Abstract
Picoeukaryotes are important members of the surface ocean microbial community with high diversity and significant temporal and spatial variations in community composition. Little is known about the picoeukaryotic biodiversity and community in the Yellow Sea, where hydrologic conditions are very different with the influence of the Yellow Sea Warm Current (YSWC). Using Illumina high throughput sequencing targeting 18S rDNA, we investigated the composition of picoeukaryotes at a permanent monitoring site in the central Yellow Sea from 2011 to 2013. Alveolata, Stramenopiles, and Archaeplastida were the main super groups found. Prasinophytes were dominant in N-YSWC (not influenced by the YSWC) samples whilst YSWC (influenced by the YSWC) samples were dominated by different groups, such as MALV-II (novel marine Alveolata), MAST-3, MAST-4 (novel marine Stramenopiles), and Dictyochophyceae. N-YSWC samples were grouped together in nMDS (non-metric multidimensional scaling) using the Bray–Curtis method. Distances between each two YSWC samples were greater. Based on indicator operational taxonomic unit (OTU) analysis (IOA), indicator species of the YSWC were represented by Pseudochattonella farcimen, Florenciella parvula within the class Dictyochophyceae, and Phaeocystis cordata within the class Prymnesiophyceae. The findings in our study suggest that picoeukaryotic communities in the central Yellow Sea differ temporally in response to changes in the YSWC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Picoeukaryotes (≤2 µm) play a fundamental role in both ecosystem functioning and biogeochemical cycles in the ocean. They are key components of “the microbial loop” (Pomeroy et al. 2007). Though small, picoeukaryotes make a significant contribution to the biomass and primary production in the oceans, especially coastal areas (Li 1994; Worden 2006; Not et al. 2008). The biodiversity of picoeukaryotes is high and they are composed of multiple metabolic types including phototrophs, phagotrophs, and parasites (de Vargas et al. 2015).
Methodologically, the small sub-unit RNA gene of eukaryotic cells, i.e. 18S SSU rDNA, is widely used as a universal gene maker because of its ubiquity and relatively low rate of evolution (Hillis and Dixon 1991). Sanger-based sequencing methods, combined with 18S rDNA as cloned genes, were initially and widely used to investigate the diversity of picoeukaryotes (Moon-van der Staay et al. 2001). Since the whole length of rDNA sequences could be retrieved, phylogenetic trees of picoeukaryotes assemblages were analyzed from many areas (Worden et al. 2004; Acosta et al. 2013; Rocke et al. 2013). However, Sanger-based sequencing methods often face the obstacle of low diversity coverage (Edgcomb et al. 2011). The recently developed high throughput sequencing (HTS) method, also called next-generation sequencing (NGS) technology, highlighted an avenue to sequence both environmental and cultivated samples in a fast and convenient way.
The Yellow Sea, located between the Chinese mainland and the Korean Peninsula, is a marginal sea of the northwestern Pacific. It links to the Bohai Sea in the north and the East China Sea in the south, with an average depth of about 50 m. A long-term study (from 1976 to 2000) has shown that the ecosystems of the Yellow Sea have responded significantly to changes in climate, especially increasing temperature (Lin et al. 2005). The Yellow Sea Warm Current (YSWC), a branch of the Kuroshio Current, flows from the south to the Northern Yellow Sea (northern part of the Yellow Sea) and can reach as far as the Bohai Sea. It is the only exogenous oceanic water flowing into the Yellow Sea in early winter and has a relatively higher temperature, higher salinity, and lower dissolved oxygen (Liu et al. 2015). As a surface flow, the YSWC plays as an important role in water circulation, heat transportation and coastal climate (Cui et al. 2005). Previous studies have mostly focused on the origins, paths, nutrients, and biogeochemical processes of the YSWC (Cui et al. 2004), while the biological process has been inadequately described and mostly based on traditional microscopic observations (Lü et al. 2013). The few studies on microbial community were mostly focused on the association with harmful algae blooms (Tang et al. 2013), and failed to identify inherently the YSWC microbial community, which can persist for several months every year. Until recently, the seasonal distribution of phytoplankton and its association with environmental parameters in the Yellow Sea was explored using flow cytometry and pigment analyses (Liu et al. 2015). Little is known about picoeukaryotes, although they contribute a large part to the primary production and many also act as grazers in the surface of the central Yellow Sea (Hu et al. 2004).
Many marine pelagic communities respond to environmental changes, but the level of the response differs among seasons (Edwards and Richardson 2004). Especially, picoeukaryotes respond to short-term (days) and long-term (years) changes in the sea (Countway et al. 2005; Medlin et al. 2006). Surface water in the central Yellow Sea is influenced by the YSWC from the beginning of every winter and reports have shown inter-annual variability on its strength (e.g. Song et al. 2009). In this study, we attempt to reveal the community composition of picoeukaryotes in the YSWC and its temporal and inter-annual variations by using an Illumina Miseq PE300 sequencing platform. Pair-wise primers were used to amplify the V4 region, which was identified as the most informative part of the 18S rDNA (Hugerth et al. 2014).
2 Materials and methods
2.1 Sample collection
Five picoplankton samples were taken during the National Natural Foundation Science of China (NFSC) cruise on board R/V “Dong Fang Hong 2” in none-YSWC period (sample names: S12 and S13) and YSWC period (sample names: W11, W12, and W13) from 2011 to 2013 (except for N-YSWC sample in 2011). The sampling site (34°0′N, 124°0′E) was located in the southern Yellow Sea within the YSWC. 2 L of surface seawater (3 m, sampling depth) was collected from the Niskin bottles mounted on a CTD frame (SBE911). Each water sample was immediately filtered through a 3 µm pore-sized polycarbonate filter followed by a 0.22 µm filter (Whatman, Piscataway, NJ, USA). Each filter, together with its size-fractionated contents, was carefully placed into a 5 ml tube with 2 mL of lysis buffer (50 mmol/L Tris–HCl, 1.0 mmol/L EDTA, 150 mmol/L NaCl, and 0.1% SDS). The tubes were then quickly frozen with liquid nitrogen and stored at −80 °C for DNA extraction (Stoeck and Epstein 2003) and further study.
2.2 DNA extraction and PCR amplification
Samples for extraction were thawed, and lysates were purified under a phenol–chloroform-isoamyl (1:1) extraction combined with precipitation and a washing procedure. The precipitated DNA assemblage was then diluted in 30 μL of ddH2O and kept at −20 °C for further analysis. For PCR amplification, bidirectional primers were designed to amplify V4 regions of the 18S SSU rDNA: 573FWD (5′-CGCGGTAATTCCAGCTCCA-3′) and 951REV (5′-TTGGYRAATGCTTTCGC-3′) (Hugerth et al. 2014). A subsequent limited-cycle amplification step was performed to add overhang adapters and library-specific barcodes to primers. The libraries were then generated with 2 μL of both forward and reverse primers (10 μM), PCR mix [36.7 μL of ddH2O, 5 μL of pyrobest buffer (10×), 0.3 μL of pyrobest DNA polymerase (2.5 u/μL, TaKaRa Code: DR005A), 4 μL of dNTPs (2.5 mM)], and 1 μL of template DNA. PCR amplification were completed at 95 °C for 5 min, followed by 25 cycles of 95 °C for 30 s, 56 °C for 40 s, 72 °C for 40 s, and a final extension of 10 min at 72 °C.
2.3 Illumina Miseq sequencing and data processing
PCR amplicons were checked by 2% agarose gels electrophoresis and purified using the AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA) and quantified using QuantiFluor™-ST (Promega, USA). The concentration of these purified DNA extracts was measured with Qubit 2.0 fluorometer (Thermo Fisher Scientific Inc., USA). The purified amplicons were then pooled in equimolar concentration for paired-end sequencing on an Illumina Miseq PE300 platform (http://www.illumina.com.cn/systems/miseq/workflow.aspx). Raw reads in fastq files with low quality (Q, minimum value <20 or length <200 bp) were discarded using QIIME (version 1.17). Tags were obtained by merging the paired reads according to their overlaps using COPE (Connecting Overlapped Pair-End, V1.2.3.3) after cutting off the sequences of barcodes and primers. High quality pair-wise sequences were obtained using the following standards: (1) bases with ASCII value below 33 were screened out; (2) a minimum overlap of 19 bp between reads; (3) no more than one mismatch was accepted while cutting off the sequences of primers. OTU clustering was performed at a minimum sequence similarity of 97% using QIIME (version 1.8.0). Chimeric sequences were screened out through UCHIME. Each representative OTU after clustering was compared against the Silva (SSU115) 18S rRNA database using a confidence threshold of 70% (Amato et al. 2013), for taxonomic affiliations. Lastly, OTUs affiliated with Metazoa (1.5% of total sequences), Archaea (0.01% of total sequences), terrestrial plants (0.4% of total sequences) and non-affiliated OTUs (0.09% of total sequences), also singletons (0.003% of total sequences) were removed from all the data set for further analysis (Duret et al. 2015). Raw reads were deposited into the NCBI Sequence Read Archive (SRA) database with accession number: SRR3635293 and SRX1825581.
2.4 Sea surface temperature (SST)
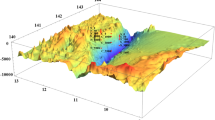
SST was used as an indicator of the presence of the YSWC (Fig. 1). Monthly rather than daily maps SST maps were drawn [using Matlab (R2010b)] as daily temperature data at some locations was not available. Additionally, SST for June 2011 was shown for comparison. Moderate Resolution Imaging Spectroradiometer (MODIS) satellite SST data was downloaded from the website: http://oceancolor.gsfc.nasa.gov/cms/#. SSTs of China’s marginal seas are shown, especially the Yellow Sea, to illustrate that the sampling site was indeed influenced by the YSWC. SSTs of adjacent months showed the same pattern and trend (data not shown).
Sampling site and seasonal SST maps. Samples were collected from a site (34°0′N, 124°0′E) at the central Yellow Sea. SST maps showed the sea surface temperature of the site and surrounding area with an additional map for June 2011 (a) (without sampling). Samples b and c are not influenced by the YSWC while d–f are YSWC samples
2.5 Statistical and phylogenetic analyses
R software (version 3.2.2, http://cran.r-project.org) was employed to show the relative abundances of OTUs. Diversity and richness indices were computed with the “Vegan” package (Dixon 2003). The Venn diagram was drawn in the “VennDiagram” package, after calculating overlapped OTUs among the five samples. OTUs at a given site were divided into three groups by relative abundance: abundant OTUs (>1%), rare OTUs (<0.01%), and intermediate OTUs (0.01–1%) (Logares et al. 2014). A heat map of locally abundant OTUs was generated by the “pheatmap” package with corresponding taxonomy to each OTU (Amaral-Zettler 2013). Hierarchical clustering, similarity profile (SIMPROF) permutation tests and non-metric multidimensional scaling (nMDS) were generated by PRIMER v6.1 (Clarke and Gorley 2006). Differences among groups which were discriminated by nMDS were tested by using permutational multivariate analysis of variance (PERMANOVA) (Anderson et al. 2008) test. Similarity percentage (SIMPER) (Merriam 1978) analysis defines OTUs that contributed to separate each two groups and was conducted between YSWC group (W11, W12, and W13) and N-YSWC group (S12 and S13). A final indicator OTUs analysis (IOA) helped to identify representative species of the YSWC, by using an “indicator value” (IV) (Dufrêne and Legendre 1997).
3 Results
3.1 Alpha diversity, super and finer groups
A total of 381 phylogenetically different OTUs with a mean length of 350 bp were recovered from the five samples (97% similarity). The numbers of paired-end sequences in each library varied from 20899 to 41125. Among the two N-YSWC samples (S12 and S13) and three YSWC samples (W11, W12, and W13), S12 yielded the fewest number of OTUs (116) while W11 had the most (248) (Table S1).
Eight super groups (Alveolata, Stramenopiles, Archaeplastida, Hacrobia, Opisthokonta, Rhizaria, Apusozoa, and Amoebozoa) and 29 finer groups at the class or phylum level (e.g. Dinoflagellata, Ciliophora, MASTs, Dictyochophyceae, Prasinophyceae, Fungi, Prymnesiophyceae, Cercozoa) were detected in the samples. Alveolata was the best represented OTU super group across the five samples, comprising about 60% of the total richness. The next most abundant was the Stramenopiles (18.67%).
3.2 Beta diversity and community composition
Percentages of relative abundance of OTUs are shown in Fig. 2. At the highest taxonomic level (Fig. 2a), the three super groups Alveolata, Stramenopiles, and Archaeplastida, were consistently abundant in all five samples. OTUs within an affiliation to Archaeplastida overwhelmingly dominated the N-YSWC community, with an average combined abundance of 71.55%, but their proportions were much lower in the YSWC samples (26.6%, average). Across the five samples Alveolates had relative abundances between 18.49 and 72.93% (34.86%, average) and were the highest in W12. In the three YSWC samples, Alveolates and Stramenopiles dominated the OTUs in W12 and W13, respectively. Archaeplastida accounted for nearly half of the total abundance of OTUs in W11, which had a more similar community composition to the N-YSWC samples. OTUs belonging to Rhizatia, Opisthokonta, and Hacrobia were present, but had low relative abundances in all five samples (0.44, 0.27, and 3.09%, respectively). Additionally, one OTU affiliated with Amoebozoa and two OTUs affiliated with Apusozoa were recovered, but not shown in the histograms due to the very few numbers of reads. At a finer taxonomic level (Fig. 2b), the class Prasinophyceae was the only dominant Archaeplastida in all samples. Marine alveolates group MALV-II, including parasitic species Amoebophrya spp., were detected with different relative abundances across all samples, varying from 0.45 to 29.45%. A large number of OTUs belonging to Ciliophora (Alveolata) were also recovered (19.93%, average), particularly in W12. Dictyochophyceae appeared to be the main Stramenopiles, comprising 56.17% in W13. “Others” referred to finer groups with relatively low abundance such as Apicomplexa (Alveolata), MASTs (Stramenopiles), Picozoa (Hacrobia), Fungi (Opisthokonta), and Cercozoa (Rhizaria).
Hierarchical clustering based on SIMPROF grouped the two N-YSWC samples S12 and S13 and separated the three YSWC samples (W11, W12, W13) at similarity level of 50% (p < 0.05, PERMANOVA test) (Fig. 3). The clustering results are also displayed in the nMDS plot where N-YSWC samples shared the nearest distance of community structure, with a similarity value of 58.51% (Fig. 3b). YSWC samples had lower similarities, not only among each other (average similarity = 15.53%), but also with the group of N-YSWC samples (average similarity = 23.18%). Only W11 had a closer distance with N-YSWC samples (similarity = 46.52%) and this is also reflected in the composition and relative abundances of OTUs (Fig. 2b). The Venn diagram shows the numbers of shared and sample-specific OTUs of the five samples (Fig. 4). All samples had sample-specific OTUs. The numbers of shared OTUs were much higher among YSWC samples of different years, and 51 OTUs were found in all samples.
3.3 Most abundant OTUs
Locally abundant OTUs (relative abundances >1% at a given site) (Mangot et al. 2013) are highlighted in the heatmap in the order of group-shared to sample-specific with corresponding taxonomies on the right (Fig. 5). OTU22, with a BLAST result of Micromonas pusilla (100% identity), was detected as the only widely spread OTU getting a relative abundance over 1% across the five samples (2.9%, average). For N-YSWC samples OTU4, belonging to Bathycoccus prasinos (100% identity), and OTU1, belonging to Ostreococcus lucimarinus (100%, identity), were the main group-shared species with much higher relative abundances than other OTUs. For the YSWC samples, OTU1 (O. lucimarinus) dominated in W11. OTU7 with 100% identity to M. pusilla and OTU6 with 92% identity to an uncultured clone within Ciliophora, dominated in W12. OTU3, with a 99% identity to Pseudochattonella farcimen (Ochrophyta, Dictyochophyceae), was the most abundant species in W13. From the similarity percentage (SIMPER) analysis (Tables 1, 2, 3), species within the genus Mamiellales (Chlorophyta, prasinophyceae) dominated the representative OTUs of N-YSWC group within which OTU1 contributed the most to the grouping of S12 and S13 (78.93%, contribution) (Table 1). While three OTUs (OTU4, OTU7, and OTU22) within the order Mamiellales (Chlorophyta, Prasinophyceae) contributed substantially to the shared OTUs from YSWC group (W11, W12, and W13) (average contribution 15.48%), OTU11, OTU31 within Syndiniales Group II (Protalveolata; Syndiniales), OTU27, OTU177 within Prymnesiales (Haptophyta, Prymnesiophyceae) and OTU39 with affiliation to Triparma strigata (Ochrophyta, Bolidophyceae) appeared to have a significant contribution value (Table 2). OTUs that separated the two groups (N-YSWC and YSWC), with contribution values over 1%, were listed, together with their taxonomies (Table 3). OTU1 made the greatest contribution (31.62%) to the separation of the two groups, followed by OTU3 (9.84%, contribution) and OTU7 (4.68%, contribution). IOA (indicator OTUs analysis) performed on these SIMPER-listed OTUs showed three OTUs belonging to Pseudochattonella farcimen (Ochrophyta, Dictyochophyceae), Florenciella parvula (Dictyochophyceae) and Phaeocystis cordata (Haptophyta, Prymnesiophyceae) had the highest IV values and could most likely be the indicator picoeukaryotes of the YSWC (Table 4).
Heatmap of OTUs occurring at a frequency of over 1% at a given sample. N-YSWC samples are the two on the left and three YSWC samples on the right, OTUs that occur at both N-YSWC and YSWC samples are listed at the top of the table while OTUs occurring at only a N-YSWC or YSWC sample are arranged below. Hot colors indicate higher percentages while cooler colors indicate lower percentages. Corresponding taxonomy of each OTU is on the right as a result of BLAST with over 97% similarity
4 Discussion
The influences of the YSWC were depicted in the SST maps comparatively for YSWC samples (W11, W12, and W13). Although one N-YSWC sample from 2011 was missing, the two N-YSWC samples (S12 and S13) here demonstrate a “background” water mass with a distinct pattern of picoeukaryotic biodiversity and community structure. Methodologically, since the range of rDNA copies for picoeukaryotes is quite small and similar to prokaryotes, 18S rDNA is appropriate as a target gene to estimate picoeukaryotic biodiversity and the relative abundances of each taxon rather than larger-sized eukaryotes (Zhu et al. 2005). Primer pairs targeting the V4 region allow discrimination with paired-end reads as short as 150 bp (with 75% accuracy at genus level), as the V4 region is identified as the most informative part of the 18S rDNA (Hugerth et al. 2014). This study provided the first insight into the picoeukaryotic biodiversity at the species and community level in the central Yellow Sea by molecular ways.
4.1 Community structure and abundant groups
Different dynamics of picoeukaryotic community structure have been studied across monthly and seasonal timescales (Countway et al. 2010; Christaki et al. 2014; Kulk et al. 2012). In the present study, the two N-YSWC samples shared a higher similarity although with big sampling temperature difference; they also had a longer distance from the YSWC samples in the nMDS (non-metric multidimensional scaling) plot. This implies that the exogenous water from the YSWC would contain a community that is distinct from the local ones. For YSWC samples W12 and W13, although they were both sampled in November at similar temperature, their community compositions were quite different. This suggests that the influences of the YSWC on local waters varies inter-annually.
Ciliates (Ciliophora) are an important group of heterotrophic protists that graze on phytoplankton and have a considerable biomass (Pierce and Turner 2009). Dinoflagellates, with Dinophyceae as a representative class, are a well-studied group, many of which are mixtrophs. They are also one of the main causative organisms of harmful algae blooms (HABs) (Stoecker 1999). These two groups contributed about half of the sequences in W12 and the ciliates were also well distributed among the five samples. Considering that the minimum cell size of these two groups is over 5 µm, the existence of these is probably due to either cell lysis during the filtration or to extracellular dissolved DNA which are most not from marine picoeukaryotes (Massana 2011). These two groups will not be discussed further.
Three commonly reported super groups, Archaeplastida, Alveolata, and Stramenopiles, dominated the samples in this study. However, they had an uneven distribution among the five samples, probably indicating community changes over the long sampling time interval (Countway et al. 2005; Christaki et al. 2014) and the impact of the exogenous water brought in by the YSWC (Liu et al. 2015). As a green algae group within Archaeplastida, prasinophytes (including three “flagship” genera Ostreococcus, Micromonas, and Bathycoccus) dominated the biomass in the N-YSWC samples. Prasinophytes have many photosynthetic lineages and make an important contribution to primary production with their relatively high growth rate (Not et al. 2004). They have been frequently reported in temperate and polar coastal samples (Not et al. 2008; Kilias et al. 2014) and show adaptations to low nutrient conditions (Vaulot et al. 2008). The dominant distribution of prasinophytes in the N-YSWC samples is consistent with a recent study focusing on the phytoplankton community in the central Yellow Sea (Liu et al. 2015), though PCR bias toward or against some groups may exist (Potvin and Lovejoy 2009).
Unlike the N-YSWC samples which were dominated by only one photosynthetic class, YSWC samples displayed abundant groups with multi-trophic strategies. Marine alveolates MALV-I and MALV-II were firstly reported as the most diverse and abundant novel groups in the aphotic zone of the Antarctic polar front (Lόpez-Garcίa et al. 2001). More new lineages, together with five clades (MALV-I, II, III, IV, V), are parasites of plankton, and they have also been detected especially in surface sea waters (Guillou et al. 2008). The two groups, MALV-I and MALV-II, were reported comprising 66.5% (25.31 and 41.19%, respectively) of the total clones along a section of the South China Sea (Wu et al. 2014), but abundances of these groups was much lower in the present samples (average 0.5 and 10.14%, respectively). MALV-II had more diverse sub-groups than MALV-I (Guillou et al. 2008). In the study by Wu et al. (2014), substantial number of MALV-II was supposed to be related to parasitism with local corals and temperature variation in other extreme environments. In the present study, the lower percentage of these two group, especially MALV-II, could be due to the higher temperature and lower abundance of their potential host Dinophyceae (2.6%, average) (Park et al. 2004). MASTs are ecologically important and branch at the base of Stramenopiles, including a large number of heterotrophic and autotrophic groups (Logares et al. 2012). Among them, MAST-3 is one of the most wide spread and abundant and MAST-4 shows a relatively low genetic diversity with only five major lineages (Rodríguez-Martínez et al. 2012). In the present samples, MAST-3 and MAST-4 were only found in W11 as abundant groups, and only five OTUs were found within MAST-4. However, the two groups were also detected in other samples although at low abundances, suggesting that they are wide spread across a broad temperature range (Massana et al. 2004; Rodríguez-Martínez et al. 2013).
OTUs belonging to the class Dictyochophyceae (Stramenopiles) dominated in W13, comprising almost 50%. Dictyochophyceae together with Pelagophyceae and Prymnesiophyceae have been reported as the dominant photosynthetic groups in oligotrophic regions such as the South Pacific Ocean (Shi et al. 2011). The genus Pseudochattonella with an affiliation to Dictyochophyceae, contains some ichthyotoxic algae such as Pseudochattonella farcimen and P. verruculosa, which can form HABs (Skjelbred et al. 2013). The three most abundant Dictyochophyceae species in W13 were P. farcimen, Florenciella parvula, and P. verruculosa. Including the Prymnesiophyceae, the community structure of W13 was similar to sites in oligotrophic regions such as the South China Sea (Wu et al. 2014). This observation is consistent with both the YSWC and the South China Sea being nutrient-poor (Liu et al. 2015). Generally, the increase of other photosynthetic groups (e.g. Prymnsiphytes), grazers (e.g. ciliates), and parasites (e.g. Amoebophyra spp.) in the YSWC samples demonstrates the complexity of the microbial systems in the marginal seas of China (Liu et al. 2015), with potentially temporal and inter-annual changes (Lin et al. 2005).
Most natural biological communities are characterized by having a few abundant taxa and many rare taxa (Logares et al. 2014). Identifying these elements is important to understand community dynamics and system functioning for both bacteria and eukaryotes. Locally abundant OTUs in this study have similar abundances in each sample and show a stable pattern of diversity (from 4.1 to 5.9%). Also, 51 OTUs were found in all samples here, most of which were locally rare species with affiliations to all the super groups. This number is more than twice the numbers of sample-specific OTUs, indicating the presence of wide spread, but rare picoeukaryotes (Logares et al. 2014). As the aim of this investigation was to find distinct OTUs that could best characterize two different environments (i.e. YSWC and central Yellow Sea) and not to discover novel species, only abundant taxa were analyzed here, and rare OTUs received less attention. However, many of the rare protistan taxa can become dominant under suitable environmental conditions (Caron and Countway 2009). If the rare picoeukaryotic taxa follow a similar pattern in the complex central Yellow Sea area, then the increase in abundance of some groups in a particular sample, e.g. Dictyochophyceae and Prymnesiophyceae in W13, are probably due to the mixing of locally cold water with warm water brought in by the YSWC.
4.2 Temporal differences represented with indicating species
The temporal succession of picoeukaryotes, which are persistent and abundant members of the microbial community, shows that these organisms can exploit a variety of life strategies (such as saprotrophy and parasitism) (Christaki et al. 2014). Among these locally abundant species, some are described as indicator species that can define specific habitats. For instance, diatom genus Pinnularia and the ciliate genus Frontonia are good indicators of acidic and alkaline conditions, respectively (Amaral-Zettler 2013). Deep insight into these indicator groups on the species level is important. Similarity percentage (SIMPER) analysis is often used to identify the OTUs that contribute the most to the separation of two groups, as well as highlight shared OTUs within a certain group. In the present study, one of the aims was to identify indicator picoeukaryotes in the YSWC. We paid more attention to the highlighted OTUs shared by YSWC groups, especially distinctive ones compared with N-YSWC group, which could be a support of the later IOA result.
For SIMPER analysis, OTU1, with affiliation to O. lucimarinus, made the highest contribution to the shared OTUs of N-YSWC samples (Table 1), also to the separation of the two groups (Table 3). O. lucimarinus, which was the most abundant species in samples S12 and S13 here, is one of the smallest free-living eukaryotes with a cell size <1 µm, with genetic adaptation to surface ocean conditions (Palenik et al. 2007). Together with Ostreococcus tauri, its genome, metabolic mechanism, and morphology have been well characterized (Lanier et al. 2008). O. lucimarinus has been observed year-round in the coastal North Pacific Ocean (Worden 2006). Within the same genus, O. tauri was firstly reported in a long-term study conducted in a French lagoon and it was found in blooms associated with a “brown tide” caused by Aureococcus anophagefferens (Stramenopiles, Pelagophyceae) (O’Kelly et al. 2003). Notably, in a previous genomic study (Palenik et al. 2007), O. lucimarinus was reported to have 19 of 20 candidate selenocysteine-encoding genes, which are vital to their adaptations to local environment, shared with O. tauri. Spring blooms, especially “brown tides” have occurred in coastal area and subsequently influence the central Yellow Sea, lasting for up to 2 months from April to May (Tang et al. 2013). Since the N-YSWC samples reported herein were collected in May and June, the dominant presence of O. lucimarinus could also be associated with a “brown tide”, probably a similar burst as O. tauri. As a result, in Table 3, with relatively much lower average percentage of abundance in YSWC group (0.08%, compared with N-YSWC), OTU1 ranked first in the separation of the two groups, suggested its potential preference for local or coastal water, rather than exogenous YSWC water transferred from pelagic ocean.
OTU3, the most abundant taxon in W13, was identified as Pseudochattonella farcimen, taking up 9.84% of the contribution to the separation of two groups (Table 3). It shared a 99% similarity with P. farcimen strain UIO 110 from the study by Edvardsen et al. (2007) where it was given the name Verrucophora farcimen (initially named Chattonella aff. Verruculosa). In that study, cultures of P. farcimen had a maximal growth rate at temperatures between 5 and 10 °C and did not tolerate temperatures >15 °C. However, in the Yellow Sea sample, W13, with a sampling temperature >17 °C, P. farcimen was the dominant picoeukaryote. It is possible that this difference may reflect differences in temperature adaptation between naturally growing picoeukaryotes and those grown in culture. P. farcimen, with an affiliation to Dictyochophyceae, is a well-studied species associated with harmful algae blooms, which cause fish mortality in European coastal areas (Dittami et al. 2012). Since algal blooms of this species have not previously been reported in the marginal seas of China, it remained unknown why Pseudochattonella farcimen was relatively the most abundant in warmer waters (for W13) of the central Yellow Sea in early winter here.
As additional representative species within the order Mamiellales (Chlorophyta, Prasinophyceae), OTU7 (M. pusilla) and OTU4 (B. prasinos) helped to separate the YSWC group from the N-YSWC group with contributions of 4.68 and 2.57%, respectively (Table 3). Micromonas spp., as the second most abundant prasinophytes in the samples, was represented by a single species (M. pusilla), which can be the dominant photosynthetic picoeukaryote in some marine ecosystems (Kilias et al. 2014; Lovejoy et al. 2007). M. pusilla, which has genetic thermal adaptions, is distributed widely in both cold and warm waters (Worden et al. 2009). In some studies, however, it has been reported as a cold-adapted ecotype (Lovejoy et al. 2007). In the present study, the only OTU that was abundant in all samples was OTU22, with 100% identification as M. pusilla strain RCC570 (Roscoff Culture Collection). Its highest relative abundance in S12 (with the lowest sampling temperature), compared with other samples, suggests that it has a growth preference for cold water (Kilias et al. 2014). Only one strain (OTU4) of B. prasinos (RCC801) was found in the samples. The Bathycoccus genus is thought to contain only a single species, B. prasinos (known as RCC1105) with a sequenced genome proving its basic position of the green lineage (Moreau et al. 2012). B. prasinos was present at all stations in a recent study conducted in the eastern North Pacific Ocean (Melinda et al. 2016) while little was known about its distributions in the Yellow Sea. The substantial distributions of M. pusilla and B. prasinos in our samples reported here confirm that prasinophytes have a widespread temporal and spatial distribution in the central Yellow Sea and the YSWC.
It is worthwhile to mention that OTU11 and OTU18, which were uncultured species within MALV-II, contributed to the separation with 1.68 and 1.1%, respectively (Table 3), while representative OTUs of the N-YSWC group were all within Prasinophyceae at the contribution cut-off value of 1% (Table 1). The two taxa (OTU11 and OTU18) both had 100% identification with clone 18S ribosomal RNA genes in samples from the subtropical-tropical South China Sea (Wu et al. 2014). Amoebophrya spp., belonging to MALV-II, are host-specific parasites that infect dinoflagellates and play a key role in controlling dinoflagellates blooms. They were reported as extremely diversiform and host specific parasites infecting various hosts including dinoflagellate, ciliate, cercozoans, and larger sized planktons. Thus, most marine platonic groups are potentially affected by these parasites, similar to viruses that control bacterial populations (Chambouvet et al. 2008). However, Amoebophrya spp. in this study were uncommon with only one taxon (OTU30) abundant in W11. OTU11 and OTU18 shared lower identifications with Amoebophrya sp. ex Gymnodinium instriatum from BLAST results (91% and 89%, respectively). This may reflect the broad genetic diversity of species within MALV-II and the potential existence of uncultured species that are better adapted to environment of YSWC than Amoebophrya spp.
Indicator OTUs analysis (IOA) helps to identify indicator OTUs for site groups with an “Indicator Value (IV)”, which is the product of the relative abundance and the constancy, ranging from 0 to 100. This method is a new way to identify indicator species for typologies of samples obtained by any hierarchical or non-hierarchical classification procedure (Dufrêne and Legendre 1997). It is a convenient way to find OTUs (or species) that can best define particular environments (Amaral-Zettler 2013). Because of limited available sampling sites during the cruises, IOA here was conducted to assess differences due to time and location, i.e. waters with and without the influence of the YSWC. In this study, IV was calculated between the N-YSWC group (S12, S13) and the YSWC group (W11, W12, W13), in order to strengthen the widely spread OTUs of the YSWC samples and to weaken the effect of abundant OTUs that were only present in a single YSWC sample. Not considering the ciliates and dinoflagellates, OTU3 (Pseudochattonella farcimen), OTU10 (Florenciella parvula), and OTU25 (Phaeocystis cordata) had IV values over 95 (p < 0.05, Monte Carlo test), representing indicator species of YSWC samples.
As discussed above, P. farcimen is an algal species that can form blooms and kill fish. Florenciella parvula (Dictyochales, Dictyochophyceae) was firstly isolated from the English Channel and is an important member of the picoplankton in polar and temperate systems (Etkrem et al. 2004). Though several Dictyochophyceae species can form HABs (Dittami et al. 2013), they have not received much attention in the Yellow Sea and neighboring marginal seas of China. Phaeocystis cordata and Chrysochromulina acantha (both belonging to the class Prymnesiophyceae) have been reported as occurring year round in North Atlantic coastal waters (Egge et al. 2015). They were detected in all the N-YSWC and YSWC samples reported herein and were abundant in W13, but rare in other samples. P. cordata, which is closely related to the colonial species P. globose, is limited by temperature during N-YSWC period (Zingone et al. 1999). The P. cordata strain (OTU25) detected here had a 91% identity with P. globosa as a result of alignment. This study has identified an increased number of indicator species and determined their growth rate at different temperatures in order to demonstrate their presence in the YSWC.
5 Conclusions
In summary, the present study revealed patterns of picoeukaryotic community structures in the central Yellow Sea. Differences between N-YSWC and YSWC samples and also within the YSWC samples showed that the YSWC plays an important role in the ecosystems of the Yellow Sea. With massive high throughput sequencing tags of the 18S rDNA V4 region, differences in the distribution of abundant and rare groups and between distinct species in the YSWC and background samples were investigated. Prasinophytes dominated the N-YSWC samples. Indicator species of the YSWC, such as Pseudochattonella farcimen, are ecologically important members of the ecosystem and also have potentially negative economic consequences through the development of HABs. Our results show that the central Yellow Sea picoeukaryotic community can be influenced by the YSWC and in response to temporal and inter-annual variability.
References
Acosta F, Ngugi DK, Stingl U (2013) Diversity of picoeukaryotes at an oligotrophic site off the Northeastern Red Sea Coast. Aquat Biosyst 9(1):16
Amaral-Zettler LA (2013) Eukaryotic diversity at pH extremes. Front Microbiol 3(2):441
Amato KR, Yeoman CJ, Kent A et al (2013) Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. ISME J 7(7):1344–1353
Anderson MJ, Gorley RN, Clarke KR (2008) PERMANOVA+ for PRIMER Guide to software and statistical methods. PRIMER-E Ltd., Plypouth
Caron DA, Countway PD (2009) Hypotheses on the role of the protistan rare biosphere in a changing world. Aquat Microb Ecol 57:227–238
Chambouvet A, Morin P, Marie D, Guillou L (2008) Control of toxic marine dinoflagellate blooms by serial parasitic killers. Science 322:1254–1256
Christaki U, Kormas KA, Genitsaris S et al (2014) Winter–summer succession of unicellular eukaryotes in a meso-eutrophic coastal system. Microb Ecol 67:13–23
Clarke KR, Gorley RH (2006) PRIMER-6 user manual/tutorial. PRIMER-E Ltd., Plymouth
Countway PD, Gast RJ, Saval P et al (2005) Protistan diversity estimates based on 18S rDNA from seawater incubations in the western North Atlantic. J Eukaryot Microbiol 52:95–106
Countway PD, Vigil PD, Schnetzer A, Moorthi SD, Carona DA (2010) Seasonal analysis of protistan community structure and diversity at the USC Microbial Observatory (San Pedro Channel, North Pacific Ocean). Limnol Oceanogr 55:2381–2396
Cui MC, Hu DX, Mo J (2004) Seasonality and causes of the Yellow Sea Warm Current. Chin J Oceanol Limnol 22:265–270
de Vargas C, Audic S, Henry N, Decelle J et al (2015) Eukaryotic plankton diversity in the sunlit ocean. Science 348:29223618–29223625
Dittami SM, Riisberg I, John U (2012) Analysis of expressed sequence tags from the marine microalga Pseudochattonella farcimen (Dictyochophyceae). Protist 163:143–161
Dittami SM, Hostyeva V, Egge ES et al (2013) Seasonal dynamics of harmful algae in outer Oslofjorden monitored by microarray, qPCR, and microscopy. Environ Sci Pollut Res 20:6719–6732
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14(6):927–930
Dufrêne M, Legendre P (1997) Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol Monogr 67:345–366
Duret MT, Pachiadaki MG, Stewart FJ et al (2015) Size-fractionated diversity of eukaryotic microbial communities in the Eastern Tropical North Pacific oxygen minimum zone. FEMS Microbiol Ecol 91:1–42
Edgcomb V, Orsi W, Bunge J et al (2011) Protistan microbial observatory in the Cariaco Basin, Caribbean. I. Pyrosequencing vs Sanger insights into species richness. ISME J 5:1344–1356
Edvardsen B, Eikrem W, Shalchian-Tabrizi K, Riisberg I et al (2007) Verrucophora farcimen gen. et sp. nov. (Dictyochophyceae, Heterokonta)—a bloom-forming ichthyotoxic flagellate from the Skagerrak. Nor J Phycol 43:1054–1070
Edwards M, Richardson AJ (2004) Impact of climate change on marine pelagic phenology and trophic mismatch. Nature 430:881–884
Egge ES, Johannessen TV, Andersen T et al (2015) Seasonal diversity and dynamics of haptophytes in the Skagerrak, Norway, explored by high-throughput sequencing. Mol Ecol 24:3026–3042
Etkrem W, Romari K, Latasa M et al (2004) Florenciella parvula gen. et sp. nov. (Dictyochophyceae, Heterokontophyta), a small flagellate isolated from the English Channel. Phycologia 43:658–668
Guillou L, Viprey M, Chambouvet A, Welsh RM, Kirkham AR, Massana R, Scanlan DJ, Worden AZ (2008) Widespread occurrence and genetic diversity of marine parasitoids belonging to Syndiniales (Alveolata). Environ Microbiol 10:3349–3365
Hillis DM, Dixon MT (1991) Ribosomal DNA: molecular evolution and phylogenetic inference. Q Rev Biol 66:441–453
Hu HG, Wan ZW, Yuan YI (2004) Simulation of seasonal variation of phytoplankton in the southern Huanghai (Yellow) Sea and analysis on its influential factors. Acta Oceanol Sin 26:74–88
Hugerth LW, Muller EE, Hu YO et al (2014) Systematic design of 18S rRNA gene primers for determining eukaryotic diversity in microbial consortia. PLoS One 9:e95567
Kilias ES, Nothig E, Wolf C, Metfies K (2014) Picoeukaryote plankton composition off West Spitsbergen at the entrance to the Arctic Ocean. J Eukaryot Microbiol 61(6):569–579
Kulk G, van de Poll W, Buma AG (2012) Temperature-dependent growth and photophysiology of prokaryotic and eukaryotic oceanic picophytoplankton. Mar Ecol Prog Ser 466:43–55
Lanier W, Moustafa A, Bhattacharya D, Comeron JM (2008) EST analysis of Ostreococcus lucimarinus, the most compact eukaryotic genome, shows an excess of introns in highly expressed genes. PLoS One 3:e2171
Li WKW (1994) Primary production of prochlorophytes, cyanobacteria, and eucaryotic ultraphytoplankton: measurements from flow cytometric sorting. Limnol Oceanogr 39:169–175
Lin C, Ning XR, Su JL, Lin Y, Xu B (2005) Environmental changes and the responses of the ecosystems of the Yellow Sea during 1976–2000. J Marine Syst 55:223–234
Liu X, Huang BQ, Huang Q, Wang L (2015) Seasonal phytoplankton response to physical processes in the southern Yellow Sea. J Sea Res 95:45–55
Logares R, Audic R, Santini S (2012) Diversity patterns and activity of uncultured marine heterotrophic flagellates unveiled with pyrosequencing. ISME J 6:1823–1833
Logares R, Audic S, Bass D et al (2014) Patterns of rare and abundant marine microbial eukaryotes. Curr Biol 24:1–9
Lovejoy C, Vincent WF, Bonilla S et al (2007) Distribution, phylogeny and growth of cold-adapted picoprasinophytes in arctic seas. J Phycol 43:78–89
Lü LG, Wang X, Wang H et al (2013) The variations of zooplankton biomass and their migration associated with the Yellow Sea Warm Current. Cont Shelf Res 64:10–19
Lόpez-Garcίa P, Rodríguez-Valera F, Pedrós C, Moreira D (2001) Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409:603–607
Mangot JF, Domaizon I, Taib N et al (2013) Short-term dynamics of diversity patterns: evidence of continual reassembly within lacustrine small eukaryotes. Environ Microbiol 15:1745
Massana R (2011) Eukaryotic picoplankton in surface oceans. Annu Rev Microbiol 65:91–110
Massana R, Castresana J, Balagué V et al (2004) Phylogenetic and ecological analysis of novel marine stramenopiles. Appl Environ Microb 70:3528–3534
Medlin LK, Metfies K, Mehl H, Wiltshire K, Valentin K (2006) Picoeukaryotic plankton diversity at the Helgoland time series site as assessed by three molecular methods. Microb Ecol 52:53–71
Melinda PS, Sebastian S, Adam M et al (2016) Abundance and biogeography of picoprasinophyte ecotypes and other phytoplankton in the Eastern North Pacific Ocean. Appl Environ Microbiol 82(6):1693–1705
Merriam DF (1978) Computational methods of multivariate analysis in physical geography. Earth Sci Rev 14(2):173–174
Moon-van der Staay SY, de Wachter R, Vaulot D (2001) Oceanic 18S rDNA sequences from picoplankton reveal unsuspected eukaryotic diversity. Nature 409:607–610
Moreau H, Verhelst B, Couloux A et al (2012) Gene functionalities and genome structure in Bathycoccus prasinos reflect cellular specializations at the base of the green lineage. Genome Biol 13:R74
Not F, Latasa M, Marie D, Cariou T, Vaulot D, Simon N (2004) A single species, Micromonas pusilla (Prasinophyceae), dominates the eukaryotic picoplankton in the Western English Channel. Appl Environ Microb 70:4064–4072
Not F, Latasa M, Scharek R et al (2008) Protistan assemblages across the Indian Ocean, with a specific emphasis on the picoeukaryotes. Deep Sea Res Pt I 55:1456–1473
O’Kelly CJ, Sieracki ME, Thier EC, Hobson IC (2003) A transient bloom of Ostreococcus (Chlorophyta, Prasinophyceae) in West Neck Bay, Long Island, New York. J Phycol 39:850–854
Palenik B, Grimwoodc J, Aerts A et al (2007) The tiny eukaryote Ostreococcus provides genomic insights into the paradox of plankton speciation. Proc Natl Acad Sci USA 104:7705–7710
Park MG, Yih W, Coats DW (2004) Parasites and phytoplankton, with special emphasis on dinoflagellate infections. J Eukaryot Microbiol 51:145–155
Pierce RW, Turner JT (2009) Ecology of planktonic ciliates in marine food webs. Rev Aquat Sci 6:139–181
Pomeroy LR, Williams PJI, Azam F, Hobbie JE (2007) The microbial loop. Oceanography 20:28–33
Potvin M, Lovejoy C (2009) PCR-Based diversity estimates of artificial and environmental 18S rRNA gene libraries. J Eukaryot Microbiol 56(2):174–181
Rocke E, Jing H, Liu H (2013) Phylogenetic composition and distribution of picoeukaryotes in the hypoxic northwestern coast of the Gulf of Mexico. MicrobiolOpen 2:130–143
Rodríguez-Martínez R, Rocap G, Logares R, Romac S, Massana R (2012) Low evolutionary diversification in a widespread and abundant uncultured protist (MAST-4). Mol Biol Evol 29:1393–1406
Rodríguez-Martínez R, Rocap G, Salarza G, Massana R (2013) Biogeography of the uncultured marine picoeukaryote MAST-4: temperature-driven distribution patterns. ISME J 7:1531–1541
Shi XL, Lepère C, Scanlan DJ, Vaulot D (2011) Plastid 16S rRNA gene diversity among eukaryotic picophytoplankton sorted by flow cytometry from the South Pacific Ocean. PLoS One 6(4):e18979
Skjelbred B, Edvardsen B, Andersen T (2013) Environmental optima for seven strains of Pseudochattonella (Dictyochophyceae, Heterokonta). J Phycol 49:54–60
Song DX, Bao XW, Wang XH et al (2009) The inter-annual variability of the Yellow Sea Warm Current surface axis and its influencing factors. Chinese J Oceanol Limnol 27(3):607–613
Stoeck T, Epstein S (2003) Novel eukaryotic lineages inferred from small sub-unit rRNA analyses of oxygen-depleted marine environments. Appl Environ Microbiol 69:2657–2663
Stoecker DK (1999) Mixotrophy among Dinoflagellates. J Eukaryn Microbiol 46:397–401
Tang QS, Su JL, Zhang J (2013) Spring blooms and the ecosystem processes: the case study of the Yellow Sea. Deep Sea Res Pt II 97:1–3
Vaulot D, Eikrem W, Viprey M, Moreau H (2008) The diversity of small eukaryotic phytoplankton (≤3 μm) in marine ecosystems. FEMS Microbiol Rev 32:795–820
Worden A (2006) Picoeukaryote diversity in coastal waters of the Pacific Ocean. Aquat Microb Ecol 43:165–175
Worden AZ, Nolan JK, Palenik B (2004) Assessing the dynamics and ecology of marine picophytoplankton: the importance of the eukaryotic component. Limnol Oceanogr 49:168–179
Worden AZ, Lee JH, Mock T et al (2009) Green evolution and dynamic adaptations revealed by genomes of the marine picoeukaryotes Micromonas. Science 423:268–272
Wu W, Huang B, Liao Y, Sun P (2014) Picoeukaryotic diversity and distribution in the subtropical-tropical South China Sea. FEMS Microbiol Ecol 89:563–579
Zhu F, Massana R, Not F et al (2005) Mapping of picoeucaryotes in marine ecosystems with quantitative PCR of the 18S rRNA gene. FEMS Microbiol Ecol 52:79–92
Zingone A, Chrétiennot-Dinet MJ, Lange M, Medlin L (1999) Morphological and genetic characterization of Phaeocystis Cordata and P. Jahnii (Prymnesiophyceae), two new species from the Mediterrranean Sea. J Phycol 35:1322–1337
Acknowledgements
This study was supported by the National Natural Foundation Science of China (NFSC) (Grants 31500339, 41676178, 41076088); Fundamental Research Funds for the Central University of Ocean University of China (Grant Numbers 201512013, 201564010 and 201512008). We are grateful to the captain and crews of the RV ‘Dong Fang Hong 2’. The authors would like to thank the editor and anonymous reviewers for their constructive comments.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, Z., Song, X., Wang, M. et al. Community patterns and temporal variation of picoeukaryotes in response to changes in the Yellow Sea Warm Current. J Oceanogr 73, 687–699 (2017). https://doi.org/10.1007/s10872-017-0425-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10872-017-0425-1