Abstract
Indoleamine 2,3-dioxygenase 1 (IDO1) is a heme-containing enzyme that catalyzes the first and rate-limiting step in catabolism of tryptophan via the kynurenine pathway, which plays a pivotal role in the proliferation and differentiation of T cells. IDO1 has been proven to be an attractive target for many diseases, such as breast cancer, lung cancer, colon cancer, prostate cancer, etc. In this study, docking-based virtual screening and bioassays were conducted to identify novel inhibitors of IDO1. The cellular assay demonstrated that 24 compounds exhibited potent inhibitory activity against IDO1 at micromolar level, including 8 compounds with IC50 values below 10 μM and the most potent one (compound 1) with IC50 of 1.18 ± 0.04 μM. Further lead optimization based on similarity searching strategy led to the discovery of compound 28 as an excellent inhibitor with IC50 of 0.27 ± 0.02 μM. Then, the structure–activity relationship of compounds 1, 2, 8 and 14 analogues is discussed. The interaction modes of two compounds against IDO1 were further explored through a Python Based Metal Center Parameter Builder (MCPB.py) molecular dynamics simulation, binding free energy calculation and electrostatic potential analysis. The novel IDO1 inhibitors of compound 1 and its analogues could be considered as promising scaffold for further development of IDO1 inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tryptophan is an essential amino acid not only used for protein synthesis, but also as a substrate to synthesize the neurotransmitter serotonin and catabolism through the kynurenine pathway (Kyn) [1,2,3,4].Over 90% of L-tryptophan depleted by humans is processed through kynurenine pathway (Kyn) [5]. The kynurenine pathway of tryptophan (Trp) catabolism is shown in Fig. 1. Indoleamine 2,3-Dioxygenase 1 (IDO1) is a heme-containing enzyme that catalyzes the oxidative cleavage of the C2-C3 double bond of the indole ring in tryptophan to provide N-formylkynurenine (NFK)[6, 7]. The dioxygenase reaction of Trptophan catalyzed by IDO1 is shown in Scheme 1. This reaction is known as the first and rate-limiting step of the kynurenine pathway [8]. The generated N-Formylkynurenine is then converted into L-kynurenine, 3-hydroxykynurenine, 3-hydroxyanthranilic acid, quinolinic acid, and picolinic acid in the metabolic pathway (Fig. 1) [9,10,11]. The tryptophan depletion results in inhibiting the proliferation of T lymphocytes, which are sensitive to low Trp levels. Production of bioactive metabolites induces the differentiation of regulatory T cells and apoptosis of effector T cells that gave rise to immunosuppression. Both are associated with the immunosuppressed state of the tumor microenvironment [12,13,14]. IDO1 overexpression has been observed in a variety of tumors, including breast [15], lung [16], colon [17, 18], prostate [19], bladder [20], melanoma and neuroblastoma [21]. IDO1 is a promising target for cancer immunotherapy [22,23,24].
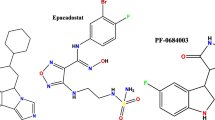
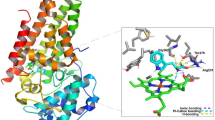
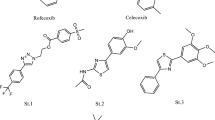
Recently, there are several classes of IDO1 inhibitors have been discovered (Fig. 2). The first IDO1 inhibitor, 1-methyl-D-tryptophan (D-1MT) with Ki of 48 mM developed by New Link Genetics has advanced into clinical development for the treatment of cancer [10, 25]. INCB024360 (also named as epacadostat) developed by Incyte shows potent inhibitory activity (IC50 = 72 nM) against IDO1 [26, 27]. Peng et al. identified an imidazoleisoindole derivative NLG919 as an IDO1 inhibitor, which was later licensed to Genentech as GDC-0919 [28]. GDC-0919 is in the phase 1 study for the treatment of advanced solid tumors [29, 30]. In addition, Shingo et al. reported an imidazothiazole derivative Amg-1(IC50 = 36 μM) as an IDO1 inhibitor, which is the first case that the detailed structural information of the pocket B has been clarified. [6]. The amide side chain of Amg-1 is located at the expanded pocket B, and the methylenedioxyphenyl group is located near the two residues Phe226 and Arg231. The crystal structure of the Amg-1-IDO1 complex (Fig. 3 shows the solid surface representation of crystal structures of Amg-1-IDO1 complex) is very important for structure-based drug design, which provides detailed structural information, where the IDO1 inhibitor interacts with the IDO1 protein at both Pocket A and Pocket B (refer to Fig. 3). Although significant progress has been made in the research mentioned above, there are currently only a few drugs in clinical trials testing IDO1 inhibition as a strategy for the treatment of cancer (e.g. epacadostat and GDC-0919). Thus, it is still a need to discover novel and structurally diverse IDO1 inhibitors with characterized therapeutic utility.
Solid surface representation of crystal structures of Amg-1-IDO1 complex (PDB ID:4PK5). The protein is colored in cyan, cofactor heme and ligand Amg-1 are shown in brown stick model, iron ions is shown in orange balls, the coordination bond are colored in purple dash. The binding sites, pocket A and pocket B, are colored in pink
With the rapid development of computational methods, virtual screening (VS) has been successfully used in many hit compound discovery [8, 31,32,33,34,35,36,37,38]. In this study, we performed a docking-based virtual screening and biological assays to identify potent lead compounds targeting IDO1. ChemDiv (version 2018) database containing 1,492,362 molecules was used to perform the screening. The formation of coordination bonds between small molecules and heme iron atom is crucial for the binding of small molecules and IDO1 protein. Therefore, we use the coordination bond as an important basis for the selection of small molecules while small molecules occupy pocket A and pocket B of IDO1 protein. The cellular assay proved that 24 molecules with potent inhibitory activity in the micro molar range were discovered in our work. Among the 24 active molecules, 8 hits show IC50 values of less than 10 μM and the best one shows an IC50 value of 1.18 ± 0.04 μM. Then we find the analogues with the core scaffold structure of three promising IDO1 inhibitors (compounds 1, 2, 8) from Chemdiv database. 14 analogues were identified and purchased for bioassays. Among the resulted analogues, 7 hits show IC50 values of less than 10 μM and the best one shows an IC50 value of 0.27 ± 0.02 μM. The interaction modes of two compounds (compound1and compound30) against IDO1 were further explored through MCPB.py molecular dynamics simulation, binding free energy calculation and electrostatic potential analysis. The results show that ligand-heme interaction took a large proportion in the binding affinity of inhibitors through metal coordination bond interaction, hydrogen bond interaction and hydrophobic interaction.
Materials and methods
Docking-based virtual screening
Compared with the crystal structure of IDO1 in complex with 4-phenylimidazole (PDB ID:2D0T), the crystal structure of IDO1 in complex with Amg-1 (PDB ID:4pk5) was chosen as the template for molecular docking because of its relatively high resolution and more structural information at pocket B [39].The protein preparation module in Schrödinger 2015 [40]. was used to assign bond orders, add hydrogens, create zero-order bond to metals, create disulfide bonds, deleting water molecules beyond 5 Å from het group, assign partial charge, assign protonation states at pH 7.0, and minimize the structure using the OPLS-2005 force field [41].The Glide 6.6 module in Schrödinger 2015 was used to generate grid, the grid was defined as a 20 × 20 × 20 Å box. In addition, one metal-coordination bond was constrained to heme iron during grid box generation. For the other parameters, the default values were assigned.
The ChemDiv database (version 2018) was used as the source for screening. These compounds were filtered by applying Lipinski’s rule of five. [42] The Ligprep 3.3 module in Schrödinger 2015 was used to generate stereoisomers, and the protonation states of ligands at pH 7.0 ± 2.0 were generated with Epik 3.1. For the other parameters, the default values were assigned.
A flowchart of docking-based virtual screening is shown in Fig. 4. All structures were docked and scored by the Glide high-throughput virtual screening (HTVS) mode, and the 10% top-ranked structures were saved. The saved structures from the previous step were redocked and scored by the Glide standard precision (SP) mode, and the 10% top-ranked structures were saved. Finally, the chosen structures by SP were redocked and scored by the Glide extra precision (XP) scoring mode. The top 3800 molecules ranked by the Glide XP scoring mode were selected and clustered into 100 groups. The compounds that form a metal-coordination with heme iron in each cluster were selected by visual inspection. Finally, 25 compounds were selected and purchased from TOPSCIENCE company for further biological assay evaluation.
HeLa cellular IDO1 assays
The activity of the purchased compounds were evaluated by HeLa cellular IDO1 assays according to the protocol introduced in the references [33, 43]. BMS-986205 was evaluated in the assays as positive compound [44].To perform the HeLa cell based IDO1 assay, HeLa cells were seeded at 50,000 cells per well into 96-well microplate in 100 μl of growth medium. Cells were incubated at 37 °C and 5% CO2 overnight. The next day 100 μl per well of diluted inhibitor in growth medium was added at a final concentration of 100 ng/mL human IFNγ. A series of dilutions are made in 0.1% DMSO in assay medium so that the final concentration of DMSO is 0.1% in all of treatments. Cells were incubated at 37 °C in a CO2 incubator for 24 h. The next day 140 μl of medium was removed into a new 96-well plate and 10 µl of 6.1 N trichloroacetic acid (TCA) was added. The plate was incubated at 50 °C for 30 min to hydrolyze N-formylkynurenine produced by IDO to kynurenine. The plate was then centrifuged at 2500 rpm for 10 min to remove sediments. 100 μl of supernatant per well was transferred to another 96-well plate and mixed with 100 μl of 2% (w/v) 4-(Dimethylamino) benzaldehyde in acetic acid. The plate was incubated at RT for 10 min, the yellow color derived from kynurenine was recorded by measuring absorbance at 480 nm using a microplate reader ((PerkinElmer, USA). The absorbance data were analyzed using the computer software, Graphpad Prism. In the absence of the compound and presence of 100 ng/mL IFNγ, the absorbance (At) in each data set was defined as 100%. The absorbance of medium blank (Ab) in each data set was defined as 0%. The percent absorbance in the presence of each compound was calculated according to the following equation: % Absorbance = (A-Ab)/(At-Ab), where A = the absorbance in the presence of the compound and IFNγ, Ab = the absorbance of medium blank, and At = the absorbance in the absence of the compounds and presence of IFNγ. The IC50 values were calculated using GraphPad Prism 6.0 software.
Similarity searching
The Canvas 2.3 module in Schrödinger 2015 was used to find the analogues of compound 1 with the core scaffold structure of promising IDO1 inhibitor from ChemDiv database.
Molecular dynamics (MD) simulations
The MD simulations of two active compounds (compound 1, 30) in complex with IDO1 were conducted to investigate how the R2 groups in the analogues of compound 1 affect the inhibitory activity against IDO1. The docked structures of the inhibitors in complex with IDO1 were used as the initial structures for MD simulations. MCPB.py [45], a python based metal center parameter builder was used to generate prmtop and coordination parameters for further molecular dynamics simulations. The charge of Fe is set as + 3 state [46].AM1-BCC charge method of antechamber module was used to generate the charges for HEM and ligand. The Gaussian 09 [47] program was used to perform quantum chemistry calculations for metal ion and surrounding atoms by using B3LYP with 6-31G* basis set. The general AMBER force field (gaff) [48] and ff14SB [49] force field were used for the inhibitor and IDO1 respectively. The systems were solvated in a 10 Å cubic box with TIP3PBOX water molecules. Chloride ions are added to neutralize the system. The molecular dynamics simulations were performed in four steps. Firstly, energy minimization was performed to remove possible steric stresses in the system. The systems were subjected to 2500 steps of the steepest descent minimization followed by 2500 steps of conjugated gradient ionization. Then each system was heated from 0 to 300 K in the NVT ensemble with a force constrained of 2.0 kcal mol−1A−2. After that, each system was equilibrated with force constant decreasing from 2.0 to 0 kcal mol−1A−2 in 1 ns. Finally, the production run of 250 ns was performed for each system in the NPT ensemble at 300 K with 1.0 atm pressure. The snapshots for all the trajectories were saved per 2 ps. all the simulations were accomplished by the pmemd module in AMBER18 [50]. The root-mean-square deviation (RMSD), The root-mean-square fluctuations (RMSF), Radius of gyration (Rg), and hydrogen bond occupancy were calculated using the cpptraj module in AMBER18.
MM-GBSA calculation
The binding free energy of compound (1, 30) to IDO1 was analyzed by molecular mechanics generalized born surface area (MM-GBSA) method [51,52,53,54]. 100 snapshots were extracted from the stable 20 ns trajectories and used for MM-GBSA calculation [55]. For each complex system, the binding free energy was calculated according to the equation below:
where Gcomplex, Greceptor, Gligand are the free energy of complex, receptor and ligand molecules, respectively. Egas is the gas-phase energy; Eint is the internal energy; Eele and EvdW are the Coulomb and van der Waals energies, Gsol is the solvation free energy and can be decomposed into polar and nonpolar contributions. GGB is the polar solvation contribution calculated by solving the GB equation. Gnonpl, sol is the nonpolar solvation contribution and was estimated by the SASA determined using a water probe radius of 1.4 Å. The surface tension constant γ was set to 0.0072 kcal/(mol·Å2) [56].T and S are the temperature and solute entropy, respectively. Entropy contributions can be estimated by classical statistical thermodynamics using normal-mode analysis [57]. The entropy cannot be calculated in this protein system because the IDO1 contains heme iron ions. The parameter settings in MM-GBSA calculation were referred to our previous works [58,59,60,61,62,63].
Results and discussion
Docking-based virtual screening and IDO1 inhibitory activity evalulation
The ChemDiv database was used as ligand database for virtual screening. These compounds were filtered by applying Lipinski’s rule of five. Then the remaining molecules were prepared to dock into the catalytic site of IDO1 (PDB ID: 4PK5). The top 3800 compounds ranked by Glide docking score were selected and clustered into 100 groups. The compound of each cluster was selected manually with the following principles: (1) Lower molecular weight; (2) Forming metal ligand interaction with the heme iron; (3) Occupying pocket A and pocket B. Finally, 25 compounds were selected and purchased from TOPSCIENCE company for further biological assay evaluation.
As shown In Table 1 and Fig. 5, among them, 24 compounds showed IC50 values in the range from 1.18 ± 0.04 μM to 35.03 ± 0.96 μM. Eight hits (refer to Table 1, compound1-8) have IC50 values less than 10 μM.The chemical structures of the 25 molecules are shown in Fig. 5. The compounds 1, 2, 8 have the same core scaffold with 1, 3, 4-oxazole ring and urea structure. The concentration-dependent inhibition of IDO1 activities for control inhibitor, BMS-986205 (a) compound 1(b), compound 2(c) and compound 8(d) are shown in Fig. 6. The concentration-dependent inhibition of IDO1 activities for the other compounds are displayed in the supporting information (Fig. S2).
Molecular docking and binding mode prediction.
To demonstrate the receptor-ligand interaction patterns, the docking poses generated by Glide XP scoring were analyzed. The monomer of IDO1 contains two pockets, pocket A and pocket B[64,65,66,67] (Fig. 3). The crystal structure of IDO1 bound to an imidazothiazole derivative,Amg-1(PDB: 4PK5) demonstrated that the nitrogen atom of the scaffold bound to the heme iron, the p-tolyl group occupied pocket A, and the methylenedioxyphenyl group was accommodated in another hydrophobic pocket at the entrance to the binding site, pocket B (Fig. 7a).The predicted binding mode of compound 1 complex with IDO1 was compared with the X-ray crystal structure of Amg-1 complex with IDO1.As shown in Fig. 7b, compound 1 displays a similar binding mode to Amg-1,and one nitrogen atom of the 1,3,4-Oxazole ring coordinated with the heme iron. The major difference between the binding modes of these two compounds is at the binding pocket B, where 3, 4-dimethoxyphenyl group of compound 1 occupies the pocket B that extends upwards. Compound 1, 2 and 8 (Fig. 7. b, c, d) share the same pockets. The size and shape of pocket B differ depending on the bound ligands [1].
The binding mode of Amg-1 (a), compound 1 (b), compound 2 (c) and compound 8 (d) in complex with IDO1. The binding modes of compounds 1, 2, and 8 were predicted by molecular docking. Key residues for ligand binding are shown as sticks. The hydrogen bond between acceptor and ligand are colored in red dash
Structure–activity relationships of the analogues
In order to understand the structure–activity relationships (SARs) of the analogues of compound 1 and discover more potent IDO1 inhibitors with scaffold of 1, 3, 4-Oxazole ring, the similarity search was employed to screen the Chemdiv database. 14 analogues (see supporting information Fig. S1) of compound 1 were identified and purchased for biological assay. The concentration-dependent inhibition of IDO1 activities for the analogues are displayed in the supporting information (Fig. S2). As shown in Table 2, the structure–activity relationship of compounds 1, 2, 8 and 14 analogues is discussed. The results suggested that the analogues of compound 1 also exhibited inhibitory activity against IDO1, which further indicated compound 1 could be characterized as a structurally novel IDO1 inhibitor. We further analyzed the SARs of compound 1 and its analogs to find clues for future structural optimization. Based on the experimental data shown in Table 2, preliminary SARs were discussed below. Our initial array focused on the substitution at R1. Their structures could be divided into three types according to the different R1 groups. When R1 is substituted by p-methylphenyl or methylenedioxyphenyl group rather than phenyl, the inhibitory activity of the compound 1(IC50 = 1.18 ± 0.04 μM) and compound 2 (IC50 = 1.51 ± 0.11 μM) increases significantly compared compound 26(no inhibitory activity).Then, we examined the influence of R2 group. The compounds (1, 27, 28, 8, 29, 30, 31, 32) sharing identical R1 groups with p-methylphenyl were compared. The electron-donating substituent groups at meta or para position of the phenyl group of R2, such as compound 1 showed stronger inhibitory activity than electron-withdrawing groups (compound 28 and 30).Moreover, the comparison between compound 28 and 30 showed that the inhibitory activity was affected by the electron-withdrawing ability, the stronger the electron-withdrawing ability (-F > -COCH3) of the group, the weaker the inhibitory activity of the compound. Besides, the lack of inhibitory effect of compound 32 may be due to the large steric hindrance caused by more substituents on the benzyl group of the R 2 group. Similarly, the compounds (33, 34, 35, 2, 36, 37, 38) sharing identical R1 groups with methylenedioxyphenyl group were compared. The electron donating group may enhance the compound’s inhibitory activity by comparing compounds 33 and 38. Finally, we compared compounds 1 and 39.We found that when 1, 3, 4-oxadiazole and urea are located in the meta position of the benzene ring, the compound has a good inhibitory activity (compound1 IC50 = 1.18 ± 0.04 μM), when 1, 3, 4—oxadiazole and the urea are in the para position of the benzene ring, the compound has a no inhibitory activity (Compound 39).
Molecular dynamics simulations
Trajectories generated from MD simulations were further analyzed. The ligand structure and the number of ligand nitrogen and oxygen atoms in the two complex systems (compound1-IDO1, compound 30-IDO1) for molecular simulations are shown in Fig. 8. The root-mean-square deviation (RMSD) value is a useful basis for quantifying conformational changes in the same protein. As shown in Fig. 9a, The RMSD plots showed that the compound 30-IDO1 systems reach a convergence after 10 ns.The compound 1-IDO1 complex finally reached a stable state after a period of 200 ns of MD trajectory. The RMSD distributions of two systems were shown in Fig. 9b. The smaller RMSD values of compound 30-IDO1 complex may be due to the smaller radius of the F atom in the R2 group, which induces a smaller B pocket. Radius of gyration (Rg) reflects the compactness of the structure. As shown in Fig. 9c. Rg values of the two complexes were stable during simulations. The root-mean-square fluctuations (RMSF) values are used to determine the flexibility of binding pocket residues during the simulation. The simulations were observed to have similar RMSF trend and more fluction was found C- and N-terminals of IDO1 protein (Fig. 9d).
The hydrogen bond analysis results for MD trajectory were shown in Table 3. It is revealed that strong H-bond interactions with high occupancies mainly involving the backbone oxygen atom in Gly251, the backbone oxygen atom in Leu 223 with nitrogen atom on the urea group of the ligand. For compound 1-IDO1 complex, Oxygen 2 atom of compound 1 as a hydrogen bond acceptor in the R2 group additionally forms a hydrogen bond with the NH group of backbone of Lys227 of IDO1 protein, which may be one of the reasons why compound 1 have better inhibition activity against IDO1.
In order to explore the binding affinity of the two ligands, the binding Enthalpy change (ΔH) was calculated using MM-GBSA method. A total of 100 snapshots were extracted from the stable 20 ns trajectory for ΔH analysis. As summarized in Table 4.The predicted ΔH for compound 1, compound 30 are − 61.74 kcal/mol, − 47.87 kcal/mol respectively. Compound 1 has stronger binding affinity to IDO1 than compound 30 according to MM-GBSA method, which is consistent with the experimental results. The ΔH components showed in Table 4 suggest that ΔEele and ΔEvdw are the major contributions for the of ΔH the two compounds, where the polar solvation energies generate the unfavorable contributions.
Clustering analysis of the IDO1-inhibitor complex structures was carried out to obtain more detailed structural information of these two systems. Each trajectory was divided into five clusters by average linkage algorithm. Five snapshots were selected as the representative structures of these five clusters and named as C0, C1, C2, and C3 and C4.For each system, the most stable cluster is C0 and summarized in Table 5, which will be used for molecular electrostatic potentials analysis. We extract the ligands in the representative structures of C0 cluster and then calculate the single point energy of these structures. The grid data of fragmental electron density potential were generated by Multiwfn [68] and visualized by VMD [69].
The results demonstrate that nitrogen 2 atom in 1, 3, 4- oxadiazole has the tendency to form coordination bond with heme iron. So we focus on the difference in electrostatic potential on the nitrogen 2 atom of 1, 3, and 4- oxadiazole. As we can see from Fig. 10, for compounds 1(a) and compound 30(b), the ESPs around the iron coordinating nitrogen 2 atom of 1,3,4- oxadiazole are -41.50 and -39.17, respectively. The large negative value of the coordinating atom of compound 1 was owning to eletron-donating substituent group (-OCH3) of R2.As the charge on the coordinating atom decreased, there was an increase in binding affinity to heme iron[70]. This suggested that ligand-heme coordination interaction take a large proportion in the binding affinity of inhibitors.
Conclusions
In this study, among the 39 compounds identified by the structure-based virtual screening and similarity search, 35 showed obvious IDO1 inhibitory activities. In total, 15 hits have the IC50 values below 10 μM, and two hits below 1 μM. Besides, SARs analysis showed that the compound had good inhibitory activity when R1 was substituted by p-methylphenyl or methylenedioxyphenyl instead of phenyl group and substituent of benzyl group on R2 is an electron-donating group. It is important that the compound has good inhibitory activity when 1, 3, 4-oxadiazole and urea are in the meta position. MD simulations were performed to study the protein–ligand interactions of above two inhibitors with IDO1 in motion Then ESP analysis was employed to provide in-depth explanations on the important role of heme during ligand binding. Ligand-heme interaction took a large proportion in the binding affinity of inhibitors through metal coordination bond interaction, hydrogen bond interaction and hydrophobic interaction The scaffolds of these inhibitors discovered in current work can be used as the starting hits for further lead optimization.
References
Cheong JE, Ekkati A, Sun L (2018) A patent review of IDO1 inhibitors for cancer. Expert Opin Ther Pat 28(4):317–330. https://doi.org/10.1080/13543776.2018.1441290
Comai S, Bertazzo A, Brughera M, Crotti S (2020) Tryptophan in health and disease. Adv Clin Chem 95:165–218. https://doi.org/10.1016/bs.acc.2019.08.005
Le Floc’h N, Otten W, Merlot E (2011) Tryptophan metabolism, from nutrition to potential therapeutic applications. Amino Acids 41(5):1195–1205. https://doi.org/10.1007/s00726-010-0752-7
Takikawa O (2005) Biochemical and medical aspects of the indoleamine 2,3-dioxygenase-initiated L-tryptophan metabolism. Biochem Biophys Res Commun 338(1):12–19. https://doi.org/10.1016/j.bbrc.2005.09.032
Yang R, Chen Y, Pan L, Yang Y, Zheng Q, Hu Y, Wang Y, Zhang L, Sun Y, Li Z, Meng X (2018) Design, synthesis and structure-activity relationship study of novel naphthoindolizine and indolizinoquinoline-5,12-dione derivatives as IDO1 inhibitors. Bioorg Med Chem 26(17):4886–4897. https://doi.org/10.1016/j.bmc.2018.08.028
Tojo S, Kohno T, Tanaka T, Kamioka S, Ota Y, Ishii T, Kamimoto K, Asano S, Isobe Y (2014) Crystal structures and structure-activity relationships of imidazothiazole derivatives as IDO1 inhibitors. ACS Med Chem Lett 5(10):1119–1123. https://doi.org/10.1021/ml500247w
Yan D, Lin YW, Tan X (2017) Heme-containing enzymes and inhibitors for tryptophan metabolism. Metallomics 9(9):1230–1240
Coluccia A, Passacantilli S, Famiglini V, Sabatino M, Patsilinakos A, Ragno R, Mazzoccoli C, Sisinni L, Okuno A, Takikawa O, Silvestri R, La Regina G (2016) New Inhibitors of Indoleamine 2,3-Dioxygenase 1: Molecular Modeling Studies, Synthesis, and Biological Evaluation. J Med Chem 59(21):9760–9773. https://doi.org/10.1021/acs.jmedchem.6b00718
Shiokawa Z, Kashiwabara E, Yoshidome D, Fukase K, Inuki S, Fujimoto Y (2016) Discovery of a novel scaffold as an indoleamine 2,3-Dioxygenase 1 (IDO1) inhibitor based on the pyrrolopiperazinone alkaloid. Longamide B ChemMedChem 11(24):2682–2689. https://doi.org/10.1002/cmdc.201600446
Brochez L, Chevolet I, Kruse V (2017) The rationale of indoleamine 2,3-dioxygenase inhibition for cancer therapy. Eur J Cancer 76:167–182. https://doi.org/10.1016/j.ejca.2017.01.011
Brant MG, Goodwin-Tindall J, Stover KR, Stafford PM, Wu F, Meek AR, Schiavini P, Wohnig S, Weaver DF (2018) Identification of potent indoleamine 2,3-Dioxygenase 1 (IDO1) inhibitors based on a phenylimidazole scaffold. ACS Med Chem Lett 9(2):131–136. https://doi.org/10.1021/acsmedchemlett.7b00488
Fang K, Dong G, Li Y, He S, Wu Y, Wu S, Wang W, Sheng C (2018) Discovery of novel indoleamine 2,3-Dioxygenase 1 (IDO1) and histone deacetylase (HDAC) dual inhibitors. ACS Med Chem Lett 9(4):312–317. https://doi.org/10.1021/acsmedchemlett.7b00487
Hornyak L, Dobos N, Koncz G, Karanyi Z, Pall D, Szabo Z, Halmos G, Szekvolgyi L (2018) The role of indoleamine-2,3-Dioxygenase in cancer development, diagnostics, and therapy. Front Immunol 9:151. https://doi.org/10.3389/fimmu.2018.00151
Fang K, Dong G, Wang H, He S, Wu S, Wang W, Sheng C (2018) Improving the potency of cancer immunotherapy by dual targeting of IDO1 and DNA. Chem Med Chem 13(1):30–36. https://doi.org/10.1002/cmdc.201700666
Hong R, Zhou Y, Tian X, Wang L, Wu X (2018) Selective inhibition of IDO1, D-1-methyl-tryptophan (D-1MT), effectively increased EpCAM/CD3-bispecific BiTE antibody MT110 efficacy against IDO1(hi)breast cancer via enhancing immune cells activity. Int Immunopharmacol 54:118–124. https://doi.org/10.1016/j.intimp.2017.10.008
Volaric A, Gentzler R, Hall R, Mehaffey JH, Stelow EB, Bullock TN, Martin LW, Mills AM (2018) Indoleamine-2,3-Dioxygenase in non-small cell lung cancer: a targetable mechanism of immune resistance frequently coexpressed with PD-L1. Am J Surg Pathol 42(9):1216–1223. https://doi.org/10.1097/PAS.0000000000001099
Lee SJ, Jun S-Y, Lee IH, Kang BW, Park SY, Kim HJ, Park JS, Choi G-S, Yoon G, Kim JG (2018) CD274, LAG3, and IDO1 expressions in tumor-infiltrating immune cells as prognostic biomarker for patients with MSI-high colon cancer. J Cancer Res Clin Oncol 144(6):1005–1014. https://doi.org/10.1007/s00432-018-2620-x
Chen IC, Lee KH, Hsu YH, Wang WR, Chen CM, Cheng YW (2016) Expression pattern and clinicopathological relevance of the indoleamine 2,3-Dioxygenase 1/Tryptophan 2,3-Dioxygenase protein in colorectal cancer. Dis Markers 2016:8169724. https://doi.org/10.1155/2016/8169724
Kolijn K, Verhoef EI, Smid M, Böttcher R, Jenster GW, Debets R, van Leenders GJLH (2018) Epithelial-mesenchymal transition in human prostate cancer demonstrates enhanced immune evasion marked by IDO1 expression. Cancer Res 78(16):4671–4679. https://doi.org/10.1158/0008-5472.can-17-3752
Zhang W, Zhang J, Zhang Z, Guo Y, Wu Y, Wang R, Wang L, Mao S, Yao X (2018) Overexpression of indoleamine 2,3-Dioxygenase 1 promotes epithelial-mesenchymal transition by activation of the IL-6/STAT3/PD-L1 pathway in bladder cancer. Transl Oncol 12(3):485–492. https://doi.org/10.1016/j.tranon.2018.11.012
Brody JR, Costantino CL, Berger AC, Sato T, Lisanti MP, Yeo CJ, Emmons RV, Witkiewicz AK (2009) Expression of indoleamine 2,3-dioxygenase in metastatic malignant melanoma recruits regulatory T cells to avoid immune detection and affects survival. Cell Cycle 8(12):1930–1934. https://doi.org/10.4161/cc.8.12.8745
Le Naour J, Galluzzi L, Zitvogel L, Kroemer G, Vacchelli E (2020) Trial watch: IDO inhibitors in cancer therapy. Oncoimmunology 9(1):1777625. https://doi.org/10.1080/2162402X.2020.1777625
Jiang T, Sun Y (2015) Research progress of indoleamine 2,3-dioxygenase inhibitors. Future Med Chem 7(2):185–201. https://doi.org/10.4155/fmc.14.151
Komiya T, Huang CH (2018) Updates in the clinical development of epacadostat and other indoleamine 2,3-Dioxygenase 1 inhibitors (IDO1) for human cancers. Front Oncol 8:423. https://doi.org/10.3389/fonc.2018.00423
Hou DY, Muller AJ, Sharma MD, DuHadaway J, Banerjee T, Johnson M, Mellor AL, Prendergast GC, Munn DH (2007) Inhibition of indoleamine 2,3-dioxygenase in dendritic cells by stereoisomers of 1-methyl-tryptophan correlates with antitumor responses. Cancer Res 67(2):792–801. https://doi.org/10.1158/0008-5472.CAN-06-2925
Yue EW, Douty B, Wayland B, Bower M, Liu X, Leffet L, Wang Q, Bowman KJ, Hansbury MJ, Liu C, Wei M, Li Y, Wynn R, Burn TC, Koblish HK, Fridman JS, Metcalf B, Scherle PA, Combs AP (2009) Discovery of potent competitive inhibitors of indoleamine 2,3-dioxygenase with in vivo pharmacodynamic activity and efficacy in a mouse melanoma model. J Med Chem 52(23):7364–7367. https://doi.org/10.1021/jm900518f
Yue EW, Sparks R, Polam P, Modi D, Douty B, Wayland B, Glass B, Takvorian A, Glenn J, Zhu W, Bower M, Liu X, Leffet L, Wang Q, Bowman KJ, Hansbury MJ, Wei M, Li Y, Wynn R, Burn TC, Koblish HK, Fridman JS, Emm T, Scherle PA, Metcalf B, Combs AP (2017) INCB24360 (Epacadostat), a Highly Potent and Selective Indoleamine-2,3-dioxygenase 1 (IDO1) Inhibitor for Immuno-oncology. ACS Med Chem Lett 8(5):486–491. https://doi.org/10.1021/acsmedchemlett.6b00391
Peng YH, Ueng SH, Tseng CT, Hung MS, Song JS, Wu JS, Liao FY, Fan YS, Wu MH, Hsiao WC, Hsueh CC, Lin SY, Cheng CY, Tu CH, Lee LC, Cheng MF, Shia KS, Shih C, Wu SY (2016) Important Hydrogen Bond Networks in Indoleamine 2,3-Dioxygenase 1 (IDO1) Inhibitor Design Revealed by Crystal Structures of Imidazoleisoindole Derivatives with IDO1. J Med Chem 59(1):282–293. https://doi.org/10.1021/acs.jmedchem.5b01390
Nayak-Kapoor A, Hao Z, Sadek R, Dobbins R, Marshall L, Vahanian NN, Jay Ramsey W, Kennedy E, Mautino MR, Link CJ, Lin RS, Royer-Joo S, Liang X, Salphati L, Morrissey KM, Mahrus S, McCall B, Pirzkall A, Munn DH, Janik JE, Khleif SN (2018) Phase Ia study of the indoleamine 2,3-dioxygenase 1 (IDO1) inhibitor navoximod (GDC-0919) in patients with recurrent advanced solid tumors. J Immunother Cancer 6(1):61. https://doi.org/10.1186/s40425-018-0351-9
Zhao Y, Wang B, Liu J, Sun P, Liu H (2018) An overview on the methods of determining the activity of indoleamine 2, 3-dioxygenase 1. J Drug Target 27(7):724–731. https://doi.org/10.1080/1061186X.2018.1523416
Xu L, Zhang Y, Zheng L, Qiao C, Li Y, Li D, Zhen X, Hou T (2014) Discovery of novel inhibitors targeting the macrophage migration inhibitory factor via structure-based virtual screening and bioassays. J Med Chem 57(9):3737–3745. https://doi.org/10.1021/jm401908w
Shen M, Tian S, Pan P, Sun H, Li D, Li Y, Zhou H, Li C, Lee SM, Hou T (2015) Discovery of novel ROCK1 inhibitors via integrated virtual screening strategy and bioassays. Sci Rep 5:16749. https://doi.org/10.1038/srep16749
Zhang G, Xing J, Wang Y, Wang L, Ye Y, Lu D, Zhao J, Luo X, Zheng M, Yan S (2018) Discovery of novel inhibitors of indoleamine 2,3-Dioxygenase 1 through structure-based virtual screening. Front Pharmacol 9:277. https://doi.org/10.3389/fphar.2018.00277
Chong CM, Kou MT, Pan P, Zhou H, Ai N, Li C, Zhong HJ, Leung CH, Hou T, Lee SM (2016) Discovery of a novel ROCK2 inhibitor with anti-migration effects via docking and high-content drug screening. Mol Biosyst 12(9):2713–2721. https://doi.org/10.1039/c6mb00343e
Tian S, Wang X, Li L, Zhang X, Li Y, Zhu F, Hou T, Zhen X (2017) Discovery of novel and selective adenosine A2A receptor antagonists for treating parkinson’s disease through comparative structure-based virtual screening. J Chem Inf Model 57(6):1474–1487. https://doi.org/10.1021/acs.jcim.7b00188
Tomek P, Palmer BD, Flanagan JU, Sun C, Raven EL, Ching LM (2017) Discovery and evaluation of inhibitors to the immunosuppressive enzyme indoleamine 2,3-dioxygenase 1 (IDO1): Probing the active site-inhibitor interactions. Eur J Med Chem 126:983–996. https://doi.org/10.1016/j.ejmech.2016.12.029
Zhong H, Wang Z, Wang X, Liu H, Li D, Liu H, Yao X, Hou T (2019) Importance of a crystalline water network in docking-based virtual screening: a case study of BRD4. Phys Chem Chem Phys 21(45):25276–25289. https://doi.org/10.1039/C9CP04290C
Wang Z, Sun H, Shen C, Hu X, Gao J, Li D, Cao D, Hou T (2020) Combined strategies in structure-based virtual screening. Phys Chem Chem Phys 22(6):3149–3159. https://doi.org/10.1039/c9cp06303j
Zou Y, Wang F, Wang Y, Guo W, Zhang Y, Xu Q, Lai Y (2017) Systematic study of imidazoles inhibiting IDO1 via the integration of molecular mechanics and quantum mechanics calculations. Eur J Med Chem 131:152–170. https://doi.org/10.1016/j.ejmech.2017.03.021
Schrödinger, version 10.1; Schrödinger, LLC: New York (2015).
Kaminski GA, Friesner RA, Tirado-Rives J, Jorgensen WL (2001) Evaluation and reparametrization of the OPLS-AA force field for proteins via comparison with accurate quantum chemical calculations on peptides. J Phys Chem 105(28):6474–6487. https://doi.org/10.1021/jp003919d
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1(4):337–341. https://doi.org/10.1016/j.ddtec.2004.11.007
Fung SP, Wang H, Tomek P, Squire CJ, Flanagan JU, Palmer BD, Bridewell DJ, Tijono SM, Jamie JF, Ching LM (2013) Discovery and characterisation of hydrazines as inhibitors of the immune suppressive enzyme, indoleamine 2,3-dioxygenase 1 (IDO1). Bioorg Med Chem 21(24):7595–7603. https://doi.org/10.1016/j.bmc.2013.10.037
Richards T, Brin E (2018) Cell based functional assays for IDO1 inhibitor screening and characterization. Oncotarget 9(56):30814–30820. https://doi.org/10.18632/oncotarget.25720
Li P, Merz KM Jr (2016) MCPBpy: a python based metal center parameter builder. J Chem Inf Model 56(4):599–604. https://doi.org/10.1021/acs.jcim.5b00674
Oda A, Yamaotsu N, Hirono S (2005) New AMBER force field parameters of heme iron for cytochrome P450s determined by quantum chemical calculations of simplified models. J Comput Chem 26(8):818–826. https://doi.org/10.1002/jcc.20221
Frisch MJ, Schlegel HB, Scuseria GE, Robb MA, Scalmani G, Barone V, Mennucci B, Petersson GA, Caricato HN, Li X, Hratchian HP, Izmaylov AF, Zheng G, Sonnenberg JL, Hada M, Ehara M, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Nakai H, Vreven T, Montgomery JA, Peralta JE, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Kobayashi R, Normand J, Raghavachari K, Burant JC, Iyengar SS, Tomasi J, Cossi M, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Cammi R, Pomelli C, Ochterski JW, Martin RL, Zakrzewski VG, Voth GA, Salvador P, Dapprich S, Daniels AD, Farkas O, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09. Gaussian Inc, Wallingford
Junmei W, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25(9):1157–1174. https://doi.org/10.1002/jcc.20035
Maier JA, Martinez C, Kasavajhala K, Wickstrom L, Hauser KE, Simmerling C (2015) ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J Chem Theory Comput 11(8):3696–3713. https://doi.org/10.1021/acs.jctc.5b00255
Case DA, Cheatham TE 3rd, Darden T, Gohlke H, Luo R, Merz KM Jr, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The Amber biomolecular simulation programs. J Comput Chem 26(16):1668–1688. https://doi.org/10.1002/jcc.20290
Xu L, Sun H, Li Y, Wang J, Hou T (2013) Assessing the performance of MM/PBSA and MM/GBSA methods 3 The impact of force fields and ligand charge models. J Phys Chem B 117(28):8408–8421. https://doi.org/10.1021/jp404160y
Sun H, Li Y, Tian S, Xu L, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods 4 Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set. Phys Chem Chem Phys 16(31):16719–16729. https://doi.org/10.1039/c4cp01388c
Hou T, Wang J, Li Y, Wang W (2011) Assessing the performance of the MM/PBSA and MM/GBSA methods 1 the accuracy of binding free energy calculations based on molecular dynamics simulations. J Chem Inf Model 51(1):69–82. https://doi.org/10.1021/ci100275a
Sun H, Li Y, Shen M, Tian S, Xu L, Pan P, Guan Y, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods 5 Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring. Phys Chem Chem Phys 16(40):22035–22045. https://doi.org/10.1039/C4CP03179B
Miller BR 3rd, McGee TD Jr, Swails JM, Homeyer N, Gohlke H, Roitberg AE (2012) MMPBSApy: An Efficient Program for End-State Free Energy Calculations. J Chem Theory Comput 8(9):3314–3321. https://doi.org/10.1021/ct300418h
Sitkoff D, Sharp KA, Honig B (1994) Accurate calculation of hydration free energies using macroscopic solvent models. J Phys Chem 98(7):1978–1988. https://doi.org/10.1021/j100058a043
Pearlman DA, Connelly PR (1995) Determination of the differential effects of hydrogen bonding and water release on the binding of FK506 to native and Tyr82–>Phe82 FKBP-12 proteins using free energy simulations. J Mol Biol 248(3):696–717
Guo J, Wang X, Sun H, Liu H, Yao X (2012) The molecular basis of IGF-II/IGF2R recognition: a combined molecular dynamics simulation, free-energy calculation and computational alanine scanning study. J Mol Model 18(4):1421–1430. https://doi.org/10.1007/s00894-011-1159-4
Yang Y, Liu H, Yao X (2012) Understanding the molecular basis of MK2-p38alpha signaling complex assembly: insights into protein-protein interaction by molecular dynamics and free energy studies. Mol Biosyst 8(8):2106–2118. https://doi.org/10.1039/c2mb25042j
Pan D, Niu Y, Xue W, Bai Q, Liu H, Yao X (2016) Computational study on the drug resistance mechanism of hepatitis C virus NS5B RNA-dependent RNA polymerase mutants to BMS-791325 by molecular dynamics simulation and binding free energy calculations. Chemometer Intell Lab Syst 154:185–193. https://doi.org/10.1016/j.chemolab.2016.03.015
Li L, Wei W, Jia WJ, Zhu Y, Zhang Y, Chen JH, Tian J, Liu H, He YX, Yao X (2017) Discovery of small molecules binding to the normal conformation of prion by combining virtual screening and multiple biological activity evaluation methods. J Comput Aided Mol Des 31(12):1053–1062. https://doi.org/10.1007/s10822-017-0086-6
Shi D, Zhou S, Liu X, Zhao C, Liu H (1862) Yao X (2018) Understanding the structural and energetic basis of PD-1 and monoclonal antibodies bound to PD-L1: A molecular modeling perspective. Biochim Biophys Acta Gen Subj 3:576–588. https://doi.org/10.1016/j.bbagen.2017.11.022
Liu H, Yao X (2010) Molecular basis of the interaction for an essential subunit PA-PB1 in influenza virus RNA polymerase: insights from molecular dynamics simulation and free energy calculation. Mol Pharm 7(1):75–85. https://doi.org/10.1021/mp90013p
Rohrig UF, Zoete V, Michielin O (2017) The binding mode of N-Hydroxyamidines to indoleamine 2,3-Dioxygenase 1 (IDO1). Biochemistry 56(33):4323–4325. https://doi.org/10.1021/acs.biochem.7b00586
RoHrig UF, Loay A, Aurélien G, Pierre L, Vincent S, Didier C, Vincenzo C, Simpson AJG, Pierre V, Eynde BTJ, Den V (2010) Rational design of indoleamine 2,3-dioxygenase inhibitors. J Med Chem 53(3):1172–1189. https://doi.org/10.1021/jm9014718
Greco FA, Bournique A, Coletti A, Custodi C, Dolciami D, Carotti A, Macchiarulo A (2016) Docking studies and molecular dynamic simulations reveal different features of IDO1 structure. Mol Inform 35(8–9):449–459. https://doi.org/10.1002/minf.201501038
Rohrig UF, Majjigapu SR, Grosdidier A, Bron S, Stroobant V, Pilotte L, Colau D, Vogel P, Van den Eynde BJ, Zoete V, Michielin O (2012) Rational design of 4-aryl-1,2,3-triazoles for indoleamine 2,3-dioxygenase 1 inhibition. J Med Chem 55(11):5270–5290. https://doi.org/10.1021/jm300260v
Lu T, Chen F (2012) Multiwfn: a multifunctional wavefunction analyzer. J Comput Chem 33(5):580–592. https://doi.org/10.1002/jcc.22885
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Gaspari P, Banerjee T, Malachowski WP, Muller AJ, Prendergast GC, DuHadaway J, Bennett S, Donovan AM (2006) Structure−activity study of brassinin derivatives as indoleamine 2,3-dioxygenase inhibitors. J Med Chem 49(2):684–692. https://doi.org/10.1021/jm0508888
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 21775060).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10822_2021_386_MOESM1_ESM.docx
Supplementary file1 Fig. S1 14 analogues of compound 1 identified by similarity searching. Fig. S2 The concentration-dependent inhibition of IDO1 activities for all active compounds. (DOCX 8571 kb)
Rights and permissions
About this article
Cite this article
Ge, H., Mao, L., Zhao, J. et al. Discovery of novel IDO1 inhibitors via structure-based virtual screening and biological assays. J Comput Aided Mol Des 35, 679–694 (2021). https://doi.org/10.1007/s10822-021-00386-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-021-00386-6