Abstract
Binding free energy calculations are increasingly used in drug discovery research to predict protein-ligand binding affinities and to prioritize candidate drug molecules accordingly. It has taken decades of collective effort to transform this academic concept into a technology adopted by the pharmaceutical and biotech industry. Having personally witnessed and taken part in this transformation, here I recount the (incomplete) list of problems that had to be solved to make this computational tool practical and suggest areas of future development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The theoretical foundation of binding free energy calculations (BFE)—free energy perturbation (FEP)—was laid down by Zwanzig [1] in the 1950s, and later refined by Bennett, who derived the optimal analysis to estimate the free energy differences from simulations [2], and others [3]. A related technique for free energy calculations, thermodynamic integration (TI), was invented by Kirkwood even earlier [4]. In the 1980s, a number of groups demonstrated that such free energy methods could be used to compute the hydration free energies of small molecule solutes [5, 6] and the binding free energies between protein receptors and small molecule ligands [7,8,9,10,11,12]. Techniques were introduced to compute either the individual binding free energy between a ligand and a receptor (by so-called “absolute” binding free energy calculations, or ABFE for short) [13, 14], or the difference in the binding free energies between two ligands against the same receptor (by “relative” binding free energy calculations, or RBFE) [15, 16]. In the early days, however, BFE calculations were hard to set up and they took a long time to run, and they seemed a long way away from commonplace utility in drug discovery.
The simple and elegant theoretical foundation for BFE belies the subtleties in performing correct and efficient calculations. The statistical precision of an FEP calculation depends on the extent of change in the equilibrium distribution of molecular configurations from the initial state to the end state of the alchemical transformations: the smaller the change, the higher the precision [17]. Key to efficient BFE calculations is to limit this change by restraining the ligand in position and in conformation during the transformations, in such a way that the restraints’ contribution to the free energies can be accounted for [13, 18]. A general set of criteria for setting the restraints can be derived by separability of integrals in the partition functions.
One by one, the technical challenges of performing correct and precise BFE calculations have been resolved by a number of, primarily academic, groups. We have learned how to avoid numerical instabilities in BFE calculations by the introduction of softcore potentials [19, 20], how to treat ligands with net charges [21, 22], how to enhance the sampling of the ligand binding pose and the conformation of the binding pocket [23,24,25,26], how to treat the non-negligible contribution of the omitted dispersion interactions between atoms beyond the cutoff distance by a mean field approximation [27], and how to best analyze the results [3, 28] and estimate the statistical errors [29]. BFE has been validated against the independent method of computing binding affinities by long molecular dynamics (MD) simulations of reversible protein-ligand binding [22]. The best practices for BFE are summarized in a recent review [30]. Academic drug hunters—and a few industrial early adopters—have developed their own BFE solutions and successfully applied BFE in identifying potent drug candidates [31,32,33,34,35,36,37]. Expertise in BFE, however, was necessary in such early successes.
To bring binding free energy calculations (BFE) from academia to the drug discovery industry [38,39,40] (so that non-experts can use them effectively), one had to implement the simple and elegant idea from physics in the messy reality of chemistry, at scale, with sufficient accuracy and throughput. An integrated tool chain has to be developed to prepare the ligands (in their correct protonation and tautomeric states), parametrize their force field, generate their binding poses, map atoms from one ligand to another in RBFE calculations, submit and monitor the many simulations in BFE, analyze the output, and report the predicted binding free energies with associated error estimates (Fig. 1).
As is common in developing an academic concept into an industrial product, one group needs to assemble in a complete solution all the puzzle pieces worked out by many—scattered in various papers, books, presentations, and personal communications—and then some.
The Tower of Binding Free Energy Calculations. Built on the simple foundations of free energy perturbation theory and enabled by advances in force field models and the readily available computing power afforded by graphic processing units (GPUs), BFE required an integrated tool chain for it to become a routine computational tool in industrial drug discovery
Here I share a brief personal account of the inception and development of the FEP+ software, probably the most widely used commercial implementation of BFE in the pharmaceutical industry today. Its intellectual seed was planted when I learned extensively about BFE in the academic research by my lab-mates in Ken Dill’s group [41], where I was a postdoc. Later, after working with my colleagues in D. E. Shaw Research (DESRES) to finish an early version of the DESMOND MD simulation program [42] in 2006, I started to develop BFE as an extension—which I called the Gibbs module—to DESMOND. In that same year, the computational chemistry software company Schrodinger expressed an interest in DESMOND, intending it to be a tool to sample the protein’s conformations in docking studies [43]. Soon that interest pivoted to developing a new software solution for BFE (to complement MCPRO+ [16]). A handful of scientific developers in DESRES and Schrodinger, in collaboration with a few academic groups, persisted through early disappointing results and prevalent skepticism (Outside the BFE experts, BFE was joked to be the most expensive random number generator). In 2013, almost three decades after the first proof-of-concept BFE calculation was published and seven years after I implemented the bare-bone functionality of BFE in DESMOND, Schrodinger started to ship the new BFE solution, bundled with Schrodinger’s OPLS3 force field [44] and branded FEP+, to customers of pharmaceutical companies. A validation study on eight different targets was published in 2015 [45]. Others have since developed their own toolboxes for running BFE calculations using a variety of MD programs, including AMBER [46, 47], OpenMM [48], and GROMACS [49, 50].
Two concurrent developments drove the adoption of BFE in drug discovery. First, graphic processing units (GPUs) became ubiquitous and a number of MD software packages implemented GPU-accelerated codes that were an order-of-magnitude faster than the CPU codes [51,52,53]. What used to take a month could now be completed in only three days, which fit in a typical weekly design-predict-make-test-analyze (DPMTA) cycle in drug discovery. Second, the force field models for both proteins and, importantly, small drug-like molecules were finally good enough to make BFE predictions adequately accurate for prioritizing the candidate molecules by their predicted affinities.
The large-scale deployment of BFE exposed unexpected problems, each requiring its own solution. For example, during the RBFE calculations, the molecular geometry may be distorted when the system is in the midst of changing between two ligands and thus its Hamiltonian does not correspond to one of a realistic molecular system, which leads to numerical instabilities. A solution to this problem was to introduce additional bonded interactions within an alchemical group that is no longer interacting with the rest of the molecular system: these extraneous interactions help maintain reasonable molecular geometries, their contributions to the BFE results canceling out because of the separability of integrals in the partition functions. Another example is RBFE calculations between enantiomers: a restraining potential is required to ensure the correct chirality as one molecule is transformed to its mirror image. For each problem encountered, a programmatic solution must be coded into the standard tool chain, so that the same problem should never have to be solved more than once.
The number of published applications of binding free energy calculations in drug discovery each year. Only journals in medicinal chemistry and drug discovery—including a few general journals—are considered (see Supporting Information for the Pubmed search query used). The empty bar represents the incomplete year of 2022. As discussed in the main text, the true numbers of publications reporting BFE applications in drug discovery may be twice as many
Despite the increasing adoption of BFE in drug discovery projects [54, 55], the number of published studies reporting discovery and optimization of small molecule drugs by BFE is—albeit growing—still relatively small. Out of more than 790 citations (per Google Scholar) garnered by Schrodinger’s landmark BFE paper [45], only 19 (2.4%) reported drug discovery efforts resulting in new chemical matters or new activities [56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74], out of which four did not report the actual use of BFE [62,63,64, 69] and one was unclear [68]. Since 1988, 3646 papers have been published that contain key words related to binding free energy calculations, but only 145 (4%) of these are published in the medicinal chemistry and drug discovery journals (Fig. 2 and Supplementary Information). Even if the number of publications reporting drug discovery employing BFE is twice the total count in Fig. 2 (7 out of the above-mentioned 19 publications citing the Schrodinger paper are counted in Fig. 2, implying an under-counting factor of \((19 - 4.5)/7 \approx 2.\)), it still represents a tiny fraction (0.4%) of the total number of publications in the medicinal chemistry and drug discovery journals (74,179 since 1988, Supplementary Information).
The following are some active areas of research that may help broaden the use of BFE in drug discovery.
The more dissimilar a pair of molecules are, the harder it is to compute their binding free energy difference by RBFE [75], but the more valuable such predictions are, because they allow larger chemical modifications—which often entails higher cost in synthesis—to be explored computationally. For example, RBFE attained much wider adoption after it accommodated scaffold hopping [76, 77]. Its domain of applicability will continue to expand as we enable RBFE to predict the binding free energy changes associated with ever larger chemical transformations.
One type of change of particular interest to drug discovery is a ligand modification associated with the displacement of a water molecule inside the binding pocket [78], as large binding affinities may be gained if the displaced water molecule is of high free energy. A number of approaches have been proposed to take into account such “water hopping” in RBFE [79,80,81,82,83]; this functionality should come standard in future BFE toolboxes.
The accuracy of BFE calculations is fundamentally limited by the accuracy of the underlying force field models. One promising avenue of research is multi-fidelity modeling: BFE first uses conventional force field models in the MD simulations, then more accurate but more computationally expensive energy models—such as QM/MM models [84, 85] or ML models trained on QM results [86,87,88,89]—are applied sparingly, so that an energy difference between the models can be computed and applied to correct the BFE results by FEP.
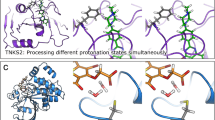
Often not all relevant molecular conformations are sampled in the simulations of BFE, and their contributions to the binding free energies are thus unaccounted for. A fruitful area of research is to combine conformational free energy calculations with BFE to incorporate the effect of receptor conformational flexibility and potentially multiple binding poses of each ligand [90, 91] into BFE [23, 92, 93]. For example, RBFE may be used to compute the difference in the binding free energies \(\Delta \Delta G^{\mathrm {bind}}_{ab,\mu }\) between two ligands a and b to each receptor conformation \(\mu \), and an enhanced conformational sampling method [94, 95] may be used to compute the conformational free energy differences between any two conformations \(\mu \) and \(\nu \) of either the apo receptor (\(\Delta \Delta G^{\mathrm {conf}}_{\mu \nu }\)) or the receptor in complex with a ligand a (\(\Delta \Delta G^{\mathrm {conf}}_{a,\mu \nu }\)), as illustrated in Fig. 3. From these results, the conformation-specific binding free energy, \(\Delta G^{\mathrm {bind}}_{a,\mu }\), of each ligand a to each receptor conformation \(\mu \) may be solved from the (over-determined) simultaneous equations
where \(\Delta G^{\mathrm {conf}}_\mu \) (or \(\Delta G^{\mathrm {conf}}_\nu \)) is the conformational free energy of the apo receptor in conformation \(\mu \) (or \(\nu \)). The collection of pairwise free energy differences in Eq. 1 may be planned and analyzed using an optimal measurement network of pairwise differences [96]. The overall binding free energy of a ligand a to the receptor is derived from the combination of the conformation-specific binding free energies:
where k is the Boltzmann constant and T the temperature. Note that in the above the binding free energies for a set of ligands are determined up to a constant (\(\Delta G_0\) in Fig. 3).
In drug discovery, many molecules need to be considered in each DPMTA cycle, which calls for an efficient plan of RBFE calculations between well-chosen pairs of molecules [97]. New computational methods have recently been published that optimize the organization of RBFE calculations for many molecules, using the theory of experimental design to minimize the total statistical uncertainty in the calculations [96, 98, 99]. Bennett’s method has also been extended to the analysis of such calculations [100].
A related and exciting area of research is to efficiently integrate BFE and other computational and experimental techniques in a seamless workflow to drastically accelerate (by 10\(\sim \)100 times) the exploration of chemical space in the DPMTA cycle. For example, starting with 10,000 molecular designs from generative models [101,102,103], one may perform BFE on 100 diverse molecules chosen by a machine-learning model of quantitative structure-activity relationship (QSAR) trained on previous experimental and computational results. The QSAR model is then updated by the new BFE results (and new experimental results when available) and guides the selection of another 100 molecules for a second round of BFE. So on and so forth. Such active learning [104] may enable tens of thousands of molecular designs to be computationally generated and ranked by a feasible number of rigorous BFE calculations each week and substantially shorten the times of hit-to-lead and lead-optimization in drug discovery.
I would like to end with a personal reflection. I was fortunate to enjoy the long-time friendship with many people who shared an unwavering interest in BFE. We believed that together we could harness our understanding of physics to make a difference in the development of medicine for patients. When there was limited acceptance of BFE in drug discovery, attending free energy workshops and being surrounded by these friends helped sustain my interest and spur me to contribute to this endeavor. It is no small comfort to see that the workshops grew bigger each year and that BFE has started to play a key role in the development of molecules currently in clinical trial [105].
References
Zwanzig RW (1954) High-temperature equation of state by a perturbation method. I. Nonpolar gases. J Chem Phys 22(8):1420–1426. https://doi.org/10.1063/1.1740409
Bennett CH (1976) Efficient estimation of free energy differences from Monte Carlo data. J Comp Phys 22(2):245–268. https://doi.org/10.1016/0021-9991(76)90078-4
Shirts MR, Chodera JD (2008) Statistically optimal analysis of samples from multiple equilibrium states. J Chem Phys 129(12):124105. https://doi.org/10.1063/1.2978177
Kirkwood JG (1935) Statistical mechanics of fluid mixtures. J Chem Phys 3(5):300–313. https://doi.org/10.1063/1.1749657
Postma JPM, Berendsen HJC, Haak JR (1982) Thermodynamics of cavity formation in water. A molecular dynamics study. Faraday Symposia Chem Soc 17:55–67. https://doi.org/10.1039/fs9821700055
Jorgensen WL, Ravimohan C (1985) Monte Carlo simulation of differences in free energies of hydration. J Chem Phys 83(6):3050–3054. https://doi.org/10.1063/1.449208
Tembe BL, McCammon JA (1984) Ligand-receptor interactions. Comput Chem 8(4):281–283
Bash PA, Singh UC, Brown FK, Langridge R, Kollman PA (1987) Calculation of the relative change in binding free energy of a protein-inhibitor complex. Science 235(4788):574–576. https://doi.org/10.1126/science.3810157
Singh UC, Benkovic SJ (1988) A free-energy perturbation study of the binding of methotrexate to mutants of dihydrofolate reductase. Proc Natl Acad Sci 85(24):9519–9523. https://doi.org/10.1073/pnas.85.24.9519
Straatsma TP, McCammon JA (1992) Computational alchemy. Annu Rev Phys Chem 43(1):407–435. https://doi.org/10.1146/annurev.pc.43.100192.002203
Rao BG, Kim EE, Murcko MA (1996) Calculation of solvation and binding free energy differences between VX-478 and its analogs by free energy perturbation and AMSOL methods. J Comput Aided Mol Des 10(1):23–30. https://doi.org/10.1007/bf00124462
Erion MD, Reddy MR (1998) Calculation of relative hydration free energy differences for heteroaromatic compounds: use in the design of adenosine deaminase and cytidine deaminase inhibitors. J Am Chem Soc 120(14):3295–3304. https://doi.org/10.1021/ja972906j
Boresch S, Tettinger F, Leitgeb M, Karplus M (2003) Absolute binding free energies: a quantitative approach for their calculation. J Phys Chem B 107(35):9535–9551. https://doi.org/10.1021/jp0217839
Woo H-J, Roux B (2005) Calculation of absolute protein-ligand binding free energy from computer simulations. Proc Natl Acad Sci USA 102(19):6825–6830. https://doi.org/10.1073/pnas.0409005102
Radmer RJ, Kollman PA (1997) Free energy calculation methods: a theoretical and empirical comparison of numerical errors and a new method qualitative estimates of free energy changes. J Comput Chem 18(7):902–919
Jorgensen WL, Tirado-Rives J (2005) Molecular modeling of organic and biomolecular systems using BOSS and MCPRO. J Comput Chem 26(16):1689–1700. https://doi.org/10.1002/jcc.20297
Shenfeld DK, Xu H, Eastwood MP, Dror RO, Shaw DE (2009) Minimizing thermodynamic length to select intermediate states for free-energy calculations and replica-exchange simulations. Phys Rev E 80:046705. https://doi.org/10.1103/PhysRevE.80.046705
S S, Roux B, Andersen OS (2000) Free energy simulations: thermodynamic reversibility and variability. J Phys Chem B 104(21):5179–5190. https://doi.org/10.1021/jp994193s
Beutler TC, Mark AE, van Schaik RC, Gerber PR, van Gunsteren WF (1994) Avoiding singularities and numerical instabilities in free energy calculations based on molecular simulations. Chem Phys Lett 222(6):529–539. https://doi.org/10.1016/0009-2614(94)00397-1
Lee T-S, Lin Z, Allen BK, Lin C, Radak BK, Tao Y, Tsai H-C, Sherman W, York DM (2020) Improved alchemical free energy calculations with optimized smoothstep softcore potentials. J Chem Theory Comput 16(9):5512–5525. https://doi.org/10.1021/acs.jctc.0c00237
Rocklin GJ, Mobley DL, Dill KA, Hünenberger PH (2013) Calculating the binding free energies of charged species based on explicit-solvent simulations employing lattice-sum methods: an accurate correction scheme for electrostatic finite-size effects. J Chem Phys 139(18):184103. https://doi.org/10.1063/1.4826261
Pan AC, Xu H, Palpant T, Shaw DE (2017) Quantitative characterization of the binding and unbinding of millimolar drug fragments with molecular dynamics simulations. J Chem Theory Comput 13(7):3372–3377. https://doi.org/10.1021/acs.jctc.7b00172 (PMID: 28582625)
Mobley DL, Chodera JD, Dill KA (2007) Confine-and-release method: obtaining correct binding free energies in the presence of protein conformational change. J Chem Theory Comput 3(4):1231–1235. https://doi.org/10.1021/ct700032n (PMID: 18843379)
Wang L, Deng Y, Knight JL, Wu Y, Kim B, Sherman W, Shelley JC, Lin T, Abel R (2013) Modeling local structural rearrangements using fep/rest: application to relative binding affinity predictions of cdk2 inhibitors. J Chem Theory Comput 9(2):1282–1293. https://doi.org/10.1021/ct300911a
Jiang W, Roux B (2010) Free energy perturbation Hamiltonian replica-exchange molecular dynamics (fep/h-remd) for absolute ligand binding free energy calculations. J Chem Theory Comput 6(9):2559–2565. https://doi.org/10.1021/ct1001768
Zhang S, Hahn DF, Shirts MR, Voelz VA (2021) Expanded ensemble methods can be used to accurately predict protein-ligand relative binding free energies. J Chem Theory Comput 17(10):6536–6547. https://doi.org/10.1021/acs.jctc.1c00513
Shirts MR, Mobley DL, Chodera JD, Pande VS (2007) Accurate and efficient corrections for missing dispersion interactions in molecular simulations. J Phys Chem B 111(45):13052–13063. https://doi.org/10.1021/jp0735987
Chodera JD (2016) A simple method for automated equilibration detection in molecular simulations. J Chem Theory Comput 12(4):1799–1805. https://doi.org/10.1021/acs.jctc.5b00784
Klimovich PV, Shirts MR, Mobley DL (2015) Guidelines for the analysis of free energy calculations. J Comput-Aided Mol Des 29(5):397–411. https://doi.org/10.1007/s10822-015-9840-9
Mey ASJS, Allen BK, Bruce McDonald HE, Chodera JD, Hahn DF, Kuhn M, Michel J, Mobley DL, Naden LN, Prasad S, Rizzi A, Scheen J, Shirts MR, Tresadern G, Xu H (2020) Best practices for alchemical free energy calculations [article v1.0]. Living J Comput Mol Sci 2(1):18378. https://doi.org/10.33011/livecoms.2.1.18378
Price MLP, Jorgensen WL (2000) Analysis of binding affinities for celecoxib analogues with COX-1 and COX-2 from combined docking and Monte Carlo simulations and insight into the COX-2/COX-1 selectivity. J Am Chem Soc 122(39):9455–9466. https://doi.org/10.1021/ja001018c
Lee T-S, Kollman PA (2000) Theoretical studies suggest a new antifolate as a more potent inhibitor of thymidylate synthase. J Am Chem Soc 122(18):4385–4393. https://doi.org/10.1021/ja9925554
Reddy MR, Erion MD (2001) Calculation of relative binding free energy differences for fructose 1,6-bisphosphatase inhibitors using the thermodynamic cycle perturbation approach. J Am Chem Soc 123(26):6246–6252. https://doi.org/10.1021/ja0103288
Reddy MR, Erion MD (2001) Free energy calculations in rational drug design. Kluwer Academic/Plenum Publishers, New York
Bollini M, Domaoal RA, Thakur VV, Gallardo-Macias R, Spasov KA, Anderson KS, Jorgensen WL (2011) Computationally-guided optimization of a docking hit to yield catechol diethers as potent anti-HIV agents. J Med Chem 54(24):8582–8591. https://doi.org/10.1021/jm201134m
Ivetac A, Swift SE, Boyer PL, Diaz A, Naughton J, Young JAT, Hughes SH, McCammon JA (2014) Discovery of novel inhibitors of HIV-1 reverse transcriptase through virtual screening of experimental and theoretical ensembles. Chem Biol Drug Des 83(5):521–531. https://doi.org/10.1111/cbdd.12277
Dziedzic P, Cisneros JA, Robertson MJ, Hare AA, Danford NE, Baxter RHG, Jorgensen WL (2015) Design, synthesis, and protein crystallography of biaryltriazoles as potent tautomerase inhibitors of macrophage migration inhibitory factor. J Am Chem Soc 137(8):2996–3003. https://doi.org/10.1021/ja512112j
Abel R, Wang L, Harder ED, Berne BJ, Friesner RA (2017) Advancing drug discovery through enhanced free energy calculations. Acc Chem Res 50(7):1625–1632. https://doi.org/10.1021/acs.accounts.7b00083
Cournia Z, Allen B, Sherman W (2017) Relative binding free energy calculations in drug discovery: recent advances and practical considerations. J Chem Inf Model 57(12):2911–2937. https://doi.org/10.1021/acs.jcim.7b00564
Sherborne B, Shanmugasundaram V, Cheng AC, Christ CD, DesJarlais RL, Duca JS, Lewis RA, Loughney DA, Manas ES, McGaughey GB, Peishoff CE, Vlijmen Hv (2016) Collaborating to improve the use of free-energy and other quantitative methods in drug discovery. J Comput-Aided Mol Des 30(12):1139–1141. https://doi.org/10.1007/s10822-016-9996-y
Mobley DL, Graves AP, Chodera JD, McReynolds AC, Shoichet BK, Dill KA (2007) Predicting absolute ligand binding free energies to a simple model site. J Mol Biol 371(4):1118–1134. https://doi.org/10.1016/j.jmb.2007.06.002
Bowers KJ, Chow DE, Xu H, Dror RO, Eastwood MP, Gregersen BA, Klepeis JL, Kolossvary I, Moraes MA, Sacerdoti FD, Salmon JK, Shan Y, Shaw DE (2006) Scalable algorithms for molecular dynamics simulations on commodity clusters. In: SC ’06: Proceedings of the 2006 ACM/IEEE conference on supercomputing, p 84. https://doi.org/10.1109/SC.2006.54
Sherman W, Day T, Jacobson MP, Friesner RA, Farid R (2006) Novel procedure for modeling ligand/receptor induced fit effects. J Med Chem 49(2):534–553. https://doi.org/10.1021/jm050540c
Harder E, Damm W, Maple J, Wu C, Reboul M, Xiang J, Wang L, Lupyan D, Dahlgren MK, Knight JL, Kaus JW, Cerutti DS, Krilov G, Jorgensen WL, Abel R, Friesner RA (2016) OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J Chem Theory Comput 12(1):281–296. https://doi.org/10.1021/acs.jctc.5b00864
Wang L, Wu Y, Deng Y, Kim B, Pierce L, Krilov G, Lupyan D, Robinson S, Dahlgren MK, Greenwood J, Romero DL, Masse C, Knight JL, Steinbrecher T, Beuming T, Damm W, Harder E, Sherman W, Brewer M, Wester R, Murcko M, Frye L, Farid R, Lin T, Mobley DL, Jorgensen WL, Berne BJ, Friesner RA, Abel R (2015) Accurate and reliable prediction of relative ligand binding potency in prospective drug discovery by way of a modern free-energy calculation protocol and force field. J Am Chem Soc 137(7):2695–2703. https://doi.org/10.1021/ja512751q
Lee T-S, Allen BK, Giese TJ, Guo Z, Li P, Lin C, McGee TD, Pearlman DA, Radak BK, Tao Y, Tsai H-C, Xu H, Sherman W, York DM (2020) Alchemical binding free energy calculations in amber20: advances and best practices for drug discovery. J Chem Inf Model 60(11):5595–5623. https://doi.org/10.1021/acs.jcim.0c00613
Zou J, Yin J, Fang L, Yang M, Wang T, Wu W, Bellucci MA, Zhang P (2020) Computational prediction of mutational effects on sars-cov-2 binding by relative free energy calculations. J Chem Inf Model 60(12):5794–5802. https://doi.org/10.1021/acs.jcim.0c00679
Kuhn M, Firth-Clark S, Tosco P, Mey ASJS, Mackey MD, Michel J (2020) Assessment of binding affinity via alchemical free energy calculations. J Chem Inf Model 60(6):3120–3130. https://doi.org/10.1021/acs.jcim.0c00165
Aldeghi M, Heifetz A, Bodkin MJ, Knapp S, Biggin PC (2017) Predictions of ligand selectivity from absolute binding free energy calculations. J Am Chem Soc 139(2):946–957. https://doi.org/10.1021/jacs.6b11467
Lemkul J (2019) From proteins to perturbed hamiltonians: a suite of tutorials for the GROMACS-2018 molecular simulation package [Article v1.0]. Living J Comput Mol Sci 1(1):5068. https://doi.org/10.33011/livecoms.1.1.5068
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E (2015) Gromacs: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Salomon-Ferrer R, Götz AW, Poole D, Le Grand S, Walker RC (2013) Routine microsecond molecular dynamics simulations with amber on gpus. 2. Explicit solvent particle mesh ewald. J Chem Theory Comput 9(9):3878–3888. https://doi.org/10.1021/ct400314y
Bergdorf M, Robinson-Mosher A, Guo X, Law K-H, Shaw DE (2021) Desmond/gpu performance as of april 2021. Technical report, New York, New York, USA. D. E. Shaw Research. https://www.deshawresearch.com/publications/Desmond-GPU_Performance_April_2021.pdf
Zhang C-H, Spasov KA, Reilly RA, Hollander K, Stone EA, Ippolito JA, Liosi M-E, Deshmukh MG, Tirado-Rives J, Zhang S, Liang Z, Miller SJ, Isaacs F, Lindenbach BD, Anderson KS, Jorgensen WL (2021) Optimization of triarylpyridinone inhibitors of the main protease of SARS-CoV-2 to low-nanomolar antiviral potency. ACS Med Chem Lett 12(8):1325–1332. https://doi.org/10.1021/acsmedchemlett.1c00326
Schindler CEM, Baumann H, Blum A, Böse D, Buchstaller H-P, Burgdorf L, Cappel D, Chekler E, Czodrowski P, Dorsch D, Eguida MKI, Follows B, Fuchß T, Grädler U, Gunera J, Johnson T, Lebrun CJ, Karra S, Klein M, Knehans T, Koetzner L, Krier M, Leiendecker M, Leuthner B, Li L, Mochalkin I, Musil D, Neagu C, Rippmann F, Schiemann K, Schulz R, Steinbrecher T, Tanzer E-M, Lopez AU, Follis AV, Wegener A, Kuhn D (2020) Large-scale assessment of binding free energy calculations in active drug discovery projects. J Chem Inf Model 60(11):5457–5474. https://doi.org/10.1021/acs.jcim.0c00900
Zhang B, D’Erasmo MP, Murelli RP, Gallicchio E (2016) Free energy-based virtual screening and optimization of RNase H inhibitors of HIV-1 reverse transcriptase. ACS Omega 1(3):435–447. https://doi.org/10.1021/acsomega.6b00123
Ciordia M, Pérez-Benito L, Delgado F, Trabanco AA, Tresadern G (2016) Application of free energy perturbation for the design of BACE1 inhibitors. J Chem Inf Model 56(9):1856–1871. https://doi.org/10.1021/acs.jcim.6b00220
Lovering F, Aevazelis C, Chang J, Dehnhardt C, Fitz L, Han S, Janz K, Lee J, Kaila N, McDonald J, Moore W, Moretto A, Papaioannou N, Richard D, Ryan MS, Wan Z, Thorarensen A (2016) Imidazotriazines: spleen tyrosine kinase (Syk) inhibitors identified by free-energy perturbation (FEP). ChemMedChem 11(2):217–233. https://doi.org/10.1002/cmdc.201500333
Cole DJ, Janecek M, Stokes JE, Rossmann M, Faver JC, McKenzie GJ, Venkitaraman AR, Hyvönen M, Spring DR, Huggins DJ, Jorgensen WL (2017) Computationally-guided optimization of small-molecule inhibitors of the Aurora A kinase-TPX2 protein-protein interaction. Chem Commun 53(67):9372–9375. https://doi.org/10.1039/c7cc05379g
Schmidt TC, Eriksson P-O, Gustafsson D, Cosgrove D, Frølund B, Boström J (2017) Discovery and evaluation of anti-fibrinolytic plasmin inhibitors derived from 5-(4-Piperidyl)isoxazol-3-ol (4-PIOL). J Chem Inf Model 57(7):1703–1714. https://doi.org/10.1021/acs.jcim.7b00255
Chen Z, Cox BD, Garnier-Amblard EC, McBrayer TR, Coats SJ, Schinazi RF, Amblard F (2017) Synthesis and anti-HCV activity of a series of \(\beta \)-d-2’-deoxy-2’-dibromo nucleosides and their corresponding phosphoramidate prodrugs. Bioorg Med Chem Lett 27(23):5296–5299. https://doi.org/10.1016/j.bmcl.2017.10.024
Carabet LA, Lallous N, Leblanc E, Ban F, Morin H, Lawn S, Ghaidi F, Lee J, Mills IG, Gleave ME, Rennie PS, Cherkasov A (2018) Computer-aided drug discovery of Myc-Max inhibitors as potential therapeutics for prostate cancer. Eur J Med Chem 160:108–119. https://doi.org/10.1016/j.ejmech.2018.09.023
Välimäki M (2018) Discovery of cardioprotective isoxazole-amide compounds targeting the synergy of transcription factors gata4 and nkx2-5. PhD thesis, University of Oulu, Faculty of Medicine. http://urn.fi/urn:isbn:9789529412525
Bassanini I, D’Annessa I, Costa M, Monti D, Colombo G, Riva S (2018) Chemo-enzymatic synthesis of ( E )-2,3-diaryl-5-styryl- trans -2,3-dihydrobenzofuran-based scaffolds and their in vitro and in silico evaluation as a novel sub-family of potential allosteric modulators of the 90 kDa heat shock protein (Hsp90). Org Biomol Chem 16(20):3741–3753. https://doi.org/10.1039/c8ob00644j
Li Z, Li X, Huang Y-Y, Wu Y, Liu R, Zhou L, Lin Y, Wu D, Zhang L, Liu H, Xu X, Yu K, Zhang Y, Cui J, Zhan C-G, Wang X, Luo H-B (2020) Identify potent SARS-CoV-2 main protease inhibitors via accelerated free energy perturbation-based virtual screening of existing drugs. Proc Natl Acad Sci 117(44):27381–27387. https://doi.org/10.1073/pnas.2010470117
Montgomery AP, Dobie C, Szabo R, Hallam L, Ranson M, Yu H, Skropeta D (2020) Design, synthesis and evaluation of carbamate-linked uridyl-based inhibitors of human ST6Gal I. Bioorg Med Chem 28(14):115561. https://doi.org/10.1016/j.bmc.2020.115561
Leger PR, Hu DX, Biannic B, Bui M, Han X, Karbarz E, Maung J, Okano A, Osipov M, Shibuya GM, Young K, Higgs C, Abraham B, Bradford D, Cho C, Colas C, Jacobson S, Ohol YM, Pookot D, Rana P, Sanchez J, Shah N, Sun M, Wong S, Brockstedt DG, Kassner PD, Schwarz JB, Wustrow DJ (2020) Discovery of potent, selective, and orally bioavailable inhibitors of USP7 with in vivo antitumor activity. J Med Chem 63(10):5398–5420. https://doi.org/10.1021/acs.jmedchem.0c00245
Kesely K, Noomuna P, Vieth M, Hipskind P, Haldar K, Pantaleo A, Turrini F, Low PS (2020) Identification of tyrosine kinase inhibitors that halt Plasmodium falciparum parasitemia. PLoS ONE 15(11):0242372. https://doi.org/10.1371/journal.pone.0242372
Fratev F, Gutierrez DA, Aguilera RJ, Tyagi A, Damodaran C, Sirimulla S (2020) Discovery of new AKT1 inhibitors by combination of in silico structure based virtual screening approaches and biological evaluations. J Biomol Struct Dyn 39(1):1–11. https://doi.org/10.1080/07391102.2020.1715835
Fushimi M, Buck H, Balbach M, Gorovyy A, Ferreira J, Rossetti T, Kaur N, Levin LR, Buck J, Quast J, Heuvel J.v.d, Steegborn C, Finkin-Groner E, Kargman S, Michino M, Foley MA, Miller M, Liverton NJ, Huggins DJ, Meinke PT (2021) Discovery of TDI-10229: a potent and orally bioavailable inhibitor of soluble adenylyl cyclase (sAC, ADCY10). ACS Med Chem Lett 12(8):1283–1287. https://doi.org/10.1021/acsmedchemlett.1c00273
Balasubramaniam M, Lakkaniga NR, Dera AA, Fayi MA, Abohashrh M, Ahmad I, Chandramoorthy HC, Nalini G, Rajagopalan P (2021) FCX-146, a potent allosteric inhibitor of Akt kinase in cancer cells: lead optimization of the second-generation arylidene indanone scaffold. Biotechnol Appl Biochem 68(1):82–91. https://doi.org/10.1002/bab.1896
Palmer MJ, Deng X, Watts S, Krilov G, Gerasyuto A, Kokkonda S, Mazouni FE, White J, White KL, Striepen J, Bath J, Schindler KA, Yeo T, Shackleford DM, Mok S, Deni I, Lawong A, Huang A, Chen G, Wang W, Jayaseelan J, Katneni K, Patil R, Saunders J, Shahi SP, Chittimalla R, Angulo-Barturen I, Jiménez-Díaz MB, Wittlin S, Tumwebaze PK, Rosenthal PJ, Cooper RA, Aguiar ACC, Guido RVC, Pereira DB, Mittal N, Winzeler EA, Tomchick DR, Laleu B, Burrows JN, Rathod PK, Fidock DA, Charman SA, Phillips MA (2021) Potent antimalarials with development potential identified by structure-guided computational optimization of a pyrrole-based dihydroorotate dehydrogenase inhibitor series. J Med Chem 64(9):6085–6136. https://doi.org/10.1021/acs.jmedchem.1c00173
Chang W, Altman MD, Lesburg CA, Perera SA, Piesvaux JA, Schroeder GK, Wyss DF, Cemerski S, Chen Y, DiNunzio E, Haidle AM, Ho T, Kariv I, Knemeyer I, Kopinja JE, Lacey BM, Laskey J, Lim J, Long BJ, Ma Y, Maddess ML, Pan B-S, Presland JP, Spooner E, Steinhuebel D, Truong Q, Zhang Z, Fu J, Addona GH, Northrup AB, Parmee E, Tata JR, Bennett DJ, Cumming JN, Siu T, Trotter BW (2022) Discovery of MK-1454: a potent cyclic dinucleotide stimulator of interferon genes agonist for the treatment of cancer. J Med Chem 65(7):5675–5689. https://doi.org/10.1021/acs.jmedchem.1c02197
Val C, Rodríguez-García C, Prieto-Díaz R, Crespo A, Azuaje J, Carbajales C, Majellaro M, Díaz-Holguín A, Brea JM, Loza MI, Gioé-Gallo C, Contino M, Stefanachi A, García-Mera X, Estévez JC, Gutiérrez-de-Terán H, Sotelo E (2022) Optimization of 2-amino-4, 6-diarylpyrimidine-5-carbonitriles as potent and selective A1 antagonists. J Med Chem 65(3):2091–2106. https://doi.org/10.1021/acs.jmedchem.1c01636
Liu S, Wang L, Mobley DL (2015) Is ring breaking feasible in relative binding free energy calculations? J Chem Inf Model 55(4):727–735. https://doi.org/10.1021/acs.jcim.5b00057
Wang L, Deng Y, Wu Y, Kim B, LeBard DN, Wandschneider D, Beachy M, Friesner RA, Abel R (2017) Accurate modeling of scaffold hopping transformations in drug discovery. J Chem Theory Comput 13(1):42–54. https://doi.org/10.1021/acs.jctc.6b00991
Zou J, Li Z, Liu S, Peng C, Fang D, Wan X, Lin Z, Lee T-S, Raleigh DP, Yang M, Simmerling C (2021) Scaffold hopping transformations using auxiliary restraints for calculating accurate relative binding free energies. J Chem Theory Comput 17(6):3710–3726. https://doi.org/10.1021/acs.jctc.1c00214
Pearlstein RA, Sherman W, Abel R (2013) Contributions of water transfer energy to protein-ligand association and dissociation barriers: watermap analysis of a series of p38 \(\alpha \) MAP kinase inhibitors. Proteins Struct Funct Bioinf 81(9):1509–1526. https://doi.org/10.1002/prot.24276
Hamelberg D, McCammon JA (2004) Standard free energy of releasing a localized water molecule from the binding pockets of proteins: double-decoupling method. J Am Chem Soc 126(24):7683–7689. https://doi.org/10.1021/ja0377908
Michel J, Tirado-Rives J, Jorgensen WL (2009) Energetics of displacing water molecules from protein binding sites: consequences for ligand optimization. J Am Chem Soc 131(42):15403–15411. https://doi.org/10.1021/ja906058w
Bergazin TD, Ben-Shalom IY, Lim NM, Gill SC, Gilson MK, Mobley DL (2021) Enhancing water sampling of buried binding sites using nonequilibrium candidate Monte Carlo. J Comput-Aided Mol Des 35(2):167–177. https://doi.org/10.1007/s10822-020-00344-8
Ben-Shalom IY, Lin Z, Radak BK, Lin C, Sherman W, Gilson MK (2020) Accounting for the central role of interfacial water in protein-ligand binding free energy calculations. J Chem Theory Comput 16(12):7883–7894. https://doi.org/10.1021/acs.jctc.0c00785
Ben-Shalom IY, Lin C, Radak BK, Sherman W, Gilson MK (2021) Fast equilibration of water between buried sites and the bulk by molecular dynamics with parallel Monte Carlo water moves on graphical processing units. J Chem Theory Comput 17(12):7366–7372. https://doi.org/10.1021/acs.jctc.1c00867
Hudson PS, Woodcock HL, Boresch S (2015) Use of nonequilibrium work methods to compute free energy differences between molecular mechanical and quantum mechanical representations of molecular systems. J Phys Chem Lett 6(23):4850–4856. https://doi.org/10.1021/acs.jpclett.5b02164
Giese TJ, York DM (2019) Development of a robust indirect approach for MM \(\rightarrow \) QM free energy calculations that combines force-matched reference potential and Bennett’s acceptance ratio methods. J Chem Theory Comput 15(10):5543–5562. https://doi.org/10.1021/acs.jctc.9b00401
Smith JS, Isayev O, Roitberg AE (2017) Ani-1: an extensible neural network potential with dft accuracy at force field computational cost. Chem Sci 8:3192–3203. https://doi.org/10.1039/C6SC05720A
Rufa DA, Bruce Macdonald HE, Fass J, Wieder M, Grinaway PB, Roitberg AE, Isayev O, Chodera JD (2020) Towards chemical accuracy for alchemical free energy calculations with hybrid physics-based machine learning / molecular mechanics potentials. bioRxiv. https://doi.org/10.1101/2020.07.29.227959
Ko TW, Finkler JA, Goedecker S, Behler J (2021) A fourth-generation high-dimensional neural network potential with accurate electrostatics including non-local charge transfer. Nat Commun 12(1):398. https://doi.org/10.1038/s41467-020-20427-2
Christensen AS, Sirumalla SK, Qiao Z, O’Connor MB, Smith DGA, Ding F, Bygrave PJ, Anandkumar A, Welborn M, Manby FR et al (2021) Orbnet denali: a machine learning potential for biological and organic chemistry with semi-empirical cost and dft accuracy. J Chem Phys 155(20):204103. https://doi.org/10.1063/5.0061990
Mobley DL, Dill KA (2009) Binding of Small-Molecule Ligands to Proteins: “What You See’’ Is Not Always “What You Get’’. Structure 17(4):489–498. https://doi.org/10.1016/j.str.2009.02.010
Xu H, Palpant T, Weinberger C, Shaw DE (2022) Characterizing receptor flexibility to predict mutations that lead to human adaptation of influenza hemagglutinin. J Chem Theory Comput 18(8):4995–5005. https://doi.org/10.1021/acs.jctc.1c01044
Lawrenz M, Baron R, Wang Y, McCammon JA (2011) Effects of biomolecular flexibility on alchemical calculations of absolute binding free energies. J Chem Theory Comput 7(7):2224–2232. https://doi.org/10.1021/ct200230v
Araki M, Kamiya N, Sato M, Nakatsui M, Hirokawa T, Okuno Y (2016) The effect of conformational flexibility on binding free energy estimation between kinases and their inhibitors. J Chem Inf Model 56(12):2445–2456. https://doi.org/10.1021/acs.jcim.6b00398
Zuckerman DM, Chong LT (2016) Weighted ensemble simulation: review of methodology, applications, and software. Annu Rev Biophys 46(1):43–57. https://doi.org/10.1146/annurev-biophys-070816-033834
Fu H, Zhang H, Chen H, Shao X, Chipot C, Cai W (2018) Zooming across the free-energy landscape: shaving barriers, and flooding valleys. J Phys Chem Lett 9(16):4738–4745. https://doi.org/10.1021/acs.jpclett.8b01994
Xu H (2019) Optimal measurement network of pairwise differences. J Chem Inf Model 59(11):4720–4728. https://doi.org/10.1021/acs.jcim.9b00528
Liu S, Wu Y, Lin T, Abel R, Redmann JP, Summa CM, Jaber VR, Lim NM, Mobley DL (2013) Lead optimization mapper: automating free energy calculations for lead optimization. J Comput-Aided Mol Des 27(9):755–70. https://doi.org/10.1007/s10822-013-9678-y
Yang Q, Burchett W, Steeno GS, Liu S, Yang M, Mobley DL, Hou X (2019) Optimal designs for pairwise calculation: an application to free energy perturbation in minimizing prediction variability. J Comput Chem 41(3):247–257. https://doi.org/10.1002/jcc.26095
Li P, Li Z, Wang Y, Dou H, Radak BK, Allen BK, Sherman W, Xu H (2021) Precise binding free energy calculations for multiple molecules using an optimal measurement network of pairwise differences. J Chem Theory Comput 18(2):650–663. https://doi.org/10.1021/acs.jctc.1c00703
Giese TJ, York DM (2021) Variational method for networkwide analysis of relative ligand binding free energies with loop closure and experimental constraints. J Chem Theory Comput 17(3):1326–1336. https://doi.org/10.1021/acs.jctc.0c01219
Segler MHS, Kogej T, Tyrchan C, Waller MP (2017) Generating focused molecule libraries for drug discovery with recurrent neural networks. ACS Cent Sci 4(1):120–131. https://doi.org/10.1021/acscentsci.7b00512
Gómez-Bombarelli R, Wei JN, Duvenaud D, Hernández-Lobato JM, Sánchez-Lengeling B, Sheberla D, Aguilera-Iparraguirre J, Hirzel TD, Adams RP, Aspuru-Guzik A (2018) Automatic chemical design using a data-driven continuous representation of molecules. ACS Cent Sci 4(2):268–276. https://doi.org/10.1021/acscentsci.7b00572
Popova M, Isayev O, Tropsha A (2018) Deep reinforcement learning for de novo drug design. Sci Adv 4(7):7885. https://doi.org/10.1126/sciadv.aap7885
Konze KD, Bos PH, Dahlgren MK, Leswing K, Tubert-Brohman I, Bortolato A, Robbason B, Abel R, Bhat S (2019) Reaction-based enumeration, active learning, and free energy calculations to rapidly explore synthetically tractable chemical space and optimize potency of cyclin-dependent kinase 2 inhibitors. J Chem Inf Model 59(9):3782–3793. https://doi.org/10.1021/acs.jcim.9b00367
Allen BK, Kulkarni MM, Chamberlain B, Dwight T, Koh C, Samant R, Jernigan F, Rice J, Tan D, Li S, Marino K, Huang H, Chiswick E, Tesar B, Sparks S, Lin Z, McGee TD, Kolossváry I, Lin C, Shechter S, Soutter H, Bastos C, Taimi M, Lai S, Petrin A, Kane T, Swann S, Gardner H, Winter C, Sherman W (2022) Design of a systemic small molecule clinical sting agonist using physics-based simulations and artificial intelligence. bioRxiv. https://doi.org/10.1101/2022.05.23.493001
Acknowledgements
I wish to acknowledge my colleagues, especially Justin Gullingsrud, Ross Lippert, Michael Bergdorf, and Sean Baxter, for their critical contributions to the BFE functionality in DESMOND. I thank Woody Sherman and Alexander Tropsha for thoughtful suggestions to this writing and Yunxing (Stella) Li for her help in preparing Fig. 1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Huafeng Xu is a shareholder of Stingthera, which is conducting clinical trials of the STING agonist described in reference [105]. Huafeng Xu is the sole author of this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10822_2022_494_MOESM1_ESM.xlsx
Supplementary file1 (XLSX 49 kb). Number of publications in medicinal chemistry and drug discovery journals reporting BFE applications
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, H. The slow but steady rise of binding free energy calculations in drug discovery. J Comput Aided Mol Des 37, 67–74 (2023). https://doi.org/10.1007/s10822-022-00494-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-022-00494-x