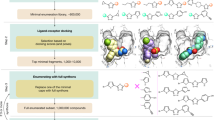
Abstract
Although the size of virtual libraries of synthesizable compounds is growing rapidly, we are still enumerating only tiny fractions of the drug-like chemical universe. Our capability to mine these newly generated libraries also lags their growth. That is why fragment-based approaches that utilize on-demand virtual combinatorial libraries are gaining popularity in drug discovery. These à la carte libraries utilize synthetic blocks found to be effective binders in parts of target protein pockets and a variety of reliable chemistries to connect them. There is, however, no data on the potential impact of the chemistries used for making on-demand libraries on the hit rates during virtual screening. There are also no rules to guide in the selection of these synthetic methods for production of custom libraries. We have used the SAVI (Synthetically Accessible Virtual Inventory) library, constructed using 53 reliable reaction types (transforms), to evaluate the impact of these chemistries on docking hit rates for 40 well-characterized protein pockets. The data shows that the virtual hit rates differ significantly for different chemistries with cross coupling reactions such as Sonogashira, Suzuki–Miyaura, Hiyama and Liebeskind–Srogl coupling producing the highest hit rates. Virtual hit rates appear to depend not only on the property of the formed chemical bond but also on the diversity of available building blocks and the scope of the reaction. The data identifies reactions that deserve wider use through increasing the number of corresponding building blocks and suggests the reactions that are more effective for pockets with certain physical and hydrogen bond-forming properties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Screening of virtual libraries of synthesizable compounds has become an increasingly important step in drug discovery [1, 2]. The surge in utilization of computational approaches has been stimulated by improvements in binding energy calculations, the growth of computational resources, advances in protein structures determination and availability of large and diverse virtual libraries of compounds [3,4,5,6,7,8,9,10,11,12,13,14,15]. However, our ability to access the vast druggable chemical space is still limited and will be impacted by the limits of computing resources for the foreseeable future [16,17,18,19]. We have the potential of generating trillions or more of virtual synthesizable molecules, but enumerating these chemical spaces, and thus converting them into screenable files is impractical if not impossible in practice. That is why fragment-based approaches that enumerate only parts of the chemical spaces, thus generating on-demand virtual combinatorial libraries are widely used [10, 20,21,22]. Most fragment-based methods identify synthetic blocks binding to sub-pocket(s) of a larger protein pocket first and then virtually “synthesize” libraries containing these blocks [8, 10, 23]. However, many different chemistries can be used for combining a particular set of blocks, and there are currently no time- and resources-saving guidelines for selection of reactions. To evaluate the impact of chemistries on the number of hits obtained through virtual docking, we have used the recently generated Synthetically Accessible Virtual Inventory (SAVI). SAVI comprises nearly 1.75 billion virtual molecules, each with a proposed synthesis scheme. It was constructed from 155,129 building blocks provided by Enamine (Kyiv, Ukraine, enamine.net) using robust chemistries encoded in 53 transforms [24]. SAVI transforms were written into rules based on an adaptation and extension of the CHMTRN/PATRAN programming languages describing chemical synthesis expert knowledge [25], which were originally developed in the context of the LHASA project [26]. We point out that the version of SAVI used for this study, SAVI-2020, consists of only single-step reactions, i.e. follows the scheme R1 + R2-> P (R1, R2: reactants; P: product). We note the terminology used here: We call the general reaction type (typically a "named reaction") a "chemistry" in the context of SAVI, whereas the individual CHMTRN/PATRAN rules are called "transforms." For example, SAVI uses the Suzuki–Miyaura cross-coupling chemistry, which is expressed in 6 different transforms (bromo, iodo, alkene cross-coupling etc.). Transforms have a descriptive name but also a four-digit number, which will frequently be used in the following. All 53 transforms can be downloaded from [27]. The cheminformatics toolkit CACTVS [28] was used to apply these rules for the virtual synthesis of the entries. By now, 169 SAVI compounds have been synthesized by us, Enamine LLC and the Medicinal Chemistry group of the National Center for Advancing Translational Sciences, NIH. These are the SAVI molecules whose syntheses we are aware of. Since the entire SAVI database can be freely downloaded without user registration, more SAVI compounds may have been synthesized elsewhere without our knowledge. SAVI’s predictions of synthetic accessibility were found to be more than 95% accurate. Enamine provides a database called REAL, which could be called a "sister" of SAVI since it is constructed from essentially the same set of building blocks. The mutual overlap between these two ultra-large databases, which in 2020 had somewhat over 1 billion molecules each, was only about 10%. Due to the essentially identical building block sets used for SAVI vs. REAL, the only plausible explanation was the difference in chemistries applied for the generation of the entries [24]. Unexpectedly, we found significant differences in the number of virtual hits when docking approximately equal-sized SAVI or REAL diversity sets (each about 3 million compounds in size) into the same protein pocket. This observation provided the impetus for this study. Since these two databases were constructed from the same building blocks, and the major differences were in the reactions used, we hypothesized that linking chemistries may favor certain pockets but not the others.
A systematic investigation of the impact of chemistries on the number of virtual hits is particularly important as the ultra-large libraries continue to grow rapidly. We have expanded the number of reliable transforms that can be used for SAVI generation to more than 120. The number of commercially available synthesis building blocks has also increased. Enamine alone has now 1 billion made-on-demand blocks (MADE) [29]. Consequently, the next version of SAVI could represent trillions of molecules. The expansion of the accessible chemical space is a welcome trend that is likely to improve and accelerate drug discovery. However, enumeration and screening of such databases and their use in their entirety will remain impossible for the foreseeable future. Enumeration of only specific parts of accessible chemical space (so-to-speak the "optimal chunks" of the space) that maximize success in screening is therefore likely to continue to be widely used in the future. Consequently, to enumerate the most appropriate part of the chemical space for a given target, it would be helpful to know not only which building blocks are better suited for the target pocket, but also which coupling chemistries are more likely to generate high-scoring virtual hits.
To evaluate the possible correlation between pocket properties and transforms used for library generation, we performed docking of a SAVI diversity set, containing about 3 million structures collected from all 53 transforms listed in Table 2, into 40 protein pockets (Table 1). We conducted the analysis around the chemistries generating the compounds rather than structural motifs they produce because of practical considerations: During the generation of SAVI, filtering by the CHMTRN reaction logic occurs, which for SAVI-2020 excluded about 50% of the initially formed reactant pairs because the proposed structures could not be made in a one-step reaction. This filtering influences the set of resulting structures and their ability to interact with proteins. Thus, the sets of compounds containing the same structural motif generated by different synthetic methods will be different because of different exclusion rules in the individual transforms. This is why we analyze the chemistries separately, focusing on the reaction rather than on the structural motif it produces.
Results and discussion
Target pockets (Table 2) were selected from PDB to represent two pocket types: small molecule (SM) pockets and protein–protein interaction (PPI) pockets. The majority of selected pockets bind well-characterized ligands that have advanced into the clinics. However, we also included several less studied but interesting and potentially impactful pockets that either are difficult to target, or which represent surfaces involved in protein–protein interactions. PPI-based inhibitors and modulators are the type of therapeutics that the scientific community is increasingly aiming at. To ensure that the structures were suitable for virtual screening, the ligands present in the chosen complexes were redocked into the corresponding pockets, and only structures that allowed for correct prediction of ligand poses were included in the analysis. For docking and virtual screens, we used the ICM-Pro software (Molsoft, San Diego, CA). Although the software has been benchmarked before [30,31,32], we have evaluated the correctness of docking poses for the pockets with known ligands (Table 1). Most of docked complexes had RMSD < 1 Å when compared to the experimental structures. In two cases where RMSD exceeded 1 Å (PDB:5i96 and 5vv0), all the differences were in the part of the molecules exposed to the solvent, while poses inside the pocket were determined with high accuracy. Although the structures used appeared to predict binding correctly when tested with “native” ligands, the presence of false positives in virtual screens is inevitable [33]. Testing binding properties experimentally for all identified virtual hits was not possible in the context of this study. We were able to verify binding for several top virtual hits for eight targets, which are currently studied in our lab, two of which have recently been published [34, 35]. It is reasonable to suggest, however, and the data generated for the eight targets confirms, that percentage of noise or false positives for a particular pocket is distributed evenly across the database and does not depend on the transform. Consequently, virtual hit rates can serve as surrogates of number of binders.
We chose the SAVI diversity set containing 2,955,416 compounds for the exploration because of practical considerations. Docking the entire SAVI database into just one pocket would take more than 280 days when running 1000 parallel processes on the NIH supercomputer cluster. Docking of the diversity set into one pocket requires around 50,000 CPU Hours, which is doable on a computer cluster. Although docking of larger sets may allow for more sensitive detection of differences between different transforms, it would require prohibitively large computational resources when used for the multiple pockets that we aimed to evaluate for this study.
Remarkably, virtual hit rates across 40 targets differed significantly between different transforms (Fig. 1, Table 2). Several transforms had to be excluded from further analysis because they were represented by too few compounds in SAVI as well as in the diversity dataset. This underrepresentation occurs due to the low number of available synthetic blocks that are needed for these "starved" transforms. The following “starved” transforms were excluded from the analysis because they produced less than 10000 compounds for the entire SAVI: Feist synthesis of pyrroles (1039), [2 + 2]-cycloaddition of allenes to alkenes (1391), synthesis of tetrazoles from azide and nitriles (2218), benzo[b]furans synthesis from 2-hydroxyphenyl acetylenes (2684), Hiyama aryl-alkenyl cross-coupling (6013), Hiyama allyl cross-coupling (6015), Hiyama cross-coupling with arylhydrazine (6017) and Liebeskind-Srogl nitrile formation (6024) (Table 1). The number of available building blocks for each transform can be found at: https://www.nature.com/articles/s41597-020-00727-4/tables/7. Several transforms had sufficient representation in the database but could not be used for reliable evaluation because they produced too few virtual hits across all tested targets and zero hits for many of them. Transforms that had to be excluded for this reason were: pyrazole synthesis from beta carbonyl carboxylic acid derivatives (1439), synthesis of fused aryl(2,3-H/R) pyridines by Pictet-Spengler reaction (2238), Kabbe synthesis of 4-chromanones (2269), Mitsunobu carbon–carbon bond formation (6032), and sulfonamide acylation (7018). Thus, they may be less valuable for the current drug discovery efforts in general (Fig. 1, Table 1). The weak performance of some of these transforms can be attributed to the small number of compounds for one of the two building blocks needed by the transform. Although instances of the second type of blocks needed (R2) could be plentiful in the building block set and the number of generated compounds therefore relatively large, the overall diversity of the products is limited if the R1 subset consists of, say, fewer than hundred compounds (and those may be structurally closely related). Transforms 1439, 2269 and 6032 are examples of such cases: https://www.nature.com/articles/s41597-020-00727-4/tables/7. Remarkably, several chemistries produced subsets with very high virtual hit rates. Suzuki–Miyaura cross-couplings (6004, 6005 and 6006) were among the most productive ones. Interestingly, Suzuki–Miyaura coupling is among the most frequently used reactions in current medicinal chemistry [36]. Our data shows that this chemistry deserves the attention it receives. However, the most frequently used reaction, amide bond formation (transform 6038) [36], was less productive with a virtual hit rate that was roughly three times lower than that for Suzuki–Miyaura cross-couplings.
Virtual hit rates for 53 transforms used for SAVI generation. The hits were identified by docking 2,955,416 compounds of SAVI diversity set into 39 well characterized protein pockets. To compensate for differences in the occurrence rate of a particular transform in the diversity set, the total number of virtual hits for each transform has been normalized by dividing it by the number of compounds produced by the transform in the screening library
This should not be interpreted as a lower general usefulness of amides in drug discovery. This particular case emphasizes the differences in the impact of chemistries between traditional medicinal chemistry that employs multi-step synthesis and virtual libraries constructed using one- or two-step reactions. One of the possible reasons for the relatively poor performance of transform 6038 is suboptimal selectivity, which led to reduced scope, and exclusion of building blocks containing hydrogen bond donors that facilitate interactions with proteins, such as amino, carboxyl, hydroxyl and sulfonamide groups. In multi-step synthesis, protection/deprotection of these groups could preserve them, thus increase the protein interaction potential of the products. Selectivity of chemistries is likely to play a role in “productivity” of other transforms and it is an additional factor that needs to be considered during generation of custom libraries. Our data also suggest additional transforms that deserve efforts in expanding. For example, Hiyama carbonylative cross-coupling should be expanded by adding more aryl triethoxysilanes into the collection of the building blocks. Expanding the collection of arylboronic acids would benefit not only Suzuki–Miyaura cross-coupling, but also the highly productive Liebeskind–Srogl heterocyclic coupling (6025). It should be emphasized that the efficacy of a transform in producing potential virtual hits can depend not only on the properties/geometry of the bond it generates but also on reaction selectivity and diversity of the building blocks available. As discussed above, selectivity of the reaction allows to preserve the functional groups of the blocks that can be beneficial for protein binding while diversity increases the chances of finding a good fit for a particular pocket. Transforms 6004–6006 produce structurally similar di-aryl compounds through Suzuki–Miyaura cross-coupling. However, the virtual hit rates for 6004, which uses bromo aryl blocks, is about 40% higher than for 6005 or and 65% higher than for 6006 that use iodo and chloro aryls, respectively. The set of building blocks used for SAVI-2020 had a 7.3 times larger number of bromo aromatic compounds than iodo-derivatives (https://www.nature.com/articles/s41597-020-00727-4/tables/7), thus allowing for higher diversity in the products of transform 6004 vs. 6005. Chloro-aromatic blocks are even more numerous than the bromo-derivatives. However, the reaction is less selective for chloro compounds, which results in a 44 times higher number of excluded products, effectively reducing the number of useful blocks for transform 6006. The diversity of the blocks that can potentially impact virtual hit rates is likely to change with time along additional synthetic efforts in building blocks generation. Thus, virtual hit rates can be improved for less productive transforms in the future.
The pocket properties evaluated for potential impact on the number of virtual hits included volume, area, radius, hydrophobicity, nonsphericity, aromaticity, buriedness, drug-like density (DLID [37]), the numbers of hydrogen bonds donors, and the number of acceptors. The hydrogen bond forming potential of each pocket was evaluated manually. The rest of the parameters were determined using the PocketFinder function of ICM-Pro. The correlation of these properties with the number of virtual hits for each transform was determined for the entire SAVI diversity set.
The binding score produced by docking for every molecule is influenced by many factors. That is why we did not expect strong dependencies for any single parameter, but rather tendencies. That is why we include correlations that have p > 0.05. For the whole database, the number of virtual hits showed a statistically significant positive correlation (with p-value < 0.05) with properties related to pocket size: volume and radius (Fig. 2). Most of the pockets with high numbers of virtual hits had volumes between 300 and 1000 Å3, and virtual hit rates were significantly lower both below and above this range. Similarly, the graphs suggest that the most productive values of the radius are between 4 and 6.2 Å and between 300 and 900 Å2 for the pocket surface area. This can be explained by the size distribution of the database entries as it contains only limited numbers of molecules with MW < 200 and > 550 [24]. The degree of hydrophobicity of the pocket did not yield any definite trends. Surprisingly, aromaticity appeared to have negative correlation, although aromatic interactions have been suggested to contribute to ligand–protein binding [38, 39]. However, the correlation was not statistically significant.
Total number of virtual hits generated by virtual docking of SAVI diversity set into protein pockets with different properties. The parameters for each property were determined using the PocketFinder function of ICM-Pro software (Molsoft). Dotted lines represent linear trends with corresponding correlation coefficient (r), Student's t-distribution, and p-values shown
Nonsphericity and buriedness demonstrated positive correlation with the number of virtual hits (Fig. 2) but it was statistically insignificant for both parameters. The number of hydrogen bond acceptors (HBA) in the pocket did not show any significant correlation. In contrast, the number of hydrogen bond donors (HBD) had significant positive correlation with the number of docking hits (Fig. 3). The observed dependencies on HBD could be caused by prefiltering of the database building blocks for “drug-like” properties. Hydrogen bond acceptors of potential drugs are widely believed to be less detrimental than hydrogen-bond donors with regards to solubility, cell permeability and bioavailability [40]. Lipinski’s rule of 5 is more restrictive to hydrogen bond donors than to hydrogen bond acceptors allowing no more than 5 HBDs vs. up to 10 HBAs [41]. Consequently, the database will have more HBA-rich compounds that prefer HBD-rich pockets.
To compare the degrees of dependencies for different transforms, we used correlation coefficients (Tables 3 and S1). Correlations with pockets’ properties differ for different transforms (Table 3) and frequently have opposite signs. The relatively small number of pockets screened does not allow one to make statistically justified conclusions for many correlations as p-values fall short, sometimes just slightly above 0.05. The data shows that those differences do exist, and additional future screens will permit to establish comprehensive correlations. Nevertheless, several dependencies could be established. Pocket sizes showed positive correlations with virtual hit rates for all transforms, with transforms 1171, 2201, 6003, 6004, 6005, 7013, 7014 and 7022 showing the strongest correlations, suggesting that they could work better for larger pockets, but not for the small ones. Although the number of virtual hits increased with an increase of pocket buriedness and nonsphericity for the majority of transforms, only transform 2267 had statistically significant correlation with buriedness in this study.
Aromaticity had negative, but insignificant, correlation for all transforms (Table 3). Although the number of hydrogen bond acceptors in the pocket did not show any definite correlation for the whole diversity set, it demonstrated strong positive correlation for transforms 2875 (copper[I]-catalyzed azide-alkyne cycloaddition), 6031 (Mitsunobu reaction) and Wittig ketone synthesis (7020). Transform 2875 produces heterocycles with hydrogen-donating properties that can explain this trend. For transform 6031 and 7020 the reason could be the properties of the blocks that they utilize. The number of hydrogen bond donors appeared to have positive correlation with the number of virtual hits for all transforms. The strongest correlations were found for transform 7021 (Horner–Wadsworth–Emmons olefination). Hydrogen bonds are strong contributors to the binding energy. Thus, hydrogen bond-forming capacity of the pocket can be expected to have a positive effect on the number of virtual hits. However, as discussed before, prefiltering of the building blocks for “drug-like” properties, which excludes hydrogen bond donor-rich compounds to avoid cell permeability and bioavailability issues, limits the number of HBD-rich compounds, making the observed dependences less pronounced. The hydrogen bond forming capacity of a transform can be impacted by reaction selectivity. For example, both transform 7013 and 7014 generate benzimidazoles from aromatic o-diamines and aldehydes. However, 7014 uses boric acid to produce a reactive intermediate while 7013 uses molecular iodine under basic conditions. Consequently, the sets of restrictions for the starting blocks are different. As a result, 7013 generated almost twice as many compounds as 7014, but has a significantly lower overall virtual hit rate (Table 1). Nevertheless, the correlations with pocket properties are similar for these two transforms. All observed trends can assist in generation of optimally targeted virtual libraries and thus, reduce time and effort required for lead identification.
Conclusions
The results show that chemistries used for preparation of virtual libraries have substantial impact on the virtual hit rates during virtual screening. The data suggest that with the ever-expanding number of synthetically accessible compounds and limited computational resources, efforts in enumeration of virtual libraries will benefit from focusing on cross-coupling reactions such as Sonogashira, Suzuki–Miyaura, Hiyama and Liebeskind–Srogl couplings as they produce the highest numbers of virtual hits for different types of protein pockets. Transforms 1439 (pyrazoles synthesis from beta carbonyl carboxylic acid derivatives), 2238 (synthesis of fused aryl(2,3-H/R) pyridines by Pictet-Spengler reaction, 2269 (Kabbe synthesis of 4-chromanones, 6032 (Mitsunobu carbon–carbon bond formation), and 7018 (sulfonamide acylation) were among the least effective ones in generation of virtual hits for all tested pockets, and thus, can be given a lower priority in generation of custom libraries.
For larger pockets, transforms 2201, 6003, 6004, 6005, 7013 and 7014 appeared to be most effective.
The DLID (drug-like density) descriptor had a positive correlation with the number of virtual hits. The correlation was statistically significant for the entire diversity set and for 10 transforms out of 53. The lack of significant correlation for most of transforms suggests that low druggability score, in its traditional definition [37] should not discourage one from attempting virtual screens for a particular pocket as exceptions to the rule are not uncommon.
Methods
Databases
The SAVI diversity set of 2,955,416 compounds was generated from the entire SAVI-2020 database (which contains 1,748,464,003 compounds), using mini-batch k-means clustering performed with RDKit [42] and scikit-learn [43]. The Tanimoto coefficient for any two compounds in the set was < 0.6. The entire SAVI database and diversity sets are available for downloading from the SAVI download page [27].
Database docking
Docking screens were conducted using the ICM-Pro software (Molsoft L.L.C., San Diego, CA) by running 590 parallel processes (5000 compounds per job) on 590 nodes of the National Institutes of Health (NIH) Biowulf cluster supercomputer [44]. Each node contained 2 CPUs. The PocketFinder software (Molsoft) was used for the identification of the pockets. Screens were run in large-scale parallel way as so-called "swarm" jobs. The cutoff score to identify a virtual hit was set to -32 for the pockets that produced redocking scores lower than -32 for the “native” ligands of the complexes from Protein Data Bank or had no available structures of the complexes (Table 1). For the structures that produced re-docking score higher than −32 (PDBs: 3odu, 5i96, 5fhz, 4xe0, 5vv0, 5kj2, and 4ixd, Table 1) their redocking scores have been used as cutoffs. Virtual hits were extracted as Excel files. Every compound in the SAVI database has an identifier (SAVI ID) with its last four digits indicating the transform number. These numbers were used for counting virtual hits produced by every transform. Correlation coefficients, Student's t-distribution and p-values were determined using the Data Analysis function of Excel (Microsoft).
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information. The databases used in the study are freely available from NCI webpage [27].
References
Nazarova AL, Katritch V (2022) It all clicks together: in silico drug discovery becoming mainstream. Clin Transl Med 12:e766. https://doi.org/10.1002/ctm2.766
Bender BJ, Gahbauer S, Luttens A, Lyu J, Webb CM, Stein RM, Fink EA, Balius TE, Carlsson J, Irwin JJ, Shoichet BK (2021) A practical guide to large-scale docking. Nat Protoc 16:4799–4832. https://doi.org/10.1038/s41596-021-00597-z
Beroza P, Crawford JJ, Ganichkin O, Gendelev L, Harris SF, Klein R, Miu A, Steinbacher S, Klingler FM, Lemmen C (2022) Chemical space docking enables large-scale structure-based virtual screening to discover ROCK1 kinase inhibitors. Nat Commun 13:6447. https://doi.org/10.1038/s41467-022-33981-8
Danel T, Leski J, Podlewska S, Podolak IT (2023) Docking-based generative approaches in the search for new drug candidates. Drug Discov Today 28:103439. https://doi.org/10.1016/j.drudis.2022.103439
Gahbauer S, Correy GJ, Schuller M, Ferla MP, Doruk YU, Rachman M, Wu T, Diolaiti M, Wang S, Neitz RJ, Fearon D, Radchenko DS, Moroz YS, Irwin JJ, Renslo AR, Taylor JC, Gestwicki JE, von Delft F, Ashworth A, Ahel I, Shoichet BK, Fraser JS (2023) Iterative computational design and crystallographic screening identifies potent inhibitors targeting the Nsp3 macrodomain of SARS-CoV-2. Proc Natl Acad Sci U S A 120:e2212931120. https://doi.org/10.1073/pnas.2212931120
Grygorenko OO, Radchenko DS, Dziuba I, Chuprina A, Gubina KE, Moroz YS (2020) Generating multibillion chemical space of readily accessible screening compounds. iScience 23:101681. https://doi.org/10.1016/j.isci.2020.101681
Lyu J, Wang S, Balius TE, Singh I, Levit A, Moroz YS, O’Meara MJ, Che T, Algaa E, Tolmachova K, Tolmachev AA, Shoichet BK, Roth BL, Irwin JJ (2019) Ultra-large library docking for discovering new chemotypes. Nature 566:224–229. https://doi.org/10.1038/s41586-019-0917-9
Muller J, Klein R, Tarkhanova O, Gryniukova A, Borysko P, Merkl S, Ruf M, Neumann A, Gastreich M, Moroz YS, Klebe G, Glinca S (2022) Magnet for the needle in haystack: “crystal structure first” fragment hits unlock active chemical matter using targeted exploration of vast chemical spaces. J Med Chem 65:15663–15678. https://doi.org/10.1021/acs.jmedchem.2c00813
Perebyinis M, Rognan D (2023) Overlap of on-demand ultra-large combinatorial spaces with on-the-shelf drug-like libraries. Mol Inform 42:e2200163. https://doi.org/10.1002/minf.202200163
Sadybekov AA, Sadybekov AV, Liu Y, Iliopoulos-Tsoutsouvas C, Huang XP, Pickett J, Houser B, Patel N, Tran NK, Tong F, Zvonok N, Jain MK, Savych O, Radchenko DS, Nikas SP, Petasis NA, Moroz YS, Roth BL, Makriyannis A, Katritch V (2022) Synthon-based ligand discovery in virtual libraries of over 11 billion compounds. Nature 601:452–459. https://doi.org/10.1038/s41586-021-04220-9
Gentile F, Yaacoub JC, Gleave J, Fernandez M, Ton AT, Ban F, Stern A, Cherkasov A (2022) Artificial intelligence-enabled virtual screening of ultra-large chemical libraries with deep docking. Nat Protoc 17:672–697. https://doi.org/10.1038/s41596-021-00659-2
Singh I, Seth A, Billesbolle CB, Braz J, Rodriguiz RM, Roy K, Bekele B, Craik V, Huang XP, Boytsov D, Pogorelov VM, Lak P, O’Donnell H, Sandtner W, Irwin JJ, Roth BL, Basbaum AI, Wetsel WC, Manglik A, Shoichet BK, Rudnick G (2023) Structure-based discovery of conformationally selective inhibitors of the serotonin transporter. Cell 186:e17. https://doi.org/10.1016/j.cell.2023.04.010
Marin E, Kovaleva M, Kadukova M, Mustafin K, Khorn P, Rogachev A, Mishin A, Guskov A, Borshchevskiy V (2023) Regression-based active learning for accessible acceleration of ultra-large library docking. J Chem Inf Model. https://doi.org/10.1021/acs.jcim.3c01661
Potlitz F, Link A, Schulig L (2023) Advances in the discovery of new chemotypes through ultra-large library docking. Expert Opin Drug Discov 18:303–313. https://doi.org/10.1080/17460441.2023.2171984
Clyde A, Liu X, Brettin T, Yoo H, Partin A, Babuji Y, Blaiszik B, Mohd-Yusof J, Merzky A, Turilli M, Jha S, Ramanathan A, Stevens R (2023) AI-accelerated protein-ligand docking for SARS-CoV-2 is 100-fold faster with no significant change in detection. Sci Rep 13:2105. https://doi.org/10.1038/s41598-023-28785-9
Lyu J, Irwin JJ, Shoichet BK (2023) Modeling the expansion of virtual screening libraries. Nat Chem Biol. https://doi.org/10.1038/s41589-022-01234-w
Warr WA, Nicklaus MC, Nicolaou CA, Rarey M (2022) Exploration of ultralarge compound collections for drug discovery. J Chem Inf Model 62:2021–2034. https://doi.org/10.1021/acs.jcim.2c00224
Kontoyianni M (2022) Library size in virtual screening: is it truly a number’s game? Expert Opin Drug Discov 17:1177–1179. https://doi.org/10.1080/17460441.2022.2130244
Popov KI, Wellnitz J, Maxfield T, Tropsha A (2024) HIt discovery using docking ENriched by GEnerative modeling (HIDDEN GEM): a novel computational workflow for accelerated virtual screening of ultra-large chemical libraries. Mol Inform 43:e202300207. https://doi.org/10.1002/minf.202300207
Andrianov GV, Gabriel Ong WJ, Serebriiskii I, Karanicolas J (2021) Efficient hit-to-lead searching of kinase inhibitor chemical space via computational fragment merging. J Chem Inf Model 61:5967–5987. https://doi.org/10.1021/acs.jcim.1c00630
Meyenburg C, Dolfus U, Briem H, Rarey M (2023) Galileo: three-dimensional searching in large combinatorial fragment spaces on the example of pharmacophores. J Comput Aided Mol Des 37:1–16. https://doi.org/10.1007/s10822-022-00485-y
Zhou H, Cao H, Skolnick J (2021) FRAGSITE: a fragment-based approach for virtual ligand screening. J Chem Inf Model 61:2074–2089. https://doi.org/10.1021/acs.jcim.0c01160
Galyan SM, Ewald CY, Jalencas X, Masrani S, Meral S, Mestres J (2022) Fragment-based virtual screening identifies a first-in-class preclinical drug candidate for Huntington’s disease. Sci Rep 12:19642. https://doi.org/10.1038/s41598-022-21900-2
Patel H, Ihlenfeldt WD, Judson PN, Moroz YS, Pevzner Y, Peach ML, Delannee V, Tarasova NI, Nicklaus MC (2020) SAVI, in silico generation of billions of easily synthesizable compounds through expert-system type rules. Sci Data 7:384. https://doi.org/10.1038/s41597-020-00727-4
Judson PN, Ihlenfeldt WD, Patel H, Delannee V, Tarasova N, Nicklaus MC (2020) Adapting CHMTRN (CHeMistry TRaNslator) for a new use. J Chem Inf Model 60:3336–3341. https://doi.org/10.1021/acs.jcim.0c00448
Corey EJH, Pensak DA (1974) Computer-assisted synthetic analysis. methods for machine generation of synthetic intermediates involving multistep look-ahead. J Am Chem Soc 96:7724–7737
Synthetically Accessible Virtual Inventory (SAVI) database, https://cactus.nci.nih.gov/download/savi_download/.
Ihlenfeldt WD, Takahashi Y, Abe H, Sasaki S (1994) Computation and management of chemical properties in CACTVS: an extensible networked approach toward modularity and compatibility. J Chem Inf Comput Sci 34:109–116. https://doi.org/10.1021/ci00017a013
Enamine MADE Building Blocks: https://enamine.net/building-blocks/made-building-blocks. Accessed 22 Jan 2024.
Lam PC, Abagyan R, Totrov M (2019) Macrocycle modeling in ICM: benchmarking and evaluation in D3R grand challenge 4. J Comput Aided Mol Des 33:1057–1069. https://doi.org/10.1007/s10822-019-00225-9
Lam PC, Abagyan R, Totrov M (2019) Hybrid receptor structure/ligand-based docking and activity prediction in ICM: development and evaluation in D3R grand challenge 3. J Comput Aided Mol Des 33:35–46. https://doi.org/10.1007/s10822-018-0139-5
Scarpino A, Ferenczy GG, Keseru GM (2018) Comparative evaluation of covalent docking tools. J Chem Inf Model 58:1441–1458. https://doi.org/10.1021/acs.jcim.8b00228
Wang L, Shi SH, Li H, Zeng XX, Liu SY, Liu ZQ, Deng YF, Lu AP, Hou TJ, Cao DS (2023) Reducing false positive rate of docking-based virtual screening by active learning. Brief Bioinform. https://doi.org/10.1093/bib/bbac626
Bonilla PA, Hoop CL, Stefanisko K, Tarasov SG, Sinha S, Nicklaus MC, Tarasova NI (2023) Virtual screening of ultra-large chemical libraries identifies cell-permeable small-molecule inhibitors of a “non-druggable” target, STAT3 N-terminal domain. Front Oncol. https://doi.org/10.3389/fonc.2023.1144153
Lu X, Sabbasani VR, Osei-Amponsa V, Evans CN, King JC, Tarasov SG, Dyba M, Das S, Chan KC, Schwieters CD, Choudhari S, Fromont C, Zhao Y, Tran B, Chen X, Matsuo H, Andresson T, Chari R, Swenson RE, Tarasova NI, Walters KJ (2021) Structure-guided bifunctional molecules hit a DEUBAD-lacking hRpn13 species upregulated in multiple myeloma. Nat Commun 12:7318. https://doi.org/10.1038/s41467-021-27570-4
Brown DG, Bostrom J (2016) Analysis of past and present synthetic methodologies on medicinal chemistry: where have all the new reactions gone? J Med Chem 59:4443–4458. https://doi.org/10.1021/acs.jmedchem.5b01409
Sheridan RP, Maiorov VN, Holloway MK, Cornell WD, Gao YD (2010) Drug-like density: a method of quantifying the “bindability” of a protein target based on a very large set of pockets and drug-like ligands from the protein data bank. J Chem Inf Model 50:2029–2040. https://doi.org/10.1021/ci100312t
Brylinski M (2018) Aromatic interactions at the ligand-protein interface: Implications for the development of docking scoring functions. Chem Biol Drug Des 91:380–390. https://doi.org/10.1111/cbdd.13084
Li S, Xu Y, Shen Q, Liu X, Lu J, Chen Y, Lu T, Luo C, Luo X, Zheng M, Jiang H (2013) Non-covalent interactions with aromatic rings: current understanding and implications for rational drug design. Curr Pharm Des 19:6522–6533. https://doi.org/10.2174/13816128113199990440
Kenny PW (2022) Hydrogen-bond donors in drug design. J Med Chem 65:14261–14275. https://doi.org/10.1021/acs.jmedchem.2c01147
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341. https://doi.org/10.1016/j.ddtec.2004.11.007
RDKit, https://www.rdkit.org/.
Scikit-learn, https://scikit-learn.org/stable/.
Biowulf: high performance computing at NIH, https://hpc.nih.gov/. Accessed 30 Jan 2024
Acknowledgements
This work utilized the computational resources of the NIH HPC Biowulf cluster [44]. We thank Megan L. Peach for generation of the SAVI diversity sets.
Funding
Open access funding provided by the National Institutes of Health. This research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, CCR.
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Song, R.X., Nicklaus, M.C. & Tarasova, N.I. Correlation of protein binding pocket properties with hits’ chemistries used in generation of ultra-large virtual libraries. J Comput Aided Mol Des 38, 22 (2024). https://doi.org/10.1007/s10822-024-00562-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10822-024-00562-4