Abstract
Geometric morphometrics and molecular methods are effective tools to study the variability of stingless bee populations and species that merit protection given their worldwide decline. Based on previous evidence of cryptic lineages within the Scaptotrigona genus, we tested the existence of multiple evolutionary lineages within the species S. mexicana and we investigated the status of S. pectoralis. By analyzing their population structure, we found differences between the Pacific and Atlantic populations of each of these species, although geometric morphometrics of the wing only confirmed these results in S. mexicana. There was a tendency towards enhanced genetic differentiation over larger distances in the Atlantic populations of both species but not in the Pacific populations. These results revealed a pattern of differentiation among evolutionary units and a specific distribution of genetic diversity within these Scaptotrigona species in Mesoamerica, suggesting the need for future taxonomic revisions, as well as activities aimed at management and conservation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stingless bees (Hymenoptera: Apidae: Meliponini) are eusocial, haplodiploid insects that carry out ecologically and economically important pollinating activity, especially in Neotropical regions (Slaa et al. 2006). Most of the diversity of the tribe Meliponini is concentrated in this region, with approximately 42 endemic genera (Camargo and Pedro 2013) that include around 400 species (Michener 2007). One of these genera is Scaptotrigona Moure, 1942, which consists of 24 species extending from Mexico to Argentina (Michener 2007). Three Scaptotrigona species are found in Mexico: S. mexicana Guérin, 1845 the distribution of which extends to Costa Rica; S. pectoralis Dalla Torre, 1896 the distribution of which reaches Panama; and the endemic S. hellwegeri Friese, 1900 (Ayala 1999; Camargo and Pedro 2013). These three species are managed for honey production (Manzo 2009; González-Acereto 2012; Ayala et al. 2013) but at present, only S. mexicana and S. pectoralis contribute to the pollination of crops like avocado (Ish-Am et al. 1999).
Given their important contribution to agriculture, the commercial use of these species is augmenting (Albores-González et al. 2011), potentially influencing the gene flow among populations due to hive translocation (Jaffé et al. 2016; Santiago et al. 2016). Like many other insects, these species are also being affected by extensive deforestation and the extension of monocrops in these areas (Freitas et al. 2009; Stout and Morales 2009). These phenomena may potentially drive a decline in stingless bee populations and they may provoke endogamous depression, an increase in homozygosity and the ensuing loss of genetic diversity (Zayed 2009). Hence, the lack of information about their richness, diversity, genetic status and distribution is an important obstacle to ensure their protection and to establish appropriate conservation programs (Brown and Paxton 2009; Freitas et al. 2009). These programs should include measures avoiding the translocation of hives and to define which populations should remain isolated to preserve genetic integrity.
The existence of cryptic species of stingless bees has been proposed (Tavares et al. 2007; Francisco et al. 2008; May-Itzá et al. 2012) and thus, the total number of species could be higher than anticipated (Michener 2007). The number of cryptic species will probably increase as more molecular methods are made available and more powerful morphological analyses are employed. In the last decade, geometric morphometric analysis of wings has proven to be a good tool to clarify discrepancies within species and to identify bees (for example honeybees: Bouga et al. 2011; da Silva et al. 2015; or bumblebees: Barkan and Aytekin 2013). In stingless bees, this approach has proved to be very sensitive to characterize species and even to differentiate populations (Ferreira et al. 2011; Francoy et al. 2011; Lima Junior et al. 2012; Nunes et al. 2013; Bonatti et al. 2014), although genetic information should be incorporated to more rigorously define species and to resolve taxonomic problems (Schlick-Steiner et al. 2010). Species and population identification using genetic markers has advanced significantly, and population parameters important for the conservation of endangered species can now be derived from the use of molecular markers (Frankham et al. 2002; Hedrick 2005). Among these, microsatellite loci have good statistical resolution to characterize demographic events within populations (Luikart and England 1999), and they have been successfully used within the Meliponini to characterize genetic variability (Arias et al. 2006; Francisco et al. 2006; Fernandes et al. 2012) and issues like the genetic characteristics of drone congregation areas (Kraus et al. 2008; Mueller et al. 2012).
Studies of morphological and/or molecular markers in the genus Scaptotrigona have suggested the existence of genetically distinct evolutionary lineages (or cryptic species). A study on S. hellwegeri found signs of ongoing speciation, indicating that two populations distributed at different altitudes along the Trans-Mexican Volcanic Belt and the Pacific coast should be treated as separate units (Quezada-Euán et al. 2012). The existence of cryptic species was also proposed within S. mexicana colonies in two regions located at opposite extremes of the species distribution range in Mexico (Veracruz and Chiapas: Hurtado-Burillo et al. 2013). Recently, up to five distinct clusters (considered as units of management and conservation) with high rates of genetic diversity were described in another Scaptotrigona species (S. xanthotricha, which is widely distributed in the Brazilian Atlantic rainforest: Duarte et al. 2014).
As such, here we aimed to resolve taxonomic issues in S. mexicana and S. pectoralis, two species that share an equivalent distribution, as well as similar biological and ecological aspects, but that are managed distinctly. Specifically, we used geometric morphometry and microsatellite markers to assess the population structure and the molecular diversity of S. mexicana and S. pectoralis, and to test the existence of various evolutionary units within S. mexicana. In addition, these tools were used to investigate the species status of S. pectoralis in the light of the diversity observed in this genus.
Materials and methods
Sampling
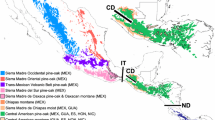
This study was carried out in Mexico and northern Guatemala, within the distribution area of S. mexicana and S. pectoralis. Colonies were sampled at 15 sites between 2008 and 2011 (Table 1; Fig. 1), and most S. mexicana samples (70 colonies in total) were provided by stingless beekeepers that maintain colonies in clay pots or in their original trunks, either on their own property or in the vicinity (meliponaries). S. pectoralis samples (33 colonies in total) were obtained from non-managed wild colonies located in native forests. Adult worker bees were collected at the entrance of each colony and preserved in absolute ethanol at −20 °C. Voucher specimens were deposited in the insect collection of the Animal Biology Laboratory at the Veterinary Faculty of the University of Murcia (Spain).
Localities in Mexico and Guatemala where specimens were collected: red dots represent S. mexicana and green dots—S. pectoralis colonies. Numbers correspond to the sampling code used in Table 1. Distribution ranges have been shadowed: in red—S. mexicana, in green—S. pectoralis and in light brown the area where the two species co-exist. (Color figure online)
Geometric morphometric analysis
One to six specimens from 70 S. mexicana and 32 S. pectoralis colonies were analysed (Table 1). Only the right wing of each specimen was used and they were photographed with a Spot Insight Firewire digital camera (Sterling Heights, USA) adapted to a Zeiss Stemi 2000C Trinocular Zoom Stereomicroscope (Thornwood, USA). We manually plotted twelve homologous landmarks of the wing vein intersections (Fig. 2) using the tpsDig software, (version 2.17: Rohlf 2013). MorphoJ software (version 1.06c: Klingenberg 2011) was used to analyze the images, which were first Procrustes aligned (Bookstein 1991) in order to identify the points of shape variation. Mean values of the specimens from each colony were used to perform the analyses at the colony level. Principal component (PCA) and canonical variate (CVA) analyses were performed with these data.
Mahalanobis distances between groups of colonies were defined by the PCA analysis and were obtained with the MorphoJ software. We also performed a Mantel test to investigate whether the Procrustes-fitted landmark co-ordinates of the wings varied with the geographic distance in each group using Past 3.08 software (Hammer et al. 2001).
DNA extraction and microsatellite amplification
DNA was extracted from the leg of one worker bee per colony (S. mexicana, N = 67; S. pectoralis N = 33: Table 1) using the DNeasy tissue kit (QIAGEN). Seven microsatellite loci were amplified in three reactions (two of them multiple), two of which were microsatellites originally described in S. postica (T4–171 and T7–5; Paxton et al. 1999), four in Melipona bicolor (Mbi278, Mbi259, Mbi254 and Mbi201; Peters et al. 1998) and one in the bumblebee Bombus terrestris (B124; Estoup et al. 1995). The T4–171, T7–5 and B124 loci have already been used successfully in S. mexicana (Kraus et al. 2008; Mueller et al. 2012) and S. hellwegeri (Quezada-Euán et al. 2012).
PCR reactions were carried out in 12.5 μl with PureTaqTM Ready-To-Go TM PCR beads (GE Healthcare) in a PTC-200 Thermal Cycler (Biorad). The amplified fragments (microsatellite alleles) were detected with an ABI prism 3100 sequencer (Applied Biosystems) and scored with Genemapper software (version 3.7: Applied Biosystems).
Molecular data analyses
The number of clusters (K) was estimated with the STRUCTURE software (version 2.2: Pritchard et al. 2000) using a Bayesian model-based method, basing the results on simulations of 80,000 burn-in steps and 1,000,000 MCMC (Markov Chain Monte Carlo) algorithm iterations. Five runs for each K-value (K = 1 − 8) were used to estimate the most likely value of K, and the runs were performed without any prior information about the origin of samples using the “Admixture Model” and considering allele frequencies correlated among populations. The number of clusters defined by the ΔK value described previously (see Evanno et al. 2005) was inferred by STRUCTURE HARVESTER (Earl and vonHoldt 2012). CLUMPP (v1.1.2: Jakobsson and Rosenberg 2007) was used to obtain the average permutated individual and population Q-matrices of the best K value. The distruct software (v1.1: Rosenberg 2004) was used to graphically display the CLUMPP results.
The dataset describing the genotypes included in each cluster was analyzed with MICRO-CHECKER (Van Oosterhout et al. 2004) to detect scoring incongruities and possible null alleles. GENALEX (Peakall and Smouse 2006) was used to calculate population genetics parameters per locus and cluster in each species (the number of detected and private alleles, and the expected heterozygosity). Given the different number of individuals sampled for each cluster, a rarefaction analysis gave an objective estimate of allelic richness (i.e. the number of alleles: Leberg 2002) and it was carried out with the HP-RARE 1.0 program (Kalinowski 2005). Differences between clusters were estimated using the two-tailed Mann–Whitney U Test (Mann and Whitney 1947).
Hardy–Weinberg and genotypic linkage equilibria, and the population differentiation using the Fisher’s exact probability test were calculated in GENEPOP (Raymond and Rousset 1995a, b). All probability values were adjusted for multiple comparison tests using sequential Bonferroni adjustments (Rice 1989).
Pairwise R ST values were calculated as a measure of genetic distance in ARLEQUIN 3.5 (Excoffier and Lischer 2010), calculating R ST pairwise values to estimate the connectivity and patterns of gene flow between clusters given that F ST values might underestimate the differentiation in highly structured populations (Balloux and Lugon‐Moulin 2002) like those of Scaptotrigona species (Quezada-Euán et al. 2012; Hurtado-Burillo et al. 2013). The correlation between genetic and geographic distances within each cluster was analyzed with a Mantel test, as implemented in GENALEX, to test whether allelic frequencies varied in function of a pattern of isolation by distance (IBD).
Results
Geometric morphometrics
PCA analysis of the 12 wing landmarks generated 20 relative warp measurements. The first 11 factors of these measurements had eigenvalues greater than one and explained 95.46 % of the total variability in the data. The first two PC values explained 64.52 % of the total variability (PC1 explained 49.31 % and PC2 15.21 %) and a PCA identified three groups, two within S. mexicana (herein named Sm1 and Sm2) and one with the S. pectoralis colonies. The graphic representation showed that the colonies from the Pacific coast (Chiapas, Sm1) were placed in quadrant 4 while the rest of the colonies from the Atlantic coast (Veracruz in Mexico and Peten in Guatemala, Sm2) were mainly placed in quadrant 3. Conversely, the samples of S. pectoralis were dispersed in quadrants 1–2 (Fig. 3).
The Mahalanobis distances between the groups confirmed the differentiation between Sm1 and Sm2 (4.847), although this value was around half of that observed between S. pectoralis and Sm1 (8.477) or Sm2 (8.171). In addition, the Mantel test showed no significant correlation between morphological and geographical distances in each putative species (Sm1: R = −0.13, p = 0.99; Sm2: R = 0.012, p = 0.3 and S. pectoralis: R = 0.010, p = 0.39).
Microsatellites
The average genotyping error rate for the seven microsatellite loci assessed in this study was <5 %. Less than four loci were amplified from two of the 67 S. mexicana specimens and thus, they were excluded from the analysis. In a first step, we looked for signs of genetic structure in each species. The highest posterior probability of the data set was detected when assuming two clusters (K = 2) in both species: S. mexicana (Sm1 and Sm2) and S. pectoralis (Sp1 and Sp2) (Fig. 4). S. mexicana clusters were congruent with the geometric morphometry data: Sm1 (colonies from Chiapas) and Sm2 (colonies from Veracruz and Petén, including in this case three more colonies, two from Puebla and one from Hidalgo). In contrast to the morphological results, S. pectoralis was divided into two clusters: Sp1 (colonies from southern Chiapas: Tapachula and Tuxtla Chico) and Sp2 (colonies from Veracruz, Yucatan and northern Chiapas: Palenque). These clusters showed a geographic pattern for both species, splitting Pacific (Sm1 and Sp1) and Atlantic populations (Sm2 and Sp2), and they were used to estimate the population genetic parameters.
Results of the Bayesian clustering showing the most probable number of clusters (K = 2) for the two species. Division of specimens into colored segments represents the assignment probability of that specimen to each of the K clusters (red—Sm1; light red—Sm2; dark green—Sp1; light green—Sp2). Numbers correspond to the sampling code used in Table 1 and acronyms to the name of the state (CHP Chiapas, PTN Petén, VRC Veracruz, HGO Hidalgo, PUE Puebla, YUC Yucatán). (Color figure online)
There was no evidence of scoring error due to stuttering or allele dropout, yet there appeared to be null alleles at locus T4-171 in Sm2, and at loci Mbi254-AAG and B124 in Sp2. These loci had a higher percentage of unsuccessfully genotyped individuals (3.5, 4.5 and 1.5 %, respectively), a fact that was taken into account to explain several population parameters.
The number of alleles of the seven microsatellite loci ranged from one to ten across all the samples (Table 2). The average allele number after rarefaction was higher in the Sm1 (2.65 ± 0.4) than in Sm2 (2.50 ± 0.3) populations, and in the Sp2 (2.41 ± 0.4) than in Sp1 (1.76 ± 0.3) populations, and fewer private alleles (i.e. alleles unique to each cluster, Npa) were evident after rarefaction. The highest Npa was detected in Sm1 within S. mexicana, (1.44 ± 0.5), whereas for S. pectoralis the highest Npa was detected in Sp2 (1.1 ± 0.3). All comparisons were not statistically significant (p > 0.05). Gene diversity was measured as the expected heterozygosity (He) and it was higher in the Sm2 than in the Sm1 population, and in the Sp2 than in the Sp1 population, although these difference were not significant.
The Sm1 cluster was in Hardy–Weinberg equilibrium before Bonferroni correction (p > 0.05 for each locus), whereas in the Sm2 population one locus (T4-171) did not show Hardy–Weinberg equilibrium even after Bonferroni correction, highlighting a marked excess of homozygotes that was probably due to the presence of null alleles. A similar result was obtained in the S. pectoralis populations as all loci were in Hardy–Weinberg equilibrium in the Sp1 population whereas Sp2 did not show such equilibrium at those loci with null alleles (Mbi254AAG and B124). The loci were generally out of linkage disequilibrium after Bonferroni correction for each cluster, with the exceptions of 4 out of the 15 pairwise comparisons in Sm2.
A Fisher’s exact test of population differentiation showed highly significant genetic differences between the clusters of each species. The pairwise R ST was higher between Sm1 and Sm2 (R ST = 0.266, p < 0.0001) than between Sp1 and Sp2 (R ST = 0.198, p < 0.0001). The IBD was evaluated with the Mantel test for each cluster and there was a significant correlation between genetic and geographic distance in the Atlantic populations of both species (Sm2 r = 0.383, p = 0.001; and Sp2 r = 0.477, p = 0.001), but no correlation in the Pacific populations (Sm1 r = −0.034, p = 0.177; and Sp1 r = 0.158, p = 0.079).
Discussion
Our results demonstrate that diversity in the Scaptotrigona genus is higher than expected in Mesoamerica. The genetic and geometric morphometric analyses fully support the existence of two evolutionary significant units (ESUs) within S. mexicana (Sm1 and Sm2), with a Pacific and Atlantic distribution, respectively. By contrast, S. pectoralis population differentiation is evident through genetic analysis (Sp1 and Sp2) but not so clearly by geometric morphometry, suggesting that S. pectoralis is in an incipient stage of population differentiation.
The clusters found within S. mexicana resembled two ESUs with different distributions, one dispersed along the Pacific coast of Mexico (Sm1) and the other along the Atlantic Mexican coast up to the North of Guatemala (Sm2). These data support previous results suggesting the existence of a cryptic species within S. mexicana based on high divergence between the populations identified by means of the barcoding method (Hurtado-Burillo et al. 2013). The results obtained here support this hypothesis, since both the molecular and morphological approaches discriminate the two ESUs. A multidisciplinary approach such as that performed here also proved to be useful in other population studies on stingless bees species (Mendes et al. 2007; Gonçalves 2010; Francoy et al. 2011; May-Itzá et al. 2012; Bonatti et al. 2014) and on honey bees, albeit at the sub-species level in the latter (Oleksa and Tofilski 2015).
The differentiation between Sm1 and Sm2 is evident through several population parameters. First, geometric mophometry of the wings indicates a significant phenotypic differentiation of two separate groups. Second, while both S. mexicana clusters share alleles, the presence of private alleles in both the Pacific Sm1 and Atlantic Sm2 populations, together with different allelic frequencies at some loci, indicate genetic differences between these two evolutionary units. Although microsatellite markers usually show less allelic diversity when used in species other than those they have been designed for (Borges et al. 2010), they yielded appropriate information in analyses of other S. mexicana populations (Kraus et al. 2008; Mueller et al. 2012) and S. hellwegeri (Quezada-Euán et al. 2012). Finally, the R ST value (0.266) points to significant divergence within S. mexicana units. While this R ST value could reflect an IBD effect that restricts gene flow among colonies, it is not conclusive of complete separation into two species. Introgression events between ESUs of S. mexicana (Sm1 and Sm2: Fig. 4) have been observed in both Atlantic and Pacific populations, possibly the result of natural gene flow. However, human influence should not be ignored as S. mexicana is one of the species traditionally managed in Mesoamerican cultures and our samples were obtained from stingless beekeepers. Although movements of hives over large distances are uncommon in Mesoamerica (May-Itzá et al. 2012; González-Acereto pers. comm.), genetic traces of human-mediated transportation have been observed elsewhere for other stingless bees (Byatt et al. 2016).
Genetic differentiation was also observed for S. pectoralis, with microsatellite analysis defining two clusters (Sp1 and Sp2) and confirming the data obtained with mitochondrial markers (Hurtado-Burillo et al. 2013). The clustering obtained previously through the barcoding approach also suggested some degree of differentiation within S. pectoralis, although the intraspecific divergence did not fully support the existence of distinct genetic lineages. Since extant geometric morphometric data does not corroborate this differentiation, the two clusters in S. pectoralis may not be ESUs but rather, units of management and conservation, as in the case of Brazilian populations of Scaptotrigona xanthotricha (Duarte et al. 2014). In S. pectoralis, the failure to detect introgression suggests there has been no recent gene flow between the two clusters. This could be due to the limited sampling (n = 33) and or the fact that the colonies sampled were wild, unmanaged colonies located in native forests.
The homogeneous genetic structure detected in distant populations of Atlantic clusters, despite the presence of three introgressed S. mexicana specimens (one from Chiapas in Sm1 and two from Petén in Sm2), and especially in the Pacific clusters of both species, may be related to the reproductive behavior of this group of bees. This behavior favors gene flow within nearby populations due to the migration of males from several colonies to form male congregation areas. This reproductive strategy is used by S. mexicana (Kraus et al. 2008) and it is an effective mechanism to avoid inbreeding (Mueller et al. 2012). Such homogeneity contrasts with the population structure observed in the Pacific and Atlantic clusters of each species, which may be explained by the influence of a geographical barrier to their distribution, the Sierra Madre, as seen in other Meliponini (May-Itzá et al. 2010, 2012), Hemiptera (Dorn et al. 2009), Coleoptera (Anducho-Reyes et al. 2008) and even in terrestrial birds within the same area (Álvarez and Morrone 2004; Yáñez-Ordóñez et al. 2008). Furthermore, molecular data indicate that recent evolutionary processes like IBD also affect the genetic diversity observed within the Sm2 and Sp2 colonies sampled at two extremes of the Atlantic axis. By contrast, the lack of IBD in Pacific Sm1 and Sp1 is probably due to the close proximity of the colonies. In addition, the lack of a significant relationship between geographic and morphometric distances suggests that wing shape may not be a neutral marker and that it is affected by selective pressures (Reed and Frankham 2001). In these stingless bees, not only the presence of geographic barriers but also the limited dispersion of the colonies influences the differentiation between populations.
Stingless bees are important elements for the preservation of ecosystems as they are the most frequent visitors of many native plants, including economically important local crops in Mesoamerica (Russell et al. 2005; Morandin and Winston 2006; Ayala et al. 2013). A major threat to stingless bees is the loss of tropical forests, which implies a reduction of potential nesting sites and foraging areas (Foley et al. 2005; Venturieri 2009; May-Itzá et al. 2010). Deforestation may drastically affect the effective size of bee populations as they remain isolated in fragments (Brown and Albrecht 2001). The effect of landscape fragmentation may be more drastic on stingless bees as these insects have some of the lowest dispersal rates, reflecting the mother–daughter colony attachment that persists in the process of swarming (Engels and Imperatriz-Fonseca 1990). This feature of stingless bee reproduction could become maladaptive when habitats are destroyed, as colonies and individuals may be unable to bridge the deteriorated landscapes (Zayed 2009). Indiscriminate use of pesticides is another factor that may affect stingless bee populations, especially those that frequently visit crops (Valdovinos-Núñez et al. 2009). The effect of human-induced translocation of colonies is also particularly relevant to species exploited for honey production (Jaffé et al. 2016), as is the case of Scaptotrigona. Avoiding the movement of colonies is crucial to preserve the genetic integrity of ecotypes adapted to particular geographic areas. Thus, it is also important to educate farmers and other individuals through conservation programs that promote the preservation of natural habitats for the reproduction of colonies.
In conclusion, the combined use of genetic and morphological techniques indicates that only the S. mexicana colonies from both coasts are distinct evolutionary significant units. A requisite to establish valid species affirmation is congruence among several independent lines of evidence (Schlick-Steiner et al. 2010). Accordingly, further studies that include data from more samples that more extensively cover the distribution area of these two ESUs and that address other aspects of their biology will be necessary to confirm their taxonomic status. In any case, the results presented here favor the promotion of conservation measures not only for the Sm1 and Sm2 populations but also, for the Sp1 and Sp2 populations, which should be treated as separate units in order to avoid inbreeding and a loss of diversity (González-Acereto et al. 2006). Programs and strategies focused on maintaining the diversity of these bees should be set-up in order to preserve their genetic diversity.
References
Albores-González ML, García Guerra TG, Durán Olguín L, Aguliar Ayón A (2011) Experiencia de la Unión de Cooperativas Tosepan en el fomento a la cría de las abejas nativas Pitsilnejmej (Scaptotrigona mexicana) in Memorias del VII Seminario Mesaomerciano sobre abejas nativas, pp 95–99
Álvarez ME, Morrone JJ (2004) Propuesta de áreas para la conservación de aves de México, empleando herramientas panbiogeográficas e índices de complementariedad. Interciencia 29:112–120
Anducho-Reyes MA, Cognato AI, Hayes JL, Zuniga G (2008) Phylogeography of the bark beetle Dendroctonus mexicanus Hopkins (Coleoptera: Curculionidae: Scolytinae). Mol Phylogenet Evol 49:930–940
Arias MC, Brito RM, Francisco FO, Moretto G, Oliveira FF, Silvestre D, Sheppard WS (2006) Molecular markers as a tool for population and evolutionary studies of stingless bee. Apidologie 37:259–274
Ayala R (1999) Revisión de las abejas sin aguijón de México (Hymenoptera, Apoidea). Folia Entomol Mex 106:1–123
Ayala R, González VH, Engels MS (2013) Mexican stingless bees (Hymenoptera: Apidae): diversity, distribution, and indigenous knowledge. In: Vit P, Pedro SRM, Roubik DW (eds) Pot-Honey: a legacy of stingless bees. Springer, New York, pp 135–152
Balloux F, Lugon-Moulin N (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165
Barkan NP, Aytekin AM (2013) Systematical studies on the species of the subgenus Bombus (Thoracobombus) (Hymenoptera: Apidae, Bombus Latreille) in Turkey. Zootaxa 3737(2):167–183
Bonatti V, Simões ZLP, Franco FF, Francoy TM (2014) Evidence of at least two evolutionary lineages in Melipona subnitida (Apidae, Meliponini) suggested by mtDNA variability and geometric morphometrics of forewings. Naturwissenschaften 101:17–24
Bookstein FL (1991) Morphometric tools for landmark data. Cambridge University Press, Cambridge
Borges AA, Campos LAO, Salomão TMF, Tavares MG (2010) Genetic variability in five populations of Partamona helleri (Hymenoptera: Apidae) from Minas Gerais State, Brazil. Genet Mol Biol 33:781–784
Bouga M, Alaux C, Bienkowska M, Büchler R, Carreck NL, Cauia E, Chlebo R, Dahle B, Dall’Olio R, De la Rúa P, Gregorc A, Ivanova E, Kence A, Kence M, Kezic N, Kiprijanovska H, Kozmus P, Kryger P, Le Conte Y, Lodesani M, Murilhas AM, Siceanu A, Soland G, Uzunov A, Wilde J (2011) A review of methods for discrimination of honey bee populations as applied to European beekeeping. J Apic Res 50(1):51–84
Brown JC, Albrecht C (2001) The effect of tropical deforestation on stingless bees of the genus Melipona (Insecta: Hymenoptera: Apidae: Meliponini) in central Rondonia, Brazil. J Biogeogr 28:623–624
Brown MJF, Paxton RJ (2009) The conservation of bees: a global perspective. Apidologie 40:410–416
Byatt MA, Chapman NC, Latty T, Oldroyd BP (2016) The genetic consequences of the anthropogenic movement of social bees. Insectes Soc 63(1):15–24
Camargo JMF, Pedro SRM (2013) Meliponini Lepeletier, 1836. In: Moure JS, Urban D, Melo GAR. (Orgs). Catalogue of bees (Hymenoptera, Apoidea) in the Neotropical Region-online version; [cited 2012 Sep 29]. http://www.moure.cria.org.br/catalogue. Accessed 23 Sept 2015
da Silva FL, Sella MLG, Francoy TM, Costa AHR (2015) Evaluating classification and feature selection techniques for honeybee subspecies identification using wing images. Comput Electron Agric 114:68–77
Dorn PL, Calderon C, Melgar S, Moguel B, Solorzano E, Dumonteil E, Rodas A, De la Rúa N, Garnica R, Monroy C (2009) Two distinct Triatoma dimidiata (Latreille, 1811) taxa are found in sympatry in Guatemala and Mexico. PLoS Negl Trop Dis 3:e393
Duarte OMP, Gaiotto FA, Costa MA (2014) Genetic differentiation in the stingless bee, Scaptotrigona xanthotricha Moure, 1950 (Apidae, Meliponini): a species with wide geographic distribution in the Atlantic rainforest. J Hered 105(4):477–484
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conser Genet Res 4(2):359–361
Engels W, Imperatriz-Fonseca VL (1990) Caste development, reproductive strategies and control of fertility in honey-bees and stingless bees. In: Engels W (ed) Social insects: an evolutionary approach to castes and reproduction. Springer, Berlin, pp 166–230
Estoup A, Scholl A, Pouvreau A, Solignac M (1995) Monoandry and polyandry in bumble bees (Hymenoptera: Bombinae) as evidenced by highly variable microsatellites. Mol Ecol 4:89–93
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Res 10(3):564–567
Fernandes CRM, Martins CF, Ferreira KM, Del Lama MA (2012) Gene variation, population differentiation and sociogenetic structure of nests of Partamona seridoensis (Hymenoptera: Apidae, Meliponini). Biochem Genet 50:325–335
Ferreira VS, Aguiar CML, Costa MA, Silva JG (2011) Morphometric analysis of populations of Centris aenea Lepeletier (Hymenoptera: Apidae) from Northeastern Brazil. Neotrop Entomol 40:97–102
Foley JA, DeFries R, Asner GP, Barford C, Bonan G, Carpenter SR, Chapin FS, Coe MT, Daily GC, Gibbs HK, Helkowski JH, Holloway T, Howard EA, Kucharik CJ, Monfreda C, Patz JA, Prentice IC, Ramankutty N, Snyder PK (2005) Global consequences of land use. Science 309:570–574
Francisco FO, Brito RM, Arias MC (2006) Allele number of heterozigosity for microsatellite loci in different stingless bee species (Hymenoptera: Apidae: Meliponini). Neotrop Entomol 35:638–643
Francisco FO, Nunes-Silva P, Francoy TM, Wittmann D, Imperatriz-Fonseca VL, Arias MC, Morgan ED (2008) Morphometrical, biochemical and molecular tools for assessing biodiversity. An example in Plebeia remota (Holmberg, 1903) (Apidae, Meliponini). Insectes Soc 55(3):231–237
Francoy TM, Grassi ML, Imperatriz-Fonseca VL, May-Itzá WdJ, Quezada-Euán JJG (2011) Geometric morphometrics of the wing as a tool for assigning genetic lineages and geographic origin to Melipona beecheii (Hymenoptera: Meliponini). Apidologie 42:499–507
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Freitas BM, Imperatriz-Fonseca VL, Medina LM, Kleinert AMP, Galetto L, Nates-Parra G, Quezada-Euán JJG (2009) Diversity, threats and conservation of native bees in the Neotropics. Apidologie 40:332–346
Gonçalves PHP (2010) Análise da variabilidade genética de uma pequena população de Frieseomelitta varia (Hymenoptera, Apidae, Meliponini) por meio de análise do DNA mitocondrial, microssatélites e morfometria geométrica das asas. Dissertation, University of São Paulo
González-Acereto JA (2012) La importancia de la meliponicultura en México, con énfasis en la Península de Yucatán. Bioagrociencias 5(1):34–41
González-Acereto JA, Quezada-Euá JJG, Medina-Medina LA (2006) New perspectives for stingless beekeeping in the Yucatan: results of an integral program to rescue and promote the activity. J Apicult Res 45(3):234–239
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:1–9
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59:1633–1638
Hurtado-Burillo M, Ruiz C, May-Itzá WdJ, Quezada-Eúan JJG, De la Rúa P (2013) Barcoding stingless bees: genetic diversity of the economically important genus Scaptotrigona in Mesoamerica. Apidologie 44:1–10
Ish-Am G, Barrientos-Priego F, Castañeda-Vildozola A, Gazit S (1999) Avocado (Persea americana Mill) pollinators in its region of origin. Rev Chapingo Ser Hortic 5:137–143
Jaffé R, Pope N, Giannini TC, Acosta AL, Alves DA, Arias MC, Francisco FO, González A, Imperatriz-Fonseca VL, Jha S, De la Rúa P, Tavares MG, Carvalheiro LG (2016) Gene flow in stingless bees is determined by geographic distance and beekeeping practices. Mol Ecol (in press)
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Kalinowski ST (2005) HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5:187–189
Klingenberg CP (2011) MorphoJ: an integrated software package for geometric morphometrics. Mol Ecol Resour 11:353–357
Kraus FB, Weinhold S, Moritz RFA (2008) Genetic structure of drone congregations of the stingless bee Scaptotrigona mexicana. Insectes Soc 55:22–27
Leberg PL (2002) Estimating allelic richness: effects of sample size and bottlenecks. Mol Ecol 11:2445–2449
Lima Junior CA, Carvalho CAL, Nunes LA, Francoy TM (2012) Population divergence of Melipona scutellaris Latreille (Hymenoptera: Meliponina) in two Restricted Areas in Bahia, Brazil. Sociobiology 59:107–122
Luikart G, England PR (1999) Statistical analysis of microsatellite DNA data. Trends Ecol Evol 14:253–255
Mann HB, Whitney DR (1947) On a test of whether one of two random variables is stochastically larger than the other. Ann Math Stat 18(1):50–60
Manzo C (2009) Informe Final de Actividades de la Huasteca. Módulo de Abejas sin Aguijón (Scaptotrigona mexicana) Municipio de Cocoxtlán S.L.P
May-Itzá WdJ, Quezada-Eúan JJG, Medina-Medina LA, Enriquez E, De la Rúa P (2010) Morphometric and genetic differentiation in isolated populations of the endangered Mesoamerican stingless bee Melipona yucatanica (Hymenoptera: Apoidea) suggest the existence of a two species complex. Conserv Genet 11:2079–2084
May-Itzá WdJ, Quezada-Eúan JJG, Ayala R, De la Rúa P (2012) Morphometric and genetic analyses differentiate Mesoamerican populations of the endangered stingless bee Melipona beecheii (Hymenoptera: Meliponidae) and support their conservation as two separate units. J Insect Conserv 16(5):723–731
Mendes MFM, Francoy TM, Nunes-Silva P, Menezes C, Imperatriz-Fonseca VL (2007) Intra-populational variability of Nannotrigona testaceicornis Lepeletier, 1836 (Hymenoptera, Meliponini) using relative warp analysis. Biosci J 23:147–152
Michener CD (2007) The bees of the world, 2nd edn. The Johns Hopkins University Press, Baltimore
Morandin LA, Winston ML (2006) Pollinators provide economic incentive to preserve natural land in agroecosystems. Agric Ecosyst Environ 116:289–292
Mueller MY, Moritz RFA, Kraus FB (2012) Outbreeding and lack of temporal genetic structure in a drone congregation of the Neotropical stingless bee Scaptotrigona mexicana. Ecol Evol 2:1304–1311
Nunes LA, Passos GB, Carvalho CAL, Araújo ED (2013) Spatial variation of size and shape of the wing in Melipona quadrifasciata anthidioides Lepeletier, 1836 (Hymenoptera; Meliponini) assessed by geometric morphometrics. Braz J Biol 73:887–893
Oleksa A, Tofilski A (2015) Wing geometric morphometrics and microsatellite analysis provide similar discrimination of honey bee subspecies. Apidologie 46(1):49–60
Paxton RJ, Weibschuh N, Quezada-Euán JJG (1999) Characterization of dinucleotide microsatellite loci for stingless bees. Mol Ecol 8:685–702
Peakall R, Smouse PE (2006) GENEALEX 6: genetic analysis in Excel, Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peters JM, Queller DC, Imperatriz-Fonseca VL, Strassmann JE (1998) Microsatellite loci for stingless bees. Mol Ecol 7:783–792
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quezada-Euán JJG, May-Itzá WdJ, Rincón M, De la Rúa P, Paxton RJ (2012) Genetic and phenotypic differentiation in endemic Scaptotrigona hellwegeri (Apidae: Meliponini): implications for the conservation of stingless bee populations in contrasting environments. Insect Conserv Divers 5(6):433–443
Raymond M, Rousset F (1995a) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Raymond M, Rousset F (1995b) An exact test for population differentiation. Evolution 49:1283–1286
Reed DH, Frankham R (2001) How closely correlated are molecular and quantitative measures of genetic variation? A meta-analysis. Evolution 55:1095–1103
Rice W (1989) Analysing tables of statistical tests. Evolution 43:223–225
Rohlf FJ (2013) TPSdig v. 2.17. NY State University at Stony Brook, Stony Brook
Rosenberg NA (2004) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Russell KN, Ikerd H, Droege S (2005) The potential conservation value of unmowed powerline strips for native bees. Biol Conserv 124:133–148
Santiago LR, Francisco FO, Jaffé R, Arias MC (2016) Genetic variability in captive populations of the stingless bee Tetragonisca angustula. Genetica 144:397–405
Schlick-Steiner BC, Steiner FM, Seifert B, Stauffer C, Christian E, Crozier RH (2010) Integrative taxonomy: a multisource approach to exploring biodiversity. Annu Rev Entomol 55:421–438
Slaa EJ, Sánchez LA, Malagodi-Braga KS, Hofstede FE (2006) Stingless bees in applied pollination: practice and perspectives. Apidologie 37:293–315
Stout JC, Morales CL (2009) Ecological impacts of invasive alien species on bees. Apidologie 40:388–409
Tavares MG, Dias LAD, Borges AA, Lopes DM, Busse AHP, Costa RG, Salomão TMF, Campos LAO (2007) Genetic divergence between populations of the stingless bee uruçu amarela (Melipona rufiventris group, Hymenoptera, Meliponini): is there a new Melipona species in the Brazilian state of Minas Gerais? Gen Mol Biol 30:667–675
Valdovinos-Núñez GR, Quezada-Euán JJG, Ancona-Xiu P, Moo-Valle H, Carmona A, Ruiz Sánchez E (2009) Comparative toxicity of pesticides to stingless bees (Hymenoptera: Apidae: Meliponini). J Econ Entomol 102:1737–1742
Van Oosterhout C, Hutchinson WF, Wills DP, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Venturieri GC (2009) The impact of forest exploitation on Amazonian stingless bees (Apidae, Meliponini). Genet Mol Res 8:684–689
Yáñez-Ordóñez O, Trujano Ortega A, Llorente Bousquets J (2008) Patrones de distribución de las especies de la tribu Meliponini (Hymenoptera: Apoidea: Apidae) en México. Interciencia 33(1):41–45
Zayed A (2009) Bee genetics and conservation. Apidologie 40:237–262
Acknowledgments
We appreciate the stingless beekeepers and Dr. Remy Vandame that provided samples. We also thank the editor and three anonymous reviewers for their comments which helped improve the manuscript and BiomedRed for English editing. This research has been funded by the Regional Government of Murcia (Fundación Séneca; Grant Number 19908/GERM/2015) and Conacyt (“CONSERVACION DE LAS ABEJAS SIN AGUIJON DE MEXICO” Grant Number 103341). Dr. De la Rúa is presently member and receives support from COST Action FA1307, Sustainable pollination in Europe: joint research on bees and other pollinators, SUPER-B (http://www.cost.eu/COST_Actions/fa/Actions/FA1307).
Author information
Authors and Affiliations
Corresponding author
Additional information
Miguel Hurtado-Burillo and Laura Jara have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Hurtado-Burillo, M., Jara, L., May-Itzá, W. et al. A geometric morphometric and microsatellite analyses of Scaptotrigona mexicana and S. pectoralis (Apidae: Meliponini) sheds light on the biodiversity of Mesoamerican stingless bees. J Insect Conserv 20, 753–763 (2016). https://doi.org/10.1007/s10841-016-9899-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10841-016-9899-1