Abstract
Understanding the factors that influence the diversity and composition of arthropod communities is a major topic in ecology. Canopy arthropod communities are a major constituent of biodiversity and show great variation in time and space according to different factors. Recently, genetic variation within tree species has attracted attention as a significant factor determining the diversity and composition of canopy arthropod communities. A major source of genetic and phenotypic novelty in plant species is interspecific hybridization, and therefore it is of interest to evaluate how this process affects the communities of associated organisms. In this study, we used microsatellite markers and geometric morphometry of leaf shape to analyze genetic and morphological variation in 45 individuals in a local hybrid zone between the oaks Quercus affinis and Q. laurina in Mexico. Individual trees were assigned to one of the parental species or to the hybrid category. The percentage of leaf area removed by herbivores was quantified in each individual and the canopies of five individuals of each categeory (two parental species and hybrids) was fogged with insecticide to assess the diversity and composition of arthropod communities. Results indicated that hybrid trees experience higher levels of herbivory than parental species and also sustain a higher abundance and richness of canopy arthropods. In general, our study supports the “hybrid susceptibility hypothesis” that predicts a higher incidence of associated arthropods on hybrid plants than in their parental species as result of the disruption of co-adapted gene complexes associated to resistance traits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tree canopies harbor great biodiversity that includes remarkably rich arthropod assemblages with several ecological functions, such as predators, pollinators, herbivores, parasites and scavengers that determine canopy structure and function (Basset et al. 2003; Hamilton et al. 2010). The diversity and composition of the arthropod communities in tree canopies have been associated with microenvironmental variables, plant architecture, vertical foliage complexity, chemical defenses and nutritional quality of hosts (Whitham et al. 2006; Bouget et al. 2011; Ulyshen 2011). Intra- and inter-specific genetic variation of host plants is also known to affect arthropod diversity and levels of damage by herbivores (Morin 2003; Wade 2003; Tovar-Sánchez et al. 2013; Maldonado-López et al. 2015; Valencia-Cuevas and Tovar-Sánchez 2015; Pérez-López et al. 2016; Cuevas-Reyes et al. 2018).
Natural hybridization of plants promotes new genetic combinations as a result of interbreeding between two different species, which can produce fertile or infertile individuals called hybrids (Rieseberg 1997; Martinsen et al. 2001). From these new genetic combinations, novel plant traits can arise in hybrids, including morphological traits or growth and chemical defense characters (Fritz 1999; Whitham et al. 1999; González-Rodríguez et al. 2004; Tovar-Sánchez and Oyama 2004; Bangert et al. 2005; Rehill et al. 2005; Cheng et al. 2011). Therefore, hybrids may represent a more extensive mosaic of resources and conditions available for arthropod communities associated to the tree canopy (Bangert and Whitham 2007; Tovar-Sánchez et al. 2013; Valencia-Cuevas and Tovar-Sánchez 2015). Consequently, hybrid zones provide an excellent scenario to study arthropod diversity and the response of herbivorous insects to the genetic variability of their host plants (Floate et al. 2016; Pérez-López et al. 2016).
Studies on plant–insect interactions in natural hybrid zones have suggested different general scenarios: (i) a higher arthropod density or herbivorous insects performance on hybrid plants compared with the parental species (“The hybrid susceptibility hypothesis”) (Fritz et al. 1994; Whitham et al. 1994; Fritz 1999); (ii) lower herbivore densities on hybrid plants than in parental species, as a result of higher levels of resistance (“The hybrid resistance hypothesis”) (Boecklen and Spellenberg 1990); (iii) intermediate densities of herbivores in hybrid plants in comparison with parental species. It is based on the idea that hybrid resistance to herbivory is the result of additive inheritance traits of both parental species (“The additive hypothesis”) (Boecklen and Spellenberg 1990; Fritz 1999); (iv) similar herbivore density in hybrids with one of the parental species, either the more susceptible or the more resistant parent (“The dominance hypothesis”) (Fritz et al. 1994; Fritz 1999); and (v) the creation of a “hybrid bridge” between host species that can facilitate host-switches by herbivores (“The hybrid bridge hypothesis”) (Floate and Whitham 1993; Pearse and Baty 2012).
When evaluating these hypotheses, it is important to consider features of the plant species such as leaf size and shape. These traits may differ between closely related species and even within species, and several studies have shown positive correlations between leaf size and herbivory levels, which suggest that larger leaves are more attractive to herbivores (Price 1991; Cuevas-Reyes et al. 2011; Garibaldi et al. 2014). It is also known that morphological attributes like leaf teeth, lobes, spines, etc. may interfere with feeding of some insects and therefore impact herbivory levels (Brown and Lawton 1991). A second aspect to consider is arthropod ontogeny, since behavioral characteristics and ecological requirements can change along the developmental stages of these organisms, and therefore their preferences for particular food or habitats (Petersen et al. 2013; Haan et al. 2018; Quintero and Bowers 2018).
The genus Quercus (Fagaceae) is a highly diverse group of woody plants that supports a great diversity of canopy arthropods (Stone and Schönrogge 2003; Tovar-Sánchez and Oyama 2006a, b). Oak species are also known for a high frequency of interspecific hybridization (González-Rodríguez et al. 2004; Tovar-Sánchez and Oyama 2004; Albarrán-Lara et al. 2010; Pérez-López et al. 2016) and depending of the introgression levels, the hybrids can express a continuum of leaf shapes (Whitham 1989). The frequent formation of hybrid zones between Quercus species in Mexico (González-Rodríguez et al. 2004; Tovar-Sánchez and Oyama 2004; Albarrán-Lara et al. 2010; Peñaloza-Ramírez et al. 2010; Valencia-Cuevas et al. 2014) offers an excellent opportunity to analyze the effects of host–plant genetic variation on the composition and structure of canopy arthropod communities and their consequences on the variation of herbivory levels. Therefore, the objective of this study was to determine the changes of structure and composition of canopy arthropod community and the patterns of herbivory in the red oak hybrid complex Quercus laurina × Quercus affinis. We addressed the following questions: (i) what are the patterns of genetic and leaf morphological variation in a local hybrid zone between Q. affinis and Q. laurina, (ii) are hybrid individuals more susceptible to consumption by herbivores than parental trees? (iii) Are there differences in the diversity and composition of canopy arthropod communities among hybrid and parental individuals? (iv) Does the frequency of the different arthropod guilds and ontogenetic stages vary among the two parental species and their hybrids?
Materials and methods
Study system
This study was conducted in Puerto del Aire, Veracruz state, Mexico (18°45′N; 97°30′W). In this site, Q. laurina Humboldt et Bonpland and Q. affinis Scheidweiler occur in sympatry at an altitude of 2380 m. Both species are red oaks (section Lobatae; subgenus Quercus). This hybrid zone was previously characterized genetically and morphologically by González-Rodríguez et al. (2004). From this, based on morphology, in the field we selected individuals of the parental species, as well as hybrid plants for all analyses. Q. laurina is a tree 10–40 m high, with dark gray and finely cracked bark and elliptical-oblanceolate to lanceolate leaves. It occurs along the Sierra Madre del Sur and the Trans-Mexican Volcanic Belt (Arizaga et al. 2009). Q. affinis is a tree that grows up to 25 m in height with elliptical to lanceolate leaves, attenuate or almost acuminate apex, bristle-tipped and cuneate base (Valencia 2004). It is distributed along the Sierra Madre Oriental and the western portion of the Trans-Mexican Volcanic Belt (González-Rodríguez et al. 2004).
Genetic analyses
In the field, we selected in total 45 trees based on their phenotype (15 trees of Q. laurina, 15 trees of Q. affinis and 15 hybrids). To verify this field assignment, individuals were genotyped using nine nuclear microsatellites (Ramos-Ortiz et al. 2016). Total DNA was extracted from 100 mg of leaf material, using a cetyltrimethyl ammonium bromide (CTAB) protocol. We used nine nuclear microsatellite loci (quru-GA-OI01, quru-GA-OM05, quru-GA-OM07, quru-GA-IC08, quru-GA-2F05, quru-GA-2M04, quru-GA-IF07,) previously designed for Quercus rubra (Aldrich et al. 2002). Primers were multiplexed in three groups (OC11 and OA01/OI01, OM05 and OM07/IC08, 1F07, 2F05 and 2M04). For PCR reactions the QIAGEN Multiplex PCR kit (QIAGEN) was used. The final volume of each reaction was 5 µL, containing 1 µL Multiplex PCR Master Mix, 2 mM each primer, deionized water, and 20 ng DNA. The thermal cycling program was run on an Applied Biosystems thermocycler. The program consisted of one cycle at 95 °C for 15 min and then 40 cycles, each with denaturation at 95 °C for 30 s, annealing at 55 °C for 90 s, and extension at 72 °C for 60 s. A final extension at 60 °C for 30 min was included. PCR products were diluted 1:1 in deionized water and run in an ABI-PRISM 3100-Avant sequencer with the GeneScan-500 LIZ size standard included (Applied Biosystems). DNA fragment final sizing was performed using Peak Scanner software, version 1.0 (Applied Biosystems).
Morphometric analysis of leaves
To determine the differences in leaf morphology and size between parental species and hybrid plants, from the same individuals selected for the genetic analyses, we collected 30 intact leaves with no apparent damage by herbivores. We obtained a digital image of each leaf and on each image, 32 anatomical marks were placed with two additional marks as size reference. All morphological marks correspond to homologous loci, which are unambiguous and repeatable marks, representing the shape of the leaves (i.e. ‘landmarks’ sensu Bookstein 1991; Cuevas-Reyes et al. 2011). We recorded the coordinates (x, y) of the 32 anatomical marks in each leaf image with the TpsDig program (Rohlf 1998). A Procrustes superimposition analysis was performed with the CoordGen6 program in the Integrated Morphometrics Package (IMP series: http://www.canisius.edu/~sheets/morphsoft.html). We considered the average configuration of all leaves as reference and then, we calculated the shape variables (Procrustes distances) based on superimposition coordinates to eliminate the effect of leaf size (Cuevas-Reyes et al. 2011). Finally, a principal components analysis (PCA) was performed to evaluate the differences in leaf morphology between the three groups of plants considering the configuration of all leaves.
Herbivory measurements
Samples were collected at the end of the rainy season after the peak of herbivore activity. In the same individuals selected for the genetic and morphometric analyses (15 trees of Q. laurina, 15 trees of Q. affinis and 15 hybrids), we randomly selected 30 leaves per tree from the top, middle and bottom strata of the canopy. Then, we took a digital image of each leaf to calculate the total leaf area and the area removed by herbivorous insects using the Image analysis software for plant disease quantification (Assess Image) (Cuevas-Reyes et al. 2013). Herbivory data were transformed as arc-sine square root (Cuevas-Reyes et al. 2018).
Canopy arthropod diversity
From the individuals selected for the genetic and morphometric analyses, we randomly chose five trees of Q. laurina, five trees of Q. affinis, and five hybrid individuals to determine canopy arthropod diversity. In each tree canopy, a Swingfog SN-50 Thermal Fogger was used to disperse a mix of synergized pyrethrins (30 g/L) and piperonyl butoxide (150 g/L). Fogging was carried out between 6:00 and 7:00 h, for a period of 5 min. The arthropod collection was done 2 h after fogging using 8 funnel-shaped trays (1 m2 in area) installed under each tree canopy (Erwin 1983). After 2 h of the first collection, a second collection was performed to capture the remaining arthropods using ropes to shake the canopy (Marques et al. 2006). The arthropods collected were stored in 95% ethanol and transported to the Agroecology Laboratory of Universidad Michoacana de San Nicolás de Hidalgo, for taxonomic identification. Arthropods were identified to family level, recording the ontogenic stage of each individual, and assigned to trophic guilds (Triplehorn et al. 2005). Considering that taxonomic knowledge is very poor, arthropod abundance and species richness were estimated using morphospecies criteria (recognizable taxonomic units) (Majer et al. 2000).
Statistical analysis
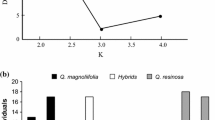
Multilocus microsatellite data were analyzed with the Structure ver. 2.3.4 software (Pritchard et al. 2000; Hubisz et al. 2009) to assign individuals as either Q. laurina, Q. affinis or hybrids (Pérez-López et al. 2016; Cuevas-Reyes et al. 2018). In the analysis the admixture model with correlated allele frequencies without prior information was used. Since previous studies (Ramos-Ortiz et al. 2016) have revealed that two is the most probable number of genetic groups in this hybrid complex, the value of k (number of potential genetic groups) was set to vary only from one to three with ten replicate runs for each value of k. Each run consisted of a burn-in of 105 steps and 106 iterations. The results were analyzed with Structure Harvester Web ver. 0.6.94 (Earl and von Holdt 2011) to confirm that k = 2 was the most probable number of genetic groups, based on the calculation of Δk (Evanno et al. 2005). After these analyses, the inferred admixture coefficient of the individuals (q value) was used to assign them as Q. laurina (q ≥ 0.8), Q. affinis (q ≤ 0.2) or hybrids (0.19 ≥ q ≤ 0.79).
To describe the genetic diversity patterns in parentals and hybrids individuals, we calculated the mean number of alleles per locus (NA), mean effective number of alleles (NE), mean observed heterozygosity (HO), mean expected heterozygosity (HE), mean fixation index (F) and their respective standard errors using GenAlEx 6.5 (Peakall and Smouse 2006).
We tested for differences in leaf area consumed by herbivorous insects between plant groups (i.e. the two parental species and the hybrids). Since leaf area consumed may depende on leaf size (Cuevas-Reyes et al. 2011; Garibaldi et al. 2014), total leaf area was included in this analysis as a covariate. The analysis of covariance (ANCOVA) was performed with the JMP ver. 8.0 software (SAS Institute). A LSMeans test was used for a posteriori comparisons among plant groups (SAS, Stokes et al. 2000).
The overall similarity level in the composition of canopy arthropod communities among the three plant groups was determined calculating a Morisita–Horn similarity matrix (Magurran 1988) and then representing the relationships with the unweighted pair-group method with arithmetic mean (UPGMA) dendrogram in the MVSP 3.2 software (Kovach 1999). Canopy arthropod abundance and species richness were compared among plant groups with a generalized linear model (GLM) analysis using a Poisson error distribution and a log link function. Plant group was used as the independent variable and arthropod abundance and richness as the response variables. A LSMeans test was used for a posteriori comparisons. Additionally, a two-way GLM was applied to analyze the differences of abundance and arthropod richness among plant groups and arthropod orders, trophic guilds and ontogenic stages. We also used a Poisson error distribution and a log link function for this case (SAS, Stokes et al. 2000).
To analyze if the arthropod community composition is related to the genetic distance among individual trees, we calculated a Bray–Curtis distance matrix (Brower and Zar 1984). The corresponding pairwise genetic distance among individuals was calculated in GenAlEx v. 6.5 (Peakall and Smouse 2006) with the method of Smouse and Peakall (1999). Mantel tests were performed between these two matrices to assess the significance of the relationship. We repeated the analysis separating arthropod communities by trophic guilds, excluding mycophages and hematophages because of the low number of individuals collected.
Results
Genetic analysis
The analysis in Structure assigned 24 individuals to Q. laurina, 13 individuals to the hybrid class, and 8 individuals to Q. affinis (Fig. 1). There was general congruence between the field assignment and the genetic assignment, but several of the trees classified in the field as Q. affinis actually belonged to the genetic hybrid class, and several of the morphologically intermediate individuals were genetically assigned as Q. laurina. In all the following analyses we used the genetic assignment to define the plant groups.
Genetic diversity levels were high and similar among the three plant groups, even though HE and HO were slightly higher in the hybrid group (Table 1). The fixation index (F) was positive and similar in the three groups, what can be explained by the presence of null alleles, as has been previously reported (Ramos-Ortiz et al. 2016).
Morphometric analysis
We found consistent differences in leaf morphology among the plant groups. The principal component analysis distinguished the three groups: individuals of Q. laurina, individuals of Q. affinis and the hybrids. In the PCA, the first two axes explained 62.9% of the variance, with 44.9% for PC1 and 18.0% for PC2 (Fig. 2a). The degree of leaf shape variation based on the mean configuration of the coordinate superimposition analysis showed that leaves of Q. laurina were more elongated and wider in comparison with leaves of hybrids and Q. affinis (Fig. 2b). The three plant groups also differed significantly in total leaf area (F = 83.2; df = 2; P < 0.0001). Mean leaf area was significantly greater in Q. laurina (67.8 ± 1.6 cm2) than in hybrids (61.1 ± 1.1 cm2) and Q. affinis (47. 53 ± 1.05 cm2).
Differences in leaf shape morphology between plant groups according to morphometric analysis. a Scatterplot of PCA that show the differences in leaf shape morphology between individuals of the three plant groups. White circles: Q. laurina. White stars: Q. affinis. Black crosses: hybrids (b). Procrustes superposition analysis of leaf morphology variation between the three plant groups: mean shape of coordinates of landmark configuration of leaves. White circles: Q. laurina. White stars: Q. affinis. Black crosses: Hybrids
According to the ANCOVA, percentage of leaf area removed by folivorous insects differed significantly among the hybrids and the parental species (F = 17.73; df = 2; P < 0.001) without a significant effect of total leaf area on leaf area removed (P = 0.36). Parental species did not show significant differences in herbivory levels (Q. laurina: 11.7 ± 3.6% and Q. affinis: 16.7 ± 4.2%) but hybrids showed on average twice the leaf area removed (29.5 ± 5.1%). We found that at least nine free-feeding insect species of different families such as Coleoptera (Chrysomelidae), Orthoptera (Acrididae) and Lepidoptera (Geometridae) were responsible for most of the apparent damage of leaves.
General description of arthropod community
A total of 2217 arthropods were collected (703 in Q. laurina, 929 in hybrids and 585 in Q. affinis), representing 520 morphospecies (193 in Q. laurina, 203 in hybrids and 155 in Q. affinis), grouped in 15 orders (Table S1). The most abundant orders were Coleoptera, Psocoptera and Hemiptera. The orders Araneae, Acari, Collembola, Hymenoptera, Diptera and Lepidoptera generally represented between 5 and 10% of arthropod individuals, while Orthoptera, Opilionidae, Thysanoptera, Neuroptera, Blattodea and Pseudoscorpinidae were rare (< 5%). The Morisita–Horn index indicated that at the level of family, individuals of Q. laurina and hybrids had a 90% of similarity in their arthropod communities, while individuals of Q. affinis and hybrids presented 88% of similarity, and the similarity between Q. affinis and Q. laurina was 84% (Fig. 3). A complete description of composition, abundance and arthropod richness between parental and hybrids is described in Table S1.
Differences in arthropod community among plant groups
We found significant differences among plant groups in arthropod abundance (χ2 = 164.7, df = 2, P = 0.0001) (Fig. 4a) and morphospecies richness (χ2 = 17.2, df = 2, P = 0.0002) (Fig. 4b). Hybrid trees harbored on average higher arthropod abundance (186 ± 6.0 individuals) than Q. laurina (140.2 ± 8.2 individuals) and Q. affinis (87.7 ± 6.1 individuals). A similar pattern was found with the mean richness of arthropod morphospecies, which was higher in hybrids (46.6 ± 3.3) than in Q. laurina (32.4 ± 2.1) and Q. affinis (24.7 ± 3.0).
Diversity patterns of canopy arthropods associated to the oak hybrid complex Q. affinis × Q. laurina in Puerto del Aire, Veracruz. a Differences in total arthropod abundance between the three plant groups. b Differences in total arthropod morphospecies richness. Different letters above bars indicate statistically significant differences (P < 0.05) according to a GLM analysis followed by a LSMeans test
The two-way GLM showed significant differences in arthropod abundance among plant groups (χ2 = 26.1, df = 2, P = 0.001) and arthropod orders (χ2 = 654.34, df = 15, P = 0.0001). Hybrids harbored higher numbers of individuals in the orders Collembola, Psocoptera, Acari and Araneae than parental species. The order Blattodea was only present in hybrids. Q. laurina showed higher abundance in the orders Psocoptera and Orthoptera than hybrids and Q. affinis. Collembola, Acari and Psocoptera were the most abundant orders in Q. affinis (Fig. 5).
We found that all trophic guilds were more abundant in hybrid individuals than in the parental species (χ2 = 269.6, df = 5, P = 0.0001). Predators, phytophages and detritivores–saprophages were the most abundant trophic guilds in the three plant groups, while mycophages and hematophages were the least abundant (Fig. 6a). In the same way, hybrids harbored the highest morphospecies richness in all trophic guilds (χ2 = 340.8, df = 5, P = 0.0001) (Fig. 6b). Finally, we found differences between ontogenetic stages in abundance (χ2 = 162.3, df = 2, P = 0.0001) (Fig. 7a), and arthropod morphospecies richness (χ2 = 16.8, df = 2, P = 0.002) (Fig. 7b). In both cases, adults had a greater abundance and species richness in comparison with nymphs and larvae.
The Mantel correlation test did not show a significant correlation between the genetic distance matrix and the arthropod community composition (r = 0.078, P = 0.29). Similarly, we did not find a significant correlation between genetic distance and community composition when separating trophic guilds (predators: r = 0.019; P = 0.4; parasitoids: r = 0.065, P = 0.3; phytophages: r = − 0.033, P = 0.4 and detritivores–saprophages: r = 0.044, P = 0.06).
Discussion
The genus Quercus is characterized by the formation of hybrid zones in several regions of Mexico (González-Rodríguez et al. 2004; Tovar-Sánchez and Oyama 2004; Peñaloza-Ramírez et al. 2010; Pérez-López et al. 2016). In this study, we confirmed the hybridization between Q. affinis and Q. laurina in a local stand using both molecular markers and morphometric analysis of leaf shape. Our results showed that when the genetic assignment of individuals is used as reference, morphometric analyses also show clear separation with little overlap between Q. affinis and Q. laurina. Interestingly, hybrid individuals were similar to Q. laurina individuals in the first axis of the PC but were separated (although with some overlap) as a third morphological group in the second axis. These results may be explained by the presence of transgressive segregation that involve the presence of different phenotypes in hybrid plants in comparison with the parental species, by gene–environment interactions and dominance, resulting in the differentiation of the hybrid group with at least one parental species (Rieseberg et al. 2003; González-Rodríguez et al. 2004; Cuevas-Reyes et al. 2018).
In particular, interspecific plant hybridization has been recognized as an important mechanism in plant speciation and diversification, and therefore, may have also consequences for arthropod diversity and their food webs associated with the arboreal canopy (Fritz et al. 1994; Fritz 1999; Whitham et al. 1999; Dungey et al. 2000; Hochwender and Fritz 2004; Wimp et al. 2004; Bangert et al. 2005; Pérez-López et al. 2016). We found that in the hybrid oak complex Q. affins × Q. laurina, hybrid individuals had higher abundance and species richness of arthropods in comparison with parental species. In addition, we found that hybrid plants had higher levels of leaf area consumed by folivorous insects than their host parental species. This can be explained by the fact that genetic variation in hybrid plants potentially can generate new phenotypes (Hunter et al. 2004; Tovar-Sánchez and Oyama 2004) with intermediate and novel physiological and biochemical traits, such as secondary metabolites, nutritional quality, phenological and morphological characters that represent potential niches to be colonized by arthropods (Floate and Whitham 1993; Arnold 1997; LeBoldus et al. 2013; González-Rodríguez et al. 2004; Cheng et al. 2011; Pérez-López et al. 2016).
Furthermore, we found a high diversity of arthropod orders, guilds and ontogenic stages, coexisting in hybrid plants. Our results suggest that hybrids include a greater niche diversity that could represent more potential sites to be colonized by arthropods (Martinsen and Whitham 1994; Wimp et al. 2005; Fritz et al. 1994; Strauss 1994; Tovar-Sánchez and Oyama 2006b). Particularly, guilds such as phytophages, predators, detritivores–saprophages and parasitoids presented higher abundance and species richness in hybrid plants. Performance and survival of phytophagous insects are regulated through “bottom-up” forces (host plant quality) (Lill and Marquis 2001; Sobek et al. 2009) and “top-down” forces (predators and parasitoids) (Price et al. 1980; Hartvigsen et al. 1995; Schönrogge et al. 2013). On one hand, several studies have indicated that host plant quality, which is influenced by genetic diversity, is positively related with herbivore diversity (Wimp et al. 2004; Tovar-Sánchez and Oyama 2006a, b, 2004; Tovar-Sánchez et al. 2013; Hunter et al. 2004) and a cascade effect occurs throughout the community, extendible to multiple trophic levels (Whitham et al. 2006). In other words, an increase in host–plant genetic diversity can promote an increase in their architectural complexity and nutritional quality (Bailey et al. 2004) which in turn, favor a greater presence of herbivorous insects (Bailey and Whitham 2006) and natural enemies such as predators and parasitoids (Sarfraz et al. 2008).
Alternatively, “top-down forces” have important influences on ecosystems through direct effects on the regulation of phytophagous arthropod population densities and indirect effects propagated to primary producers (i.e., Marquis and Whelan 1994; Morin 1999; Van Bael et al. 2003; Mäntylä et al. 2011). For example, predators that feed selectively on different competitively superior species reduce the interspecific competition among surviving prey species enhancing the number of prey species that are able to coexist (Murakami and Nakano 2000). Specifically, arthropod communities associated with oak canopies respond to predation by natural enemies as parasites or parasitoids (Tovar-Sánchez et al. 2013). In this way, the high diversity of predators and parasitoids found in our study could be explained by the great abundance of phytophagous arthropods. Detritivores presented a high abundance and richness in hybrid individuals. This guild is a key group within the ecosystem processes since they are responsible for degrading and ingesting the great majority of organic matter, together with fungi and microorganisms (Speight et al. 1999; Chapin et al. 2002).
Arthropod adults and larvae were the ontogenic stages with higher diversity in hybrid individuals. This differential diversity of ontogenic stages between hybrids and parental individuals could indicate a preference of the arthropods in relation to the choice of their host plants; since the vast majority of the arthropods spend much of their life cycle associated with plants, either using them as food resources or shelter to carry out their life cycle (Southwood 1973). Therefore, a greater diversity of ontogenic stages may indicate that arthropods are able to recognize changes in the plant phenotype (consequence of genotypic variation) resulting from hybridization processes.
Overall, our results support the “hybrid susceptibility hypothesis” (Fritz et al. 1994; Whitham et al. 1994; Fritz 1999) also called the “hybrids-as-sinks hypothesis” that predicts a higher incidence of herbivorous insects or arthropods on hybrid plants than in their parental species as result of the disruption of co-adapted gene complexes associated to resistance traits (Whitham 1989). Our results agree with other ecological studies that have shown more herbivore diversity or higher herbivore performance or fitness on host hybrid plants than in parental species (Whitham 1989). The genetic structure of the Q. affinis × Q. laurina hybrid complex significantly affects arthropod diversity and herbivory patterns by folivorous insects, increasing in the hybrid plants. This result suggests the appearance of new niches that can potentially generate an increase in arthropod diversity in hybrid zones, making them important sites of biological activity in terrestrial ecosystems. Hybrid zones have been already described as important centers of arthropod diversity (Whitham et al. 1999; Tovar-Sánchez et al. 2006a, b) and also microorganisms (Strauss 1994; Whitham et al. 1999). Considering that oak species present a high frequency of hybridization, they are very important reservoirs of canopy arthropods and potential sites to analyze ecological and evolutionary process of multiple trophic levels.
References
Albarrán-Lara AL, Mendoza-Cuenca L, Valencia-Avalos S, González-Rodríguez A, Oyama K (2010) Leaf fluctuating asymmetry increases with hybridization and introgression between Quercus magnoliifolia and Quercus resinosa (Fagaceae) through an altitudinal gradient in Mexico. Int J Plant Sci 171:310–322
Aldrich PR, Michler CH, Sun W, Romero-Severson J (2002) Microsatellites markers for northern red aok (Fagaceae: Quercus rubra). Mol Ecol Notes 2:472–474
Arizaga S, Martínez-Cruz, Salcedo-Cabrales M, Bello-González MA (2009) Manual de la biodiversidad de encinos michoacanos. SEMARNAT, INE, México, 147 pp
Arnold ML (1997) Natural hybridization and evolution. Oxford University Press, New York
Bailey JP, Whitham T (2006) Interactions between cottonwood and beavers positively affect sawfly abundance. Ecol Entomol 31:294–297
Bailey JK, Bangert RK, Schweitzer JA, Trotter RT III, Shuster SM, Whitham TG (2004) Fractal geometry is heritable in trees. Evolution 59:2100–2102
Bangert RK, Whitham TG (2007) Genetic assembly rules and community phenotypes. Evol Ecol 21:549–560
Bangert RK, Turek RJ, Martinsen GD, Wimp GM, Bailey JK, Whitham TG (2005) Benefits of conservation of plant genetic diversity on arthropod diversity. Conserv Biol 19:379–390
Basset Y, Novotny V, Miller SE, Kitching RL (2003) Arthropods of tropical forests. Spatio-temporal dynamics and resource use in the canopy. Cambridge University Press, Cambridge
Boecklen WJ, Spellenberg R (1990) Structure of herbivore communities in two oak (Quercus spp.) hybrid zones. Oecologia 85:92–100
Bookstein FL (1991) Morphometric tools for landmark data: geometry and biology. Cambridge, University Press, New York
Bouget C, Brin B, Brustel H (2011) Exploring the “last biotic frontier”: Are temperate forest canopies special for saproxylic beetles? For Ecol Manag 261:211–220
Brower JE, Zar JH (eds) (1984) Community similarity. In: Field and laboratory methods for general ecology, 3rd edn. W.C. Brown Publishers, Dubuque
Brown VK, Lawton JH (1991) Herbivory and the evolution of leaf size and shape. Phil Trans R Soc Lond B 333:265–272
Chapin FS III, Matson PA, Mooney HA (2002) Principles of terrestrial ecosystem ecology. Springer, New York
Cheng D, Vrieling K, Klinkhammer PGL (2011) The effect of hybridization on secondary metabolites and herbivore resistance: Implications for the evolution of chemical diversity in plants. Phytochem Rev 110:07–117
Cuevas-Reyes P, Oyama K, González-Rodríguez A, Fernandes GW, Mendoza-Cuenca L (2011) Contrasting herbivory patterns and leaf fluctuating asymmetry in Heliocarpus pallidus between different habitat types within a Mexican tropical dry forest. J Trop Ecol 27:383–391
Cuevas-Reyes P, Gilverti L, González-Rodríguez A, Fernándes GW (2013) Patterns of herbivory and fluctuating asymmetry in Solanum lycocarpum St. Hill (Solanaceae) along an urban gradient in Brazil. Ecol Indic 24:557–561
Cuevas-Reyes P, Canché-Delgado A, Maldonado-López Y, Fernandes GW, Oyama K, González-Rodríguez A (2018) Patterns of herbivory and leaf morphology in two Mexican hybrid oak complexes: importance of fluctuating asymmetry as indicator of environmental stress in hybrid plants. Ecol Indic 90:164–170
Dungey HS, Potts BM, Whitham TG, Li H-F (2000) Plant genetics affects arthropod community richness and com- position: evidence from a synthetic eucalypt hybrid population. Evolution 54:1938–1946
Earl DA, von Holdt BM (2011) Structure harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Gen Resour 4:359–361
Erwin TL (1983) Beetles and other insects of tropical forest canopies at Manaus, Brazil, sampled by insecticidal fogging. In: Sutton SL, Whitmore TC, Chadwick AC (eds) Tropical rain forest ecology and management. Blackwell Scientific Publications, Oxford
Evanno G, Regnaut S, Goudet (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Floate KD, Whitham TG (1993) The ‘‘hybrid bridge’’ hypothesis: host shifting via plant hybrid swarms. Am Nat 141:651–662
Floate KD, Godbout J, Lau MK, Isabel N, Whitham TG (2016) Plant-herbivore interactions in a trispecific hybrid swarm of Populus: assessing support for hypotheses of hybrid bridges, evolutionary novelty and genetic similarity. New Phytol 209:832–844
Fritz RS (1999) Resistance of hybrid plants to herbivores: genes, environment both? Ecology 80:382–391
Fritz RS, Nichols-Orians CM, Brunsfeld SJ (1994) Interspecific hybridization of plants and resistance to herbivores: hypotheses, genetics, and variable responses in a diverse herbivore community. Oecologia 97:106–117
Garibaldi LA, Kitzberger T, Ruggiero A (2014) Latitudinal decrease in folivory within Nothofagus pumilio forests: dual effect of climate on insect density and leaf traits? Glob Ecol Biogeogr 20:609–619
González-Rodríguez A, Arias DM, Valencia S, Oyama K (2004) Morphological and RAPD analysis of hybridization between Quercus affinis and Q. laurina (Fagaceae), two Mexican red oaks. Am J Bot 91:401–409
Haan NL, Bakker JD, Dunwiddie PW, Linders MJ (2018) Instar-specific effects of host plants on survival of endangered butterfly larvae. Ecol Entomol. https://doi.org/10.1111/een.12656
Hamilton AJ, Basset Y, Benke KK, Grinbacher PS, Miller SE, Samuelson GA, Stork NE, Weiblen GD, Yen JD (2010) Quantifying uncertainty in estimation of tropical arthropod species richness. Am Nat 176:90–95
Hartvigsen G, Wait DA, Coleman JS (1995) Tri-trophic interactions influenced by resource availability: predator effects on plant performance depend on plant resources. Oikos 74:463–468
Hochwender CG, Fritz RS (2004) Plant genetic differences influence herbivore community structure evidence from a hybrid willow system. Oecologia 138:547–557
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 9:1322–1332
Hunter JJ, Volschenk CG, Le Roux DJ, Fouché GW, Adams L (2004) Plant material quality. A compilation of research. ARC Infruitec-Nietvoorbij, Stellenbosch, pp 1–18
Kovach WL (1999) MVSP. A multivariate statistical package for windows, ver. 3.1. Kovach Computing Services. Pentraeth
LeBoldus JM, Isabel N, Floate KD, Blenis P, Thomas BR (2013) Testing the ‘hybrid susceptibility’ and ‘phenological sink’ hypotheses using the P. balsamifera–P. deltoides hybrid zone and Septoria leaf spot [Septoria musiva]. PLoS ONE 8:e84437
Lill JT, Marquis RJ (2001) The effects of leaf quality on herbivore performance and attack from natural enemies. Oecologia 126:418–428
Magurran AE (1988) Ecological diversity and its measurement. Princeton University Press, Princeton
Majer JD, Recher HF, Ganesh S (2000) Diversity patterns of eucalypt canopy arthropods in eastern and western of Australian. Ecol Entomol 25:295–306
Maldonado-López Y, Cuevas-Reyes P, González-Rodríguez A, Pérez-López G, Acosta-Gómez C, Oyama K (2015) Relationships among plant genetics, phytochemistry and herbivory patterns in Quercus castanea across a fragmented landscape. Ecol Res 30:133–143
Mäntylä E, Klemola T, Laaksonen T (2011) Birds help plants: a meta-analysis of top-down trophic cascades caused by avian predators. Oecologia 165:143–152
Marques MI, Adis J, dos Santos GB, Battirola LD (2006) Terrestrial arthropods from tree canopies in the Pantanal of Mato Grosso, Brazil. Rev Bras Entomol 50:257–267
Marquis RJ, Whelan CJ (1994) Insectivorous birds increase growth of white oak through consumption of leaf chewing insects. Ecology 75:2007–2014
Martinsen GD, Whitham TG (1994) More birds nest in hybrid Cottonwood trees. Wilson Bull 106:474–481
Martinsen GD, Whitham TG, Turek RJ, Kim P (2001) Hybrid populations selectively filter gene introgression between species. Evolution 55:1325–1335
Morin PJ (1999) Community ecology. Blackwell Science, Oxford
Morin PJ (2003) Community ecology and the genetics of interacting species. Ecology 84:577–580
Murakami M, Nakano S (2000) Species-specific bird functions in a forest-canopy food web. P Roy Soc Lond B 267:1597–1601
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel.Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pearse IS, Baty JH (2012) The predictability of traits and ecological interactions on 17 different crosses of hybrid oaks. Oecologia 169:489–497
Peñaloza-Ramírez JM, González-Rodríguez A, Mendoza-Cuenca L, Caron H, Kremer A, Oyama K (2010) Interspecific gene flow in a multispecies oak hybrid zone in the sierra Tarahumara of Mexico. Ann Bot 105:389–399
Pérez-López G, González-Rodríguez A, Oyama K, Cuevas-Reyes P (2016) Effects of plant hybridization on the structure and composition of a highly rich community of cynipid gall wasps: the case of the oak hybrid complex Quercus magnoliifolia x Quercus resinosa in Mexico. Biodivers Conserv 25:633–651
Petersen MJ, Seto M, Peck D (2013) Linking the spatio-temporal distribution of an edaphic crane fly to its heterogeneous soil environment. Ecol Entomol 38:585–595
Price PW (1991) The plant vigor hypothesis and herbivore attack. Oikos 62:244–251
Price PW, Bouton CE, Gross P, McPheron BA, Thompson JN, Weis AE (1980) Interactions among three trophic levels: influence of plants on interactions be tween insect herbivores and natural enemies. Ann Rev Ecol System 11:41–65
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quintero C, Bowers MD (2018) Plant and herbivore ontogeny interact to shape the preference, performance and chemical defense of a specialist herbivore. Oecologia 187:401–412
Ramos-Ortiz S, Oyama K, Rodríguez-Correa H, González-Rodríguez A (2016) Geographic structure of genetic and phenotypic variation in the hybrid zone between Quercus affinis and Q. laurina in Mexico. Plant Species Biol 31:219–232. https://doi.org/10.1111/1442-1984.12109
Rehill B, Clauss A, Wieczorek L, Whitham T, Lindroth R (2005) Foliar phenolic glycosides from Populus fremontii, Populus angustifolia, and their hybrids. Biochem Syst Ecol 33:125–131
Rieseberg LH (1997) Hybrid origins of plant species. Annu Rev Ecol Syst 28:359–389
Rieseberg LH, Raymond O, Rosenthal DM, Lai Z, Livingstone K, Nakazato T, Durphy JL, Schwarzbach AE, Donovan LA, Lexer C (2003) Major ecological transitions in wild sunflowers facilitated by hybridization. Science 301:1211–1216
Sarfraz M, Dosdall LM, Keddie BA (2008) Host plant genotype of the herbivore Plutella xylostela (Lepidopetera: Plutellidae) affects the performance of its parasitoid Diadegma insulare (Hymenoptera: Ichneumonidae). Biol Control 44:42–51
Schönrogge K, Begg T, Stone GN (2013) Native birds and alien insects: spatial density dependence in songbird predation of invading oak gall wasps. PLoS ONE 8:e53959
Smouse PE, Peakall R (1999) Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82:561–573
Sobek S, Steffan-Dewenter I, Scherber C, Tschamtke T (2009) Spatio temporal changes of beetle communities across a tree diversity gradient. Divers Distrib 15:660–670
Southwood TRE (1973) The insect/plant relationship-an evolutionary perspective. In van Emden HF (ed) Insect/plant relationship, No. 6. Symposia of the Royal Entomological Society, London, pp. 3–30
Speight MR, Hunter MD, Watt AD (1999) Ecology of insects: concepts and applications. Blackwell Science, Oxford
Stokes ME, Davis CS, Koch GG (2000) Categorical data analysis using the SAS system, 2nd edn. SAS, Cary
Stone GN, Schönrogge K (2003) The adaptive significance of insect gall morphology. Tree 18:512–522
Strauss SY (1994) Levels of herbivory and parasitism in host hybrid zones. Trends Ecol Evol 9:209–214
Tovar-Sánchez E, Oyama K (2004) Natural hybridization and hybrid zones between Quercus crassifolia and Quercus crassipes (Fagaceae) in Mexico: morphological and molecular evidence. Am J Bot 91:1352–1363
Tovar-Sánchez E, Oyama K (2006a) Community structure of canopy arthropods associated to Quercus crassifolia × Quercus crassipes complex. Oikos 112:370–381
Tovar-Sánchez E, Oyama K (2006b) Effect of hybridization of the Quercus crassifolia x Quercus crassipes complex on the community structure on endophagous insects. Oecologia 147:702–713
Tovar-Sánchez E, Valencia-Cuevas L, Castillo-Mendoza E, Mussali-Galante P, Pérez- Ruíz RV, Mendoza A (2013) Association between individual genetic diversity of two oak host species and canopy arthropod community structure. Eur J For Res 132:165–179
Triplehorn CA, Johnson NF, Borror DJ (2005) An introduction to the study of insects. Thompson Brooks/Cole, Belmont
Ulyshen MD (2011) Arthropod vertical stratification in temperate deciduous forests: Implications for conservation-oriented management. For Ecol Manag 261:1479–1489
Valencia S (2004) Diversidad del género Quercus (Fagaceae) en México. Bot Soc Bot Mex 75:33–53
Valencia-Cuevas L, Tovar-Sánchez E (2015) Oak canopy arthropod communities: which factors shape its structure? Rev Chil Hist Nat 88:15
Valencia-Cuevas L, Piñero D, Mussali-Galante P, Valencia-Ávalos S, Tovar-Sánchez E (2014) Effect of a red oak species gradient on genetic structure and diversity of Quercus castanea (Fagaceae) in Mexico. Tree Genet Genom 10:641–652
Van Bael SA, Brawn JD, Robinson SK (2003) Birds defend trees from herbivores in a Neotropical forest canopy. Proc Natl Acad Sci USA 100:8304–8307
Wade MJ (2003) Community genetics and species interactions. Ecology 84:583–585
Whitham TG (1989) Plant hybrid zones as sinks for pests. Science 244:1490–1493
Whitham TG, Morrow PA, Potts BM (1994) Plant hybrid zones as centers of biodiversity—the herbivore community of endemic Tasmanian eucalypts. Oecologia 97:481–490
Whitham TG, Martinsen GD, Floate KD, Dungey H, Potts BM, Keim P (1999) Plant hybrid zones affect biodiversity: tools for a genetic-based understanding of community structure. Ecology 80:416–428
Whitham TG, Bailey JK, Scheweitzer JA, Shuster SM, Bangert RK, LeRoy CJ, Lonsdorf EV, Allan GJ, DiFazio SP, Potts BM, Fischer DC, Gehring CA, Lindroth RL, Marks JC, Hart SC, Wimp GM, Wooley SC (2006) A framework for community and ecosystem genetics: form genes to ecosystems. Nature 7:510–523
Wimp GM, Young PW, Woolbright SA, Martinsen GD, Keim P, Whitham TG (2004) Conserving plant genetic diversity for dependent animal communities. Ecol Lett 7:776–780
Wimp GM, Martinsen GD, Floate KD, Bangert RK, Whitham TG (2005) Plant genetic determinants of arthropod community structure and diversity. Evolution 59:61–69
Yang LH, Gratton C (2014) Insects as drivers of ecosystem processes. Curr Opin Insect Sci 2:26–32
Acknowledgements
This project was supported by CONACYT project CB105755 and DGAPA-PAPIIT-UNAM project RV201015. Cuevas-Reyes P thanks Coordinación de la Investigación Científica UMSNH (Grant No. 111) for their generous support. We also thank Fidel Anguiano for editing figures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Research involving human participants and/or animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Maldonado-López, Y., Vaca-Sánchez, M.S., González-Rodríguez, A. et al. Hybridization increases canopy arthropod diversity in the Quercus affinis × Quercus laurina complex. J Insect Conserv 22, 781–793 (2018). https://doi.org/10.1007/s10841-018-0103-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10841-018-0103-7