Note that by \(\rho (x)_l, \rho (x)_r\in V(S)\) we denote the children of \(\rho (x)\in V(S)\backslash L(S)\).
Definition 6
(Speciation) Let \({\mathfrak {R}}\) be a semireconciliation, \(x\in V(G')\), and \(\rho (x)\in V(S)\backslash L(S)\). If the children of x in \(G'\) can be labeled \(x'_l,x'_r\) so that \(\rho (x)_l \le \rho (x'_l) < \rho (x)\), \(\rho (x)_r \le \rho (x'_r) < \rho (x)\), then x is called a speciation. The set of all speciations is denoted by \(\varSigma ({\mathfrak {R}})\) or \(\varSigma \).
After introducing the remaining evolutionary events, we will see that \(x\in V(G')\) is a speciation if and only if \(\rho (x)\in V(S)\).
Gene duplication is an event where one gene is replaced by two identical gene copies. In Fig. 3b duplications are \(x_3\) and \(x_6\).
Refer to Fig. 2 and the comment after Definition 2 for \(v_e\).
Definition 7
(Duplication) Let \({\mathfrak {R}}\) be a semireconciliation, \(x\in V(G')\), \(x'_l,x'_r\) be the children of x in \(G'\), \(\rho (x)=e\in E(S')\). If \(v_e\le \rho (x'_l)\), \(v_e\le \rho (x'_r)\), \(deg(x'_l) > 2\) or \(x'_l\in L(G')\), and \(deg(x'_r) > 2\) or \(x'_r\in L(G')\), then x is called a duplication. The set of all duplications is denoted by \(\varDelta ({\mathfrak {R}})\) or \(\varDelta \).
From now on, we will assume \(\tau (x)=\tau (\rho (x))\) for all \(x\in V(G')\).
Gene transfer is explained in Sect. 1 and Fig. 1. In Fig. 3c the edges \((x_{10},x_{11})\) and \((x_{12},x_{13})\) are transfers. Vertices \(x_{10}\) and \(x_{12}\) are transfer parents, where \(x_{11}\) and \(x_{13}\) are transfer children.
Definition 8
(Transfer) Let \({\mathfrak {R}}\) be a semireconciliation, \(x\in V(G')\), \(x'_l,x'_r\) be the children of x in \(G'\), \(\rho (x)=e\in E(S')\). For one of the \(x'_l, x'_r\) (say \(x'_l\)) holds \(v_{e}\le \rho (x'_l)\) and \(deg(x'_l)>2\). For the other one (i.e. \(x'_r\)) we have \(\rho (x'_r)=e'\in E(S')\), \(\tau (e')\le \tau (e)\), \(deg(x'_r)=2\), and \(v_{e'}\le \rho (x''_r)\), where \(x''_r\) is the only child of \(x'_r\) in \(G'\). Then x is called a transfer parent, \(x'_r\) is a transfer child, and the edge \((x,x'_r)\in E(G')\) is a transfer. If \(\tau (x'_r)=\tau (x)\), the transfer is a horizontal transfer, and if \(\tau (x'_r) < \tau (x)\), the transfer is a diagonal transfer or transfer to the future. The set of all transfers is denoted by \(\varTheta ({\mathfrak {R}})\) or \(\varTheta \).
We say that transfer \((x,x'_r)\)belongs to edge\((a,b)\in E(G)\) if x and \(x'_r\) belong to (a, b), i.e. \(b<_{G'}x'_r<_{G'}x\le _{G'}a\).
In Fig. 3c nodes \(l_8,l_9,l_{10},l_{11}\) are gene losses.
Definition 9
(Loss) Let \({\mathfrak {R}}\) be a semireconciliation, and \(x\in L(G')\backslash L(G)\). Then x is called a loss. The set of all losses is denoted by \(\varLambda ({\mathfrak {R}})\) or \(\varLambda \).
The next two events that we define are created by pairing some of the previously defined events with a loss.
The replacement transfer is explained in Sect. 1 and Fig. 1. In Fig. 3c a replacement transfer is \((x_{10},x_{11})\), i.e. \(x_{11}\) replaces \(l_9\).
Definition 10
(Replacement transfer) Let \({\mathfrak {R}}\) be a semireconciliation, \(\delta _T: \varTheta \rightarrow \varLambda \) be an injective partial function such that for all transfers \(e=(x_1,x_2)\in \delta ^{-1}_T(\varLambda )\) holds \(\rho (x_2)=\rho (\delta _T(e))\), and \(x_2\) and \(\delta _T(e)\) are incomparable in \(G'\). If \(e\in \delta ^{-1}_T(\varLambda )\), then e is called a replacement transfer or transfer with replacement, and \(\delta _T(e)\) is its associate loss. The set of all replacement transfers is denoted by \(\varTheta '({\mathfrak {R}})\) or \(\varTheta '\), and the set of all associate losses by \(\varLambda '_T({\mathfrak {R}})\) or \(\varLambda '_T\).
In the previous definition, the mapping \(\delta _T\) pairs transfer e (or we can say the transfer child \(x_2\)) with the loss \(l=\delta _T(e)\) (see Fig. 1). In this way, we get that the gene \(x_2\) is replacing l, hence the name transfer with replacement. Requirement \(\rho (x_2)=\rho (l)\) is necessary if \(x_2\) replaces l. In our example (Fig. 3c) \(\delta _T(x_{10},x_{11})=l_9\).
Conversion is to duplication what replacement transfer is to transfer. Node \(x_6\) is a conversion (Fig. 3c).
Definition 11
(Conversion) Let \({\mathfrak {R}}\) be a semireconciliation, \(\delta _D: \varDelta \rightarrow \varLambda \) be an injective partial function such that \(\rho (x)=\rho (\delta _D(x))\), and x and \(\delta _D(x)\) be incomparable in \(G'\) for all \(x\in \delta ^{-1}_D(\varLambda )\). If \(x\in \delta ^{-1}_D(\varLambda )\), then x is called a conversion, and \(\delta _D(x)\) is its associate loss. The set of all conversions is denoted by \(\varDelta '({\mathfrak {R}})\) or \(\varDelta '\), and the set of associate losses by \(\varLambda '_D({\mathfrak {R}})\) or \(\varLambda '_D\).
We have \(\delta _D(x_6)=l_{11}\) (Fig. 3c).
The elements of \(\varLambda '=\varLambda '_T\cup \varLambda '_D\) are called free losses, because we will assign them zero cost (see the comment after Definition 13). The set of all (evolutionary) events is \(\{S, D, T, L, C, T_R\}\). Free losses in Fig. 3c are \(l_9\) and \(l_{11}\), while \(l_8\) and \(l_{10}\) are non-free losses.
Intuitively, a reconciliation between gene tree G and species tree S is a placement of G inside S. The most important part is the function \(\rho \), which defines the placement. If events that replace existing genes (like conversions or transfer with replacement) are included in the reconciliation, a reconstruction of \(G'\) is also important. Figure 3b–c depicts reconciliations. The following definition is an extension of the definition from Hasić and Tannier (2019).
Definition 12
(Reconciliation) Let \((G,G',S,\phi ,\rho , \tau )\) be a semireconciliation, and \(X\subseteq \{D, T, L, C, T_R\}\). To every node from \(V(G')\backslash L(G)\) some event from \(X\cup \{S\}\) is attached, and \(\varLambda '_T \cap \varLambda '_D=\emptyset \). Then \({\mathfrak {R}}=(G,G',S,\phi ,\rho ,\tau ,\delta _T,\delta _D,\)A) is called an X reconciliation.
If the set X is not important in the context, or it is known, then we will use just reconciliation instead of X reconciliation. If we wish to emphasize that \(G'\) and \(\rho \) are from reconciliation \({\mathfrak {R}}\), then we write \(G'_{{\mathfrak {R}}}\) and \(\rho _{{\mathfrak {R}}}\).
If the transfers with replacement, or conversions are not included in a reconciliation, then \(\delta ^{-1}_T(\varLambda )=\emptyset \), or \(\delta ^{-1}_D(\varLambda )=\emptyset \). Note that if \(x\in V(G')\) and \(deg(x)=2\), then x is a transfer child.
Speciations, duplications, transfers, losses, conversions, and transfers with replacement are called evolutionary events. A reconciliation can allow only some of these events. For example, if a reconciliation \({\mathfrak {R}}\) allows speciations, duplications and losses, we will call it a DL reconciliation. If \({\mathfrak {R}}\) also allows transfers, we call it a DTL reconciliation. Speciations are assumed to be allowed in every reconciliation, so they are not emphasized in the type of a reconciliation. If transfers are not allowed in a reconciliation, then the date function is not necessary, and can be disregarded. Note that if \(X\subseteq Y\), then any X reconciliation is also a Y reconciliation. If conversions or transfers with replacement are included in a reconciliation, then we assume that free losses are allowed. Therefore \(T_R\) reconciliation allows speciations, replacement transfers, and free losses, while \(T_RL\) reconciliations additionally allow non-free losses.
Not every semireconciliation can produce a reconciliation. For example, if a node from \(G'\) is mapped under its LCA (Last/Lowest Common Ancestor—see Goodman et al. 1979; Chauve and El-Mabrouk 2009) position, then the transfers must be allowed as an event in order to obtain a reconciliation. Figure 3b depicts an LCA reconciliation.
We introduce the weight of a reconciliation as a way to compare them.
Definition 13
(Weighted reconciliation) Let \({\mathfrak {R}}\) be an X reconciliation, and \(X=\{a_1,\ldots ,a_k\}\). If \(c_i\ge 0\) are associated with the events \(a_i\)\((i=1,\ldots ,k)\), then \(\omega ({\mathfrak {R}})=\sum c_i\cdot |a_i|\) is called the weight or cost of \({\mathfrak {R}}\), where \(|a_i|\) denotes the number of nodes in \(G'\) that are associated with the event \(a_i\) (\(i=1,\ldots ,k\)).
The speciations do not affect the weight of a reconciliation, thus their weight is 0. In this paper, the free losses (losses assigned to a conversion or replacement transfer) have weight 0. The rest of the evolutionary events have weight 1.
Definition 14
(Minimum X Reconciliation problem) Let G and S be a gene and a species trees. The problem of finding an X reconciliation with the minimum weight we call the Minimum X Reconciliation .
The next definition introduces the weight of a subtree of \(G'\). We need this to estimate the weight of a reconciliation by decomposing \(G'\) into subtrees and evaluating the weight of every subtree.
Definition 15
(The weight of a subtree) Let \({\mathfrak {R}}\) be a reconciliation and T be a subtree of \(G'\). By \(\omega _{{\mathfrak {R}}}(T)\) or \(\omega (T)\) we denote the sum of weights of all events assigned to the nodes and edges of T.
Let \(T_1\) be a subtree of \(G'\) (Fig. 3c), where \(V(T_1)=\{x_2,x_{10},x_{11},l_1,l_{12}\}\). It has one transfer with replacement, hence \(\omega (T_1)=tr\), where tr is the weight of a replacing transfer.
Let \(T_2\) be a subtree with \(V(T_2)=\{x_4,x_5,x_{12},x_{13},x_6,l_4,l_5,l_6,l_{13}\}\). Then \(\omega (T_2)=t+c\), where t and c are the weights assigned to a transfer and a conversion.
3 Finding a minimum \(\mathbf {DTLCT_R}\) reconciliation is NP-hard
In this section, and the rest of the paper, we assume that all events have weight 1, except the speciations and free losses, which have weight 0. We prove that finding a minimum reconciliation that includes transfers with replacement is NP-hard. We first prove the NP-hardness of the problem of finding a minimum reconciliation that includes all events (duplication, transfer, loss, conversion, transfer with replacement).
Let F be a logical expression/formula, or simply formula, in conjuctive normal form with (logical) variables \(x_1,\ldots , x_n\) . If x is a variable, then x is called a positive literal, and \(\lnot x\) is a negative literal. Literals \(x^1_i,x^2_i,x^3_i\) are assigned to the variable \(x_i\). We can assume that \(x^1_i\) and \(x^2_i\) have the same logical value, which is different from the logical value of \(x^3_i\). Variables can be true or false. A literal is true if it is positive and the variable is true, or if it is negative and the variable is false. Similarly, a literal is false if it is positive and the variable is false, or if it is negative and the variable is true.
We will use a reduction from the Max 2-Sat.
Max 2-Sat:
Input:\(F=C_1 \wedge C_2 \wedge \ldots \wedge C_m\); \(C_j=x'_{j_1} \vee x'_{j_2}\), \(j=1,\ldots ,m\); \(K\le m\).
Output: Is there a truth assignment for logical variables \(x_1,\ldots ,x_n\) such that there are at least K true clauses.
This problem is NP-hard (Garey et al. 1976; Garey and Johnson 1979), solvable in polynomial time if \(K=m\) (Even et al. 1976; Garey and Johnson 1979). It remains NP-hard even if every variable appears in at most three clauses (Raman et al. 1998). We assume that every variable appears in exactly three clauses, and both positive and negative literals are present (Lemma 1). We also assume the optimization version of this problem that asks for the minimum number of false clauses.
The next lemma proves the NP-completeness of the reduced problem.
Lemma 1
The Max 2-Sat is NP-hard if every variable appears in exactly three clauses, and both positive and negative literal of every variable is present.
Proof
See Appendix. \(\square \)
3.1 Variable and clause gadgets
In order to construct a polynomial reduction from the Max 2-Sat to the Minimum\(DTLCT_R\)Reconciliation, suppose we have a logical formula F as an instance of the Max 2-Sat, with n variables and m clauses, such that each variable appears exactly three times as a literal, and both positive and negative literals are present. We will construct a species tree, gene tree, and function \(\phi \) mapping the gene tree leaves to the species tree leaves as an instance of the reconciliation problem.
First, we introduce the border line that corresponds to some date, depicted by horizontal dashed line in Figs. 4, 5, 6, 7, and 8. Some nodes of the constructed gene tree will be assigned to literals of \(x_i\) (\(i=1,\ldots ,n\)), and in a minimum reconciliation, their mapping above or under this border line will decide if the literals are true or false. In consequence, the positive and negative version of a same variable must be mapped on the opposite sides of the border line in reconciliations.
For each variable and each clause we define a piece of a gene tree and a piece of a species tree with appropriate function \(\phi \).
A variable gadget denoted by \({\mathcal {G}}_{x_i}\). It is composed of \(S_{x_i}\) (a part of the species tree S) and \(G_{x_i}\) (a part of the gene tree G). The border line is the horizontal, dashed line. Nodes \(A^i_1,\ldots , A^i_{28}\) are the leaves of \(S_{x_i}\). Nodes \(C'_{i,1}, \ldots , C'_{i,12}\) are some of the inner nodes of \(S_{x_i}\), and \(s^0_{x_i}\) is the root of \(S_{x_i}\). The rest of the labels denote some of the nodes of \(G_{x_i}\). Variable \(x_i\) has two true (represented by \(x^1_i, x^2_i\in V(G)\)) and one false literal (represented by \(x^3_i\in V(G)\)). Edges (transfers) incident with \(x^1_i,x^2_i,x^3_i\) are also included in \(G_{x_i}\)
A gadget for a variable \(x_i\) is illustrated in Fig. 4. The species subtree \(S_{x_i}\) consists in 28 leaves named \(A_1^i,\dots ,A_{28}^i\), organized in two subtrees. Seven cherry trees are under the border line on each part, and then linked by two combs, one fully above and one fully under the border line.
The gene subtree \(G_{x_i}\) is also organized in two subtrees, each consisting in 7 cherry trees linked by a comb. One of the subtrees is identified as the “true literal subtree” and the other as the “false literal subtree”.
Let \((l_{i,k}, r_{i,k})\) be the kth cherry of the gene tree \(G_{x_i}\) (i.e. \(l_{i,k}, r_{i,k}\) are the children of \(b^k_i\) assigned to the left and right subtree of \(S_{x_i}\), and \(k=1,\ldots ,14\)). The function \(\phi \), mapping the leaves of the gene tree to the leaves of the species tree, is such that \(\phi (l_{i,k})=A^i_{2k-1}\) and \(\phi (r_{i,k})=A^i_{29-2k}\) (\(k\in \{1,\ldots ,7\}\)); \(\phi (l_{i,k})=A^i_{30-2k}\) and \(\phi (r_{i,k})=A^i_{2k}\) (\(k\in \{8,\ldots ,14\}\)).
Edges/transfers incident with \(x^1_i,x^2_i,x^3_i\) (horizontal full lines in Fig. 4) are also included in \(G_{x_i}\).
Next, both trees G and S (Fig. 5) are anchored by an outgroup comb of size P(n), a polynomial with sufficient size, with respective leaf sets \(a^1_i,\dots ,a^{P(n)}_i\) and \(A_{i,1},\dots ,A_{i,P(n)}\), and \(\phi (a^k_i)=A_{i,1}\) (\(k=1,\ldots ,P(n)\)).
Figure 6 illustrates a gadget for a clause \(C_j=x'_{j1} \vee x'_{j2}\). A subtree \(S_{C_j}\) of the species tree is a fully balanced binary tree with 8 leaves, denoted by \(B_1^j,\dots ,B_8^j\). The internal nodes of the subtree leading to \(B_1^j,\dots ,B_4^j\) are all above the border line, while the internal nodes of the subtree leading to \(B_5^j,\dots ,B_8^j\) are all under the border line.
To each literal from the clause corresponds a fully balanced gene tree with a root of degree two, and four leaves, respectively mapped by the function \(\phi \) to \(((B^j_1,B^j_7),(B^j_2,B^j_6))\) and \(((B^j_3,B^j_5),(B^j_4,B^j_8))\) (which is an arbitrary way of mapping each cherry to the two different species subtrees). The roots of the two gene subtrees are respectively labeled \(r_{j1}\) and \(r_{j2}\). The internal nodes are \(r_{j1}^0\), \(r_{j1}^1\), and \(r_{j2}^0\), \(r_{j2}^1\). The forest of these two gene subtrees is denoted by \(F_j\).
A clause gadget that corresponds to a clause \(C_j=x'_{j_1} \vee x'_{j_2}\). a The literal \(x'_{j_1}\) is true, and \(x'_{j_2}\) is false. b The literal \(x'_{j_2}\) is true, and \(x'_{j_1}\) is false. c Both literals are true. d Both literals are false, hence the clause is false. In this case we have an extra transfer, which is incident with \(r_{j2}\)
The clause and variable gadgets are linked to form the full trees G and S (Fig. 7). Observe the gadget for \(C_j=x'_{j1} \vee x'_{j2}\) (Fig. 6). It has two gene tree subtrees rooted at \(r_{j1},r_{j2}\). These subtrees represent literals \(x'_{j1},x'_{j2}\), and are linked to \(x'_{j1}\) and \(x'_{j2}\) that are literals of variables \(x_{i1}\) and \(x_{i2}\), i.e. \(x'_{j1}\in \{x^1_{i1},x^2_{i1},x^3_{i1}\}\), \(x'_{j2}\in \{x^1_{i2},x^2_{i2},x^3_{i2}\}\). Hence the gene tree subtrees rooted at \(r_{j1}\) and \(r_{j2}\) are linked to \(G_{x_{i1}}\) and \(G_{x_{i2}}\).
The species subtrees and the gene subtrees are linked by the comb containing all variables and clauses in the order \(x_1,\dots ,x_n,C_1,\dots ,C_m\).
A variable and clause gadget are merged, and G and S are formed. a A variable gadget with anchor. For \(i=n\), \(d_i\) (i.e. \(d_n\)) does not exist. b A clause gadget. c A proper reconciliation. Nodes \(s^\alpha _\beta \) (\(\alpha \in \{0,1\}\), \(\beta \in \{x_1,\ldots ,C_m\}\)) belong to the species tree
3.2 Proper reconciliation
Now that we have constructed an instance for the reconciliation problem from a logical formula, we need to be able to translate a reconciliation into an assignment of the variables. This is possible for a type of reconciliation named proper. Proper reconciliations are illustrated in Figs. 7 and 8 .
A proper reconciliation assigned to formula \(F_1=(x_1\vee \lnot x_2)\wedge (x_1\vee x_2)\wedge (\lnot x_1 \vee x_2)\), with the values \(x_1=1, x_2=0\). Some other formulas are also possible, like \(F_2=(\lnot x_1\vee \lnot x_2)\wedge (\lnot x_1\vee x_2)\wedge (x_1 \vee x_2)\), with the values \(x_1=0, x_2=0\). Clauses \(C_1\) and \(C_2\) are true, and clause \(C_3\) is false
Trees G and S on the one hand, and their proper reconciliation, on the other hand are formed in the same way—by merging all gadgets. For the sake of formalism, we first introduced G and S, and now we introduce the proper reconciliation.
In the next definition, \(B^j_{1,2}=lca(B^j_1,B^j_2)\), i.e. \(B^j_{1,2}\) is the minimal node (in S) that is an ancestor of \(B^j_1\) and \(B^j_2\). Similarly, \(B^j_{5,6,7,8}=lca(B^j_5,B^j_6,B^j_7,B^j_8)\) is the minimal node (in S) that is an ancestor of \(B^j_5\), \(B^j_6\), \(B^j_7\), and \(B^j_8\).
Definition 16
(Proper reconciliation) Let G and S be a gene and species tree constructed from a logical formula. We call a reconciliation \({\mathfrak {R}}=(G,G',\)\(S,\phi ,\rho ,\tau ,\)\(\delta _T,\delta _D,\{T_R\})\) a proper reconciliation if:
-
all transfers are horizontal;
-
in the variable gadgets, the gene tree vertices in the anchor comb are mapped by \(\rho \) to the species tree vertices in the anchor comb, that is, \(\rho (c^0_i)=s^0_{x_i}\), \(\rho (d^k_i)=D^i_k\) (for all \(k\in \{1,\ldots , P(n)\}\)), \(\rho (d_i)=s^1_{x_i}\);
-
in the variable gadgets, the two gene tree comb internal vertices (\(c_i^k\) in Fig. 4) are mapped to the two species tree combs (vertices \(C'_{i,k}\) in Fig. 4), in one of the two possible combinations (the two gene tree combs may map to either species tree combs).
-
in the clause gadgets, the mapping \(\phi \) corresponds to one of the four cases drawn in Fig. 6, that is:
-
(Fig. 6a) \(\rho (r_{j_1})=B^j_{1,2}\), \(\rho (r_{j_2})=B^j_{5,6,7,8}\), \(\rho (r^0_{j_1})\in (B^j_{1,2}, B^j_1)\), \(\rho (r^1_{j_1})\in (B^j_{1,2}, B^j_2)\), \(\rho (r^0_{j_2})\in (B^j_{5,6}, B^j_5)\), \(\rho (r^1_{j_2})\in (B^j_{7,8}, B^j_8)\);
-
(Fig. 6b) \(\rho (r_{j_1})=B^j_{5,6,7,8}\), \(\rho (r_{j_2})=B^j_{3,4}\), \(\rho (r^0_{j_1})\in (B^j_{5,6}, B^j_6)\), \(\rho (r^1_{j_1})\in (B^j_{7,8}, B^j_7)\), \(\rho (r^0_{j_2})\in (B^j_{3,4}, B^j_3)\), \(\rho (r^1_{j_2})\in (B^j_{3,4}, B^j_4)\);
-
(Fig. 6c) \(\rho (r_{j_1})=B^j_{1,2}\), \(\rho (r_{j_2})=B^j_{3,4}\), \(\rho (r^0_{j_1})\in (B^j_{1,2}, B^j_1)\), \(\rho (r^1_{j_1})\in (B^j_{1,2}, B^j_2)\), \(\rho (r^0_{j_2})\in (B^j_{3,4}, B^j_3)\), \(\rho (r^1_{j_2})\in (B^j_{3,4}, B^j_4)\);
-
(Fig. 6d) \(\rho (r_{j_1})=B^j_{5,6,7,8}\), \(\rho (r_{j_2})\in (B^j_{3,4},B^j_3)\), \(\rho (r^0_{j_1})\in (B^j_{5,6}, B^j_6)\), \(\rho (r^1_{j_1})\in (B^j_{7,8}, B^j_7)\), \(\rho (r^0_{j_2})\in (B^j_{3,4}, B^j_3)\), \(\rho (r^1_{j_2})\in (B^j_{3,4}, B^j_4)\).
-
-
the only transfers are the one depicted by variable and clause gadgets.
Note that a proper reconciliation is a \(T_R\) reconciliation, i.e. the only events are speciations, replacement transfers, and free losses. Hence the weight of a proper reconciliation is the number of transfers.
Let F be a logical formula and G, S be the gene and species tree assigned to F, as previously described. There is an obvious relation between a value assignment to logical variables and a proper reconciliation between G and S.
Lemma 2
Let F be a 2-Sat formula with n variables and m clauses. Trees G and S are the gene and species tree assigned to F. Let \({\mathfrak {R}}\) be a proper reconciliation between G and S. There is an assignment of the logical variables which satisfies exactly \(17n+5m - \omega ({\mathfrak {R}})\) clauses.
Proof
The assignment is constructed from the proper reconciliation according to the positions of the corresponding vertices above or under the border line.
The definition of a proper reconciliation ensures that two opposite literals are always on the opposite side of the border line.
Every variable gadget has 17 transfers (counting the ones incident with \(x^1_i,x^2_i,x^3_i\)), and the total number of transfers generated by these gadgets is 17n.
Clause gadget has 5 or 4 transfers (not counting the incoming transfers, because they are already counted in the variable gadgets), depending if the clause’s literals are both false or not. Let f be the number of the false clauses. Then we have f clause gadgets with 5 transfers corresponding to false clauses.
Hence the number of transfers, generated in the clause gadgets, is \(4(m-f)+5f=4m+f\). This yields \(\omega ({\mathfrak {R}})=17n+4m+f\), so there are \(m-f = m - (\omega ({\mathfrak {R}})-17n-4m)=17n+5m-\omega ({\mathfrak {R}})\) true clauses. \(\square \)
We see that, if we minimize the cost of a proper reconciliation, we also minimize the number of false clauses in the logical formula. We say that, for a given G and S, \({\mathfrak {R}}\) is a proper reconciliation of minimal weight if it has the minimal weight among all proper reconciliations between G and S.
As an immediate consequence of Lemma 2, we have the next lemma.
Lemma 3
To a proper reconciliation of minimal weight corresponds an optimal 2-Sat formula.
In order to prove NP-hardness, we need to show that there is a minimum reconciliation that is also proper, which can be easily (in polynomial time) obtained from an arbitrary minimum (\(DTLCT_R\)) reconciliation.
3.3 Proper minimum reconciliation
If a minimum reconciliation is also a proper, then we will call it a proper minimum reconciliation. At this point, we do not know if this reconciliation always exists. In this section, we describe how to construct this reconciliation, given a a minimum reconciliation, thus proving its existence. After this, the notions of a proper reconciliation of minimal weight and a proper minimum reconciliation become identical. Again, we always assume that G and S are constructed from a given 2-Sat formula.
We say that a node \(x\in V(G')\) is placed in \(s\in V(S)\cup E(S')\) if \(\rho (x)=s\).
If \({\mathfrak {R}}\) is a proper reconciliation, then \(\omega (G_{x_i})=17\) (here we also count three transfers incident with \(x^1_i,x^2_i,x^3_i\)), and \(\omega (F_j)\in \{4,5\}\) for all variable and clause gadgets.
Lemma 4
Let \({\mathfrak {R}}\) be a minimum X reconciliation between G and S, where G and S are a gene and a species trees constructed from a logical formula, and \(i\in \{1,\ldots , n\}\), \(j \in \{1,\ldots , m\}\). Then we have:
-
(a)
\(\omega (F_j)\ge 4\);
-
(b)
if both \(r_{j_1}\) and \(r_{j_2}\) are under the border line, then \(\omega (F_j) \ge 5\);
-
(c)
\(\omega (G_{x_i})\ge 17\);
-
(d)
if \(x^1_i\) or \(x^2_i\) is on the same side of the border line as \(x^3_i\), then \(\omega (G_{x_i})\ge 19\).
Proof
See Appendix. \(\square \)
The proof of next theorem describes a polynomial algorithm that transforms a minimum reconciliation \({\mathfrak {R}}\) into a reconciliation \({\mathfrak {R}}'\) that is both minimum and proper.
Theorem 1
Let G and S be a gene and a species tree constructed from a logical formula. There is a minimum \(DTLCT_R\) reconciliation between G and S that is a proper reconciliation.
Proof
See Appendix. \(\square \)
Theorem 2
The Minimum\(DTLCT_R\)Reconciliation is NP-hard.
Proof
We will use a reduction from the optimization version of Max 2-Sat. Let \(F=C_1 \wedge C_2 \wedge \ldots \wedge C_m\), \(C_j=x'_{j_1} \vee x'_{j_2}\), \(j=1,\ldots ,m\) be an instance of Max 2-Sat. Trees S and G can be obtained from F in polynomial time. After obtaining a minimum reconciliation between S and G as an output of the Minimum\(DTLCT_R\)Reconciliation, we can (in polynomial time) obtain a proper minimum reconciliation (the proof of Theorem 1), and from it an optimal logical formula, i.e. a logical formula F with the minimum number of false clauses (Lemma 2). \(\square \)
Since a proper reconciliation is a \(T_R\) reconciliation, i.e. it has only transfers with replacement and all losses are free, then the next theorem can be proved in the same manner as Theorem 2.
Theorem 3
Let \(X\subseteq \{D,T,L,C,T_R\}\) and \(T_R\in X\). Then the Minimum X Reconciliation is NP-hard.
Proof
We will reduce the Max 2-Sat to the Minimum X reconciliation. Let F be an instance of the Max 2-Sat, and G and S be a gene and a species tree obtained from F (as described earlier).
Let \({\mathfrak {R}}_1\) be a minimum X reconciliation, and \({\mathfrak {R}}_2\) be a minimum \(DTLCT_R\) reconciliation between G and S. Then \({\mathfrak {R}}_1\) is a \(DTLCT_R\) reconciliation, hence \(\omega ({\mathfrak {R}}_2)\le \omega ({\mathfrak {R}}_1)\).
Let \({\mathfrak {R}}_3\) be a minimum proper reconciliation obtained from \({\mathfrak {R}}_2\), as described in the proof of Theorem 1. Then \(\omega ({\mathfrak {R}}_3)=\omega ({\mathfrak {R}}_2)\).
Since \({\mathfrak {R}}_3\) is a proper reconciliation, it is a \(T_R\) reconciliation, hence an X reconciliation, and therefore \(\omega ({\mathfrak {R}}_3)\ge \omega ({\mathfrak {R}}_1)\).
From the previous inequalities, we obtain \(\omega ({\mathfrak {R}}_1)=\omega ({\mathfrak {R}}_2)=\omega ({\mathfrak {R}}_3)\), i.e. \({\mathfrak {R}}_1\) is a minimum \(DTLCT_R\) reconciliation.
Now we can repeat the earlier procedure. From \({\mathfrak {R}}_1\) we construct a proper minimum reconciliation, which we can use to find an optimal assignment for F. \(\square \)
Note that in the proof of Theorem 3 we have that a minimum X reconciliation is also a minimum \(DTLCT_R\) reconciliation. This claim is true in our case, when G and S are obtained from F. It does not hold for an arbitrary G and S.
4 The Minimum \(\mathbf {T_R}\) Reconciliation is fixed-parameter tractable
We will give a branch and bound algorithm that solves the Minimum\(T_R\)Reconciliation, with complexity O(f(k)p(n)), where p is a polynomial, k is a parameter representing an upper bound for the reconciliation’s weight, and f is a (computable) function.
We give some basic properties of the (minimum) \(T_R\) reconciliations that will be used in the proofs.
Lemma 5
Let \({\mathfrak {R}}\) be a minimum \(T_R\) reconciliation, and \(e\in E(G')\backslash E(G)\). Then e cannot be a transfer.
Proof
Assume the opposite, i.e. e is a transfer. Figure 9a depicts the construction of a reconciliation with a smaller weight, which contradicts the minimality of \({\mathfrak {R}}\). \(\square \)
a A minimum (\(T_R\)) reconciliation cannot have transfers that are not in G. Edge \((x_1,x_2)\) is a transfer, and \(l_1\) is a loss (assigned to the transfer). First, remove \((x_1,x_2)\), connect vertices \(x_2\) and \(l_1\), then suppress \(x_1,x_2,l_1\). In this way we obtain a reconciliation with a smaller weight. b Loss extension. A loss \(l_1\) is assigned to \(E_1\in E(S)\). Insert new vertices \(l_2,l_3\) such that \(\rho (l_2)=E_2\), and \(\rho (l_3)=E_3\). Connect \(l_2\) and \(l_3\) with \(l_1\), and take \(\rho (l_1)=X_1\). In this way we extend the loss \(l_1\)
Every edge of S contains exactly one lineage from \(G'\). To state and prove this fact more formally, we introduce the notion of aligned edges. Intuitively, two edges \(e_1,e_2\in E(G')\) are aligned if they are inside a same edge \(E_1\in E(S')\), and are connected, in some way, by a sequence of vertices of \(G'\) contained in \(E_1\) (Fig. 10).
Definition 17
(Aligned edges) Let \({\mathfrak {R}}\) be an X reconciliation, \(E_1=(s_1,s_2)\in E(S')\), \(e_1=(x_1,x_2)\), \(e_2=(y_1,y_2)\in E(G')\), \(s_1\ge \rho (x_1)\ge \rho (x_2)\ge s_2\), \(s_1\ge \rho (y_1)\ge \rho (y_2)\ge s_2\), and \(e_1\ne e_2\) (Fig. 10). If there are \(a_0,\ldots ,a_k\in V(G')\) (\(k\ge 0\)) such that:
-
\(s_1\ge \rho (a_i)\ge s_2\) (\(i=0,\ldots ,k\));
-
\(a_0=x_2\) and \(a_k=y_1\), or \(a_0=y_2\) and \(a_k=x_1\);
-
\((a_i,a_{i+1})\in E(G')\) or \(a_i\) is a loss assigned to \(a_{i+1}\) (\(i=0,\ldots ,k-1\));
then we say that \(e_1\) and \(e_2\) are edges aligned inside\(E_1\), or just aligned edges.
Aligned edges. For edges \(e_1=(x_1,x_2),e_2=(y_1,y_2)\in E(G')\) we have \(s_1\ge \rho (x_1)\ge \rho (x_2)\ge s_2\), and \(s_1\ge \rho (y_1)\ge \rho (y_2)\ge s_2\), where \(E_1=(s_1,s_2)\in E(S')\). a–c Edges \(e_1\) and \(e_2\) are not aligned. d Here we have a transfer leaving \(E_1\), and returning later. Although there is a sequence \(a_0,\ldots ,a_k\) between \(y_2\) and \(x_1\), edges \(e_1\) and \(e_2\) are not aligned, since \(\rho (a_2)\not \in \{s_1,E_1,s_2\}\). e Edges \(e_1\) and \(e_2\) are aligned. Sequence \(a_0,\ldots ,a_k\) contains \(a_1\) and l. f, g Here, \(e_1\) and \(e_2\) are also aligned
The next lemma basically states that every edge of S contains exactly one lineage of \(G'\). Equivalently, if we remove all the transfers from \(G'\), and suppress all the nodes of degree 2, the obtained tree is identical to S.
Lemma 6
Let \({\mathfrak {R}}\) be a \(T_R\) reconciliation, \((s_1,s_2)\in E(S')\), \(e_1=(x_1,x_2),e_2=(y_1,y_2)\in E(G')\) such that \(s_1\ge \rho (x_1)\ge \rho (x_2)\ge s_2\), and \(s_1\ge \rho (y_1)\ge \rho (y_2)\ge s_2\). Then \(e_1\) and \(e_2\) are aligned edges.
Proof
Assume the opposite. Let \(E_1\) be a maximal edge from \(E(S')\) that contains two unaligned edges \(e_1,e_2\in E(G')\) (Fig. 11a). Observe two cases.
Case 1. Assume that \(E_1\ne root_E(S)\). Let \(E_2\in E(S')\) be the parent of \(E_1\). Then \(E_2\) does not contain two unaligned edges from \(E(G')\). This is possible only if \(e_1\) or \(e_2\) are neighbours to a transfer. But, every transfer has a loss assigned, therefore we again have two unaligned edges (counting the lost ones) in \(E_2\), which is a contradiction.
Case 2. Assume that \(E_1=root_E(S)\). Since the only way to obtain two unaligned edges is a duplication or transfer, it is obvious that this case is also impossible. \(\square \)
Lemma 7
If \({\mathfrak {R}}\) is a \(T_R\) reconciliation of a gene tree G and a species tree S, then every extant species has exactly one extant gene assigned, i.e. for every \(s\in L(S)\) there is exactly one \(x\in L(G)\) such that \(\phi (x)=s\).
Proof
Let us prove that every extant species has at least one extant gene assigned. Assume the opposite. Let \(s_1 \in L(S)\) be an extant species with no assigned extant gene. Let \(f\in E(S')\cup V(S)\) be the minimal element satisfying \(s_1 < f\), and there is \(x\in V(G')\) such that \(\rho (x)=f\). As a result of the minimality of f, we have that x is not a speciation. Therefore \(f\in E(S')\). Assume that x is a minimal element of \(V(G')\) assigned to f. Then x is a loss, and it is not assigned to a transfer, i.e. x is a non-free loss. Since \({\mathfrak {R}}\) is a \(T_R\) reconciliation, it cannot have non-free losses. A contradiction.
Now we will prove the lemma’s claim. Assume the opposite. Let \(s_2\in L(S)\) be a species with at least two genes assigned (say \(x_1,x_2\in L(G)\)). Let \(E_2\in E(S')\) be the edge incident with \(s_2\). Then \(E_2\) contains at least two edges \(e_1,e_2\in E(G)\) (edges incident with \(x_1,x_2\)). This contradicts Lemma 6. \(\square \)
a There are no unaligned edges in a \(T_R\) reconciliation, because if there is an edge \(E_1\in E(S')\) with two unaligned edges from \(G'\), then the parent edge \(E_2\) also contains two unaligned edges from \(G'\). b If we raise a speciation, then we obtain a new transfer. Therefore, by raising a speciation we cannot obtain a minimal \(T_R\) reconciliation
We need the notion of extending losses in order to explain some of the properties of \(T_R\) reconciliation. The loss extension is depicted in Fig. 9b.
Definition 18
(Loss extension) Let \(l_1\in E(G')\) be a loss assigned to \(E_1\in E(S')\), \(E_2,E_3\in E(S')\) be the children of \(E_1\), and \(X_1\in V(S)\) be the common vertex of \(E_1,E_2,E_3\). Insert new vertices \(l_2,l_3\) into \(V(G')\), and connect them with \(l_1\). Next, take \(\rho (l_1)=X_1\), \(\rho (l_2)=E_2\), \(\rho (l_3)=E_3\). This procedure we call a loss extension, and we say that the loss \(l_1\) is extended.
The next lemma states that if we have only transfers, losses, speciations (i.e. we have a TL reconciliation), and every edge of \(S'\) has at most one lineage from \(G'\), then we can extend these losses in a unique way to obtain a \(T_R\) reconciliation (this is not necessarily minimum \(T_R\) reconciliation). Figure 12 depicts this fact.
Lemma 8
Let \({\mathfrak {R}}\) be a TL reconciliation such that every extant species has exactly one extant gene assigned, every \(E_1\in E(S')\) contains at most one edge from \(E(G')\), and every lost subtree has only one edge. Then there is a \(T_R\) reconciliation \({\mathfrak {R}}'\), such that \(G'_{{\mathfrak {R}}'}\) is an extension of \(G'_{{\mathfrak {R}}}\), \(\rho _{{\mathfrak {R}}} = \rho _{{\mathfrak {R}}'}/G'_{{\mathfrak {R}}}\). Among all \(T_R\) reconciliations obtained in this way, there is only one of minimal weight, and it is obtained by extending losses.
Proof
Let l be a loss, and \(\rho (l)=E_1\in E(S')\). If there is a transfer child in \(E_1\), then assign l to the transfer child.
If there is no transfer child in \(E_1\), then extend l.
Repeat the previous process until all losses are assigned to the transfer children. In this way, by extending losses, we obtain a \(T_R\) reconciliation \({\mathfrak {R}}'\). Since every extant species has exactly one extant gene assigned, during the process of extension every loss will encounter a lineage from G, i.e. a transfer child. Therefore every (extended) loss can be assigned to a transfer child.
Since all transfers in \({\mathfrak {R}}\) are from G, they are also transfers in \({\mathfrak {R}}'\). Hence the weight of \({\mathfrak {R}}'\) is not less than the number of transfers. On the other hand, all losses in \({\mathfrak {R}}'\) are free, therefore \(\omega ({\mathfrak {R}}')\) equals to the number of transfers, i.e. \({\mathfrak {R}}'\) is a \(T_R\) reconciliation of minimal weight obtainable from \({\mathfrak {R}}\).
If \({\mathfrak {R}}''\) is another \(T_R\) reconciliation of minimal weight obtained from \({\mathfrak {R}}\), then \({\mathfrak {R}}''\) does not contain extra transfers, i.e. all lost subtree of \(G'_{{\mathfrak {R}}''}\) contain only speciations, i.e. all lost subtrees are obtained by extending losses.
Since a \(T_R\) reconciliation does not contain more than one edge of \(G'\) per edge in \(S'\) (Lemma 6), there is a unique reconciliation obtained by extending losses. With this we conclude the proof. \(\square \)
From the previous lemmas we have that, when observing \(T_R\) reconciliations without additional lost transfers (the minimum reconciliations are among them), it is enough to observe G instead of \(G'\), i.e. we are interested only in positions of V(G) inside \(S'\).
4.1 Normalized reconciliation
In order to reduce the search space of the algorithm that searches for a minimum \(T_R\) reconciliation, we introduce the notion of normalized reconciliation. The algorithm we give can output every normalized reconciliation with a non-null probability. Every minimum reconciliation can be transformed into a normalized one, without changing its cost, by the operations we call transfer adjustment, node raising, and node translocation. Formally, we perform these operations by modifying \(G'\) and \(\rho \).
The next definition introduces transfer adjustment (Fig. 13). The purpose of this operation is to obtain that all transfer parents are in V(G).
Definition 19
(Transfer adjustment) Let \({\mathfrak {R}}\) be a minimum \(T_R\) reconciliation, \((x_0,x'_1)\) be a transfer and \(x_0\notin V(G)\), \(x={{\textsc {p}}}^{{\textsc {v}}}_G(x_0)\), \(x_1,\ldots ,x_k\in V(G')\) such that \(x_0<x_1<\cdots<x_k<x\), \(x'_2\) be another child of x in \(G'\) (\(x'_2\ne x_k\)), and \(x''_i\) be the child of \(x'_i\) (\(i=1,2\)).
Let \({\mathfrak {R}}'\) be a \(T_R\) reconciliation obtained from \({\mathfrak {R}}\) by modifying \(G'\) and \(\rho \) in the following way.
If x is a transfer parent, let \(f_i=\rho (x'_i)\in E(S')\) (\(i=1,2\)); \(f_{min}\in \{f_1,f_2\}\) such that \(\tau (f_{min})=min\{\tau (f_1),\tau (f_2)\}\); \(f_{max}\in \{f_1,f_2\}\) such that \(f_{max}\ne f_{min}\); \(x'={{\textsc {p}}}^{{\textsc {v}}}_{G'}(x)\). Transform \(G'\) as follows: remove all the edges (from \(G'\)) incident with x; remove the edges \((x'_{max},x''_{max})\), \((x_0,x'_1)\); suppress \(x_0\), and insert \(x''\); connect x with \(x'_{max}\), \(x''_{max}\), and \(x'_{min}\); connect \(x''\) with \(x_k\), \(x'\), and \(x'_{max}\). Define \(\rho _{{\mathfrak {R}}'}\): \(\rho _{{\mathfrak {R}}'}(x'')=\rho _{{\mathfrak {R}}}(x)\); \(\rho _{{\mathfrak {R}}'}(x)=\rho _{{\mathfrak {R}}}(x'_{max})\); \(\rho _{{\mathfrak {R}}'}(y)=\rho _{{\mathfrak {R}}}(y)\) for all \(y\in V(G'_{{\mathfrak {R}}})\backslash \{x_0\}\).
If x is a speciation, let \(s_0=\rho _{{\mathfrak {R}}}(x)\); \(f\in E(S')\) be the parent edge of \(s_0\). Transform \(G'\) as follows: remove the edges \((x_0,x'_1)\), \((x,x_k)\), and \((x,x'_2)\); suppress \(x_0\), and insert \(x_{k+1}\); connect x and \(x'_1\); connect \(x_{k+1}\) with x, \(x_k\), and \(x'_2\). Define \(\rho _{{\mathfrak {R}}'}\): \(\rho _{{\mathfrak {R}}'}(x)=f\); \(\rho _{{\mathfrak {R}}'}(x_{k+1})=s_0\); \(\rho _{{\mathfrak {R}}'}(y)=\rho _{{\mathfrak {R}}}(y)\) for all \(y\in V(G'_{{\mathfrak {R}}})\backslash \{x_0\}\).
Transfer adjustment. Transfer \((x_0,x'_1)\) has the transfer parent \(x_0\notin V(G)\). Next, \(x_1,x_2,x_3\notin V(G)\) and \(x_0<x_1<x_2<x_3<x\), where \(x={{\textsc {p}}}^{{\textsc {v}}}_G(x_0)\). a Node x is a transfer parent, and \((x,x'_2)\) is a transfer. After the adjustment, the new transfer \((x'',x'_2)\) needs adjusting. b Node x is a speciation. After the adjustment, x becomes a transfer parent, and no new transfer needs adjusting
In Fig. 13a we have \(f_{min}=f_1\), \(f_{max}=f_2\), and \(k=3\). After adjusting \((x_0,x'_1)\) (Fig. 13a), the new transfer \((x'',x'_2)\) needs adjusting.
Since the number of transfers is not changed by transfer adjustments, the next lemma is obvious, and we omit a proof.
Lemma 9
If \({\mathfrak {R}}'\) is obtained from a minimum \(T_R\) reconciliation by transfer adjustments, then \({\mathfrak {R}}'\) is also a minimum \(T_R\) reconciliation.
The next notion is depicted in Fig. 14.
Definition 20
(Node raising) Let \({\mathfrak {R}}\) be a \(T_R\) reconciliation with all transfer parents from V(G), and \(x\in V(G')\) be a transfer parent or transfer child. We can create a new reconciliation \({\mathfrak {R}}'\) in the following way.
Assume that \(x \in V(G)\) is a transfer parent, \((x,x')\) is a transfer, \(x''\) is the minimal node such that \(x<x''\) and (\(x''={{\textsc {p}}}^{{\textsc {v}}}_G(x)\) or \(x''\) is a transfer child). Let f be the maximal edge from \(E(S')\) such that \(\rho (x) \le f\le \rho (x'')\). Let \(x_1,\ldots ,x_k \in V(G')\backslash V(G)\) such that \(x<x_1<\cdots<x_k<x''\).
Define \(\rho _{{\mathfrak {R}}'}\) as \(\rho _{{\mathfrak {R}}'}(x)=f\), \(\rho _{{\mathfrak {R}}'}(y)=\rho _{{\mathfrak {R}}}(y)\) for all \(y\in V(G')\backslash \{x\}\), \((x,x')\) is also a transfer in \({\mathfrak {R}}'\). Reattach \(x_1,\ldots ,x_k\) below x so that \(x_1<\cdots<x_k<x<x''\).
Now, assume that \(x\in V(G')\) is a transfer child, \(l\in V(G')\) is a loss assigned to x, and l is a leaf of a lost subtree \(T_l\). Let f be the maximal edge from \(E(S')\) such that \(\rho (x)\le f<\rho (root(T_l))\), and \(\tau (f)\le \tau ({{\textsc {p}}}^{{\textsc {v}}}_{G'}(x))\). Also \(x_1,\ldots ,x_k \in V(G')\backslash V(G)\) such that \(l<x_1<\cdots<x_k<root(T_l)\) and \(\rho (x_k)\le f\).
Define \(\rho _{{\mathfrak {R}}'}\) as \(\rho _{{\mathfrak {R}}'}(x)=f\), \(\rho _{{\mathfrak {R}}'}(y)=\rho _{{\mathfrak {R}}}(y)\) for all \(y\in V(G')\backslash \{x\}\), \(({{\textsc {p}}}^{{\textsc {v}}}_{G'}(x),x)\) is a transfer in \({\mathfrak {R}}'\). Reattach \(x_1,\ldots ,x_k\) below x so that \(x_1<\cdots<x_k<x\).
Node raising (\(x\in V(G')\) is raised). a Node x is a transfer parent, \(x_1,x_2,x_3\in V(G')\backslash V(G)\), \(x''=min\{y\mid x<y \wedge (y\in V(G) \vee y \text { is a transfer child})\}\). Node x cannot be raised higher than \(x''\). b Now \(x\in V(G')\backslash V(G)\) is a transfer child, l is the loss assigned to x, and \(T_l\) is the lost subtree with a leaf l. Node x cannot be raised higher than \({{\textsc {p}}}^{{\textsc {v}}}_{G'}(x)\), and it must stay below \(root(T_l)\)
Note that we do not raise speciations, because it would create a new transfer (Fig. 11b) and increase the weight of a reconciliation.
We do not place a raised node in a speciation from S. If we raise a transfer parent x, then it remains a transfer parent, i.e. adjusted transfers remain adjusted.
From the previous comments we have the following lemma.
Lemma 10
If \({\mathfrak {R}}'\) is obtained from a minimum \(T_R\) reconciliation by raising nodes, then \({\mathfrak {R}}'\) is also a minimum \(T_R\) reconciliation.
Proof
Note that in a \(T_R\) reconciliation, \(\omega ({\mathfrak {R}})\) is the number of transfers. By raising nodes, we do not raise speciations, hence we do not create new transfers. Therefore \(\omega ({\mathfrak {R}}')\le \omega ({\mathfrak {R}})\). Since \({\mathfrak {R}}\) is a minimum \(T_R\) reconciliation, we obtain \(\omega ({\mathfrak {R}}')=\omega ({\mathfrak {R}})\), i.e. \({\mathfrak {R}}'\) is a minimum \(T_R\) reconciliation. \(\square \)
We cannot raise a node if its parent from G is in the same time interval, or its parent (from G) is a speciation in the next time interval.
Sometimes, nodes of \(G'\) can be raised only if moved to another edge of \(S'\) in the same time interval (Fig. 15). That is why we introduce node translocation.
Definition 21
(Node translocation) Let \({\mathfrak {R}}\) be a \(T_R\) reconciliation, \((x,x')\) be a horizontal transfer, \(x\in V(G)\), \(y'\in V(G')\) be a transfer child, \(\rho _{{\mathfrak {R}}}(x)=\rho _{{\mathfrak {R}}}(y')=E_1\in E(S')\), \(x_i\in V(G)\) such that \(x<x_i<y'\) (\(i=1,\ldots ,k\)), \(\rho (x')=E_2\in E(S')\). Let \({\mathfrak {R}}'\) be a \(T_R\) reconciliation obtained from \({\mathfrak {R}}\) by modifying \(G'\) and \(\rho \) as follows. Insert a new node \(y''\) into \(G'\), suppress \(x'\), remove edge \((x,x'')\), where \(x''\) is a child of x in \(G'\) and \(x''\ne x'\). Insert edges \((x,y'')\) and \((y'',x'')\). Define \(\rho _{{\mathfrak {R}}'}(x)=\rho _{{\mathfrak {R}}'}(x_i)=E_2\) (\(i=1,\ldots ,k\)), \(\rho _{{\mathfrak {R}}'}(y'')=E_1\), \(\rho _{{\mathfrak {R}}'}(y)=\rho _{{\mathfrak {R}}}(y)\) for all \(y\in V(G')\backslash \{x,y',x_1,\ldots ,x\}\). This operation we call a node translocation, and we say that nodes x and \(x_i\) (\(i=1,\ldots ,k\)) are translocated to\(E_2\).
Node translocation. Node \(y'\) is a diagonal transfer child. Transfer \((x,x')\) is horizontal. Nodes x and \(x_1\) are horizontal transfer parents, and \(x_2\) is a diagonal transfer parent. a We cannot raise nodes \(x,x_1,x_2\). b We move \(x,x_1,x_2\), and \(y'\) to \(E_2\). The number of transfers does not change. Now we can raise x, \(x_1\), and \(x_2\)
Note that if all transfers incident with x and \(x_i\) (\(i=1,\ldots ,k\)) are diagonal, then we cannot translocate the nodes. Also, if \(y'\) is a diagonal transfer child, then we can raise x and \(x_i\) (\(i=1,\ldots ,k\)) after translocation.
Node translocation does not change the number of transfers. Hence, the next lemma is obvious, and we do not prove it.
Lemma 11
If \({\mathfrak {R}}'\) is obtained from a minimum \(T_R\) reconciliation by node translocations, then \({\mathfrak {R}}'\) is also a minimum \(T_R\) reconciliation.
The next definition introduces the notion of normalized reconciliation, which represents the output of the main algorithm. The purpose of introducing this type of reconciliation is to reduce the search space of the branch and bound algorithm that we will to give.
Basically, a minimum reconciliation is normalized if we cannot raise any node, i.e. all nodes are “as high as possible”, and all transfer parents are from V(G). Nodes are grouped. Every group has one highest node (say y), that “blocks” the other nodes. Node y can be a speciation, transfer child, or root(G).
Figure 16 depicts a normalized reconciliation. In the group with \(x_2\) and \(x_4\), y is a speciation. In the group with \(x_3\), y is root(G). Nodes \(x_1\) and \(x_5\) are blocked by non-labeled transfer children.
Definition 22
(Normalized reconciliation) Let \({\mathfrak {R}}\) be a minimum \(T_R\) reconciliation that satisfies the following conditions. Every transfer parent is from V(G). For every transfer \((x,x')\) and \(y\in V(G')\) that is the maximal element such that \(x\le y\), \(\rho (x)\le \rho (y)\), and \(\tau (y) \le \tau (x)+1\), one of the following conditions is satisfied:
-
\(\tau (y)=\tau (x)\) and y is a horizontal transfer child;
-
\(\tau (y)=\tau (x)\), y is a diagonal transfer child, and \((x,x')\) is a diagonal transfer;
-
\(\tau (y)=\tau (x)+1\) and \(y\in V(G)\) is a speciation;
-
\(\tau (y)=\tau (x)+1\) and \(y=root(G)\).
For every diagonal transfer \((x,x')\), we have \(|E(T_l)|=1\) and \(\tau (l)=\tau (root(T_l))-1\), where l is a loss assigned to \(x'\), and \(T_l\) is a lost subtree with a leaf l. Then the reconciliation \({\mathfrak {R}}\) is called a normalized reconciliation.
A normalization of a minimum reconciliation. a Nodes \(x_1,x_4\in V(G)\) are transfer parents. We have two more transfer parents that are not from V(G), which makes 4 transfers in total. Node \(x_5\) is a transfer child, and \(x_2,x_3\in V(G)\) are speciations. b A reconciliation obtained after adjusting transfers incident with \(x_1\) and \(x_2\), and raising nodes \(x_2,x_3,x_4,x_5\). c A normalized reconciliation obtained after translocating and raising node \(x_1\)
Note that any normalized reconciliation is also minimum, and has all transfers adjusted. The proof of Theorem 4 describes how to construct a normalized reconciliation from an arbitrary minimum reconciliation.
Theorem 4
Let \({\mathfrak {R}}\) be a minimum \(T_R\) reconciliation. Then there is a normalized reconciliation \({\mathfrak {R}}'\) that can be obtained from \({\mathfrak {R}}\) by adjusting transfers, and raising and translocating nodes.
Proof
See Appendix. \(\square \)
4.2 Random normalized reconciliation
In this subsection we describe an FPT algorithm that returns a normalized reconciliation with weight not greater than k, if there is one.
The problem definition follows.
k-Minimum \(T_R\) Reconciliation:
Input:S, G, k
Output: A normalized \(T_R\) reconciliation \({\mathfrak {R}}\) such that \(\omega ({\mathfrak {R}})\le k\), or a message that there is no such reconciliation.
We are given S, G and \(\phi \), which is, in this particular case, a bijection between the leaves of G and the leaves of S (Fig. 17a). Let \(A_i\) be extant species (leaves of S), and \(a_i\) be extant genes (leaves of G) \((i=1,\ldots , n)\). We will maintain, during the execution of the algorithm, a set of active edges which initially contains the terminal edges of G, i.e. the edges with a leaf as an extremity. Some of the active edges might be lost edges, but initially none is.
We will first describe the algorithm, give appropriate figures, and in the next subsection we will give pseudocodes.
The description of the algorithm for one time slice-Cases 1 and 2. a The initialization part. To every extant gene \(a_i\) (\(i=1,\ldots ,n\)) an active edge that is incident wiht \(a_i\) is assigned. b Speciation \(s_0\in V(S)\) is in the current time slice. Edges \(e_1,e_2,e_3\) are active edges. c If at least one of the edges \(e_1\) and \(e_2\) is lost, then they coalesce at \(s_0\), and the non-lost edge is propagated to the next time slice, as well as all other edges from the current time slice. d If \(e_1\) and \(e_2\) are incident, then they coalesce at \(s_0\)
At the beginning, to every extant gene \(a_i\), where \(\phi (a_i)=A_i\) (Fig. 17a), an edge from E(G) that is incident with \(a_i\), is assigned. At this moment, these are the active edges.
We repeat the next part for every time slice, going bottom-up.
Observe one time slice. Let \(s_0\) be the internal node of S in this time slice. Let \(E_1, E_2\) be the edges from S incident with \(s_0\), and \(e_1, e_2\) active edges that belong to \(E_1, E_2\) (Fig. 17b). We have several cases.
Case 1. At least one of the edges \(e_1\) or \(e_2\) is lost. Then coalesce them at \(s_0\) (meaning the lca of the two edges in G is mapped to \(s_0\) by \(\rho \)), and the edge that is not lost propagate to the next time slice. If both edges \(e_1\) and \(e_2\) are lost, then their parent (which is also a lost edge) is propagated to the next time slice. All other edges are propagated to the next time slice as well (Fig. 17c), where they remain active.
Case 2. Edges \(e_1\) and \(e_2\) are incident in G. Then coalesce them at \(s_0\) (Fig. 17d). All other active edges propagate to the next time slice, where they remain active. The parent edge of \(e_1\) and \(e_2\) is also an active edge in the next time slice.
Case 3. Edges \(e_1\) and \(e_2\) are neither lost nor incident. Branch and bound tree is branching into three subtrees (subcases \((a_1)\), \((a_2)\), and (b)).
Case 3\(a_1\). Put \(e_1\)on hold (Fig. 18a). This means that \(e_1\) is not propagated into the next time slice, but stays active as long as it does not become a (diagonal) transfer (see Case 3b). Edge \(e_2\) and all other active edges from the current time slice are propagated into the next time slice.
The description of the algorithm for one time slice-Case 3. a Edge \(e_1\) is put on hold (staying active), waiting to become a (diagonal) transfer. Edge \(e_2\) is propagated to the next time slice, as well as all other active edges from the current time slice. b Let x be the minimal ancestor of \(e_1\) and \(e_2\) in V(G). Assign x to \(s_0\), and randomly expand all nodes between x and \(e_1\), and between x and \(e_2\)
Case 3\(a_2\). The same as Case 3\(a_1\), but \(e_2\) is on hold instead of \(e_1\).
Case 3b. Let x (Fig. 18b) be the minimum node in V(G) that is an ancestor of both \(e_1\) and \(e_2\), \(x'_1,\ldots , x'_{k_1}\in V(G)\) be the vertices in the path from \(e_1\) to x, and \(x''_1,\ldots , x''_{k_2}\in V(G)\) be the vertices in the path from \(e_2\) to x. Take \(\rho (x)=s_0\). Observe \(x'_1\). Let \(e_3\) be a random active edge with maximal \(\tau \)-value that is a descendant of \(x'_1\), \(E_3\) be the edge from E(S) that contains \(e_3\), and \(x^1_1,\ldots , x^{m1}_1\) (\(m_1\ge 0\)) be the vertices in the path from \(e_3\) to \(x'_1\). Add these vertices and corresponding edges to \(E_3\), as well as transfer \((x'_1, x^1_1)\). Repeat the process for every child of \(x'_1\). It is possible that some of the added transfers are diagonal. We say that \(x'_1\) is randomly expanded. In \(E_3\) add a lost edge \(e'_3\). If \(e_3\) was on hold, then coalesce \(e'_3\) with the other lineage at \(s_0\). Otherwise propagate \(e'_3\) to the next time slice as an active edge. Next, randomly expand the remaining nodes \(x'_2,\ldots , x'_{k_1}\), \(x''_1,\ldots , x''_{k_2}\).
The condition of maximality of \(\tau (e_3)\) is necessary, because we cannot have transfers to the past.
Case 4. We reached \(root_E(S)\). Then randomly expand all the remaining nodes from V(G) and \(\rho (root(G))=root(S)\).
The rest of the procedure is a standard branch and bound. When we reach the first solution (reconciliation) with at most k transfers, we denote it by \({\mathfrak {R}}^*\). If \({\mathfrak {R}}\) is some other reconciliation, obtained in the branch and bound process, such that \(\omega ({\mathfrak {R}})<\omega ({\mathfrak {R}}^*)\), then we take \({\mathfrak {R}}^*={\mathfrak {R}}\). If \(\omega ({\mathfrak {R}})=\omega ({\mathfrak {R}}^*)\), then we randomly take \({\mathfrak {R}}^*={\mathfrak {R}}\).
If during the branch and bound procedure, we obtain a (partial) reconciliation with more than k transfers, then we do not branch, and go one step back.
4.3 Pseudocodes and properties
In this section we give pseudocodes, prove some properties of the algorithm, and give a proof that k-Minimum\(T_R\)Reconciliation is fixed-parameter tractable.




Theorem 5
Let \({\mathfrak {R}}\) be a normalized reconciliation and \(\omega ({\mathfrak {R}})\le k\). Then \({\mathfrak {R}}\) is a possible output of Algorithm 1.
Proof
See Appendix. \(\square \)
Theorem 6
If Algorithm 1 returns a reconciliation \({\mathfrak {R}}\), then \(\omega ({\mathfrak {R}})\le k\) and \({\mathfrak {R}}\) is a normalized reconciliation.
Proof
See Appendix. \(\square \)
Theorem 7
Time complexity of Algorithm 1 is \(O(3^k n)\).
Proof
Branching in the algorithm occurs only when we add transfers, i.e. with every branching we add at least one transfer. Therefore we can have the branch depth at most k. Since the reconciliation is normalized (hence minimal), one of 3a1, 3a2, 3b holds. Given that we branch to three cases (\(a_1, a_2\), and b), the size of the branch and bound tree is \(O(3^k n)\). \(\square \)
Theorem 8
Minimum\(T_R\)Reconciliation is fixed-parameter tractable with respect to the parameter that represents an upper bound for the reconciliation’s weight.
Proof
Follows directly from Theorems 4, 5, 6, and 7 . \(\square \)
5 The Minimum Dated SPR Scenario is NP-hard and FPT
Finally, we prove that a constrained version of the well known SPR (subtree prune and regraft) distance problem, the Minimum Dated SPR Scenario, mentioned in Song (2006), is equivalent to the Minimum\(T_R\)Reconciliation problem.
Definition 23
(Dated SPR operation) Let T be a dated, fully binary, rooted tree, \(e_1=(a_2, a_1)\), \(e_2=(b_2, b_1)\in E(T)\), where \(a_2={{\textsc {p}}}^{{\textsc {v}}}_T(a_1)\), \(b_2={{\textsc {p}}}^{{\textsc {v}}}_T(b_1)\), and \(\tau (a_1) < \tau (b_2)\). Delete \(e_1\), suppress \(a_2\), subdivide \(e_2\) with a node \(a'_2\), where \(\tau (a_1) \le \tau (a'_2)\), connect \(a_1\) and \(a'_2\). Denote the obtained tree by \(T'\). We say that \(T'\) is obtained from T by a dated SPR (subtree prune and regraft) operation.
We will denote this SPR operation by \(spr((a_2,a_1),(b_2,b_1))=a'_2\), or by \(T\rightarrow T'\). Note that if \(spr((a_2,a_1),(b_2,b_1))=a'_2\), \(spr((a_2,a_1),(b_2,b_1))=a''_2\), and \(\tau (a'_2)\ne \tau (a''_2)\), then these two SPR operations are different.
Definition 24
(Minimum Dated SPR Scenario problem) Let T and \(T'\) be rooted, fully binary trees, where T is a dated and \(T'\) is undated tree. Assigning dates to \(V(T')\), and finding a minimum number (over all possible date assignments to \(V(T')\)) of SPR operations that transform T into \(T'\) is called the Minimum Dated SPR Scenario problem. The number of SPR operations is called the length of SPR scenario.
Now we introduce a parametrized version of the problem we are interested in.
k-Minimum Dated SPR Scenario:
Input:T—dated, \(T'\)—undated, full binary, rooted trees; \(k\ge 0\) is a parameter;
Output: A minimum dated SPR scenario with the length not greater than k, or a message that a scenario like this does not exist.
Lemma 12
The k-Minimum Dated SPR Scenario is (polynomially) equivalent to the k-Minimum\(T_R\)Reconciliation.
Proof
See Appendix. \(\square \)
Theorem 9
The Minimum Dated SPR Scenario is NP-hard.
Proof
Since there is a polynomial reduction from Minimum\(T_R\)Reconciliation to Minimum Dated SPR Scenario (Lemma 12) and Minimum\(T_R\)Reconciliation is NP-hard (Theorem 3), then Minimum Dated SPR Scenario is NP-hard. \(\square \)
Theorem 10
The Minimum Dated SPR Scenario is fixed-parameter tractable with respect to parametrized distance.
Proof
Since there is a polynomial reduction (which is also an FPT reduction) from k-Minimum Dated SPR Scenario to k-Minimum\(T_R\)Reconciliation (Lemma 12) and Minimum\(T_R\)Reconciliation is FPT (Theorem 8), then Minimum Dated SPR Scenario is FPT. \(\square \)
6 Conclusion
We propose an integration of two ways of detecting lateral gene transfers, and more generally to construct gene histories and handle the species tree gene tree discrepancies. On one side, SPR scenarios model transfers with replacements and are limited by computational complexity issues, the difficulty to include time constraints and other gene scale events like transfers without replacement, duplications, conversions and losses. On the other side, reconciliation algorithms usually work with dynamic programming, necessitating an independence hypothesis on different gene tree lineages, incompatible with replacing transfers.
It is a big issue for biological models, because the results can depend on the type of methodology which is chosen, leading to simplification hypotheses. Moreover, algorithms are often tested with simulations containing the same hypotheses as the inference models. This is why it can be important to explore methodological issues at the edge of both methods, which is what we do here.
Future work include imagining a way to include transfer with replacement in standard reconciliation software. This will require more integration and probably more efficient algorithms so that it does not harm the computing time.
References
Abby SS, Tannier E, Gouy M, Daubin V (2012) Lateral gene transfer as a support for the tree of life. Proc Natl Acad Sci USA 109(13):4962–4967. https://doi.org/10.1073/pnas.1116871109
Allen BL, Steel M (2001) Subtree transfer operations and their induced metrics on evolutionary trees. Ann Comb 5(1):1–15. https://doi.org/10.1007/s00026-001-8006-8
Bansal MS, Alm EJ, Kellis M (2012) Efficient algorithms for the reconciliation problem with gene duplication, horizontal transfer and loss. Bioinformatics 28(12):283–291. https://doi.org/10.1093/bioinformatics/bts225
Bansal MS, Alm EJ, Kellis M (2013) Reconciliation revisited: handling multiple optima when reconciling with duplication, transfer, and loss. J Comput Biol 20(10):738–754. https://doi.org/10.1089/cmb.2013.0073
Beiko RG, Hamilton N (2006) Phylogenetic identification of lateral genetic transfer events. BMC Evol Biol 6(1):15. https://doi.org/10.1186/1471-2148-6-15
Bonet ML, John KS (2009) Efficiently calculating evolutionary tree measures using SAT. vol 5584. LNCS. Springer, Berlin. pp 4–17. https://doi.org/10.1007/978-3-642-02777-2_3
Bordewich M, Semple C (2005) On the computational complexity of the rooted subtree prune and regraft distance. Ann Comb 8(4):409–423. https://doi.org/10.1007/s00026-004-0229-z
Chan Y, Ranwez V, Scornavacca C (2015) Exploring the space of gene/species reconciliations with transfers. J Math Biol 71(5):1179–1209. https://doi.org/10.1007/s00285-014-0851-2
Chauve C, El-Mabrouk N (2009) New perspectives on gene family evolution: losses in reconciliation and a link with supertrees. Lecture Notes in Computer Science (including subseries lecture notes in artificial intelligence and lecture notes in bioinformatics) 5541 LNBI, pp 46–58. https://doi.org/10.1007/978-3-642-02008-7_4
Chen ZZ, Fan Y, Wang L (2015) Faster exact computation of rSPR distance. J Comb Optim 29(3):605–635. https://doi.org/10.1007/s10878-013-9695-8
Chen J, Shi F, Wang J (2016) Approximating maximum agreement forest on multiple binary trees. Algorithmica 76(4):867–889. https://doi.org/10.1007/s00453-015-0087-6
Choi SC, Rasmussen MD, Hubisz MJ, Gronau I, Stanhope MJ, Siepel A (2012) Replacing and additive horizontal gene transfer in streptococcus. Mol Biol Evol 29(11):3309–3320. https://doi.org/10.1093/molbev/mss138
Dasgupta B, Ferrarini S, Gopalakrishnan U, Paryani NR (2006) Inapproximability results for the lateral gene transfer problem. J Comb Optim 11(4):387–405. https://doi.org/10.1007/s10878-006-8212-8
Doyon JP, Scornavacca C, Ranwez V, Berry V (2010) An efficient algorithm for gene/species trees parsimonious reconciliation with losses, duplications, and transfers. In: Comparative genomics: international workshop, RECOMB-CG 2010, Ottawa, Canada, October 9–11, 2010 Proceedings (October), pp 93–108. https://doi.org/10.1007/978-3-642-16181-0_9
Doyon JP, Ranwez V, Daubin V, Berry V (2011) Models, algorithms and programs for phylogeny reconciliation. Briefings Bioinf 12(5):392–400. https://doi.org/10.1093/bib/bbr045
Even S, Itai A, Shamir A (1976) On the complexity of timetable and multicommodity flow problems. SIAM J Comput 5(4):691–703. https://doi.org/10.1137/0205048
Garey MR, Johnson DS (1979) Computers and Intractability: a guide to the theory of NP-completeness. W. H. Freeman & Co., New York
Garey M, Johnson D, Stockmeyer L (1976) Some simplified NP-complete graph problems. Theor Comput Sci 1(3):237–267. https://doi.org/10.1016/0304-3975(76)90059-1
Goodman M, Czelusniak J, Moore GW, Romero-Herrera AE, Matsuda G (1979) Fitting the gene lineage into its species lineage, a parsimony strategy illustrated by cladograms constructed from globin sequences. Syst Biol 28(2):132–163. https://doi.org/10.1093/sysbio/28.2.132
Hallett MT, Lagergren J (2001) Efficient algorithms for lateral gene transfer problems. In: Proceedings of the fifth annual international conference on computational biology. RECOMB ’01. ACM, New York, pp 149–156. https://doi.org/10.1145/369133.369188
Hasić D, Tannier E (2019) Gene tree species tree reconciliation with gene conversion. J Math Biol. https://doi.org/10.1007/s00285-019-01331-w
Hein J, Jiang T, Wang L, Zhang K (1996) On the complexity of comparing evolutionary trees. Discrete Appl Math 71(1–3):153–169. https://doi.org/10.1016/S0166-218X(96)00062-5
Hickey G, Dehne F, Rau-Chaplin A, Blouin C (2008) SPR distance computation for unrooted trees. Evol Bioinform 4:17–27. https://doi.org/10.4137/EBO.S419
Keeling PJ, Palmer JD (2008) Horizontal gene transfer in eukaryotic evolution. Nat Rev Genet 9:605–618. https://doi.org/10.1038/nrg2386
Linz S, Semple C (2011) A cluster reduction for computing the subtree distance between phylogenies. Ann Comb 15(3):465–484. https://doi.org/10.1007/s00026-011-0108-3
Merkle D, Middendorf M, Wieseke N (2010) A parameter-adaptive dynamic programming approach for inferring cophylogenies. BMC Bioinf 11(1):S60. https://doi.org/10.1186/1471-2105-11-S1-S60
Nakhleh L (2012) Computational approaches to species phylogeny inference and gene tree reconciliation. Biophys Chem 34(1):13–23. https://doi.org/10.1016/j.immuni.2010.12.017
Raman V, Ravikumar B, Rao S (1998) A simplified NP-complete MAXSAT problem. Inf Process Lett 65(1):1–6. https://doi.org/10.1016/S0020-0190(97)00223-8
Rice DW, Palmer JD (2006) An exceptional horizontal gene transfer in plastids: gene replacement by a distant bacterial paralog and evidence that haptophyte and cryptophyte plastids are sisters. BMC Biol 4(1):31. https://doi.org/10.1186/1741-7007-4-31
Scornavacca C, Paprotny W, Berry V, Ranwez V (2013) Representing a set of reconciliations in a compact way. J Bioinform Comput Biol 11(02):1250025. https://doi.org/10.1142/S0219720012500254
Shi F, Feng Q, Chen J, Wang L, Wang J (2013) Distances between phylogenetic trees: a survey. Tsinghua Sci Technol 18(5):490–499. https://doi.org/10.1109/TST.2013.6616522
Shi F, Feng Q, You J, Wang J (2016) Improved approximation algorithm for maximum agreement forest of two rooted binary phylogenetic trees. J Comb Optim 32(1):111–143. https://doi.org/10.1007/s10878-015-9921-7
Song YS (2006) Properties of subtree-prune-and-regraft operations on totally-ordered phylogenetic trees. Ann Comb 10(1):147–163. https://doi.org/10.1007/s00026-006-0279-5
Suchard MA (2005) Stochastic models for horizontal gene transfer: taking a random walk through tree space. Genetics 170(1):419–431. https://doi.org/10.1534/genetics.103.025692
Szöllősi GJ, Tannier E, Lartillot N, Daubin V (2013) Lateral gene transfer from the dead. Syst Biol 62(3):386–397. https://doi.org/10.1093/sysbio/syt003
Szöllősi GJ, Tannier E, Daubin V, Boussau B (2015) The inference of gene trees with species trees. Syst Biol 64(1):42–62. https://doi.org/10.1093/sysbio/syu048
Tofigh A, Hallett M, Lagergren J (2011) Simultaneous identification of duplications and lateral gene transfers. IEEE ACM Trans Comput Biol Bioinform 8(2):517–535. https://doi.org/10.1109/TCBB.2010.14
Whidden C, Matsen F (2018) Calculating the unrooted subtree Prune-and-Regraft distance. IEEE ACM Trans Comput Biol Bioinform: 1–1. https://doi.org/10.1109/TCBB.2018.2802911
Whidden C, Beiko RG, Zeh N (2010) Fast FPT algorithms for computing rooted agreement forests: theory and experiments. Springer, Berlin, pp 141–153. https://doi.org/10.1007/978-3-642-13193-6_13
Whidden C, Beiko RG, Zeh N (2016) Fixed-parameter and approximation algorithms for maximum agreement forests of multifurcating trees. Algorithmica 74(3):1019–1054. https://doi.org/10.1007/s00453-015-9983-z
Wu Y (2009) A practical method for exact computation of subtree prune and regraft distance. Bioinformatics 25(2):190–196. https://doi.org/10.1093/bioinformatics/btn606
Acknowledgements
ET was supported by the French Agence Nationale de la Recherche (ANR) through Grant No. ANR-10-BINF-01–01 ‘Ancestrome’.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix: Proofs
Appendix: Proofs
Here we give proofs that are omitted in the main text.
Proof of Lemma 1
We will use a reduction from the Max 2-Sat, where every variable appears in at most three clauses. This problem is NP-hard (Raman et al. 1998).
Let F be an instance of the Max 2-Sat, and every variable appears in at most three clauses. If there is a variable x that appears exactly once, and it belongs to a clause C, then we can assign it a value and make C a true clause. Similarly, if there is a variable y that has only positive, or only negative literals, then we can assign it a value to make the corresponding clauses true.
In this way we eliminate all the variables that appear exactly once, or have only positive or only negative literals. Therefore, we can assume that F has variables that appear two or three times, and have both positive and negative literals.
Let \(x_0\) be a variable that appears in exactly two clauses. After inserting \((x_0\vee x^0_1)\wedge (x^0_1\vee x^0_2)\wedge (\lnot x^0_1\vee x^0_3)\wedge (\lnot x^0_2\vee x^0_3)\wedge (x^0_2\vee \lnot x^0_3)\) into F, we obtain a logical formula with \(x_0\) in exactly three clauses. The new variables \(x^0_1,x^0_2,x^0_3\) also appear in exactly three clauses, and they have positive and negative literal present.
In this way we obtain a logical formula \(F'\) that has every variable in exactly three clauses, with both positive and negative literal present. If the added variables are true, then all the new clauses will be true. Therefore the number of true clauses in F is maximized if and only if the number of true clauses in \(F'\) is maximized.
The previous reduction is obviously polynomial in n and m, where n is the number of variables, and m is the number of clauses in F. \(\square \)
Proof of Lemma 4
To prove (a) and (b), we identify two cases, according to the positions of \(r_{j_1}\) and \(r_{j_2}\).
Case 1. Node \(r_{j_1}\) or \(r_{j_2}\) is above the border line. In order to obtain \(\omega (F_j)<4\), we need that some of the nodes \(r^0_{j_1}, r^0_{j_2}, r^1_{j_1}, r^1_{j_2}\) is neither a duplication nor incident with a transfer. The only way to have this is if some of them is placed in \(s^0_{C_j}=root(S_{C_j})\).
Let us take \(\rho (r^0_{j_1}) = s^0_{C_j}\). Then \(\rho (r_{j_1}) > s^0_{C_j}\), or \(\rho (r_{j_1})\) and \(s^0_{C_j}\) are incomparable. If \(\rho (r^1_{j_1}) < s^0_{C_j}\), then \((r_{j_1},r^1_{j_1})\) is a transfer, and the weight of \(F_j\) is not decreased. If \(\rho (r^1_{j_1}) = s^0_{C_j}\), then \(r_{j_1}\) is a duplication, or one of the edges \((r_{j_1}, r^0_{j_1})\) and \((r_{j_1}, r^1_{j_1})\) contains a transfer. In this way we eliminate two transfers (that were incident with \(r^0_{j_1}\) and \(r^1_{j_1}\)), and obtain one transfer or duplication. But we generate at least one non-free loss in \(S_{C_j}\). Similar considerations apply to the other nodes of \(F_j\). Hence we cannot obtain \(\omega (F_j)<4\).
Case 2. Both nodes \(r_{j_1}\) and \(r_{j_2}\) are under the border line. Then none of the nodes \(r^0_{j_1}, r^1_{j_1}, r^0_{j_2}, r^1_{j_2}\) is placed in \(s^0_{C_j}\), therefore every one of them is incident with at least one transfer. If we wish to eliminate transfers starting at \(r_{j_1}\) or \(r_{j_2}\), then we need to place them both in \(lca(B_5,B_6,B_7,B_8)\), i.e. in the minimal node in \(S_{C_j}\) that is ancestor of \(B_5\), \(B_6\), \(B_7\), and \(B_8\) (Fig. 19). In this case we increase the number of non-free losses. Whichever placement we choose, we have \(\omega (F_j)\ge 5\).
Lemma 4a, b, Case 2. When \(r_{j1}\) and \(r_{j2}\) are positioned in \(lca(B_5,B_6,B_7,B_8)\) we obtain extra non-free losses
(c) The proof is similar in spirit to the proof of (a). See Figs. 4 and 5 . The idea is to see what happens if some of the 17 transfers, present in a proper reconciliation that belongs to \(G_{x_i}\), is not present in some other reconciliation.
First, note that if some of the nodes \(d^j_i\) are not placed in \(D^i_j\) (\(j=1,\ldots ,P(n)\)), then we would have transfers that are not present in a proper reconciliation. Also, if none of the nodes \(d^j_i\) is placed in \(D^i_j\) (\(j=1,\ldots ,P(n)\)), then we would have a reconciliation more expensive than any proper reconciliation. Hence we can assume that, for the anchoring nodes \(d^1_i\), we have \(\rho (d^1_i)=D^i_j\) (\(j=1,\ldots ,P(n)\)).
In a proper reconciliation, there are 14 transfers incident with \(b^s_i\)\((s=1,\ldots , 14)\). In an arbitrary reconciliation, we can achieve that no transfer or a duplication is incident with \(b^s_i\) only if \(\rho (b^s_i)=s^0_{x_i}\). Then a parent of \(b^s_i\) (i.e. \(c^{s-1}_i\)), as well as \(c^0_i\), is a duplication, or is incident with a transfer, and two or more non-free losses are created. Therefore by having \(\rho (b^s_i)=s^0_{x_i}\), for some values of s, does not give \(\omega (G_{x_i})<17\).
Assume that the nodes \(b^s_i\) (\(s=1,\ldots ,14\)) are placed as in the proper reconciliation. Observe nodes \(x^1_i, x^2_i\), and assume that they are not incident with a transfer, and the edge \((x^2_i, x^1_i)\) does not contain a transfer. Then we have at least two transfers at the edges \((c^4_i, x^2_i)\) and \((x^1_i, c^3_i)\), or at some other edges leading to some of the \(b^s_i\). Similar considerations apply for \(x^3_i\). Therefore, in this case too we cannot decrease the number of transfers.
Can we have less than 17 transfers if take \(\rho (b^s_i)=s^0_{x_i}\), for some values of s, and eliminate transfers incident with \(x^1_i, x^2_i\)? Let us take \(\rho (b^7_i)=s^0_{x_i}\). Then the nodes \(c^6_i\) and \(c^0_i\) are not placed as in the proper reconciliation. Hence we have at least two transfers or duplications, and non-free losses not present in the proper reconciliation. Also, if the nodes \(x^1_i,x^2_i\) are not placed as in the proper reconciliation, we have a transfer, different from the previous two, that is not present in the proper reconciliation. Therefore, we have at least three evolutionary events not present in the proper reconciliation, and we cannot obtain less than 17 events.
(d) Let us take that \(x^1_i\) and \(x^3_i\) are under the border line. Then at least three of the nodes \(c^1_i,\ldots , c^{12}_i\) are not on the gadgets positions. Some of these nodes are \(c^1_i,c^2_i,c^3_i\), because they are descendants of \(x^1_i\) in G. The paths \((c^1_i,b^2_i, A^3_i)\), \((c^2_i,b^3_i, A^5_i)\), \((c^3_i,b^4_i, A^7_i)\) generate extra three transfers. An extra transfer is created on the edge \((x^2_i,x^1_i)\), or on some other edge that is an ancestor of \(x^2_i\). Even if we we eliminate the two transfers incident with \(x^1_i\) and \(x^2_i\), we gain 4 more. Hence \(\omega (G_{x_i})\ge 19\). \(\square \)
Proof of Theorem 1
Let \({\mathfrak {R}}\) be a minimum \(DTLCT_R\) reconciliation. We use \({\mathfrak {R}}\) to construct \({\mathfrak {R}}'\) that is both minimum and proper.
The construction of a proper reconciliation is described earlier. The only thing that we need to specify in \({\mathfrak {R}}'\) is the positions of \(x^1_i,x^2_i\), and \(x^3_i\) with respect to the border line, as well as the positions of \(r_{j_1}\) and \(r_{j_2}\).
If \(x^1_i\) and \(x^2_i\) are not on the same side of the border line as \(x^3_i\) (in \({\mathfrak {R}}\)), then they are on the same side in \({\mathfrak {R}}'\) as in \({\mathfrak {R}}\). If \(x^1_i\) or \(x^2_i\) is on the same side as \(x^3_i\) (in \({\mathfrak {R}}\)), then \(x^1_i\) and \(x^2_i\) are above, and \(x^3_i\) is under the border line (in \({\mathfrak {R}}'\)).
Next, the vertices of \(F_j\) are placed in \(S_{C_j}\) as in the description of the proper reconciliation (Definition 16), so that the nodes \(r_{j_1}\) and \(r_{j_2}\) are placed on the same side of the border line as \(x'_{j_1}\) and \(x'_{j_2}\) (in \({\mathfrak {R}}'\)), respectively. A reconciliation, obtained in this way, we denote by \({\mathfrak {R}}'\). By construction, it is a proper reconciliation. Let us prove that it is a minimum reconciliation.
We have \(\omega _{{\mathfrak {R}}}(G_{x_i})\ge 17 = \omega _{{\mathfrak {R}}'}(G_{x_i})\), \(\omega _{{\mathfrak {R}}}(F_j)\ge 4\), and \(\omega _{{\mathfrak {R}}'}(F_j)\in \{4, 5\}\) (Lemma 4).
Let \(i\in \{1,\ldots ,n\}\), \(x^1_i,x^2_i,x^3_i\) be connected with \(r_{a_1}\in V(F_a), r_{b_1}\in V(F_b),\)\(r_{c_1}\in V(F_c)\) via transfers. We introduce a notation \(\varOmega _{{\mathfrak {R}}}(i)=\omega _{{\mathfrak {R}}}(G_{x_i})+\omega _{{\mathfrak {R}}}(F_a)+\omega _{{\mathfrak {R}}}(F_b)+\omega _{{\mathfrak {R}}}(F_c)\).
Case 1. Assume that \(\omega _{{\mathfrak {R}}}(F_a)\ge \omega _{{\mathfrak {R}}'}(F_a)\), \(\omega _{{\mathfrak {R}}}(F_b)\ge \omega _{{\mathfrak {R}}'}(F_b)\), \(\omega _{{\mathfrak {R}}}(F_c)\ge \omega _{{\mathfrak {R}}'}(F_c)\). Then \(\omega _{{\mathfrak {R}}}(G_{x_i})+\omega _{{\mathfrak {R}}}(F_a)+\omega _{{\mathfrak {R}}}(F_b)+\omega _{{\mathfrak {R}}}(F_c) \ge \omega _{{\mathfrak {R}}'}(G_{x_i})+\omega _{{\mathfrak {R}}'}(F_a)+\omega _{{\mathfrak {R}}'}(F_b)+\omega _{{\mathfrak {R}}'}(F_c)\), i.e.\(\varOmega _{{\mathfrak {R}}}(i) \ge \varOmega _{{\mathfrak {R}}'}(i)\).
Case 2. Assume that \(\omega _{{\mathfrak {R}}}(F_a)=4\), \(\omega _{{\mathfrak {R}}'}(F_a)=5\), \(\omega _{{\mathfrak {R}}}(F_b)\ge \omega _{{\mathfrak {R}}'}(F_b)\), \(\omega _{{\mathfrak {R}}}(F_c)\ge \omega _{{\mathfrak {R}}'}(F_c)\). Since \(\omega _{{\mathfrak {R}}'}(F_a)=5\), we have that \(x^1_i\) is under the border line (in \({\mathfrak {R}}'\)). Because of the transformation rules, at the beginning of the proof, we have that \(x^1_i\), \(x^2_i\) are under the border line (in \({\mathfrak {R}}\) and \({\mathfrak {R}}'\)), while \(x^3_i\) is above the line (in \({\mathfrak {R}}\) and \({\mathfrak {R}}'\)).
Let \(y_1\) be a literal of variable \(x_s\) (i.e.\(y_1\in \{x^1_s,x^2_s,x^3_s\}\)) connected with \(r_{a_2}\in V(F_a)\) via transfer. Since \(\omega _{{\mathfrak {R}}}(F_a)=4\), \(\omega _{{\mathfrak {R}}'}(F_a)=5\), we have that \(y_1\) is above the border line in \({\mathfrak {R}}\), and under the line in \({\mathfrak {R}}'\), hence \(y_1=x^3_s\).
Assume that \(F_{a'},F_{b'}\) are connected with \(x^1_s,x^2_s\) via transfers. Then \(\omega _{{\mathfrak {R}}'}(F_{a'})=\omega _{{\mathfrak {R}}'}(F_{b'})=4\), \(\omega _{{\mathfrak {R}}}(G_{x_s}) \ge 19\). We have \(\omega _{{\mathfrak {R}}}(F_{a'})\ge 4=\omega _{{\mathfrak {R}}'}(F_{a'})\) and \(\omega _{{\mathfrak {R}}}(F_{b'})\ge 4=\omega _{{\mathfrak {R}}'}(F_{b'})\).
From the previous arguments, \(\omega _{{\mathfrak {R}}}(G_{x_s})+\omega _{{\mathfrak {R}}}(F_a)+\omega _{{\mathfrak {R}}}(F_a) \ge \)\(19+4+4 =\)\(17+5+5 =\)\(\omega _{{\mathfrak {R}}'}(G_{x_s})+\omega _{{\mathfrak {R}}'}(F_a)+\omega _{{\mathfrak {R}}'}(F_a)\).
Finally, \(\big ( \omega _{{\mathfrak {R}}}(G_{x_i})+\omega _{{\mathfrak {R}}}(F_a)+\omega _{{\mathfrak {R}}}(F_b)+\omega _{{\mathfrak {R}}}(F_c) \big ) + \)\(\big ( \omega _{{\mathfrak {R}}}(G_{x_s})+\omega _{{\mathfrak {R}}}(F_{a'})+\omega _{{\mathfrak {R}}}(F_{b'})+\omega _{{\mathfrak {R}}}(F_{a}) \big ) \ge \)\(\big ( \omega _{{\mathfrak {R}}'}(G_{x_i})+ \omega _{{\mathfrak {R}}'}(F_a) +\omega _{{\mathfrak {R}}'}(F_b)+\omega _{{\mathfrak {R}}'}(F_c) \big ) + \)\(\big ( \omega _{{\mathfrak {R}}'}(G_{x_s}) + \omega _{{\mathfrak {R}}'}(F_{a'}) +\omega _{{\mathfrak {R}}'}(F_{b'})+\omega _{{\mathfrak {R}}'}(F_{a}) \big )\), i.e. \(\varOmega _{{\mathfrak {R}}}(i)+\varOmega _{{\mathfrak {R}}}(s) \ge \varOmega _{{\mathfrak {R}}'}(i)+\varOmega _{{\mathfrak {R}}'}(s)\).
The next cases use the approach of Case 2.
Case 3. Assume that \(\omega _{{\mathfrak {R}}}(F_b)=4\), \(\omega _{{\mathfrak {R}}'}(F_b)=5\), \(\omega _{{\mathfrak {R}}}(F_a)\ge \omega _{{\mathfrak {R}}'}(F_a)\), \(\omega _{{\mathfrak {R}}}(F_c)\ge \omega _{{\mathfrak {R}}'}(F_c)\). This case is analogous to Case 2.
Case 4. Assume that \(\omega _{{\mathfrak {R}}}(F_c)=4\), \(\omega _{{\mathfrak {R}}'}(F_c)=5\), \(\omega _{{\mathfrak {R}}}(F_a)\ge \omega _{{\mathfrak {R}}'}(F_a)\), \(\omega _{{\mathfrak {R}}}(F_b)\ge \omega _{{\mathfrak {R}}'}(F_b)\). Then \(x^3_i\) is under, and \(x^1_i,x^2_i\) are above the border line in \({\mathfrak {R}}'\). We have two subcases.
Case 4.1. Assume that \(x^1_i\) or \(x^2_i\) was on the same side of the line as \(x^3_i\) (in \({\mathfrak {R}}\)). Then \(\omega _{{\mathfrak {R}}}(G_{x_i})\ge 19\). Hence \(\omega _{{\mathfrak {R}}}(G_{x_i})+\omega _{{\mathfrak {R}}}(F_a)+\omega _{{\mathfrak {R}}}(F_b)+\omega _{{\mathfrak {R}}}(F_c) \ge \)\(19 + \omega _{{\mathfrak {R}}'}(F_a)+\omega _{{\mathfrak {R}}'}(F_b)+ 4>\)\(17 + \omega _{{\mathfrak {R}}'}(F_a)+\omega _{{\mathfrak {R}}'}(F_b)+ 5 =\)\(\omega _{{\mathfrak {R}}'}(G_{x_i})+\omega _{{\mathfrak {R}}'}(F_a)+\omega _{{\mathfrak {R}}'}(F_b)+\omega _{{\mathfrak {R}}'}(F_c)\), i.e. \(\varOmega _{{\mathfrak {R}}}(i) > \varOmega _{{\mathfrak {R}}'}(i)\).
Case 4.2. Assume that \(x^1_i\) and \(x^2_i\) were not on the same side of the line as \(x^3_i\) (in \({\mathfrak {R}}\)). Then \(x^3_i\) is under the line (in \({\mathfrak {R}}\) and \({\mathfrak {R}}'\)). Now we proceed similar to Case 2.
Let \(y_3\in \{x^1_l,x^2_l,x^3_l\}\) and it is connected with \(r_{c_2}\in V(F_c)\) via transfer. From \(\omega _{{\mathfrak {R}}}(F_c)=4\), \(\omega _{{\mathfrak {R}}'}(F_c)=5\), we have that \(y_3\) in \({\mathfrak {R}}\) was above the line, and in \({\mathfrak {R}}'\) is under the line, hence \(y_3=x^3_l\), \(\omega _{{\mathfrak {R}}}(G_{x_l}) \ge 19\), \(\omega _{{\mathfrak {R}}'}(F_{a''}) = \omega _{{\mathfrak {R}}'}(F_{b''}) = 4\), where \(F_{a''}\) and \(F_{b''}\) are connected with \(x^1_l\) and \(x^2_l\) via transfers.
It follows that \(\omega _{{\mathfrak {R}}}(G_{x_l}) + \omega _{{\mathfrak {R}}}(F_c) + \omega _{{\mathfrak {R}}}(F_c) \ge 19+4+4 =\)\(17+5+5=\)\(\omega _{{\mathfrak {R}}'}(G_{x_l}) + \omega _{{\mathfrak {R}}'}(F_c) + \omega _{{\mathfrak {R}}'}(F_c)\).
Next, \(\big ( \omega _{{\mathfrak {R}}}(G_{x_i})+\omega _{{\mathfrak {R}}}(F_a)+\omega _{{\mathfrak {R}}}(F_b)+\omega _{{\mathfrak {R}}}(F_c) \big ) + \)\(\big ( \omega _{{\mathfrak {R}}}(G_{x_l})+\omega _{{\mathfrak {R}}}(F_{ a''})+\omega _{{\mathfrak {R}}}(F_{b''})+\omega _{{\mathfrak {R}}}(F_{c}) \big ) \ge \)\(\big ( \omega _{{\mathfrak {R}}'}(G_{x_i})+\omega _{{\mathfrak {R}}'}(F_a)+\omega _{{\mathfrak {R}}'}(F_b)+\omega _{{\mathfrak {R}}'}(F_c) \big ) + \)\(\big ( \omega _{{\mathfrak {R}}'}(G_{x_l})+\omega _{{\mathfrak {R}}'}(F_{a''})+\omega _{{\mathfrak {R}}'}(F_{b''})+\omega _{{\mathfrak {R}}'}(F_{c}) \big )\), i.e. \(\varOmega _{{\mathfrak {R}}}(i)+\varOmega _{{\mathfrak {R}}}(l) \ge \varOmega _{{\mathfrak {R}}'}(i)+\varOmega _{{\mathfrak {R}}'}(l)\).
Case 5. Assume that \(\omega _{{\mathfrak {R}}}(F_a)=\omega _{{\mathfrak {R}}}(F_b)=4\), \(\omega _{{\mathfrak {R}}'}(F_a)=\omega _{{\mathfrak {R}}'}(F_b)=5\), and \(\omega _{{\mathfrak {R}}}(F_c)\ge \omega _{{\mathfrak {R}}'}(F_c)\). By a similar argument as in the previous cases, we have that \(x^1_i,x^2_i\) are under the line (in \({\mathfrak {R}}\) and \({\mathfrak {R}}'\)), while \(x^3_i\) is above the line (in \({\mathfrak {R}}\) and \({\mathfrak {R}}'\)). Let \(y_1\in \{x^1_r,x^2_r,x^3_r\}\) be connected with \(r_{a_2}\in V(F_a)\), and \(y_2\in \{x^1_t,x^2_t,x^3_t\}\) be connected with \(r_{b_2}\in V(F_b)\). As in the previous cases, we have \(y_1=x^3_r\), \(y_2=x^3_t\), and they were above the line in \({\mathfrak {R}}\), and under the line in \({\mathfrak {R}}'\). Hence \(\omega _{{\mathfrak {R}}}(G_{x_r}) \ge 19\) and \(\omega _{{\mathfrak {R}}}(G_{x_t}) \ge 19\). Let \(x^1_r,x^2_r,x^1_t,x^2_t\) be connected with \(F_{a_r},F_{b_r},F_{a_t},F_{b_t}\). Then \(\omega _{{\mathfrak {R}}'}(F_{a_r})=\omega _{{\mathfrak {R}}'}(F_{b_r})=\omega _{{\mathfrak {R}}'}(F_{a_t})=\omega _{{\mathfrak {R}}'}(F_{b_t})=4\).
Therefore \(\omega _{{\mathfrak {R}}}(G_{x_r}) + \omega _{{\mathfrak {R}}}(G_{x_t}) + \omega _{{\mathfrak {R}}}(F_a) + \omega _{{\mathfrak {R}}}(F_a) + \omega _{{\mathfrak {R}}}(F_b) + \omega _{{\mathfrak {R}}}(F_b) \ge \)\(19+19+4+4+4+4 = 17+17+5+5+5+5 = \)\(\omega _{{\mathfrak {R}}'}(G_{x_r}) + \omega _{{\mathfrak {R}}'}(G_{x_t}) + \omega _{{\mathfrak {R}}'}(F_a) + \omega _{{\mathfrak {R}}'}(F_a) + \omega _{{\mathfrak {R}}'}(F_b) + \omega _{{\mathfrak {R}}'}(F_b)\).
Hence \(\big ( \omega _{{\mathfrak {R}}}(G_{x_i})+\omega _{{\mathfrak {R}}}(F_a)+\omega _{{\mathfrak {R}}}(F_b)+\omega _{{\mathfrak {R}}}(F_c) \big ) +\)\(\big ( \omega _{{\mathfrak {R}}}(G_{x_r})+\omega _{{\mathfrak {R}}}(F_{a_r})+\omega _{{\mathfrak {R}}}(F_{b_r})+\omega _{{\mathfrak {R}}}(F_a) \big ) +\)\(\big ( \omega _{{\mathfrak {R}}}(G_{x_t})+\omega _{{\mathfrak {R}}}(F_{a_t})+\omega _{{\mathfrak {R}}}(F_{b_t})+\omega _{{\mathfrak {R}}}(F_b) \big ) \ge \)\(\big ( \omega _{{\mathfrak {R}}'}(G_{x_i})+\omega _{{\mathfrak {R}}'}(F_a)+\omega _{{\mathfrak {R}}'}(F_b)+\omega _{{\mathfrak {R}}'}(F_c) \big ) +\)\(\big ( \omega _{{\mathfrak {R}}'}(G_{x_r})+\omega _{{\mathfrak {R}}'}(F_{a_r})+\omega _{{\mathfrak {R}}'}(F_{b_r})+\omega _{{\mathfrak {R}}'}(F_a) \big ) +\)\(\big ( \omega _{{\mathfrak {R}}'}(G_{x_t})+\omega _{{\mathfrak {R}}'}(F_{a_t})+\omega _{{\mathfrak {R}}'}(F_{b_t})+\omega _{{\mathfrak {R}}'}(F_b) \big )\), i.e. \(\varOmega _{{\mathfrak {R}}}(i) + \varOmega _{{\mathfrak {R}}}(r) + \varOmega _{{\mathfrak {R}}}(t) \ge \)\(\varOmega _{{\mathfrak {R}}'}(i) + \varOmega _{{\mathfrak {R}}'}(r) + \varOmega _{{\mathfrak {R}}'}(t)\).
The next three cases are not possible, because \(x^3_i\) cannot be on the same side of the line as \(x^1_i\) or \(x^2_i\) in \({\mathfrak {R}}'\).
Case 6. Assume that \(\omega _{{\mathfrak {R}}}(F_a)=\omega _{{\mathfrak {R}}}(F_c)=4\), \(\omega _{{\mathfrak {R}}'}(F_a)=\omega _{{\mathfrak {R}}'}(F_c)=5\), \(\omega _{{\mathfrak {R}}}(F_b) \ge \omega _{{\mathfrak {R}}'}(F_b)\).
Case 7. Assume that \(\omega _{{\mathfrak {R}}}(F_b)=\omega _{{\mathfrak {R}}}(F_c)=4\), \(\omega _{{\mathfrak {R}}'}(F_b)=\omega _{{\mathfrak {R}}'}(F_c)=5\), \(\omega _{{\mathfrak {R}}}(F_a) \ge \omega _{{\mathfrak {R}}'}(F_a)\).
Case 8. Assume that \(\omega _{{\mathfrak {R}}}(F_a)=\omega _{{\mathfrak {R}}}(F_b)=\omega _{{\mathfrak {R}}}(F_c)=4\), \(\omega _{{\mathfrak {R}}'}(F_a)=\omega _{{\mathfrak {R}}'}(F_b)=\omega _{{\mathfrak {R}}'}(F_c)=5\).
Every \(i\in \{1,\ldots , n\}\) belongs to exactly one case. Variables s (from Cases 2 and 3), l (Case 4.2), t and r (Case 5) are equal to some \(i\in \{1,\ldots , n\}\), but are different among themselves, i.e. there is no value that repeats itself among variables s, l, r, t. Let \(A_1\) be the set of all values of i from the Case 1 that are different from all s, l, r, t. In a similar manner we introduce sets \(A_{2,3}\), \(A_{4.1}\), \(A_{4.2}\), \(A_{5}\)
We will use the previous cases to prove \(\omega ({\mathfrak {R}}) \ge \omega ({\mathfrak {R}}')\). We have \(2\cdot \omega ({\mathfrak {R}}) = \sum _{i}\omega _{{\mathfrak {R}}}(G_{x_i}) + \sum _{i}\varOmega _{{\mathfrak {R}}}(i) = \)\(\sum _{i}\omega _{{\mathfrak {R}}}(G_{x_i})\)\(+\)\(\sum _{A_1} \varOmega _{{\mathfrak {R}}}(i)\)\(+\)\(\sum _{A_{2,3}}\big ( \varOmega _{{\mathfrak {R}}}(i) + \varOmega _{{\mathfrak {R}}}(s) \big )\)\(+\)\(\sum _{A_{4.1}} \varOmega _{{\mathfrak {R}}}(i)\)\(+\)\(\sum _{A_{4.2}}\big ( \varOmega _{{\mathfrak {R}}}(i) + \varOmega _{{\mathfrak {R}}}(l)\big )\)\(+\)\(\sum _{A_5}\big ( \varOmega _{{\mathfrak {R}}}(i) + \varOmega _{{\mathfrak {R}}}(r) + \varOmega _{{\mathfrak {R}}}(t) \big )\)\(\ge \)\(\sum _{i}\omega _{{\mathfrak {R}}'}(G_{x_i})\)\(+\)\(\sum _{A_1} \varOmega _{{\mathfrak {R}}'}(i)\)\(+\)\(\sum _{A_{2,3}}\big ( \varOmega _{{\mathfrak {R}}'}(i) + \varOmega _{{\mathfrak {R}}'}(s) \big )\)\(+\)\(\sum _{A_{4.1}} \varOmega _{{\mathfrak {R}}'}(i)\)\(+\)\(\sum _{A_{4.2}}\big ( \varOmega _{{\mathfrak {R}}'}(i) + \varOmega _{{\mathfrak {R}}'}(l)\big )\)\(+\)\(\sum _{A_5}\big ( \varOmega _{{\mathfrak {R}}'}(i) + \varOmega _{{\mathfrak {R}}'}(r) + \varOmega _{{\mathfrak {R}}'}(t) \big )\)\(=\)\(2\cdot \omega ({\mathfrak {R}}')\).
Finally, \(\omega ({\mathfrak {R}}) \ge \omega ({\mathfrak {R}}')\). Therefore \({\mathfrak {R}}'\) is a minimum reconciliation. \(\square \)
Proof of Theorem 4
We will transform \({\mathfrak {R}}\) into \({\mathfrak {R}}'\) in two steps. First, we adjust all transfers that need to be adjusted, and then we alternately raise and translocate nodes.
Note that the definitions of the operations give the sufficient conditions for executing them. We assume that we perform these operations only if the conditions are satisfied. Additionally, we will translocate nodes only if \(y'\) (\(y'\) is from Definition 21) is a diagonal transfer child. This results in a raisable node after translocation.
Step 1. We adjust all transfers, with the transfer parent not from G, in an arbitrary order. This procedure will end in polynomial number of steps.
Indeed, we have two situations. In the first situation (Fig. 13a) we obtain another transfer that needs adjusting (transfer \((x'',x'_2)\)), but the transfer parent \(x''\) is positioned above x in \(G'\). In the second situation (Fig. 13b), the total number of transfers waiting for adjusting decreases by 1. Hence the effect of adjusting transfer is either positioning transfer parent higher in \(G'\), or reducing the number of unadjusted transfer. Since we are bounded by the size of \(G'\) and the number of transfers, the number of adjustments is finite and it is polynomial in size of G and S.
Therefore all transfers will be adjusted.
Step 2. Take an arbitrary transfer parent or a transfer child \(x\in V(G')\) that we can raise, and raise it. Repeat the previous procedure, as long as there is a node that we can raise.
After there are no more transfer parents or children that can be raised, translocate some node, it there is such a node. Then, again, raise all nodes that can be raised. Note that we translocate a node only if it results in a raisable node (see the second paragraph of this proof).
Repeat the previous procedure of raising and translocating nodes as long as possible.
This procedure will end in a polynomial number of steps. Indeed, by raising a node x, \(\tau (x)\) increases. Since \(\tau (x)<\tau (root(G))\), we have that Step 2 must end in polynomial number of steps, i.e. we will obtain a reconciliation in which no transfer parent or child can be raised.
By applying Steps 1 and 2 we obtain a reconciliation \({\mathfrak {R}}'\). Since the number of transfers is not changed (Lemmas 9, 10, and 11 ), we have \(\omega ({\mathfrak {R}}')=\omega ({\mathfrak {R}})\), i.e. \({\mathfrak {R}}'\) is a minimum reconciliation. We need to prove that \({\mathfrak {R}}'\) is a normalized reconciliation.
Let \((x,x')\in E(G')\) be a transfer, \(y\in V(G')\) be the maximal element such that \(x \le y\), \(\rho (x) \le \rho (y)\), and \(\tau (y)\le \tau (x)+1\).
Let us prove that y is not a transfer parent. Assume the opposite. Then \(y\in V(G)\). Let \(x''\) be the element as described in Definition 20 obtained by raising y. Then \(\tau (y)=\tau (x'')\) and \(x''\) is a transfer parent or child; or \(\tau (y)=\tau (x'')-1\) and \(x''\) is a speciation or root(G). In both cases we obtain a contradiction with the maximality of y.
Let y be a transfer child. We need to prove that \(\tau (x)=\tau (y)\). Assume the opposite, i.e. \(\tau (x)<\tau (y)\). Let us take the maximal \(x_1\) such that \(x\le x_1\le y\) and \(\tau (x_1)=\tau (x)\). Since all the transfers are adjusted, we have \(x_1\in V(G)\) and \(x_1\) is a transfer parent. Since \(\tau (x_1)<\tau (y)\), node \(x_1\) can be raised, which is a contradiction with Step 2, where we raise and translocate nodes as long as possible. Therefore \(\tau (x)=\tau (y)\).
Let y be a diagonal transfer child. We need to prove that \((x,x')\) is a diagonal transfer. Assume the opposite, i.e. \(\tau (x)=\tau (x')\). Let \(\rho (y)=E_1\in E(S')\) and \(\rho (x')=E_2\in E(S')\). We have \(\tau (E_1)=\tau (E_2)=\tau (x)\). Since there are no speciations from S with the same date (see a comment after Definition 1), one of the edges \(E_1\) or \(E_2\) is not incident with a speciation from S. If \(E_1\) is not incident with a speciation, then we can raise y. If \(E_2\) is not incident with a speciation, then we can translocate x to \(E_2\) and raise x. In both situations we have a contradiction with Step 2. Therefore, \((x,x')\) is a diagonal transfer.
Let y be a speciation, or \(y=root(G)\). Then \(\tau (x)<\tau (y)\). Since \(\tau (y)\le \tau (x)+1\), we have \(\tau (y) = \tau (x)+1\).
Let \((x,x')\) be a diagonal transfer, l a loss assigned to \(x'\), \(T_l\) a lost subtree with a leaf l. From Step 2 we have that we cannot raise \(x'\). This is possible only if \(\tau (T_l)=\tau (l)+1\), and therefore \(T_l\) has only one edge.
We proved that the properties of a normalized reconciliation are satisfied. Hence \({\mathfrak {R}}'\) is a normalized reconciliation.\(\square \)
Proof of Theorem 5
Since \({\mathfrak {R}}\) is a normalized reconciliation, it is also, by definition, minimum. If I is a time slice, then \(R_I\) denotes the partial reconciliation induced by I, i.e. the part of \({\mathfrak {R}}\) that is inside I, and all other time slices before I, where “before” refer to those lower in the tree. We will prove that the algorithm constructs \({\mathfrak {R}}_I\) during the execution. We will use mathematical induction on I.
Let \(I_0\) be the first time slice (i.e. the lowest time slice), and \(s_0\in V(S)\) be a speciation such that \(\tau (s_0)\in I_0\) (Fig. 17), \(E_1, E_2\in E(S)\) are incident with \(s_0\). Next, \(e_1,e_2\) are the minimal edges of \(G'\) contained in \(E_1\) and \(E_2\). More precisely, \(e_1=(x_1,x_2)\), \(e_2=(y_1,y_2)\) are the minimal edges of \(G'\) such that \(\rho (x_2)\le E_1 \le \rho (x_1)\) and \(\rho (y_2)\le E_2 \le \rho (y_1)\). Edges \(e_1\) and \(e_2\) are unique (Lemma 6).
Let us prove that we can obtain \({\mathfrak {R}}_{I_0}\) during the execution of the algorithm. We have several cases.
Case 1. Edges \(e_1\) and \(e_2\) are incident. Let us prove that \(e_1\) and \(e_2\) coalesce at \(s_0\). Assume the opposite, \(\rho (x)\ne s_0\), where \(x\in V(G)\) is incident with both \(e_1\) and \(e_2\). Then \(e_1\) or \(e_2\) is a transfer, hence we can construct a reconciliation with smaller weight by placing x in \(s_0\), which contradicts the minimality of \({\mathfrak {R}}\). Figure 20 depicts a more detailed argumentation.
Situations that cannot occur in a normalized reconciliation. Time slice \(I_k\) contains \(E_1,E_2\), and \(s_0\). Edges \(e_1,e_2\) are active edges, and are incident with x. Edges \(E_1,E_2 \in E(S')\) are incident with \(s_0\). Node \(x\in V(G)\) must be positioned in \(s_0\in V(S)\), i.e. \(\rho (x)=s_0\). a If \(\rho (x)=E_1\), then the reconciliation is not minimal, since we can eliminate one transfer by taking \(\rho (x)=s_0\). Here, \(x''={{\textsc {p}}}^{{\textsc {v}}}_{G'}(x)\) is a transfer child, and \(x'={{\textsc {p}}}^{{\textsc {v}}}_{G'}(x'')\) is a transfer parent. b In a similar way we obtain a reconciliation with smaller weight. c, d These situaions are not possible, since the reconciliation is normalized, and we cannot have a transfer \((x_1,x_2)\) with \(x_1\notin V(G)\). Also, we could obtain a reconciliation with smaller weight, by taking \(\rho (x)=s_0\). e Here we have \(\tau (x')=\tau (x)\). Since the reconciliation is normalized, and we have only one speciation from S (i.e. \(s_0\)) in the time slice \(I_k\), then we have \(\tau (x')>\tau (s_0)\), hence this situation is not possible
Case 2. Edges \(e_1\) and \(e_2\) are not incident. We will investigate subcases. Some subcases are not obtainable by the algorithm. For them, we will prove they cannot occur in \({\mathfrak {R}}\). Let x be the minimal element from V(G) that is an ancestor of \(e_1\) and \(e_2\).
Case 2.1. Let \(\rho (x)=s_0\), \(\rho (x'_{i_1})=E_1\), \(\rho (x''_{i_2})=E_2\)\((i_1=1,\ldots ,k_1\); \(i_2=1,\ldots ,k_2.)\), where \(x'_{i_1}\) and \(x''_{i_2}\) are explained in Sect. 4.2. This case refers to Case 3b of Sect. 4.2.
We will prove that there is a random choice such that the random expansion of \(x'_1\) produces placement of the nodes identical to the one in \({\mathfrak {R}}\).
Assume the opposite, there is no such random choice. This means that we cannot obtain a situation depicted by Fig. 18b. Then there are descendants of \(x'_1\), denoted by \(y'_j\) (\(j=1,\ldots ,k\)) (Fig. 21) such that \(y'_1,\ldots ,y'_{k-1}\in V(G)\), \(y'_j={{\textsc {p}}}_{G'}(y'_{j+1})\) (\(j=1,\ldots ,k-1\)), \(y'_k\) is a transfer parent, \(y'_k \in V(G')\backslash V(G)\), and \(\rho (y'_1)=\ldots =\rho (y'_k)=E_3\). Let \(E_4\in E(S')\) be the edge that contains \(y'_{k+1}\), which is a child of \(y'_k\). By translocating \(y'_1,\ldots y'_k\) to \(E_4\) we obtain a reconciliation with one transfer less, which contradicts the minimality of \({\mathfrak {R}}\). Another reason why this case is not possible is that transfer \((y'_k,y'_{k+1})\) is not adjusted, which contradicts the fact that \({\mathfrak {R}}\) is a normalized reconciliation.
a A situation impossible for an optimal \(T_R\) reconciliation. Node \(y'_1\) is a transfer child, and its descendant \(y'_k\in V(G')\backslash V(G)\) is a transfer parent. b If we translocate \(y'_1,\ldots ,y'_k\) to \(E_4\), we eliminate one transfer, hence obtain a reconciliation with smaller weight
Therefore, we can obtain the expansion of \(x'_1\). The same reasoning applies for \(x'_2,\ldots ,x'_{k_1}\) and \(x''_1,\ldots ,x''_{k_2}\) and their children.
Case 2.2. Assume that \(E_i\in E(S')\) receives a diagonal transfer and \(e_{3-i}\in E(G)\) is propagated to the next time slice (\(i=1\) or \(i=2\)). This case is also obtainable by the algorithm (Cases 3\(a_1\) and 3\(a_2\)).
Case 2.3. Both \(e_1\) and \(e_2\) are propagated to the next time slice. Then \(s_0\) contains two unaligned edges from G, which is impossible for a \(T_R\) reconciliation (Lemma 6). Therefore this case cannot occur.
Case 2.4 We have \(\rho (x)=s_0\), and there is \(y_1\in \{x'_1,\ldots ,x'_{k_1}\}\), or \(y_2\in \{x''_1,\ldots ,x''_{k_2}\}\) such that \(\rho (y_1)\ne E_1\), or \(\rho (y_2)\ne E_2\). Since \(s_0\) is the only speciation in S in the current time slice, then all \(x'_1,\ldots ,x'_{k_1}\) and \(x''_1,\ldots ,x''_{k_2}\) are transfers.
Let \({\mathfrak {R}}'\) be a reconciliation such that \(\rho _{{\mathfrak {R}}'}(x'_1)=\ldots =\rho _{{\mathfrak {R}}'}(x'_{k_1})=E_1\), \(\rho _{{\mathfrak {R}}'}(x''_1)=\ldots =\rho _{{\mathfrak {R}}'}(x''_{k_2})=E_2\), and \(\rho _{{\mathfrak {R}}'}(y)=\rho _{{\mathfrak {R}}}(y)\) for all the remaining \(y\in V(G)\). Then \({\mathfrak {R}}'\) is a reconciliation with smaller weight than \({\mathfrak {R}}\), which contradicts the optimality of \({\mathfrak {R}}\).
Case 2.5. Assume that \(\tau (x) > \tau (s_0)\), \(\tau (y_1)\le \tau (s_0)\), and \(\tau (y_2)\le \tau (s_0)\) for some \(y_1\in \{x'_1,\ldots ,x'_{k_1}\}\), \(y_2\in \{x''_1,\ldots ,x''_{k_2}\}\). Then \({\mathfrak {R}}\) is not a normalized reconciliation. Hence this case is not possible.
Case 2.6. If x is in \(I_0\) and \(\rho (x) \ne s_0\), then by taking \(\rho (x)=s_0\) we get a reconciliation with fewer transfers (similarly to Case 1 and Fig. 20), contrary to the minimality of \({\mathfrak {R}}\).
For the inductive hypothesis part, assume that the statement is true for time slices \(I_0, I_1,\ldots , I_{k-1}\). Let us prove that it is true for \(I_k\). Proving the statement for \(I_k\) is the same as for \(I_0\), therefore we will not repeat it.
Hence \({\mathfrak {R}}_I\) is obtainable by the procedure. Since \({\mathfrak {R}}_I={\mathfrak {R}}\) for the final time slice I, \({\mathfrak {R}}\) is also obtainable by the algorithm. Since it is a minimal reconciliation, \({\mathfrak {R}}\) is a possible output of the algorithm. \(\square \)
We will use the next lemma in the proof of Theorem 6. Basically, it states that it is not important which random choice we select in Case 3b of the algorithm (see Sect. 4.2).
Lemma 13
The random choice in Case 3b of the algorithm does not affect the weight of an output reconciliation.
Proof
Let \(I_k\) be the observed time interval, and \(I_0,\ldots ,I_{k-1}\) be the time intervals before \(I_{k}\) (Fig. 22).
Let \(x'_1\) be a node that we randomly expand, and assume we have more than one choice for a random active edge with maximal \(\tau \)-value that is a descendant of \(x'_1\). Let \(e_{31}\) and \(e_{32}\) be two of those edges.
We will use notations from Sect. 4.2. When constructing \({\mathfrak {R}}_{I_k}\) from \({\mathfrak {R}}_{I_{k-1}}\), we are adding some new nodes from G. Only x is a speciation, and all other nodes are transfer parents. Since every transfer has a parent from V(G), we obtain that the number of newly added transfers is equal to the number of newly added nodes from V(G) minus one. Therefore \(\omega ({\mathfrak {R}}_{I_k})\) is not affected by a choice of \(e_3\).
Note that the active edges in \(I_{k+1}\) are not affected by a choice of \(e_3\).\(\square \)
Case 3b of the algorithm from Sect. 4.2. When randomly expanding \(x'_1\), we can have more than one candidate for \(e_3\) (denoted by \(e_{31}\) and \(e_{32}\)). The choice of \(e_3\) does not affect minimality of an output reconciliation. a We choose \(e_{31}\). b We choose \(e_{32}\). We can obtain this choice from (a) by translocating \(x^1_1\). A transfer is lost, and another is gained. Hence the number of transfers is unchanged
Proof of Theorem 6
It is obvious that \(\omega ({\mathfrak {R}})\le k\), because the algorithm cuts an edge of the branch and bound tree if \(t>k\), where t is the number of transfers in a partially constructed reconciliation.
Now we will prove that the conditions of Definition 22 are satisfied. Let \((x, x')\in E(G')\) be a transfer in \({\mathfrak {R}}\), and \(y\in V(G')\) be the maximal element such that \(x\le y\), \(\rho (x)\le \rho (y)\), \(\tau (y)\le \tau (x)+1\).
In the algorithm, transfers are created when nodes are randomly expanded. Since only nodes in V(G) are randomly expanded, every transfer starts in a node from V(G). Hence \(x\in V(G)\).
Transfers are constructed in Cases 3b and 4 (see Sect. 4.2). Therefore, y can be a speciation from G, transfer child, or root(G). If y is a speciation, then \(x\in \{x'_1,\ldots , x'_{k_1}, x''_1,\ldots , x''_{k_2}\}\), where \(x'_1,\ldots , x'_{k_1}, x''_1,\ldots , x''_{k_2}\) are explained in Sect. 4.2, and \(\tau (y)=\tau (x)+1\). If y is a transfer child, then \(\tau (y)=\tau (x)\). If y is root(G), then \(\tau (y)=\tau (x)+1\).
Let y be a diagonal transfer child. Diagonal transfers are made by using edges from G that were on hold. From Case 3a we have that a loss l, assigned to y, belongs to a lost subtree \(T_l\) with one edge and \(\tau (root(T_l))=\tau (y)+1\). Also, \((x,x')\) cannot be a horizontal transfer, because when we put an edge on hold all other edges are propagated to the next time slice, leaving no room for accepting a transfer.
Now we will prove that \({\mathfrak {R}}\) is a minimal reconciliation. The algorithm given in Sect. 4.2 is branch and bound, and it exhaustively observes every case possible for a normalized reconciliation, which we stated in the proof of Theorem 5. Also, which random option it takes in Case 3b does not affect the optimality of an output (Lemma 13). The algorithm always chooses a reconciliation of a smaller weight, if it finds one. Therefore, if it returns a reconciliation as a output, then it is a minimal reconciliation. i.e. \({\mathfrak {R}}\) is a minimal reconciliation. \(\square \)
Proof of Lemma 12
Note that if \((a_2,a_1)\in E(G)\), then there is a path in \(G'\)\((a_2, b_1, \ldots , b_s, a_1)\). The length of this path is at least 1, i.e. \(s\ge 0\). Hence every edge from G is a path in \(G'\). Also, \((a_2, a_1)\) can contain a transfer. In this proof we assume that all transfers are adjusted (as described by Definition 19 and Fig. 13), i.e. all transfers start in V(G).
We introduce a coloring of edges and nodes that were involved in some SPR operation. Let \(spr((a_2, a_1), (b_2, b_1)) = a'_2\) be the i-th SPR operation \(T_i \rightarrow T_{i+1}\). Then we color the edge \((a'_2, a_1)\) and node \(a'_2\) with color \(C_i\). If the edge \((b_2,b_1)\) was colored, then edges \((b_2,a'_2)\) and \((a'_2,b_1)\) are colored with the same color. Let \(c_1\) be the child of \(a_2\) (in \(T_i\)) different from \(a_1\), and \(c_2\) be the parent of \(a_2\) (in \(T_i\)). Then \(c_2\) is the parent of \(c_1\) (in \(T_{i+1}\)). If edge \((c_2,a_2)\) was colored with a color, then the edge \((c_2,c_1)\) is colored with the same color.
To a minimum SPR scenario we will assign a minimum \(T_R\) reconciliation. Colored edges will represent transfers, colored nodes will be transfer parents, non-colored edges will coincide with the edges of the species tree, and non-colored nodes will be speciations.
Let us first demonstrate the reduction from k-Minimum\(T_R\)Reconciliation to k-Minimum Dated SPR Scenario. Let S and G be a species and gene tree, \(S=T_0\rightarrow T_1\rightarrow \cdots \rightarrow T_k=G\) be a minimum SPR scenario transforming S into G. Using this minimum SPR scenario, we will construct a minimum \(T_R\) reconciliation.
Note that in \(T_k\) we have at most k nodes that are colored. Also, colored edges form (colored) subtrees of \(T_k\) with colored roots and inner nodes, while the leaves of these trees are not colored.
If \(a\in V(T_k)\) is a non-colored node, then it can be observed as a node from S and node from G. Take \(\rho (a)=a\in V(S)\), for all non-colored nodes \(a\in V(T_k)=V(G)\). Non-colored paths connect non-colored nodes. All non-colored edges from \(T_k=G\) place inside S so that they contain no transfer. Note that the leaves of \(T_k\) are non-colored.
Now, inside S we will place colored nodes and colored edges. Let \(T_c\) be an arbitrary colored tree, and \(c_0\) be its root. Then \(c_0\) is on a non-colored path of G, and we will leave it there in S. Next, the inner nodes of \(T_c\) we place inside S. Let \(L(T_c)=\{l_1,\ldots , l_s\}\), and \(\tau (l_1)\ge \cdots \ge \tau (l_s)\). Assume that \(c^1_1, c^1_2, \ldots , c^1_{i_1}\) are inner nodes of \(T_c\) in the path from \(l_1\) to \(c_0\) whose placement inside S is not defined. Then place these nodes in the edge of \(S'\) just above \(l_1\), i.e. \(\rho (c^1_1) = \ldots =\rho (c^1_{i_1})={{\textsc {p}}}^{{\textsc {e}}}_{S'}(\rho (l_1))\). Repeat the previous process for leaves \(l_2, \ldots l_s\). In this way we obtain a reconciliation with transfers, and every edge of S at any moment contains at most one lineage from \(G'\), hence if we extend losses we obtain a \(T_R\) reconciliation. Since a transfer can start only at a colored node, we have at most k transfers, i.e. \(\omega ({\mathfrak {R}}) \le k\).
After the next reduction, we will prove that \({\mathfrak {R}}\) is a minimum reconciliation.
In the second part, we demonstrate a reduction from k-Minimum Dated SPR Scenario to k-Minimum\(T_R\)Reconciliation. Let T be a dated and \(T'\) is an undated binary rooted tree. We need a minimum dated SPR scenario \(T=T_0\rightarrow T_1\rightarrow \cdots \rightarrow T_k=T'\).
Take \(S=T\) and \(G=T'\). Let \({\mathfrak {R}}\) be a minimum \(T_R\) reconciliation, and \(\omega ({\mathfrak {R}})=k\). We will prove that the length of minimum dated SPR scenario is k, and reconstruct it using \({\mathfrak {R}}\).
First, let us construct a scenario of the length k. Adjust all transfers in \({\mathfrak {R}}\), so they start at the nodes from V(G), just like in the first step of the proof of Theorem 4 (Definition 19, Fig. 13).
Take \(T_k=T'\), \(G_k=G\), \(G'_k=G'\), and \({\mathfrak {R}}_k={\mathfrak {R}}\). Let \((x_2, x_1)\) be an arbitrary transfer, \(x'_1\) be the child of \(x_1\) in \(G'\), l be the loss assigned to \(x_1\), and \(l_0=root(T_l)\), where \(T_l\) is a lost subtree such that \(l\in L(T_l)\). Let \(p_k = (l_0, l_1, \ldots , l_{s-1}, l_s=l)\) be a path in \(G'\) (i.e. in \(T_l\)), and therefore a lost path. Remove \((x_2, x_1)\) from \(G'_k\), suppress \(x_2\), include the path \(p_k\) into \(G_k\) (\(p_k\) is not a lost path anymore), suppress \(x_1\). Thus we eliminate one transfer, and obtain \(G_{k-1}, G'_{k-1}, {\mathfrak {R}}_{k-1}\), where \(\omega ({\mathfrak {R}}_{k-1})=\omega ({\mathfrak {R}}_k)-1\). By repeating this procedure, we obtain an SPR scenario \(T'=T_k\rightarrow T_{k-1}\rightarrow \cdots \rightarrow T_0=T\), i.e. \(T=T_0\rightarrow T_1\rightarrow \cdots \rightarrow T_k=T'\).
Since the transfers can be horizontal or diagonal, corresponding SPR operations are dated. We proved that optimal dated SPR scenario transforming T into \(T'\) has the length at most k.
Let us prove that the previous reductions construct a minimum reconciliation (the first reduction) and a minimum SPR scenario (the second reduction). Let \(T_1 \rightarrow \ldots T_k\) be a minimum SPR scenario. Take \(S=T_1, G=T_k\) and \({\mathfrak {R}}\) is a reconciliation obtained in the first reduction. We have \(k'=\omega ({\mathfrak {R}})\le k\). Now, let \(T_1=T'_1\rightarrow T'_2\rightarrow \cdots \rightarrow T'_{k''}=T_k\) be a SPR scenario obtained from G and S in the second reduction. Then \(k''\le k' \le k\). Since there is no SPR scenario, transforming \(T_1\) into \(T_k\), with the length less than k, we have \(k''=k'=k\). \(\square \)
Rights and permissions
About this article
Cite this article
Hasić, D., Tannier, E. Gene tree reconciliation including transfers with replacement is NP-hard and FPT. J Comb Optim 38, 502–544 (2019). https://doi.org/10.1007/s10878-019-00396-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10878-019-00396-z